Abstract
The inflammatory response is an energy-intensive process. Consequently, metabolism is closely associated with immune function. The autophagy machinery plays a role in metabolism by providing energy but may also be used to attack invading pathogens (xenophagy). The autophagy machinery may function to protect against not only the threats of infection but also the threats of the host’s own response acting on the central immunological tolerance and the negative regulation of innate and inflammatory signaling. The balance between too little and too much autophagy is critical for the survival of immune cells because autophagy is linked to type 2-cell death programmed necrosis and apoptosis. Changes in inflammatory cells are driven by extracellular signals; however, the mechanisms by which cytokines mediate autophagy regulation and govern immune cell function remain unknown. Certain cytokines increase autophagy, whereas others inhibit autophagy. The relationship between autophagy and inflammation is also important in the pathogenesis of metabolic, non-communicable diseases. Inflammation per se is not the cause of obesity-associated diseases, but it is secondary to both the positive energy balance and the specific cellular responses. In metabolic tissues, the suppression of autophagy increases inflammation with the overexpression of cytokines, resulting in an activation of autophagy. The physiological role of these apparently contradictory findings remains uncertain but exemplifies future challenges in the therapeutic modulation of autophagy in the management of disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
6.1 Basic Concepts and Background
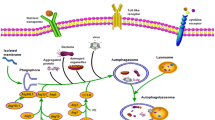
Disturbances in the normal function of cells (cellular stress) result in the accumulation of damaged molecules and dysfunctional organelles that may be deleterious. To mitigate this damage, cells clean up the accumulated end products using the following two major systems: the ubiquitin proteasome system and the autophagy (from the Greek for “self-digestion” or “self-eating”) lysosome system. The key elements in the recognition and processing of ubiquitin-protein conjugates have been recently summarized [1], but the exact mechanisms by which cells regulate self-cannibalism are still under investigation. Chaperone-mediated autophagy, microautophagy, and macroautophagy (hereafter referred to as autophagy) are currently considered distinctive forms [2] (Fig. 6.1). Each form differs from the others in its physiological functions and the mode and nature of cargo delivery.
Autophagy was first described in mammalian cells over 50 years ago, but the molecular basis for this process has not been completely elucidated. The following three autophagy forms have been identified: chaperon-mediated autophagy, microautophagy and macroautophagy (autophagy). LAMP lysosomal associated membrane protein
The overall process of autophagy can be separated into the following seven discrete steps: induction or selection/packaging of cargo, nucleation, vesicle expansion, completion, fusion, degradation, and export (Fig. 6.2). A discussion of the known (and supposed) molecular mechanisms is beyond the scope of this manuscript, but it appears that autophagy is generally induced to provide an alternate source of certain basic building blocks required for cell survival (the non-selective pathway). The mechanistic target of rapamycin (MTOR) kinase coordinates nutrient availability and cell growth. When the supply is sufficient, MTOR is active and phosphorylates important proteins for cellular growth. When nutrients are in a limited supply, MTOR is inactivated and limits protein translation, focusing on those required for cell survival [3]. This mechanism might be easily modulated by the action of common and inexpensive drugs, such as metformin [4] and chloroquine [5], but might also be regulated by selected stimuli (i.e., the selective pathway initiated from the cargo itself).
6.1.1 Autophagy: A Tightly Regulated Process
Autophagy is a simple process and is used to provide energy under starvation conditions, but it must be tightly regulated because it has the capacity to be harmful (i.e., it can degrade entire organelles). In mammals, as opposed to yeasts, the regulation of this process is exquisitely complex because it is also used, among other functions, for the purposes of development; consequently, the mechanisms of autophagy must be initiated at precise times. The targeted degradation of a cellular component requires signals derived from either the cargo selected for degradation or the particular function of the cell. The regulation of this process is poorly understood, but it is likely that certain selective autophagy pathways are relevant to disease.
For example, autophagy of mitochondria (mitophagy) is involved in the response of the cell to pernicious stimuli and, particularly, in the avoidance of effects from inflammation or oxidative stress (details are summarized in Fig. 6.3). This response is clearly observed in conditional knockout mouse models [6], which accumulate deformed mitochondria in the liver under defective autophagy conditions. It has also been observed that chemical inhibitors of autophagy and mitochondrial depolarization (cyclosporine) jeopardize the defensive role of mitophagy in hepatocytes [7]. The autophagy of peroxisomes (pexophagy) has only been explored in yeasts, and it is difficult to differentiate the process from peroxisomal biogenesis; however, a role in mammals is likely to exist. To further complicate the issue, there is cross communication between the two major pathways of protein degradation, i.e., substrates that are normally degraded by the ubiquitin-proteasome pathway can be selectively degraded by autophagy under certain circumstances [8]. The induction of autophagy is also critical in the maintenance of glycogen stores. Glycogen autophagy in the liver, heart and diaphragm is a selective and highly regulated process that is most likely due to the abrupt increases in the energy requirement under critical conditions, such as birth [9].
Mitochondria, apoptosis, autophagy and inflammation are closely related. Autophagy limits both inflammation and apoptosis (cell death). Conversely, the loss of autophagy (mitophagy) leads to the accumulation of damaged mitochondria, which promotes inflammation. In healthy mitochondria, BCL2 and BCL-XL bind to Beclin 1, which is inhibited. PTEN-induced putative kinase 1 (PINK1) is a mitochondrial serine/threonine-protein kinase that apparently protects cells from stress-induced mitochondrial dysfunction. PINK1 is imported into healthy mitochondria, according to the mitochondrial transmembrane potential (Δψm), where it is degraded by the protease PARL (presenilins-associated rhomboid-like protein). Mitophagy is activated by a number of factors. For example, Beclin 1 may be displaced by hypoxia. In response to the uncoupling, the mitochondrial permeability transition (MPT) or the mitochondrial damage, there is an accumulation of full-length PINK1 and an excess amount of ROS. Autophagy or increased mitochondrial turnover may be the consequence of AMP accumulation (decreased ATP), leading to the activation by AMPK of proteins that function specifically in mitophagy. Other lethal stimuli lead to BAX- or BAK-mediated mitochondrial outer membrane permeabilization (MOMP) or to the activation of the permeability transition pore complex (PTPC). The net consequence is the release of intermembrane space proteins (IMSP) and other conjugates with pro-apoptotic and/or pro-inflammatory action. Consequently, mitophagy acts as a defensive mechanism to alleviate inflammation, cell death and mitochondrial dysfunction
6.2 The Relevance of Autophagy in Disease
6.2.1 Autophagy Is Important in the Pathogenesis of Cancer, Neurodegeneration and Myopathies
Autophagy has been implicated in the pathogenesis of several diseases, primarily as the consequence of an imbalance in its dual role of adapting to cellular stresses and/or contributing to cell death. Mitochondria are particularly important in the process.
There are at least three different types of programmed cell death as follows: type I (nuclear or apoptotic), type II (autophagic), and type III (cytoplasmic). The autophagic and apoptotic pathways are closely related [10], although autophagy is relatively unknown compared to apoptosis, and it is unclear whether autophagy represents a failed effort to preserve cell viability. This observation is particularly evident in cancer. Several findings have led to the hypotheses that BECN1 (beclin-1) and other genes involved in autophagy may function as tumor suppressors and that defects in autophagy may promote cancer. Similarly, the PI3K (phosphoinositide 3 kinase)/AKT (protein kinase B)/MTOR pathway is frequently activated in response to mutations in genes encoding negative regulators, thereby constraining autophagy in certain tumors [5, 11–13]. Many signals promoting unrestricted cell proliferation also inhibit autophagy, which is normally induced to sustain cells under conditions of nutrient limitation. Interestingly, the uncoupling of the cellular response to the nutrient availability renders cells more susceptible to a metabolic catastrophe; thus, the concept of “oncometabolite” and the requirement for further insights in cancer metabolism are rapidly evolving. This concept may be a first step in understanding why the defensive metabolic and inflammatory responses to tumor necrosis may promote rather than entangle an increase in the overall tumor burden [14]. Cancer in adults, but not in children, arises under chronic inflammation circumstances, in a tumor microenvironment that is characterized by hypoxia, glycolysis, perpetual autophagy and the resultant necrosis under conditions of stress. The regulation of autophagy in tumor cells provides possible therapeutic strategies, although given the complexity of the signaling pathways, these should be tailored to the different classes of tumors. For example, in tumors that do not outgrow their food supply, the inhibition of autophagy may render cells more susceptible to conventional treatments. Conversely, the stimulation of autophagy in tumors with constitutive activation of the PI3K/AKT pathway can render them more susceptible to apoptosis.
Autophagic activity is essential for the normal function of the nervous system. A deficiency of basal autophagy in the mouse brain (with a knockout of autophagic proteins) results in neurodegeneration [15]. Although the role of autophagy may vary in different neurodegenerative diseases, age-related neurodegenerative diseases are generally characterized by the accumulation of protein aggregates in the affected brain. These protein aggregates may be degraded by autophagy [16], suggesting that a regulated and specific increase in autophagy may become a protective response in both Alzheimer’s and Parkinson’s diseases. Lysosomal storage disorders, which result in alterations of the lysosomal function, should inhibit autophagosome maturation and the subsequent accumulation of autophagic vacuoles in the affected cells. Inefficient autophagolysosomal recycling of mitochondria may generate fragmented mitochondria and an increased sensitivity to apoptosis, thus complicating the course of the disease [17]. Many congenital myopathies are characterized by the presence of myofibrillar disorganization and the accumulation of autophagic vacuoles. In this context, increased autophagy appears to be involved in pathways leading to muscle wasting, an effect that should be considered in the pharmacological modulation of this process [18].
6.2.2 The Role of Autophagy in Infection and Inflammatory Disorders
Recent studies have demonstrated the role of autophagy in processes affecting the immune system, including the coordination of metabolic signals, immune cell differentiation, secretion of cytokines and both innate and adaptive immune defenses against pathogens. Therefore, it is tempting to speculate that detailed insights into the function of autophagy may provide novel therapeutic strategies in the management of inflammatory disorders.
6.2.2.1 The Relationship Between Autophagy and Immunity
Autophagosomes play an important role in the major histocompatibility complex (MHC) class II antigen presentation. The endocytosis of protein antigens from the extracellular space is followed by autophagy and subsequent lysosomal degradation and transfer to the MHC class II loading compartment prior to transport to the cell surface [19]. Other studies have indicated that dendritic cells (DCs) require autophagy to efficiently process and present antigens on the MHC class II molecules. This process is not limited to certain antigens, and autophagy also appears to be important in the MHC class I antigen presentation [20]. Other studies have implicated the signaling lymphocyte activation molecule, a cell adhesion protein, as a bacterial sensor that regulates the degradation of endocytosed gram-negative bacteria in macrophages via NADPH oxidase activity and autophagy [21]. Receptor ligation is also important in the defense against other pathogens and in the function of other cells, such as neutrophils, to induce autophagy, a potential mechanism for clearing pathogens that has also been associated with facilitating the recognition of pathogen-associated molecular patterns. Autophagy is involved in the mechanisms by which neutrophils capture microbes through neutrophil extracellular traps (NETs) that induce a form of cell death, termed NETosis [22]. Several autophagy proteins also play a role in B cell development and survival. Beclin 1–deficient chimeric mice have reduced numbers of lymphoid progenitor cells in the bone marrow, indicating a role for Beclin 1 in early B cell development. These and other available results demonstrate that autophagy plays a role in several aspects of B cell physiology, development, and maintenance, MHC class II presentation, and co-stimulation through surface receptors [23]. Moreover, autophagic pathways intersect with a variety of T cell functions, including the development of CD4+ T cell self-tolerance. Similar to B cells, multiple autophagy proteins are essential for the development, maintenance, and survival of T cells. An investigation of autophagy-deficient peripheral T cells showed that they had excess mitochondria and elevated reactive oxygen species (ROS) production. Autophagy maintains mitochondrial homeostasis, and mitophagy is responsible for removing damaged mitochondria and preventing their accumulation (Fig. 6.3). Proliferating T cells clear the mitochondria as they shift to a more glycolytic metabolism after activation, and blocking autophagy can impair ATP production in T cells [24]. Other results indicate that autophagy plays a role in regulating the endoplasmic reticulum (ER) and calcium homeostasis in T cells and that impaired autophagy can lead to ER stress and impaired calcium homeostasis. Alternatively, autophagy can be induced by ER stress–signaling pathways independent of the unfolded protein response that activates NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells). Therefore, these roles in maintaining the ER and mitochondrial homeostasis and in controlling the ROS levels could explain the importance of autophagy proteins in the survival of immune cells [25, 26].
6.2.2.2 The Role of Autophagy Against Pathogens (Xenophagy)
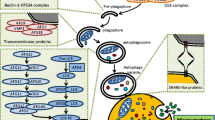
The mechanisms by which autophagy mediates resistance to infection are not fully understood but are likely to involve a combination of the effects summarized in Fig. 6.4. The process of removing large amounts of protein aggregates may be selective and may be used to clear intracellular bacteria, parasites, and viruses. Autophagy appears to be the only pathway to address this function, and if it fails, the only other option for infected cells is cell death and clearance by phagocytes.
The first line of defense during pathogen infection is the activation of innate immune responses. The autophagy machinery is also used to attack invading pathogens (first described in 2004, [27]). This fact and other findings could explain the evolution in certain pathogens of mechanisms that aim to evade, inhibit, and usurp the host cell autophagy to promote survival and replication.
6.2.2.2.1 The Antibacterial Role of Autophagy
Bacteria may enter mammalian cells by mechanisms that are different for gram-negative and gram-positive bacteria [28]. Once inside the host cells, the predominant strategy against bacterial pathogens appears to involve compartmentalization within membrane-bound vacuoles, including autophagosomes [29]. The cellular autophagic response differs according to the nature of the intracellular bacterial pathogen and is involved in sequestering and eliminating intracellular bacteria such as Group A Streptococcus, Salmonella typhimurium, Shigella flexneri, and Listeria monocytogenes. The exact mechanisms are unknown, but the role of autophagy in macrophages compared with that in epithelial cells differs. In epithelial cells, autophagy limits bacterial growth, and in macrophages, mitophagy leads to cell death [30]. Alveolar macrophages ingest the M. tuberculosis bacilli and enclose them in phagosomes, which are arrested in maturation. The bacilli survive and grow within the phagosomes until the macrophages die. Alternatively, the activation of infected macrophages by an interferon induces autophagy, and phagosomes fuse with autophagosomes with the subsequent degradation of the bacilli [31]. In contrast, infection with Legionella pneumophila or Porphyromonas gingivalis activates autophagy, a mechanism that provides a replicative, noxious niche. At least in vitro, the suppression of autophagy results in the trafficking of internalized bacteria to phagolysosomes and degradation of the organism [32].
6.2.2.2.2 The Autophagic Responses to Fungal Infections
Although knowledge in this area is limited, the chemical or genetic disruption of autophagy in murine macrophages resulted in decreased Cryptococcus neoformans uptake, replication, and escape from host cells [33]. The goal of the ongoing studies is to understand the mechanisms by which autophagy proteins determine the ability of C. neoformans to establish a replicative niche and how these functions are coordinated with other host factors.
6.2.2.2.3 The Autophagic Responses to Parasites
Knowledge regarding the autophagic responses to parasites is also limited. However, following infection in macrophages, Toxoplasma gondii, an obligate intracellular protozoan parasite, replicates in specialized vacuoles that are protected from fusion with lysosomes. Experiments in animal models have indicated a role for autophagy involving cell-cell receptor-ligand interactions in activated macrophages that eliminate the parasite [34].
6.2.2.2.4 The Autophagic Responses to Viruses
In this particular type of infection, the virus and the host cell type determine the functions of the autophagy-mediated responses. For example, autophagy is antiviral in response to herpes simplex virus type 1, but viruses have evolved mechanisms to perturb autophagy, most likely using Beclin 1 as a preferential target. Antagonistic interactions between viral proteins and Beclin 1 can subvert autophagosome formation and ultimately promote virulence. Certain viruses, including hepatitis B, hepatitis C, and the Dengue viruses, promote autophagy as a means to encourage viral replication. The exact mechanisms are poorly understood but have been extensively studied concerning infection with human immunodeficiency virus (HIV-1). For example, HIV-1 blocks autophagy in dendritic cells by activating the MTOR pathway, most likely via the interaction of envelope glycoproteins with the CD4 receptor, which leads to an increased cellular viral content, an increased transfer of HIV-1 to CD4+ T cells, an increased virus survival and reduced antigen presentation [35, 36]. Other studies have indicated a major role for autophagy in HIV-1-mediated cell death in uninfected CD4+ T cells. HIV-1 inhibits autophagy in infected CD4+ T cells by downregulating Beclin 1 at the transcriptional level and induces autophagy in uninfected CD4 + T cells [37, 38]. Therefore, HIV-1 manipulates autophagy during infection to evade the immune response and illustrates the risks and benefits induced by the exogenous manipulation of the host autophagy [39, 40].
6.2.2.3 Autophagy and Immunity
Autophagy may also be implicated in autoimmunity by some of the mechanisms listed in Fig. 6.4 (i.e., promotion of the MHC class II presentation of antigens, control of T lymphocyte homeostasis, induction by Th1 cytokines and possibly by facilitation of specific serum autoantibodies) [41]. The role of autophagy in the innate immune system has been extensively studied but remains poorly understood. Pathogen recognition and intracellular killing can be controlled by autophagy, but this process might also act beyond pathogen control and be related to the general function of the clearance of dead cells. In the absence of autophagy, apoptotic cells fail to signal for efficient removal [42], and LC3 II (Microtubule-associated protein 1 light chain 3 alpha)-associated phagocytosis is a requirement for dead cell clearance [43]. This defect may be linked to susceptibility to autoimmunity in mice [42, 43], and it is clinically evident that a number of autoimmune diseases are precipitated or aggravated after infections.
Induced autophagy may exacerbate the process of presentation on MHC-II of peptides from intracellular sources [44]. Dendritic cell-mediated antigen presentation in the context of MHC-II is most likely an area in which autophagy might influence immune diseases because this is the major activator of the adaptive immune cells. Dendritic cells are also important in the pathogenesis of Crohn’s disease because impaired autophagy facilitates the replication of adherent-invasive Escherichia coli, an important factor in the pathogenesis of the disease [45].
Diseases resulting from increased activation of the immune system comprise the following two different categories: autoinflammatory diseases and autoimmune diseases in which the adaptive immune cells target the self-antigens. Inflammasome activation modulates cytokine secretion, and autophagy plays a negative role with respect to inflammasome activation; autophagy deficiency leads to the increased production of IL-1β and IL-18. The IL-1β receptor blockade has beneficial effects in rheumatoid arthritis and has been suggested as a therapy for autoinflammatory diseases [46]. Autophagy is also involved in a number of secretory processes in immune cells (and non-immune cells). Proteins with a signal peptide are secreted through a canonical pathway involving the endoplasmic reticulum. A secretion of proteins without a signal peptide appears to be mediated by autophagy [47]. Interestingly, autophagy is involved in interferon (IFN) secretion. Elevated levels of IFN-α are the hallmark of systemic lupus erythematosus, and ongoing clinical trials are testing the effectiveness of monoclonal antibodies against IFN-α [48]. The paucity of data in humans using agents that modulate autophagy is intriguing. However, the antimalarial drug chloroquine has been used for decades as an anti-inflammatory agent for the treatment of systemic lupus erythematosus and rheumatoid arthritis. A mechanistic explanation has been recently reported indicating that chloroquine promotes the transrepression of proinflammatory cytokines by the glucocorticoid receptor. The fact that glucocorticoid signaling is regulated by lysosomes provides a mechanistic basis for treating autoimmune diseases with a combination of glucocorticoids and lysosomal inhibitors [49].
6.3 Nutritional Stress, Autophagy and Inflammation
6.3.1 Influence of Changes in the Metabolic State on the Immune System
Nutritional factors (i.e., obesity, metabolic syndrome, nutritional restriction, among others) have an important regulatory role in modulating the function of the immune system. Possible mechanisms include changes in secretion of immune mediators by metabolic tissues, alterations in the composition of resident populations of T cells or macrophages, direct effects of metabolites on immune cells and intrinsic defects in signaling pathways. Changes in the autophagic activity of different cell populations are likely to influence the quality of the immune responses that occur in response to nutritional stress or nutritional intervention, particularly in the context of a high fat diet [50–54]. Metabolic complications are associated with a state of subclinical chronic inflammation that has been identified as a pathogenic risk. Because inflammation is not the cause of obesity, we speculate that it may represent a superimposed mechanism that may be defensive and is related to autophagy, oxidation, dietary factors and the consequent metabolic response in specific types of cells.
6.3.2 Chemokine (C-C motif) Ligand 2, Autophagy and Inflammation
Obese adipose tissue, characterized by adipocyte hyperplasia and hypertrophy, is a mediator of low-grade inflammation of the white adipose tissue that is due to a chronic activation of the innate immune system leading to an increase in cytokines, which mediate the recruitment of macrophages and T cells. Certain cytokines, primarily chemokine (C-C motif) ligand 2 (CCL2), also support vasculogenesis.
Autophagy modifies pancreatic beta cell functions regulating glucose homeostasis [55]. Because ER stress is involved in insulin resistance, autophagy might also be involved in insulin resistance by modulating the ER stress response [56]. Inflammation apparently represents an indicator of the failure of adipose tissue in its major function of energy storage. We have recently developed an animal model overexpressing CCL2 that is designed to characterize the relationship between autophagy and cytokine gene expression. The preliminary data suggest that metabolism (i.e., dietary variations) plays a contributory factor; animals overexpressing CCL2 displayed an increased number of autophagosomes and a differential rate of mitophagy as compared with CCL2 deficient animals (Fig. 6.5) [57]. However, the role of CCL2 may vary in response to different environments [58]. Ongoing studies in our model may add information about the regulation of autophagy by cytokines, which is based on relatively limited data obtained in mammalian cells. IFN-γ, a Th1 cytokine, induces autophagy [59], but Th2 cytokines (IL-4 and IL-13) act as suppressors of autophagy and most likely activate the MTOR cascade [60]. More importantly, in vascular smooth muscle cells (VSMC) from atherosclerotic plaques, the expression of autophagy genes in VSMC was controlled by insulin growth factor like-1 (IGF-1) and TNF-α; TNF-α may induce autophagy [61]. This result suggests that IGF-1 can induce cell survival by inhibiting autophagy in plaque-associated VSMC [62, 63]. This effect could be important to provide novel strategies to increase stability of atherosclerotic plaques.
The ingestion of highly caloric, fat-rich, food elicited significant differences in weight increase among mice overexpressing CCL2 when compared to CCL2 knockout animals (a). Additionally, dietary conditions were also responsible for significant changes in the fusion-fission balance in mitochondrial biogenesis (b) and the amount of autophagosomes in the liver (c)
This finding is also relevant in obesity, a condition associated with a frightening disease burden. Although the search for a simple and acceptable nutritionally based therapeutic approach continues, other strategies are clearly needed. The effect on liver morphology and function of obesity and the consequent development of fatty liver disease has received substantial attention in our laboratory [64–66]. It has been repeatedly invoked that autophagy may play a dual role in cell fate, depending on the stimuli and cell type. Therefore, it is essential to consider the effect of autophagy in every metabolic tissue involved in obesity-associated disturbances. Most likely, the effect has been predominantly investigated in adipose tissue and the liver, especially in nonalcoholic fatty liver disease (NAFLD). Intriguingly, the degree of steatosis is heterogeneous among obese patients and a substantial number of patients do not develop this complication. Obese patients with steatosis display a higher number of autophagosomes in the liver and toroidal mitochondria are constantly observed in obese patients without steatosis (Fig. 6.6). Apparently, in the liver, these effects are not related to inflammation but autophagy affects the inflammatory status of the adipocyte. The reported data are conflicting and even counterintuitive. Patients with steatosis displayed higher adipocytes than those without and may constitute a predictive factor, which is associated with circulating C-reactive protein (Fig. 6.6).
The degree of steatosis is considerably heterogeneous among obese patients and this may be associated with the presence of autophagy (mostly mitophagy) (a). In addition, the adipocytes in subcutaneous adipose tissue are significantly higher in patients with steatosis (b). It remains to be ascertained whether this measurement could be qualified as a clinical biomarker but it is apparently associated with circulating C-reactive protein concentration (c)
The absence of autophagy in adipocytes, as identified in adipocyte-specific ATG7 knockout mice, or the mitigation observed in CCL2 deficient mice, protect against high-fat diet-induced obesity and insulin resistance in mice [67–69]. However, other data report that autophagy may function to limit excessive inflammation in adipose tissue during obesity [70]. In addition, obese patients do not necessarily display insulin resistance. The inhibition of autophagic action by insulin could partly explain an expected increase of autophagy in obese patients with insulin resistance, and consequently steatosis should be more likely associated with metabolic disturbances. In these patients, however, changes in autophagy could also play a role in both inflammatory gene expression and secretion of cytokines. This role may not be exactly the case for other types of cells (i.e., macrophages) that are present in expanded adipose tissue. Therefore, it remains plausible that the inhibition of autophagy in adipose tissue might worsen insulin sensitivity via its effects on inflammatory cytokine production, and that inflammation is associated with autophagy as part of the induced effects of obesity. Other findings [71, 72] suggest that obesity-associated inflammation in adipose tissue up-regulates autophagy to mitigate the production of proinflammatory cytokines (i.e., autophagy could be a consequence rather than a cause of obesity-induced adipose tissue inflammation).
6.3.3 Lipophagy and the Lipolytic Role of Macrophages in Adipose Tissue
The function of autophagy in regulating intracellular lipid droplets (LDs) is also relevant for obesity, especially in tissues that are not designed for energy storage, such as the liver or muscle (lipophagy). Whether impaired lipophagy is a sufficient condition to cause NAFLD is debatable [73]. Among other data, the existence of lipophagy has been suggested, via different mechanisms, to degrade LDs through autophagy, indicating that lysosomes do not fuse directly with LDs but with LD-containing autophagosomes [74]. In the context of NAFLD, we have previously shown the synergic roles of both dietary energy ingestion and the tissue CCL2 response [75–77]. LDs were initially viewed as storage lipid depots, but it is now known that these organelles may participate in different cell functions [78]. Moreover, when induced by oxidized lipids, LDs are active in the arachidonic acid conversion into leukotrienes through the 5-LO pathway, a key component in the pathogenesis of atherosclerosis [79–81]. Prostaglandins and leukotrienes are important inflammation mediators; consequently, LDs may act in the coordination of immune responses. Different subpopulations of LDs that may differ in their lipid and protein concentrations have been observed in different type cells within the same cell and/or in response to different stimuli, suggesting maturation and integration in multiple cellular processes [82]. Curiously, emerging evidence suggests a linkage between LDs and the regulation of immune responses in the context of host-pathogen interactions, which have been previously reviewed [83]. The regulatory network of autophagy in the development of NAFLD may also involve signaling through AMP-activated protein kinase (AMPK), which is a central regulator of cellular metabolism [84]. AMPK regulates metabolism through its effects on glucose homeostasis, lipid metabolism, protein synthesis, and oxidative metabolism. However, ongoing studies in a model of hyperlipidemia suggest that the modulation of AMPK by metformin may have different effects according to the baseline metabolic context. AMPK may up-regulate glycolysis and decrease the synthesis of glycogen [85]. The control of lipid metabolism by AMPK is most likely the result of both a decrease in lipogenesis and the stimulation of mitochondrial fatty acid oxidation. The possible mechanisms remain unclear, but the former is most likely due to the inactivation of 3-hydroxy-3-methylglutaryl-coenzyme A reductase and acetyl CoA carboxylase 1. The ability of AMPK to promote fatty acid oxidation may be due to the decreased synthesis of malonyl CoA or a decrease in transcription of lipogenic genes via the inhibition of the sterol regulatory element-binding protein-1c. AMPK most likely regulates PGC-1α, which subsequently increases mitochondrial biogenesis [86]. All of these findings are particularly important because metformin has been considered an agent against fatty liver disease. Moreover, the invoked mechanisms of action are continuously revisited [87–89]. A certain alteration in the mitochondrial function is likely and deserves future research. Metformin may also attenuate inflammatory responses by suppressing the production of TNF-α, suppressing the expressions of scavenger receptors in macrophages [90], and exacerbating the allergic eosinophilic inflammation in high fat-diet-induced obesity in mice [91].
To further complicate the issue, macrophages in adipose tissue (ATMs) play a role in lipolysis that is independent of inflammatory action. In an attempt to identify the cellular functions of ATMs that are regulated by adiposity, a program of obesity-associated lysosome biogenesis has been recently identified, which suggests that ATM lysosomes are important in lipid metabolism [92]. A class of antimalarial drugs, including chloroquine, inhibits the acidification of lysosomes (i.e., decreased autophagy), consequently leading to the accumulation of LDs in macrophages. Macrophages are cells that possess distinct noninflammatory trophic functions that are regulated by the cellular and/or metabolic context. These new findings indicate that ATMs, which are the dominant immune cell in AT, respond to increases in adiposity and are closely coupled to lipid accumulation (i.e., the net effect is to buffer increased concentrations of lipids). Moreover, the trafficking of lipids differs between lean and obese individuals. In large adipocytes, the released lipid is taken up by macrophages and targeted to lysosomes for metabolism. This process may be central to the development of metabolic diseases [93]; however, attempts to target autophagy and lysosome function require further extensive research (Fig. 6.7).
Despite their apparent complexity and the possible variations among metabolic tissues, inflammation, lipolysis and autophagy are closely related in metabolic diseases. Treatment with available drugs may require further elucidation of the uptake of lipids by adipose tissue and cells in the environment, particularly macrophages, the signaling effects of lipid sensing, the fate of lysosomal products of lipid hydrolysis, and the anabolic factors released by autophagy
6.4 Conclusion and Perspectives
The crosstalk between the signaling pathways and autophagy, particularly the link between MTOR downregulation and autophagy induction, requires further exploration considering the possible regulatory effects of inflammation. Caloric restriction apparently delays the onset of chronic diseases, which are more common later in life. Caloric restriction also favors autophagy, which provides energy to cells from self-degraded components and removes harmful compounds that play a role in oxidative stress and immune function. In older organisms, autophagic activity decreases, which may possibly favor the genesis of chronic diseases and partially explain the relationship between these diseases and an excessive energy intake. Although caloric restriction does not appear to be feasible in the human population, the induced decrease in chronic systemic low-grade inflammation may be focal in the development of many typical Western diseases centered on obesity. For example, epidemiological studies indicate that the consumption of polyphenol-rich foods, which may have a caloric restriction mimetic effect, may be associated with a lower chronic disease risk.
Autophagy research may lead to either the discovery of future therapeutic targets or to the efficient use of either marketed or newly developed drugs under safe and beneficial conditions. It should be particularly important to uncover the multiple interactions between autophagy and cytokines, suggesting important roles for autophagy in the control of inflammation and fine-tuning of the immune response. In this setting, atherosclerosis appears to be a relatively novel target. Certain experiments in mice demonstrate defective autophagy in macrophages of atherosclerotic plaques. Complete deficiency of macrophage autophagy resulted in a pro-inflammatory state and enhanced plaque progression via an inflammasome-dependent mechanism mediated by cholesterol crystals [94]. The inflammasomes, recently discovered multi-protein complexes, seem to be crucial to understand how the immune system sense perturbations of cellular homeostasis, and at which stage of metabolic disease the subsequent interactions impact disease progression.
Knowledge on the mechanisms and pathways involved in autophagy might reveal a considerable potential for pharmacological modulation of the process in the management of inflammation and related conditions. Noninflammatory functions of immune cells should also be considered in the context of potential improvements in the therapeutic approach for the devastating metabolic diseases.
References
Finley D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu Rev Biochem. 2009;78:477–513.
Kundu M, Thompson CB. Autophagy: basic principles and relevance to disease. Annu Rev Pathol. 2008;3:427–55.
Kim J, Guan KL. Amino acid signaling in TOR activation. Annu Rev Biochem. 2011;80:1001–32.
Menendez JA, Cufí S, Oliveras-Ferraros C, Vellon L, Joven J, Vazquez-Martin A. Gerosuppressant metformin: less is more. Aging (Albany NY). 2011;3:348–62.
Cufí S, Vazquez-Martin A, Oliveras-Ferraros C, Corominas-Faja B, Cuyàs E, López-Bonet E, et al. The anti-malarial chloroquine overcomes primary resistance and restores sensitivity to trastuzumab in HER2-positive breast cancer. Sci Rep. 2013;3:2469. doi:10.1038/srep02469.
Komatsu M, Waguri S, Ueno T, Iwata J, Murata S, Tanida I, et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J Cell Biol. 2005;169:425–34.
Rodriguez-Enriquez S, Kim I, Currin RT, Lemasters JJ. Tracker dyes to probe mitochondrial autophagy (mitophagy) in rat hepatocytes. Autophagy. 2006;2:39–46.
Li D. Selective degradation of the IkappaB kinase (IKK) by autophagy. Cell Res. 2006;16:855–6.
Mizushima N, Yamamoto A, Matsui M, Yoshimori T, Ohsumi Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell. 2004;15:1101–11.
Yousefi S, Perozzo R, Schmid I, Ziemiecki A, Schaffner T, Scapozza L, et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat Cell Biol. 2006;8:1124–32.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H, Levine B. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature. 1999;402:672–6.
Hay N. The Akt-mTOR tango and its relevance to cancer. Cancer Cell. 2005;8:179–83.
Joven J, Rull A, Rodriguez-Gallego E, Camps J, Riera-Borrull M, Hernández-Aguilera A, et al. Multifunctional targets of dietary polyphenols in disease: a case for the chemokine network and energy metabolism. Food Chem Toxicol. 2013;51:267–79.
Jin S, DiPaola RS, Mathew R, White E. Metabolic catastrophe as a means to cancer cell death. J Cell Sci. 2007;120:379–83.
Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature. 2006;441:885–9.
Taylor JP, Hardy J, Fischbeck KH. Toxic proteins in neurodegenerative disease. Science. 2002;296:1991–5.
Jennings Jr JJ, Zhu JH, Rbaibi Y, Luo X, Chu CT, et al. Mitochondrial aberrations in mucolipidosis type IV. J Biol Chem. 2006;281:39041–50.
Tanaka Y, Guhde G, Suter A, Eskelinen EL, Hartmann D, et al. Accumulation of autophagic vacuoles and cardiomyopathy in LAMP-2-deficient mice. Nature. 2000;406:902–6.
Schmid D, Pypaert M, Munz C. Antigen-loading compartments for major histocompatibility complex class II molecules continuously receive input from autophagosomes. Immunity. 2007;26:79–92.
English L, Chemali M, Duron J, Rondeau C, Laplante A, et al. Autophagy enhances the presentation of endogenous viral antigens on MHC class I molecules during HSV-1 infection. Nat Immunol. 2009;10:480–7.
Berger SB, Romero X, Ma C, Wang G, Faubion WA, et al. SLAM is a microbial sensor that regulates bacterial phagosome functions in macrophages. Nat Immunol. 2010;11:920–7.
Brinkmann V, Reichard U, Goosmann C, Fauler B, Uhlemann Y, et al. Neutrophil extracellular traps kill bacteria. Science. 2004;303:1532–5.
Watanabe K, Tsubata T. Autophagy connects antigen receptor signaling to costimulatory signaling in B lymphocytes. Autophagy. 2009;5:108–10.
Hubbard VM, Valdor R, Patel B, Singh R, Cuervo AM, Macian F. Macroautophagy regulates energy metabolism during effector T cell activation. J Immunol. 2010;185:7349–57.
Hildeman DA, Mitchell T, Teague TK, Henson P, Day BJ, et al. Reactive oxygen species regulate activation-induced T cell apoptosis. Immunity. 1999;10:735–44.
Cardenas C, Miller RA, Smith I, Bui T, Molgo J, et al. Essential regulation of cell bioenergetics by constitutive InsP3 receptor Ca2+ transfer to mitochondria. Cell. 2010;142:270–83.
Nakagawa I, Amano A, Mizushima N, Yamamoto A, Yamaguchi H, Kamimoto T, et al. Autophagy defends cells against invading group A Streptococcus. Science. 2004;306:1037–40.
Inohara N, Chamaillard M, McDonald C, Nunez G. NOD-LRR proteins: role in host-microbial interactions and inflammatory disease. Annu Rev Biochem. 2005;74:355–83.
Delbridge LM, O’Riordan MX. Innate recognition of intracellular bacteria. Curr Opin Immunol. 2007;19:10–6.
Rich KA, Burkett C, Webster P. Cytoplasmic bacteria can be targets for autophagy. Cell Microbiol. 2003;5:455–68.
Deretic V, Singh S, Master S, Harris J, Roberts E, et al. Mycobacterium tuberculosis inhibition of phagolysosome biogenesis and autophagy as a host defence mechanism. Cell Microbiol. 2006;8:719–27.
Amer AO, Swanson MS. Autophagy is an immediate macrophage response to Legionella pneumophila. Cell Microbiol. 2005;7:765–78.
Qin QM, Luo J, Lin X, Pei J, Li L, et al. Functional analysis of host factors that mediate the intracellular lifestyle of Cryptococcus neoformans. PLoS Pathog. 2011;7:e1002078.
Portillo JA, Okenka G, Reed E, Subauste A, Van Grol J, et al. The CD40-autophagy pathwayis needed for host protection despite IFN-γ-dependent immunity and CD40 induces autophagy via control of P21 levels. PLoS One. 2010;5:e14472.
Kyei GB, Dinkins C, Davis AS, Roberts E, Singh SB, et al. Autophagy pathway intersects with HIV-1 biosynthesis and regulates viral yields in macrophages. J Cell Biol. 2009;186:255–68.
Joven J, Menéndez JA, Fernandez-Sender L, Espinel E, Rull A, Beltrán-Debón R, et al. Metformin: a cheap and well-tolerated drug that provides benefits for viral infections. HIV Med. 2013;14:233–40.
Denizot M, Varbanov M, Espert L, Robert-Hebmann V, Sagnier S, et al. HIV-1 gp41 fusogenic function triggers autophagy in uninfected cells. Autophagy. 2008;4:998–1008.
Zhou D, Spector SA. Human immunodeficiency virus type-1 infection inhibits autophagy. AIDS. 2008;22:695–9.
Paton NI, Goodall RL, Dunn DT, Franzen S, Collaco-Moraes Y, Gazzard BG, et al. Effects of hydroxychloroquine on immune activation and disease progression among HIV-infected patients not receiving antiretroviral therapy: a randomized controlled trial. JAMA. 2012;308:353–61.
Alonso-Villaverde C, Menéndez JA, Joven J. Metabolic stress in infected cells may represent a therapeutic target for human immunodeficiency virus infection. Med Hypotheses. 2013;81:125–30.
Deretic V. Autophagy: an emerging immunological paradigm. J Immunol. 2012;189:15–20.
Qu X, Zou Z, Sun Q, Luby-Phelps K, Cheng P, et al. Autophagy gene-dependent clearance of apoptotic cells during embryonic development. Cell. 2007;128:931–46.
Martinez J, Almendinger J, Oberst A, Ness R, Dillon CP, Fitzgerald P, et al. Microtubule-associated protein1 light chain 3 alpha (LC3)-associated phagocytosis is required for the efficient clearance of dead cells. Proc Natl Acad Sci U S A. 2011;108:17396–401.
Dengjel J, Schoor O, Fischer R, Reich M, Kraus M, et al. Autophagy promotes MHC class II presentation of peptides from intracellular source proteins. Proc Natl Acad Sci U S A. 2005;102:7922–7.
Lapaquette P, Glasser AL, Huett A, Xavier RJ, Darfeuille-Michaud A. Crohn’s disease-associated adherent-invasive E.coli are selectively favoured by impaired autophagy to replicate intracellularly. Cell Microbiol. 2010;12:99–113.
Goldbach-Mansky R. Blocking interleukin-1 in rheumatic diseases. Ann N Y Acad Sci. 2009;1182:111–23.
Giuliani F, Grieve A, Rabouille C. Unconventional secretion: a stress on GRASP. Curr Opin Cell Biol. 2011;23:498–504.
Lichtman EI, Helfgott SM, Kriegel MA. Emerging therapies for systemic lupus erythematosus–focus on targeting interferon-alpha. Clin Immunol. 2012;143:210–21.
He Y, Xu Y, Zhang C, Gao X, Dykema KJ, Martin KR, et al. Identification of a lysosomal pathway that modulates glucocorticoid signaling and the inflammatory response. Sci Signal. 2011;4:ra44.
Dorshkind K, Swain S. Age-associated declines in immune system development and function: causes, consequences, and reversal. Curr Opin Immunol. 2009;21:404–7.
Olefsky JM, Glass CK. Macrophages, inflammation, and insulin resistance. Annu Rev Physiol. 2010;72:219–46.
Wen H, Gris D, Lei Y, Jha S, Zhang L, Huang MT, et al. Fatty acid-induced NLRP3-ASC inflammasome activation interferes with insulin signaling. Nat Immunol. 2011;12:408–15.
O’Neill LA, Hardie DG. Metabolism of inflammation limited by AMPK and pseudo-starvation. Nature. 2013;493:346–55.
Rull A, Camps J, Alonso-Villaverde C, Joven J. Insulin resistance, inflammation, and obesity: role of monocyte chemoattractant protein-1 (or CCL2) in the regulation of metabolism. Mediators Inflamm. 2010;2010:pii: 326580. doi: 10.1155/2010/326580.
Ebato C, Uchida T, Arakawa M, Komatsu M, Ueno T, Komiya K, et al. Autophagy is important in islet homeostasis and compensatory increase of beta cell mass in response to high-fat diet. Cell Metab. 2008;8:325–32.
Quan W, Lim YM, Lee MS. Role of autophagy in diabetes and endoplasmic reticulum stress of pancreatic beta cells. Exp Mol Med. 2012;44:81–8.
Rodríguez-Gallego E, Riera-Borrull M, Hernández-Aguilera A, Mariné-Casadó R, Rull A, Beltrán-Debón R, et al. Ubiquitous transgenic overexpression of C-C Chemokine Ligand 2: a model to assess the combined effect of high energy intake and continuous low-grade inflammation. Mediat Inflamm. 2013;2013:953841. doi: 10.1155/2013/953841.
Tous M, Ferré N, Rull A, Marsillach J, Coll B, Alonso-Villaverde C, Camps J, Joven J. Dietary cholesterol and differential monocyte chemoattractant protein-1 gene expression in aorta and liver of apo E-deficient mice. Biochem Biophys Res Commun. 2006;340:1078–84.
Gutierrez MG, Master SS, Singh SB, Taylor GA, Colombo MI, Deretic V. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell. 2004;119:753–66.
Wright K, Ward SG, Kolios G, Westwick J. Activation of phosphatidylinositol 3-kinase by interleukin-13. An inhibitory signal for inducible nitric-oxide synthase expression in epithelial cell line HT-29. J Biol Chem. 1997;272:12626–33.
Jia G, Cheng G, Gangahar DM, Agrawal DK. Insulin-like growth factor-1 and TNF-alpha regulate autophagy through c-jun N-terminal kinase and Akt pathways in human atherosclerotic vascular smooth cells. Immunol Cell Biol. 2006;84:448–54.
Hall JL, Gibbons GH, Chatham JC. IGF-I promotes a shift in metabolic flux in vascular smooth muscle cells. Am J Physiol Endocrinol Metab. 2002;283:E465–71.
Coll B, Alonso-Villaverde C, Joven J. Monocyte chemoattractant protein-1 and atherosclerosis: is there room for an additional biomarker? Clin Chim Acta. 2007;383:21–9.
Must A, Spadano J, Coakley EH, Field AE, Colditz G, Dietz WH. The disease burden associated with overweight and obesity. JAMA. 1999;282:1523–9.
Joven J, Micol V, Segura-Carretero A, Alonso-Villaverde C, Menendez JA. Polyphenols and the modulation of gene expression pathways: can we eat our way out of the danger of chronic disease? Crit Rev Food Sci Nutr. 2014;54(8):985–1001. doi: 10.1080/10408398.2011.621772.
Segura-Carretero A, Puertas-Mejía MA, Cortacero-Ramírez S, Beltrán R, Alonso-Villaverde C, Joven J, Dinelli G, Fernández-Gutiérrez A. Selective extraction, separation, and identification of anthocyanins from Hibiscus sabdariffa L. using solid phase extraction-capillary electrophoresis-mass spectrometry (time-of-flight/ion trap). Electrophoresis. 2008;29:2852–61.
Goldman S, Zhang Y, Jin S. Autophagy and adipogenesis: implications in obesity and type II diabetes. Autophagy. 2010;6:179–81.
Singh R, Xiang Y, Wang Y, Baikati K, Cuervo AM, Luu YK, et al. Autophagy regulatesadipose mass and differentiation in mice. J Clin Invest. 2009;119:3329–39.
Zhang Y, Goldman S, Baerga R, Zhao Y, Komatsu M, Jin S. Adipose-specific deletion of autophagy-related gene 7 (atg7) in mice reveals a role in adipogenesis. Proc Natl Acad Sci U S A. 2009;106:19860–5.
Jansen HJ, van Essen P, Koenen T, Joosten LA, Netea MG, Tack CJ, Stienstra R. Autophagy activity is up-regulated in adipose tissue of obese individuals and modulates proinflammatory cytokine expression. Endocrinology. 2012;153:5866–74.
Yoshizaki T, Kusunoki C, Kondo M, Yasuda M, Kume S, Morino K, et al. Autophagy regulates inflammation in adipocytes. Biochem Biophys Res Commun. 2012;417:352–7.
Marsillach J, Camps J, Ferré N, Beltran R, Rull A, Mackness B, Mackness M, Joven J. Paraoxonase-1 is related to inflammation, fibrosis and PPAR delta in experimental liver disease. BMC Gastroenterol. 2009;9:3. doi:10.1186/1471-230X-9-3.
Liu K, Czaja MJ. Regulation of lipid stores and metabolism by lipophagy. Cell Death Differ. 2013;20:3–11.
Singh R, Kaushik S, Wang Y, Xiang Y, Novak I, Komatsu M, et al. Autophagy regulates lipid metabolism. Nature. 2009;458:1131–5.
Vinaixa M, Rodríguez MA, Rull A, Beltrán R, Bladé C, Brezmes J, et al. Metabolomic assessment of the effect of dietary cholesterol in the progressive development of fatty liver disease. J Proteome Res. 2010;9:2527–38.
Rull A, Rodríguez F, Aragonès G, Marsillach J, Beltrán R, Alonso-Villaverde C, Camps J, Joven J. Hepatic monocyte chemoattractant protein-1 is upregulated by dietary cholesterol and contributes to liver steatosis. Cytokine. 2009;48:273–9.
Joven J, Espinel E, Rull A, Aragonès G, Rodríguez-Gallego E, Camps J, et al. Plant-derived polyphenols regulate expression of miRNA paralogs miR-103/107 and miR-122 and prevent diet-induced fatty liver disease in hyperlipidemic mice. Biochim Biophys Acta. 1820;2012:894–9.
Bozza PT, Magalhães KG, Weller PF. Leukocyte lipid bodies – biogenesis and functions in inflammation. Biochim Biophys Acta. 2009;1791:540–51.
Silva AR, Pacheco P, Vieira-de-Abreu A, Maya-Monteiro CM, D’Alegria B, Magalhães KG, et al. Lipid bodies in oxidized LDL-induced foam cells are leukotriene-synthesizing organelles: a MCP-1/CCL2 regulated phenomenon. Biochim Biophys Acta. 2009;1791:1066–75.
Spanbroek R, Grabner R, Lotzer K, Hildner M, Urbach A, Ruhling K, et al. Expanding expression of the 5-lipoxygenase pathway within the arterial wall during human atherogenesis. Proc Natl Acad Sci U S A. 2003;100:1238–43.
Ferré N, Martínez-Clemente M, López-Parra M, González-Périz A, Horrillo R, Planagumà A, et al. Increased susceptibility to exacerbated liver injury in hypercholesterolemic ApoE-deficient mice: potential involvement of oxysterols. Am J Physiol Gastrointest Liver Physiol. 2009;296:G553–62.
Hodges BD, Wu CC. Proteomic insights into an expanded cellular role for cytoplasmic lipid droplets. J Lipid Res. 2010;51:262–73.
Saka HA, Valdivia R. Emerging roles for lipid droplets in immunity and host-pathogen interactions. Annu Rev Cell Dev Biol. 2012;28:411–37.
Hardie DG. AMP-activated protein kinase: an energy sensor that regulates all aspects of cell function. Genes Dev. 2011;25:1895–908.
Jørgensen SB, Nielsen JN, Birk JB, Olsen GS, Viollet B, Andreelli F, et al. The alpha2-5′AMP-activated protein kinase is a site 2 glycogen synthase kinase in skeletal muscle and is responsive to glucose loading. Diabetes. 2004;53:3074–81.
Narkar VA, Downes M, Yu RT, Embler E, Wang YX, Banayo E, et al. AMPK and PPARdelta agonists are exercise mimetics. Cell. 2008;134:405–15.
Zhou G, Myers R, Li Y, Chen Y, Shen X, Fenyk-Melody J, et al. Role of AMP-activated protein kinase in mechanism of metformin action. J Clin Invest. 2001;108:1167–74.
Zang M, Zuccollo A, Hou X, Nagata D, Walsh K, Herscovitz H, et al. AMP-activated protein kinase is required for the lipid-lowering effect of metformin in insulin-resistant human HepG2 cells. J Biol Chem. 2004;279:47898–905.
Menendez JA, Joven J. One-carbon metabolism: an aging-cancer crossroad for the gerosuppressant metformin. Aging (Albany NY). 2012;4:894–8.
Hyun B, Shin S, Lee A, Lee S, Song Y, Ha NJ, Cho KH, Kim K. Metformin down-regulates TNF-α secretion via suppression of scavenger receptors in macrophages. Immune Netw. 2013;13:123–32.
Calixto MC, Lintomen L, André DM, Leiria LO, Ferreira D, Lellis-Santos C, et al. Metformin attenuates the exacerbation of the allergic eosinophilic inflammation in high fat-diet-induced obesity in mice. PLoS One. 2013;8:e76786.
Xu X, Grijalva A, Skowronski A, van Eijk M, Serlie MJ, Ferrante Jr AW. Obesity activates a program of lysosomal-dependent lipid metabolism in adipose tissue macrophages independently of classic activation. Cell Metab. 2013;18:816–30.
Camps J, Rodríguez-Gallego E, García-Heredia A, Triguero I, Riera-Borrull M, Hernández-Aguilera A, Luciano-Mateo F, Fernández-Arroyo S, Joven J. Paraoxonases and chemokine (C–C Motif) ligand-2 in noncommunicable diseases. Adv Clin Chem. 2014;63:247–308.
Razani B, Feng C, Coleman T, Emanuel R, Wen H, Hwang S, et al. Autophagy links inflammasomes to atherosclerotic progression. Cell Metab. 2012;15:534–44.
Acknowledgements
We would like to acknowledge the contribution of the staff members that assisted in the clinical management, laboratory measurements, statistical assessment, and data collection during the last years. The Unitat de Recerca Biomèdica is currently being supported by the program of consolidated groups from the Universitat Rovira iVirgili and grants from the Fondo de Investigación Sanitaria (FIS PI08/1032, PI08/1381 and PI11/00130). E. Rodríguez-Gallego is the recipient of a fellowship from the Generalitat de Catalunya (2012FI B 00389).
Conflict of Interest
The authors declare no competing interests.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Joven, J., Guirro, M., Mariné-Casadó, R., Rodríguez-Gallego, E., Menéndez, J.A. (2014). Autophagy Is an Inflammation-Related Defensive Mechanism Against Disease. In: Camps, J. (eds) Oxidative Stress and Inflammation in Non-communicable Diseases - Molecular Mechanisms and Perspectives in Therapeutics. Advances in Experimental Medicine and Biology, vol 824. Springer, Cham. https://doi.org/10.1007/978-3-319-07320-0_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-07320-0_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-07319-4
Online ISBN: 978-3-319-07320-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)