Abstract
During mRNA synthesis and maturation, the introduction of errors can strongly influence the expression of certain genes and/or the activity of the proteins for which they encode. To minimise these defects, eukaryotic cells have evolved several cytoplasmic and translation-dependent quality control pathways aimed at detecting and degrading mRNAs that would lead to the production of aberrant proteins. The nonsense-mediated mRNA decay pathway (NMD) clears cells from mRNAs harbouring premature in-frame stop codons. Two other pathways (NSD for nonstop decay and NGD for No-Go decay) degrade mRNAs on which ribosomes have stalled during elongation. In this chapter, we describe the current knowledge on the biological roles and molecular mechanisms of these surveillance pathways, which were mainly unravelled using baker’s yeast as model system.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Premature Termination Codon
- Nascent Peptide
- Apurinic Site
- Stall Ribosome
- Translation Termination Factor eRF1
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
In eukaryotes, the production of functional translatable mRNAs requires several maturation steps (splicing, capping, polyadenylation, export, …), all of which offer the possibility for introducing errors. The translation of such faulty mRNAs would produce aberrant proteins, which could have dramatic effects and lead to diseases or even cell death. However, these aberrant mRNAs are rarely translated as eukaryotic cells have evolved numerous surveillance (or QC for quality control) pathways dedicated to the detection and the rapid degradation of these mRNAs and to the concomitant clearance of nascent proteins derived from these mRNAs. The most extensively described process is the nonsense-mediated mRNA decay pathway (known as NMD), which clears cells from mRNAs containing in-frame premature termination codons (PTC) (Kervestin and Jacobson 2012; Losson and Lacroute 1979). Other QC pathways specialised in the rapid degradation of mRNAs responsible for translation elongation stalls have been described more recently. The nonstop (or NSD) and No-Go (or NGD) mRNA decay pathways degrade mRNAs lacking stop codons (Frischmeyer et al. 2002; van Hoof et al. 2002) or mRNAs inducing strong translational stalls (Doma and Parker 2006), respectively. These evolutionarily conserved mechanisms have been discovered and deeply characterised using Saccharomyces cerevisiae yeast. This chapter presents an overview of these cytoplasmic and translation-dependent mRNA decay pathways as a wealth of information obtained within the last years offers a more detailed understanding of these QC mechanisms.
The Nonsense-Mediated mRNA Decay Pathway
NMD Substrates
NMD rids cells from mRNAs harbouring a PTC and thereby prevents the accumulation of potentially harmful truncated proteins. PTC resulting in NMD activation can occur in mRNAs due to genetic mutations, transcription and/or mRNA maturation errors, especially splicing defaults (Kervestin and Jacobson 2012; Mitrovich and Anderson 2000). NMD substrates also include bicistronic mRNAs, pseudogene-derived transcripts, mRNAs subjected to leaky scanning leading to translation initiation errors or to frameshifting, or mRNAs with upstream reading frames (uORFs) (He et al. 2003; Ruiz-Echevarria and Peltz 2000; Welch and Jacobson 1999). In mammals, they also arise from alternative splicing of mRNAs (Hansen et al. 2009) or are produced by genes undergoing programmed rearrangement such as those encoding antibodies, B and T cell receptors (Li and Wilkinson 1998). NMD is also activated by a stop codon followed by normal or biologically regulated long 3′ UTRs, which mimic a premature termination context (Kebaara and Atkin 2009; Muhlrad and Parker 1999).
Beyond mRNA QC, NMD also modulates the cellular level of up to 10 % of normal genes in S. cerevisiae, D. melanogaster and humans and hence directly regulates the expression of many physiological transcripts (He et al. 2003; Mendell et al. 2004; Rehwinkel et al. 2005; Wittmann et al. 2006).
NMD Factors
NMD is activated when the stop codon present in the ribosomal A-site is recognised as premature. This pathway relies on the NMD specific factors and in particular, the three conserved Upf proteins: Upf1, Upf2, and Upf3 initially identified in S. cerevisiae and C. elegans (Smg2, 3 and 4, respectively) (Cui et al. 1995; Leeds et al. 1992; Pulak and Anderson 1993). Mutations of UPF genes lead to a specific stabilisation of PTC-containing transcripts (He et al. 1997).
The Upf proteins interact together at the premature stop codon and form the surveillance UPF complex, where Upf1 is assumed to be the key effector of NMD while Upf2 and Upf3 act as essential regulators of its function.
Upf1
Upf1 is a large cytoplasmic protein composed of two functional domains: an N-terminal Cysteine- and Histidine-rich zinc-finger domain (CH domain) and a larger C-terminal helicase domain from the SF1 family (de la Cruz et al. 1999). The CH domain has a RING-box architecture and exhibits U3 ubiquitin-ligase activity that may be involved in the elimination of the aberrant peptide by the proteasome (Kadlec et al. 2006; Takahashi et al. 2008). This domain is also involved in Upf1 interaction with Rps26 from the ribosomal 40S subunit (Min et al. 2013). The helicase domain consists of two canonical RecA-like subdomains with two additional inserted subdomains (called 1B and 1C) and exhibits ATPase and RNA unwinding activities (Fig. 8.1). Both activities are essential for NMD and are downregulated by the CH domain (Bhattacharya et al. 2000; Chamieh et al. 2008; Czaplinski et al. 1995; Weng et al. 1996).
The architecture of the Upf complex. a The modular organisation of Upf1 (top), Upf2 (middle) and Upf3 (bottom) proteins. The interacting domains of Upf1, Upf2 and Upf3 are connected by black lines. b The X-ray structure of yeast Upf1 in complex with a poly(U)9 RNA (orange and an ATP analog (ADP-AlF4 −, brown). The CH domain is coloured in cyan, the RecA1 and RecA2 domains are coloured in dark blue and the 1B and the 1C insertions are coloured in light blue
Upf3
Upf3 is a small protein with a conserved central RNA recognition motif (RRM), which is unable to bind RNA in vitro (Kadlec et al. 2004). Upf3 C-terminal domain harbours a functional Nuclear localisation signal (NLS) allowing its shuttling between the nucleus and the cytoplasm (Fig. 8.1) (Lee and Culbertson 1995; Shirley et al. 1998).
Upf2
Upf2 is the largest protein of the surveillance complex. It harbours three conserved mIF4G-like domains followed by an acidic linker and a small C-terminal domain (Fig. 8.1) (Chakrabarti et al. 2011; Clerici et al. 2009; He et al. 1997; Kadlec et al. 2006). Upf2 is generally considered as the scaffold protein within the UPF complex as it bridges Upf1 to Upf3. Indeed, Upf2 interacts with the Upf3 RRM domain through its third mIF4G domain and with the Upf1 CH domain via its C-terminal domain.
However, beyond its scaffolding role, Upf2 also enhances Upf1 enzymatic activities (Chamieh et al. 2008). Indeed, Upf2 binding to the Upf1 CH domain displaces it by 120° relative to the Upf1 helicase domain, thus releasing its cis-inhibitory effect on helicase and ATPase activities (Chakrabarti et al. 2011; Clerici et al. 2009).
The role of the two first mIF4G domains remains unclear, although their deletion abolishes NMD in yeast without affecting the formation of the UPF complex (He et al. 1997). The first mIF4G domain was proposed to harbour a putative conserved “NLS” whose deletion provokes severe NMD defects in yeast. However, yeast Upf2 is cytoplasmic and the NMD defects observed upon “NLS” depletion are unlikely to be caused by Upf2 mislocalisation (He and Jacobson 1995). This first mIF4G domain is phosphorylated in yeast but the precise role of this post-translational modification in NMD remains unclear (Wang et al. 2006).
Other Yeast NMD Factors
Beyond the central UPF complex, other yeast proteins have been suggested to play secondary roles in NMD but their precise role is still controversial.
Hrp1 is an essential nucleocytoplasmic protein that stabilises the mRNA 3′ poly(A) tail thus contributing to the polyadenylation process (Kessler et al. 1997). Hrp1 also interacts with Upf1 and promotes NMD activation by recognising specific sequences located downstream of the PTC (called DSE for Downstream Sequence Element) (Gonzalez et al. 2000). Hrp1 was thus suggested to be a “marker” protein displaced by the translating ribosome when the stop codon is ‘normal’, but remaining tethered to PTC-containing mRNA, thereby triggering NMD. However, these DSE were found in several NMD mRNA reporters (PGK1, HIS4, ADE3 and GCN4) but share a weak sequence consensus, while other NMD substrates are free of DSE (Hagan et al. 1995; Ruiz-Echevarria et al. 1998; Zhang et al. 1995). Hence, this model could not be adopted as a generalised NMD activation process.
The deletion of the EBS1 gene provokes a slight but consistent stabilisation of several NMD substrates (Luke et al. 2007). The Ebs1 protein was proposed to harbour an N-terminal 14-3-3 domain and to be a putative orthologue of human SMG7, which is involved in UPF1 dephosphorylation (Luke et al. 2007; Ohnishi et al. 2003). Although Upf1 is phosphorylated in yeast (Lasalde et al. 2013; Wang et al. 2006), no clear evidence implicates Ebs1 in the sensing of Upf1 phosphorylation status. Ebs1 may rather influence NMD by inhibiting translation (Ford et al. 2006).
The DEAD-box RNA helicase Dbp2 associates with Upf1 and is involved both in NMD and rRNA processing in yeast (Bond et al. 2001; He and Jacobson 1995). Its human orthologue, p68 (Ddx5), associates with Upf3b and activates the NMD-mediated regulation of several specific genes, including its own gene (Bond et al. 2001; He and Jacobson 1995). Hence, Dbp2 is probably the most interesting factor whose role in NMD should be clarified.
NMD Mechanism
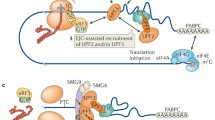
NMD is classically considered as a three-step mechanism, where the first step consists in the recruitment of the canonical translation termination machinery (eRF1 and eRF3) upon entry of a stop codon in the ribosomal A-site. The second step is the discrimination between mRNAs harbouring PTC versus those with normal stops. The final step consists in the rapid decay of the faulty mRNA by the cytoplasmic degradation machinery (Fig. 8.2). Accordingly, both the first and third steps involve the cellular effectors of the canonical translation termination (see Chap. 5 for details) and mRNA degradation (see Chap. 7 for details) pathways, respectively. Only the second step is carried out by the NMD specific Upf factors.
A unified model for NMD pathway. If the recognition of a stop codon in the ribosomal A-site is accompanied by the enrichment of Upf1 on an abnormally long 3′ UTR, the Upf2 and Upf3 proteins will be recruited to form the surveillance complex, thereby signalling for the presence of a premature stop codon. This complex will enhance the degradation of the faulty mRNA as well as the proteasomal decay of the truncated peptide
In this section, we will mainly focus on the last two steps as the recognition of stop codons has already been described in Chap. 5.
PTC Recognition and NMD Activation
Following the entry of a stop codon in the ribosomal A-site, several elements will sense the termination context and if it is detected as aberrant, will trigger a cascade of events ending in the degradation of the faulty mRNA.
It is now assumed that most of the NMD events in yeast can be explained by the ‘faux 3′ UTR model’, where the abnormally long sequence downstream the PTC constitutes an NMD activating signal (Amrani et al. 2004; Kervestin and Jacobson 2012). It was initially speculated that the Pab1 failure to interact with eRF3, due to the remoteness of the 3′ poly(A) tail, was the NMD triggering signal. Thus, eRF3 would bind Upf1, implying that Pab1 and Upf1 compete for eRF3 binding, thus handling the balance between normal and premature termination events (Kervestin et al. 2012). Strong support for this hypothesis came from experiments showing that artificial tethering of yeast Pab1 in the vicinity of the PTC cause stabilisation of the corresponding mRNA (Amrani et al. 2004). In addition, artificial shortening of the ‘faux 3′ UTR’ by deleting the region downstream of a PTC stabilises the faulty mRNA (Hagan et al. 1995; Peltz et al. 1993). Some aspects of this model have been validated, but others have not. For instance, the requirement of the Pab1-eRF3 interaction to antagonise NMD has been discarded, as the absence of Pab1 or the deletion of the Pab1-interacting region from eRF3 do not convert a normal mRNA into an NMD substrate (Kervestin et al. 2012; Meaux et al. 2008). Rather, the key requirement for NMD activation by a ‘faux 3′ UTR’ context would be a proper interaction between eRF3 and Upf1. The lower efficiency of a premature termination event could indeed cause inefficient release of eRF3 thus prompting Upf1 recruitment to the PTC (Amrani et al. 2004; Kervestin and Jacobson 2012). Recent studies in human cells showed that Upf1 binds specifically to the 3′ UTR region in a length-dependent manner (Hogg and Goff 2010; Hwang et al. 2010; Kurosaki and Maquat 2013; Shigeoka et al. 2012; Zund et al. 2013). Accordingly, Upf1 is believed to sense the 3′ UTR length and to associate with long 3′ UTR-containing mRNAs, thus targeting them to NMD. This is further supported by the observation that yeast Upf1 associates preferentially with NMD substrates rather than normal mRNAs (Johansson et al. 2007). Upf1 could then recruit Upf2 and Upf3 to PTC-containing transcripts and then signal these as aberrant mRNAs to be degraded (Fig. 8.2).
This model reconciles several discrepancies between yeast and higher eukaryotes and corresponds to the most elaborated manner to interpret the differences between a normal and a ‘premature’ termination event. However, some twilight zones still exist and require further studies to be properly elucidated.
Faulty mRNA Degradation
The predominant mRNA decay pathway involved in yeast NMD is the 5′-3′ decay pathway (Hagan et al. 1995). Compared to normal mRNA decay, the deadenylation step is skipped in NMD and the PTC-containing transcripts undergo rapid decapping followed by subsequent exonucleolytic degradation by Xrn1 (Fig. 8.2) (Muhlrad and Parker 1994). Upf1 was proposed to recruit the decapping enzyme Dcp2 through the decapping activators Pat1 and Edc3 (He and Jacobson 2001; Swisher and Parker 2011). However, in the absence of the 5′-3′ degradation pathway, the faulty mRNA can undergo a slower 3′-5′ degradation involving Ski7 and the exosome (Mitchell and Tollervey 2003).
Proteasomal Decay of the Truncated Peptide
Beyond faulty mRNA decay, NMD also activates the rapid degradation of the truncated polypeptide by the ubiquitin-proteasome pathway (Kuroha et al. 2013; Kuroha et al. 2009). The truncated nascent peptide will be released from the ribosome through the action of the eRF1-eRF3 translation termination factors. Upf1 seems to enhance the degradation of this truncated peptide by acting as an E3 ubiquitin ligase. Indeed, its N-terminal CH domain is structurally homologous to E3 RING finger domains and associates to the E2 enzyme Ubc3 (Takahashi et al. 2008).
NMD Factors in Higher Eukaryotes
In metazoa, NMD is a more sophisticated process and hence relies on the involvement of additional factors that are absent in yeast. The description of these factors is beyond the scope of this book, which focuses on fungi, but one can briefly mention some of these. Indeed, several SMG proteins (SMG-1 and SMG-5 to SMG-9) are involved in the regulation of UPF1 phosphorylation status. In addition, SMG-6 endonucleolytically cleaves NMD substrates in human and D. melanogaster (Eberle et al. 2009; Gatfield and Izaurralde 2004). Finally, the exon junction complex (EJC) is involved in the degradation of a subset of human NMD substrates (Buhler et al. 2006; Sauliere et al. 2010).
NMD Importance in Human: Involvement in Genetic Diseases and in Some Cancers
Although this chapter has almost exclusively focused on yeast NMD, this QC pathway is conserved in eukaryotes and has biological implications in human health. Indeed, it is estimated that PTC-containing mRNAs are responsible for about one third of inherited genetic disorders such as Duchenne muscular dystrophy or some forms of cystic fibrosis as well as many forms of cancer. In some instances, the truncated proteins produced by these mRNAs may be very harmful or have a dominant negative effect. In some other cases, such truncated proteins may be partially active and could, when properly expressed, decrease the disease severity. Hence, NMD is not always beneficial and could rather be seen as a double-edged sword preventing cells from producing truncated proteins that could do damage but also eliminating mRNAs encoding truncated proteins that could function normally.
It has been reported that the NMD is specifically repressed in some cancers and that this repression provokes the anarchic proliferation of the tumour cells (Gardner 2010; Wang et al. 2011a; Wang et al. 2011b). Conversely, growing evidence shows that inhibiting NMD in the tumour could play a preventive role against cancer. Indeed, NMD inhibition by siRNA (short interfering RNAs)-mediated silencing of SMG-11 or UPF1 has proved to be efficient for tumour regression by inducing an immune response against new antigens expressed in the tumour (Gilboa 2013; Pastor et al. 2010). In the case of the genetic disorders caused by PTC, the healing strategy is rather based on inducing selective PTC read-through (for a recent review, see Bidou et al. 2012). Aminoglycosides, especially gentamicin, were first used as PTC read-through inducers to restore CFTR expression in several cases of cystic fibrosis (Bedwell et al. 1997; Wilschanski et al. 2003). However, due to their toxicity and their random efficiency, aminoglycosides are now replaced by a new molecule called PTC124, which proved to be less toxic and more efficient in inducing specific PTC read-through (Welch et al. 2007).
Quality Control Pathways Dealing with Translation Elongation Arrests
In-frame stop codons are not only crucial for correct translation termination but also for proper recycling of ribosomes and subsequent rounds of translation (see Chap. 5). Hence, mRNAs lacking stop codons or inducing strong translational stalls would trap translating ribosomes and the accumulation of these mRNAs would deplete cells from functional ribosomes. To avoid this, cells have evolved two other QC pathways (Graille and Seraphin 2012). The nonstop decay (or NSD) detects and degrades mRNAs lacking stop codons. The No-Go decay (NGD) pathway clears cells from mRNAs causing ribosomal stalls during elongation. Both pathways also trigger the degradation of the polypeptide derived from these aberrant mRNAs. These pathways have been initially identified in yeast using artificial reporters but natural substrates were later identified, rationalising their biological importance. Finally, the molecular mechanisms of these pathways have been largely deciphered very recently using yeast as a model system.
Nonstop mRNA Decay or NSD
Poly(A)+ NSD Substrates
The absence of in-frame stop codon within mRNAs (hereafter named nonstop mRNAs) can arise from single-point mutants converting a stop codon into a sense codon. Besides, under some circumstances that decrease stop codon recognition efficiency, ribosomes can also perform stop codon read-through and synthesise longer proteins. These two classes of NSD substrates are poly(A)+ mRNAs and it is commonly considered that ribosomes translating these mRNAs will be stalled on the 3′ poly(A) tail and produce a polylysine extension at the C-terminal extremity of these extended proteins that will remain covalently bound to the P-site tRNA. However, such substrates are rather rare as 3′ UTRs are generally rich in in-frame stop codons.
The poly(A)+ NSD mRNAs are strongly destabilised both in yeast and mammals (Frischmeyer et al. 2002; van Hoof et al. 2002) through the exo- and endonucleolytic activities of the Rrp44/Dis3 exosome catalytic subunit as well as the SKI complex and their associated factor Ski7 (a yeast-specific protein and a member of the eEF1A translational GTPase family; Schaeffer and van Hoof 2011; van Hoof et al. 2002). In human cells, the role played by Ski7 in yeast NSD is performed by Hbs1, another member of eEF1A translational GTPase family (Saito et al. 2013). Concomitant to the accelerated mRNA decay, the corresponding nonstop proteins are not detected in the cells suggesting a translational repression mechanism, a higher instability or both (Dimitrova et al. 2009; Inada and Aiba 2005; Ito-Harashima et al. 2007). The levels of nonstop proteins but not nonstop mRNAs are strongly decreased by proteins linked to the proteasome such as the Ltn1 RING-domain-type E3 ubiquitin ligase (Bengtson and Joazeiro 2010; Wilson et al. 2007). These proteins ubiquitinylate nascent nonstop proteins, further triggering their degradation by the proteasome (Bengtson and Joazeiro 2010). Altogether, the expression of nonstop poly(A)+ mRNAs exhibits three levels of regulation: mRNA stability, translational repression and nonstop protein stability.
Poly(A)-less NSD Substrates
Another type of nonstop mRNAs can arise in vivo following endonucleolytic cleavage of mRNAs (i.e. in the case of NGD, see section "No-Go decay or NGD"). These are poly(A)-less mRNAs and lead to ribosomes stalled at the 3′ end of these mRNAs, unable to recycle, thereby producing a shorter polypeptide chain remaining attached to the P-site tRNA. These mRNAs are also highly unstable but several discrepancies exist when compared to the decay pathway described for poly(A)+ nonstop mRNAs. In particular, while the latter requires both the N and C-terminal domains of Ski7 to be degraded, the decay of poly(A)-less nonstop mRNAs does not require the Ski7 C-terminal domain (Meaux and Van Hoof 2006). In addition, proteins derived from these nonstop poly(A)-less mRNAs are produced at a low level. This could be caused by a reduced translation of the nonstop mRNA and/or to a decreased protein stability due to defects in peptide release from the ribosome because of the lack of stop codon. Again, the Ltn1 protein, together with Cdc48 and the RQC complex, address these nonstop proteins to the proteasome for degradation (Brandman et al. 2012; Defenouillere et al. 2013). It was also shown that the protein production from poly(A)-less nonstop mRNA is dependent on the Dom34 and Hbs1 proteins (Kobayashi et al. 2010; see below for details on these two proteins).
No-Go Decay or NGD
A third class of aberrant mRNAs induce translational stalls due to the presence of a stable stem loop, pseudoknot, rare codons, stretch of consecutive identical residues (either K12 or R12) or apurinic sites (Dimitrova et al. 2009; Doma and Parker 2006; Gandhi et al. 2008; Kuroha et al. 2010). These are degraded by the NGD pathway, which endonucleolytically cleaves these mRNAs close to the stalling site prior to their degradation by classical exonucleases (Xrn1 and the exosome). The Dom34 and Hbs1 proteins, which share significant similarity to translation termination factors eRF1 and eRF3, respectively, are important but not essential for the endonucleolytic cleavage observed in NGD (Doma and Parker 2006; Kuroha et al. 2010). As a result, the levels of nascent proteins produced by these mRNAs are lower than expected (Dimitrova et al. 2009; Kuroha et al. 2010). It has recently been proposed that the S. cerevisiae Asc1 protein (or RACK1), a core component of the small ribosomal 40S subunit, which binds to the exit of the mRNA channel, might stimulate translational arrest, thereby leading to nascent protein degradation by the Not4 and Ltn1 E3 ubiquitin ligase proteins (Bengtson and Joazeiro 2010; Dimitrova et al. 2009; Kuroha et al. 2010; Panasenko et al. 2006).
Dom34 and Hbs1, Central Factors of these mRNA QC Pathways
During the last years, several studies have unravelled the mechanisms of NSD and NGD. Dom34 and Hbs1 appear to play a central role in these processes.
Dom34 displays strong structural similarity with class I translation termination factor eRF1 (Graille et al. 2008; Lee et al. 2007). Indeed, Dom34 is composed of three distinct domains: N-terminal, central and C-terminal domains. These domains are spatially arranged so as to mimic a tRNA with the N-terminal and central domains corresponding to the anticodon loop and amino acyl acceptor arm of the tRNAs, as observed for eRF1. Despite this structural similarity, Dom34 and eRF1 proteins display some important differences. First, the universally conserved GGQ motif from the eRF1 central domain, which enters into the ribosomal peptidyltransferase centre to catalyse the hydrolysis of the peptidyl-tRNA bond, is absent in the Dom34 central domain, indicating that Dom34 should not induce release of the nascent peptide. Second, the Dom34 N-terminal domain is structurally radically different from eRF1 N-terminal domain. In Dom34, this domain adopts an Sm/Lsm like fold, suggesting a role in RNA binding by analogy with other Sm/Lsm domains (Wilusz and Wilusz 2005).
Hbs1 belongs to the translational GTPases family encompassing bacterial and eukaryotic elongation factors EF-Tu and eEF1A as well as the eukaryotic class II release factor eRF3 and the yeast-specific Ski7 protein (Atkinson et al. 2008). Hbs1 is mainly composed of a GTPase domain followed by two β-barrels domains (II and III) (van den Elzen et al. 2010). GTP binding to Hbs1 is required for its biological function and for its roles in mRNA QC pathways (Carr-Schmid et al. 2002; Kobayashi et al. 2010; van den Elzen et al. 2010).
Hbs1 and Dom34 proteins interact together to form a stable complex, which structurally mimics both eRF1-eRF3 and EF-Tu-tRNA complexes (Graille et al. 2008; Kobayashi et al. 2010; Kobayashi et al. 2012; Nissen et al. 1995). The similarity with the EF-Tu–tRNA complex is further reinforced by the binding mode of Dom34–Hbs1 to the ribosomal A-site of stalled ribosomes (Becker et al. 2011).
Mechanism of NSD and NGD QC Pathways
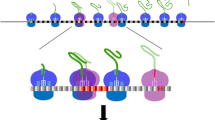
Based on all the information gathered in vivo and in vitro within recent years, there is growing evidence indicating that NGD and NSD pathways function in a very similar manner. It is now possible to propose the following mechanism for these QC pathways that degrade mRNAs impeding translation elongation by the ribosome (Fig. 8.3).
(1) The Dom34-Hbs1 complex in its GTP form is recruited to the A-site of ribosomes stalled in translation. Contrary to the eRF1-eRF3 complex that specifically recognises a stop codon in the A-site, Dom34-Hbs1 binding to the ribosome is independent of the codon present in the ribosomal A-site (Shoemaker et al. 2010). (2) The mRNA associated with the stalled ribosome is endonucleolytically cleaved mainly upstream of the stalled ribosome (Tsuboi et al. 2012). It is noteworthy that following this cleavage, when the ribosomes located upstream of the cleavage site will reach the 3′ end of the truncated mRNA, no stop codon will be present and hence, at this point NGD and NSD meet together. (3) After cleavage, the ribosome stimulates Hbs1 GTPase activity, which could be accompanied by a large conformational change of the intrinsically flexible Dom34 central domain. Dom34 will then adopt a conformation similar to the tRNA “A/A” state observed for EF-Tu-tRNA bound to the bacterial ribosome (Becker et al. 2012; Schmeing et al. 2009), with its central domain oriented towards the peptidyltransferase centre. GTP hydrolysis could also induce a rearrangement of the Hbs1 GTPase domain relative to domains II and III to adopt a conformation similar to that of the S. pombe Dom34-Hbs1 complex and hence lead to Hbs1 dissociation from the ribosome (Chen et al. 2010). (4) The highly conserved and essential Rli1 protein (a member of the ABC family known as ABCE1 in human) is recruited to the ribosome and binds to the same sites as Hbs1 both on the ribosome and on Dom34 (Becker et al. 2012). (5) ATP hydrolysis by Rli1 will result in ribosome dissociation (Pisarev et al. 2010) followed by mRNA degradation by the Xrn1 exonuclease and the exosome (Doma and Parker 2006). (6) The peptidyl-tRNA bound to the P-site should be released from the ribosome. In the case of ribosomes stalled after a few rounds of elongation, the nascent peptide attached to the P-site tRNA should be short and the peptidyl-tRNA could drop-off easily from the ribosome. However, in the case of ribosomes stopped after several rounds of elongation, the nascent peptide will be longer, already deeply engaged into the ribosomal exit tunnel. Despite the structural similarity between Dom34 and eRF1, Dom34 does not catalyse the release of the newly synthesised protein. It has recently been shown that the RQC complex (formed by the Ltn1, Tae2 and Rqc1 proteins) together with the Cdc48 AAA+ ATPase and its cofactors (Npl4 and Ufd1) and Not4 (in some cases) address the nascent proteins derived from NGD and NSD substrates to the proteasome for degradation (Bengtson and Joazeiro 2010; Brandman et al. 2012; Defenouillere et al. 2013; Dimitrova et al. 2009). (7) The 5′ and 3′ fragments from the defective mRNA should now be eliminated. The 5′–>3′ exonuclease Xrn1 will degrade the 3′ fragment, which does not contain a cap structure at its 5′ extremity. The 5′ fragment still contains ribosomes engaged in translation. Since the stalling site has been removed, one can imagine that translation by these ribosomes should be resumed until they reach the 3′ end of this mRNA fragment, which does not contain stop codon. This 5′ fragment then becomes an NSD substrate and ribosomes stalled at the 3′ end of this fragment should be removed by reiteration of steps 1 to 6. If there is no ribosome left on this mRNA fragment, this one can then be degraded through the action of the exosome.
Biological Implications of NSD and NGD QC Pathways
Although mainly studied in budding yeast, the NSD and NGD pathways are evolutionarily conserved, supporting that they play important biological functions. Indeed, these two QC pathways have been described for D. melanogaster and human cells, where they also involve the Dom34 and Hbs1 proteins (Frischmeyer et al. 2002; Passos et al. 2009; Saito et al. 2013).
The NSD pathway relies on the absence of in-frame stop codons. Mutations of the stop codon into a sense codon thereby resulting in the absence of in-frame stop codons have been documented to be responsible for two human diseases: 2, 8-dihydroxyadenine urolithiasis and hypogonadotrophic hypogonadism (Seminara et al. 2003; Taniguchi et al. 1998). In both cases, the levels of nonstop mRNA and the resulting protein are significantly reduced, suggesting that these nonstop mRNAs are cleared from cells by the NSD pathway. However, mutations in normal termination codons or stop codon read-through would not routinely initiate NSD due to the frequent occurrence of in-frame stop codons in the 3′ UTR but would rather result in C-terminally extended proteins. Hence, the evolutionary pressure that has resulted in maintenance of NSD eukaryotes should result from the presence of a non-negligible number of endogenous mRNA NSD substrates. In particular, eukaryotic genes can contain consensus sequences for 3′ end processing (i.e. cleavage and polyadenylation) within their coding region. This is the case for approximately 0.7–0.8 % of yeast (such as CBP1 and RNA14) and human genes (Frischmeyer et al. 2002; Mayer and Dieckmann 1991; Sparks and Dieckmann 1998). Prematurely polyadenylated truncated forms of the yeast CBP1 and chicken Growth hormone receptor (for GHR) mRNAs are indeed NSD substrates. Hence, the NSD pathway could, under certain physiological conditions, regulate the abundance of some mRNAs.
NGD substrates are probably more frequent than NSD substrates in cells as various events can cause translational stalls. For instance, S-adenosyl-l-Methionine rules the stability of the A. thaliana CGS1 mRNA encoding cystathione γ-synthase by inducing translation elongation arrest followed by mRNA endonucleolytic cleavage (Onouchi et al. 2005). Furthermore, bioinformatics searches for yeast genes containing signals susceptible to enforce ribosome pausing (i.e. stretches of at least 10 consecutive basic residues, stable stem loops or pseudo-knots, …) have identified potential NGD substrates (Dimitrova et al. 2009; Jacobs et al. 2007). Some of these were experimentally characterised. The JJJ1, MAP2 and RMP1 mRNAs induce translational arrest and release of nonstop protein products. Similarly, upon DOM34 deletion, the steady state levels of mRNA encoding for Est2 and Bub3 are strongly stabilised while mRNA encoding for Spr6 is stabilised by two-fold (Belew et al. 2010).
The Dom34 and Hbs1 proteins have also been involved in the degradation of mRNAs containing apurinic sites, which can cause elongation stalls due to imperfect mRNA codon–tRNA anticodon base pairing (Gandhi et al. 2008). The occurrence of apurinic sites caused by chemical compounds is well characterised in DNA as well as the associated repair mechanisms such as base excision repair (BER), which allow regenerating an intact copy of the genetic information (Robertson et al. 2009). The chemical damages underwent by RNAs are much less characterised but growing evidences suggests that mRNAs as well as non-coding RNAs can be oxidised, alkylated and damaged by other means (reviewed in Wurtmann and Wolin 2009). For instance, the oxidation of mRNAs has been shown to lead to translation elongation stalls and reduction of the production of the corresponding proteins (Shan et al. 2007). This could be due to the action of NGD and NSD pathways. Compared to damaged DNA molecules, which have to be repaired to reduce spreading of errors during cell division, one can imagine that degradation of damaged mRNAs is less energy-consuming for cells than repair, in particular for transient molecules with short half-lives such as mRNAs. Hence, the NGD pathway may be one of the mechanisms used by eukaryotic cells to degrade subsets of damaged mRNAs enforcing ribosomes to stall during translation elongation.
Finally, several observations suggest that a biologically relevant function of the Dom34-Hbs1 complex is probably related to the degradation of immature or non-functional, small ribosomal subunits that cannot elongate properly (Cole et al. 2009; LaRiviere et al. 2006; Soudet et al. 2010; Strunk et al. 2012). Hence, the Dom34-Hbs1 complex could play a predominant role by detecting and inducing the degradation of non-functional ribosomes that have passed successfully through all the check points, are able to initiate translation but are unable to proceed in elongation.
Conclusion
Baker’s yeast undeniably played a central and key role for the identification and the description of these conserved translation-dependent eukaryotic mRNA QC pathways. Some twilight zones still persist and although studies performed with human cells become accessible to more laboratories, yeast will undoubtedly continue to play a major role in the future description of the still unknown steps of these processes. Among these terra incognita to be explored, the exact NMD mechanism responsible for the discrimination between premature and normal termination events remains to be clarified. Similarly, the molecular connection between the Upf factors and the decapping machinery remains fuzzy. The potential implication in NMD of other yet unidentified factors has also to be addressed. Regarding NSD and NGD, the description of the molecular mechanisms of these two processes has been very successful within the last 5 years and these studies also raised the veil on the mechanism of termination of the translation. However, further studies are clearly needed to decipher the physiological roles of these eukaryotic pathways. Finally, there is growing evidence linking these mRNA QC pathways with ribosome biogenesis process. Indeed, the Dbp2 rRNA processing factor is involved in NMD (Bond et al. 2001; Geissler et al. 2013), while Dom34 has been shown to play a role in a late GC checkpoint during 40S maturation (Soudet et al. 2010; Strunk et al. 2012). The relationships between these processes will have to be addressed in the future.
References
Amrani N, Ganesan R, Kervestin S, Mangus DA, Ghosh S, Jacobson A (2004) A faux 3′-UTR promotes aberrant termination and triggers nonsense-mediated mRNA decay. Nature 432(7013):112–118
Atkinson GC, Baldauf SL, Hauryliuk V (2008) Evolution of nonstop, no-go and nonsense-mediated mRNA decay and their termination factor-derived components. BMC Evol Biol 8:290
Becker T, Armache JP, Jarasch A, Anger AM, Villa E, Sieber H, Motaal BA, Mielke T, Berninghausen O, Beckmann R (2011) Structure of the no-go mRNA decay complex Dom34-Hbs1 bound to a stalled 80S ribosome. Nat Struct Mol Biol 18(6):715–720
Becker T, Franckenberg S, Wickles S, Shoemaker CJ, Anger AM, Armache JP, Sieber H, Ungewickell C, Berninghausen O, Daberkow I, Karcher A, Thomm M, Hopfner KP, Green R, Beckmann R (2012) Structural basis of highly conserved ribosome recycling in eukaryotes and archaea. Nature 482(7386):501–506
Bedwell DM, Kaenjak A, Benos DJ, Bebok Z, Bubien JK, Hong J, Tousson A, Clancy JP, Sorscher EJ (1997) Suppression of a CFTR premature stop mutation in a bronchial epithelial cell line. Nat Med 3(11):1280–1284
Belew AT, Advani VM, Dinman JD (2010) Endogenous ribosomal frameshift signals operate as mRNA destabilizing elements through at least two molecular pathways in yeast. Nucleic Acids Res 39(7):2799–2808
Bengtson MH, Joazeiro CA (2010) Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature 467(7314):470–473
Bhattacharya A, Czaplinski K, Trifillis P, He F, Jacobson A, Peltz SW (2000) Characterization of the biochemical properties of the human Upf1 gene product that is involved in nonsense-mediated mRNA decay. RNA 6(9):1226–1235
Bidou L, Allamand V, Rousset JP, Namy O (2012) Sense from nonsense: therapies for premature stop codon diseases. Trends Mol Med 18(11):679–688
Bond AT, Mangus DA, He F, Jacobson A (2001) Absence of Dbp2p alters both nonsense-mediated mRNA decay and rRNA processing. Mol Cell Biol 21(21):7366–7379
Brandman O, Stewart-Ornstein J, Wong D, Larson A, Williams CC, Li GW, Zhou S, King D, Shen PS, Weibezahn J, Dunn JG, Rouskin S, Inada T, Frost A, Weissman JS (2012) A ribosome-bound quality control complex triggers degradation of nascent peptides and signals translation stress. Cell 151(5):1042–1054
Buhler M, Steiner S, Mohn F, Paillusson A, Muhlemann O (2006) EJC-independent degradation of nonsense immunoglobulin-mu mRNA depends on 3′ UTR length. Nat Struct Mol Biol 13(5):462–464
Carr-Schmid A, Pfund C, Craig EA, Kinzy TG (2002) Novel G-protein complex whose requirement is linked to the translational status of the cell. Mol Cell Biol 22(8):2564–2574
Chakrabarti S, Jayachandran U, Bonneau F, Fiorini F, Basquin C, Domcke S, Le Hir H, Conti E (2011) Molecular mechanisms for the RNA-dependent ATPase activity of Upf1 and its regulation by Upf2. Mol Cell 41(6):693–703
Chamieh H, Ballut L, Bonneau F, Le Hir H (2008) NMD factors UPF2 and UPF3 bridge UPF1 to the exon junction complex and stimulate its RNA helicase activity. Nat Struct Mol Biol 15(1):85–93
Chen L, Muhlrad D, Hauryliuk V, Cheng Z, Lim MK, Shyp V, Parker R, Song H (2010) Structure of the Dom34-Hbs1 complex and implications for no-go decay. Nat Struct Mol Biol 17(10):1233–1240
Clerici M, Mourao A, Gutsche I, Gehring NH, Hentze MW, Kulozik A, Kadlec J, Sattler M, Cusack S (2009) Unusual bipartite mode of interaction between the nonsense-mediated decay factors, UPF1 and UPF2. EMBO J 28(15):2293–2306
Cole SE, LaRiviere FJ, Merrikh CN, Moore MJ (2009) A convergence of rRNA and mRNA quality control pathways revealed by mechanistic analysis of nonfunctional rRNA decay. Mol Cell 34(4):440–450
Cui Y, Hagan KW, Zhang S, Peltz SW (1995) Identification and characterization of genes that are required for the accelerated degradation of mRNAs containing a premature translational termination codon. Genes Dev 9(4):423–436
Czaplinski K, Weng Y, Hagan KW, Peltz SW (1995) Purification and characterization of the Upf1 protein: a factor involved in translation and mRNA degradation. RNA 1(6):610–623
de la Cruz J, Kressler D, Linder P (1999) Unwinding RNA in Saccharomyces cerevisiae: DEAD-box proteins and related families. Trends Biochem Sci 24(5):192–198
Defenouillere Q, Yao Y, Mouaikel J, Namane A, Galopier A, Decourty L, Doyen A, Malabat C, Saveanu C, Jacquier A, Fromont-Racine M (2013) Cdc48-associated complex bound to 60S particles is required for the clearance of aberrant translation products. Proc Natl Acad Sci USA 110(13):5046–5051
Dimitrova LN, Kuroha K, Tatematsu T, Inada T (2009) Nascent peptide-dependent translation arrest leads to Not4p-mediated protein degradation by the proteasome. J Biol Chem 284(16):10343–10352
Doma MK, Parker R (2006) Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 440(7083):561–564
Eberle AB, Lykke-Andersen S, Muhlemann O, Jensen TH (2009) SMG6 promotes endonucleolytic cleavage of nonsense mRNA in human cells. Nat Struct Mol Biol 16(1):49–55
Ford AS, Guan Q, Neeno-Eckwall E, Culbertson MR (2006) Ebs1p, a negative regulator of gene expression controlled by the Upf proteins in the yeast Saccharomyces cerevisiae. Eukaryot Cell 5(2):301–312
Frischmeyer PA, van Hoof A, O’Donnell K, Guerrerio AL, Parker R, Dietz HC (2002) An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science 295(5563):2258–2261
Gandhi R, Manzoor M, Hudak KA (2008) Depurination of Brome mosaic virus RNA3 in vivo results in translation-dependent accelerated degradation of the viral RNA. J Biol Chem 283(47):32218–32228
Gardner LB (2010) Nonsense-mediated RNA decay regulation by cellular stress: implications for tumorigenesis. Mol Cancer Res 8(3):295–308
Gatfield D, Izaurralde E (2004) Nonsense-mediated messenger RNA decay is initiated by endonucleolytic cleavage in Drosophila. Nature 429(6991):575–578
Geissler V, Altmeyer S, Stein B, Uhlmann-Schiffler H, Stahl H (2013) The RNA helicase Ddx5/p68 binds to hUpf3 and enhances NMD of Ddx17/p72 and Smg5 mRNA. Nucleic Acids Res 41(16):7875–7888
Gilboa E (2013) Expression of new antigens on tumor cells by inhibiting nonsense-mediated mRNA decay. Immunol Res 57:44–51
Gonzalez CI, Ruiz-Echevarria MJ, Vasudevan S, Henry MF, Peltz SW (2000) The yeast hnRNP-like protein Hrp1/Nab4 marks a transcript for nonsense-mediated mRNA decay. Mol Cell 5(3):489–499
Graille M, Chaillet M, van Tilbeurgh H (2008) Structure of yeast Dom34: a protein related to translation termination factor Erf1 and involved in No-Go decay. J Biol Chem 283(11):7145–7154
Graille M, Seraphin B (2012) Surveillance pathways rescuing eukaryotic ribosomes lost in translation. Nat Rev Mol Cell Biol 13(11):727–735
Hagan KW, Ruiz-Echevarria MJ, Quan Y, Peltz SW (1995) Characterization of cis-acting sequences and decay intermediates involved in nonsense-mediated mRNA turnover. Mol Cell Biol 15(2):809–823
Hansen KD, Lareau LF, Blanchette M, Green RE, Meng Q, Rehwinkel J, Gallusser FL, Izaurralde E, Rio DC, Dudoit S, Brenner SE (2009) Genome-wide identification of alternative splice forms down-regulated by nonsense-mediated mRNA decay in Drosophila. PLoS Genet 5(6):e1000525
He F, Brown AH, Jacobson A (1997) Upf1p, Nmd2p, and Upf3p are interacting components of the yeast nonsense-mediated mRNA decay pathway. Mol Cell Biol 17(3):1580–1594
He F, Jacobson A (1995) Identification of a novel component of the nonsense-mediated mRNA decay pathway by use of an interacting protein screen. Genes Dev 9(4):437–454
He F, Jacobson A (2001) Upf1p, Nmd2p, and Upf3p regulate the decapping and exonucleolytic degradation of both nonsense-containing mRNAs and wild-type mRNAs. Mol Cell Biol 21(5):1515–1530
He F, Li X, Spatrick P, Casillo R, Dong S, Jacobson A (2003) Genome-wide analysis of mRNAs regulated by the nonsense-mediated and 5′ to 3′ mRNA decay pathways in yeast. Mol Cell 12(6):1439–1452
Hogg JR, Goff SP (2010) Upf1 senses 3′UTR length to potentiate mRNA decay. Cell 143(3):379–389
Hwang J, Sato H, Tang Y, Matsuda D, Maquat LE (2010) UPF1 association with the cap-binding protein, CBP80, promotes nonsense-mediated mRNA decay at two distinct steps. Mol Cell 39(3):396–409
Inada T, Aiba H (2005) Translation of aberrant mRNAs lacking a termination codon or with a shortened 3′-UTR is repressed after initiation in yeast. EMBO J 24(8):1584–1595
Ito-Harashima S, Kuroha K, Tatematsu T, Inada T (2007) Translation of the poly(A) tail plays crucial roles in nonstop mRNA surveillance via translation repression and protein destabilization by proteasome in yeast. Genes Dev 21(5):519–524
Jacobs JL, Belew AT, Rakauskaite R, Dinman JD (2007) Identification of functional, endogenous programmed −1 ribosomal frameshift signals in the genome of Saccharomyces cerevisiae. Nucleic Acids Res 35(1):165–174
Johansson MJ, He F, Spatrick P, Li C, Jacobson A (2007) Association of yeast Upf1p with direct substrates of the NMD pathway. Proc Natl Acad Sci USA 104(52):20872–20877
Kadlec J, Guilligay D, Ravelli RB, Cusack S (2006) Crystal structure of the UPF2-interacting domain of nonsense-mediated mRNA decay factor UPF1. RNA 12(10):1817–1824
Kadlec J, Izaurralde E, Cusack S (2004) The structural basis for the interaction between nonsense-mediated mRNA decay factors UPF2 and UPF3. Nat Struct Mol Biol 11(4):330–337
Kebaara BW, Atkin AL (2009) Long 3′-UTRs target wild-type mRNAs for nonsense-mediated mRNA decay in Saccharomyces cerevisiae. Nucleic Acids Res 37(9):2771–2778
Kervestin S, Jacobson A (2012) NMD: a multifaceted response to premature translational termination. Nat Rev Mol Cell Biol 13(11):700–712
Kervestin S, Li C, Buckingham R, Jacobson A (2012) Testing the faux-UTR model for NMD: analysis of Upf1p and Pab1p competition for binding to eRF3/Sup35p. Biochimie 94(7):1560–1571
Kessler MM, Henry MF, Shen E, Zhao J, Gross S, Silver PA, Moore CL (1997) Hrp1, a sequence-specific RNA-binding protein that shuttles between the nucleus and the cytoplasm, is required for mRNA 3′-end formation in yeast. Genes Dev 11(19):2545–2556
Kobayashi K, Kikuno I, Kuroha K, Saito K, Ito K, Ishitani R, Inada T, Nureki O (2010) Structural basis for mRNA surveillance by archaeal Pelota and GTP-bound EF1alpha complex. Proc Natl Acad Sci USA 107(41):17575–17579
Kobayashi K, Saito K, Ishitani R, Ito K, Nureki O (2012) Structural basis for translation termination by archaeal RF1 and GTP-bound EF1a complex. Nucleic Acids Res 40(18):9319–9328
Kuroha K, Akamatsu M, Dimitrova L, Ito T, Kato Y, Shirahige K, Inada T (2010) Receptor for activated C kinase 1 stimulates nascent polypeptide-dependent translation arrest. EMBO Rep 11(12):956–961
Kuroha K, Ando K, Nakagawa R, Inada T (2013) The Upf Factor Complex Interacts with Aberrant Products Derived from mRNAs Containing a Premature Termination Codon and Facilitates Their Proteasomal Degradation. J Biol Chem 288(40):28630–28640
Kuroha K, Tatematsu T, Inada T (2009) Upf1 stimulates degradation of the product derived from aberrant messenger RNA containing a specific nonsense mutation by the proteasome. EMBO Rep 10(11):1265–1271
Kurosaki T, Maquat LE (2013) Rules that govern UPF1 binding to mRNA 3′ UTRs. Proc Natl Acad Sci USA 110(9):3357–3362
LaRiviere FJ, Cole SE, Ferullo DJ, Moore MJ (2006) A late-acting quality control process for mature eukaryotic rRNAs. Mol Cell 24(4):619–626
Lasalde C, Rivera AV, Leon AJ, Gonzalez-Feliciano JA, Estrella LA, Rodriguez-Cruz EN, Correa ME, Cajigas IJ, Bracho DP, Vega IE, Wilkinson MF, Gonzalez CI (2014) Identification and functional analysis of novel phosphorylation sites in the RNA surveillance protein Upf1. Nucleic Acids Res 42(3):1916–1929
Lee BS, Culbertson MR (1995) Identification of an additional gene required for eukaryotic nonsense mRNA turnover. Proc Natl Acad Sci USA 92(22):10354–10358
Lee HH, Kim YS, Kim KH, Heo I, Kim SK, Kim O, Kim HK, Yoon JY, Kim HS, Kim do J, Lee SJ, Yoon HJ, Kim SJ, Lee BG, Song HK, Kim VN, Park CM, Suh SW (2007) Structural and functional insights into Dom34, a key component of No-Go mRNA decay. Mol Cell 27(6):938–950
Leeds P, Wood JM, Lee BS, Culbertson MR (1992) Gene products that promote mRNA turnover in Saccharomyces cerevisiae. Mol Cell Biol 12(5):2165–2177
Li S, Wilkinson MF (1998) Nonsense surveillance in lymphocytes? Immunity 8(2):135–141
Losson R, Lacroute F (1979) Interference of nonsense mutations with eukaryotic messenger RNA stability. Proc Natl Acad Sci USA 76(10):5134–5137
Luke B, Azzalin CM, Hug N, Deplazes A, Peter M, Lingner J (2007) Saccharomyces cerevisiae Ebs1p is a putative ortholog of human Smg7 and promotes nonsense-mediated mRNA decay. Nucleic Acids Res 35(22):7688–7697
Mayer SA, Dieckmann CL (1991) Yeast CBP1 mRNA 3′ end formation is regulated during the induction of mitochondrial function. Mol Cell Biol 11(2):813–821
Meaux S, Van Hoof A (2006) Yeast transcripts cleaved by an internal ribozyme provide new insight into the role of the cap and poly(A) tail in translation and mRNA decay. RNA 12(7):1323–1337
Meaux S, van Hoof A, Baker KE (2008) Nonsense-mediated mRNA decay in yeast does not require PAB1 or a poly(A) tail. Mol Cell 29(1):134–140
Mendell JT, Sharifi NA, Meyers JL, Martinez-Murillo F, Dietz HC (2004) Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat Genet 36(10):1073–1078
Min EE, Roy B, Amrani N, He F, Jacobson A (2013) Yeast Upf1 CH domain interacts with Rps26 of the 40S ribosomal subunit. RNA 19(8):1105–1115
Mitchell P, Tollervey D (2003) An NMD pathway in yeast involving accelerated deadenylation and exosome-mediated 3′ –> 5′ degradation. Mol Cell 11(5):1405–1413
Mitrovich QM, Anderson P (2000) Unproductively spliced ribosomal protein mRNAs are natural targets of mRNA surveillance in C. elegans. Genes Dev 14(17):2173–2184
Muhlrad D, Parker R (1994) Premature translational termination triggers mRNA decapping. Nature 370(6490):578–581
Muhlrad D, Parker R (1999) Aberrant mRNAs with extended 3′ UTRs are substrates for rapid degradation by mRNA surveillance. RNA 5(10):1299–1307
Nissen P, Kjeldgaard M, Thirup S, Polekhina G, Reshetnikova L, Clark BF, Nyborg J (1995) Crystal structure of the ternary complex of Phe-tRNAPhe, EF-Tu, and a GTP analog. Science 270(5241):1464–1472
Ohnishi T, Yamashita A, Kashima I, Schell T, Anders KR, Grimson A, Hachiya T, Hentze MW, Anderson P, Ohno S (2003) Phosphorylation of hUPF1 induces formation of mRNA surveillance complexes containing hSMG-5 and hSMG-7. Mol Cell 12(5):1187–1200
Onouchi H, Nagami Y, Haraguchi Y, Nakamoto M, Nishimura Y, Sakurai R, Nagao N, Kawasaki D, Kadokura Y, Naito S (2005) Nascent peptide-mediated translation elongation arrest coupled with mRNA degradation in the CGS1 gene of Arabidopsis. Genes Dev 19(15):1799–1810
Panasenko O, Landrieux E, Feuermann M, Finka A, Paquet N, Collart MA (2006) The yeast Ccr4-Not complex controls ubiquitination of the nascent-associated polypeptide (NAC-EGD) complex. J Biol Chem 281(42):31389–31398
Passos DO, Doma MK, Shoemaker CJ, Muhlrad D, Green R, Weissman J, Hollien J, Parker R (2009) Analysis of Dom34 and Its Function in No-Go Decay. Mol Biol Cell 20(13):3025–3032
Pastor F, Kolonias D, Giangrande PH, Gilboa E (2010) Induction of tumour immunity by targeted inhibition of nonsense-mediated mRNA decay. Nature 465(7295):227–230
Peltz SW, Brown AH, Jacobson A (1993) mRNA destabilization triggered by premature translational termination depends on at least three cis-acting sequence elements and one trans-acting factor. Genes Dev 7(9):1737–1754
Pisarev AV, Skabkin MA, Pisareva VP, Skabkina OV, Rakotondrafara AM, Hentze MW, Hellen CU, Pestova TV (2010) The role of ABCE1 in eukaryotic posttermination ribosomal recycling. Mol Cell 37(2):196–210
Pulak R, Anderson P (1993) mRNA surveillance by the Caenorhabditis elegans smg genes. Genes Dev 7(10):1885–1897
Rehwinkel J, Letunic I, Raes J, Bork P, Izaurralde E (2005) Nonsense-mediated mRNA decay factors act in concert to regulate common mRNA targets. RNA 11(10):1530–1544
Robertson AB, Klungland A, Rognes T, Leiros I (2009) DNA repair in mammalian cells: Base excision repair: the long and short of it. Cell Mol Life Sci 66(6):981–993
Ruiz-Echevarria MJ, Gonzalez CI, Peltz SW (1998) Identifying the right stop: determining how the surveillance complex recognizes and degrades an aberrant mRNA. EMBO J 17(2):575–589
Ruiz-Echevarria MJ, Peltz SW (2000) The RNA binding protein Pub1 modulates the stability of transcripts containing upstream open reading frames. Cell 101(7):741–751
Saito S, Hosoda N, Hoshino S (2013) The Hbs1-Dom34 protein complex functions in non-stop mRNA decay in mammalian cells. J Biol Chem 288(24):17832–17843
Sauliere J, Haque N, Harms S, Barbosa I, Blanchette M, Le Hir H (2010) The exon junction complex differentially marks spliced junctions. Nat Struct Mol Biol 17(10):1269–1271
Schaeffer D, van Hoof A (2011) Different nuclease requirements for exosome-mediated degradation of normal and nonstop mRNAs. Proc Natl Acad Sci USA 108(6):2366–2371
Schmeing TM, Voorhees RM, Kelley AC, Gao YG, FVt Murphy, Weir JR, Ramakrishnan V (2009) The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326(5953):688–694
Seminara SB, Messager S, Chatzidaki EE, Thresher RR, Acierno JS Jr, Shagoury JK, Bo-Abbas Y, Kuohung W, Schwinof KM, Hendrick AG, Zahn D, Dixon J, Kaiser UB, Slaugenhaupt SA, Gusella JF, O’Rahilly S, Carlton MB, Crowley WF Jr, Aparicio SA, Colledge WH (2003) The GPR54 gene as a regulator of puberty. N Engl J Med 349(17):1614–1627
Shan X, Chang Y, Lin CL (2007) Messenger RNA oxidation is an early event preceding cell death and causes reduced protein expression. FASEB J 21(11):2753–2764
Shigeoka T, Kato S, Kawaichi M, Ishida Y (2012) Evidence that the Upf1-related molecular motor scans the 3′-UTR to ensure mRNA integrity. Nucleic Acids Res 40(14):6887–6897
Shirley RL, Lelivelt MJ, Schenkman LR, Dahlseid JN, Culbertson MR (1998) A factor required for nonsense-mediated mRNA decay in yeast is exported from the nucleus to the cytoplasm by a nuclear export signal sequence. J Cell Sci 111(Pt 21):3129–3143
Shoemaker CJ, Eyler DE, Green R (2010) Dom34:Hbs1 promotes subunit dissociation and peptidyl-tRNA drop-off to initiate no-go decay. Science 330(6002):369–372
Soudet J, Gelugne JP, Belhabich-Baumas K, Caizergues-Ferrer M, Mougin A (2010) Immature small ribosomal subunits can engage in translation initiation in Saccharomyces cerevisiae. EMBO J 29(1):80–92
Sparks KA, Dieckmann CL (1998) Regulation of poly(A) site choice of several yeast mRNAs. Nucleic Acids Res 26(20):4676–4687
Strunk BS, Novak MN, Young CL, Karbstein K (2012) A Translation-Like Cycle Is a Quality Control Checkpoint for Maturing 40S Ribosome Subunits. Cell 150(1):111–121
Swisher KD, Parker R (2011) Interactions between Upf1 and the decapping factors Edc3 and Pat1 in Saccharomyces cerevisiae. PLoS ONE 6(10):e26547
Takahashi S, Araki Y, Ohya Y, Sakuno T, Hoshino S, Kontani K, Nishina H, Katada T (2008) Upf1 potentially serves as a RING-related E3 ubiquitin ligase via its association with Upf3 in yeast. RNA 14(9):1950–1958
Taniguchi A, Hakoda M, Yamanaka H, Terai C, Hikiji K, Kawaguchi R, Konishi N, Kashiwazaki S, Kamatani N (1998) A germline mutation abolishing the original stop codon of the human adenine phosphoribosyltransferase (APRT) gene leads to complete loss of the enzyme protein. Hum Genet 102(2):197–202
Tsuboi T, Kuroha K, Kudo K, Makino S, Inoue E, Kashima I, Inada T (2012) Dom34:hbs1 plays a general role in quality-control systems by dissociation of a stalled ribosome at the 3′ end of aberrant mRNA. Mol Cell 46(4):518–529
van den Elzen AM, Henri J, Lazar N, Gas ME, Durand D, Lacroute F, Nicaise M, van Tilbeurgh H, Seraphin B, Graille M (2010) Dissection of Dom34-Hbs1 reveals independent functions in two RNA quality control pathways. Nat Struct Mol Biol 17(12):1446–1452
van Hoof A, Frischmeyer PA, Dietz HC, Parker R (2002) Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295(5563):2262–2264
Wang D, Wengrod J, Gardner LB (2011a) Overexpression of the c-myc oncogene inhibits nonsense-mediated RNA decay in B lymphocytes. J Biol Chem 286(46):40038–40043
Wang D, Zavadil J, Martin L, Parisi F, Friedman E, Levy D, Harding H, Ron D, Gardner LB (2011b) Inhibition of nonsense-mediated RNA decay by the tumor microenvironment promotes tumorigenesis. Mol Cell Biol 31(17):3670–3680
Wang W, Cajigas IJ, Peltz SW, Wilkinson MF, González CI (2006) Role for Upf2p phosphorylation in Saccharomyces cerevisiae nonsense-mediated mRNA decay. Mol Cell Biol 26(9):3390–3400
Welch EM, Barton ER, Zhuo J, Tomizawa Y, Friesen WJ, Trifillis P, Paushkin S, Patel M, Trotta CR, Hwang S, Wilde RG, Karp G, Takasugi J, Chen G, Jones S, Ren H, Moon YC, Corson D, Turpoff AA, Campbell JA, Conn MM, Khan A, Almstead NG, Hedrick J, Mollin A, Risher N, Weetall M, Yeh S, Branstrom AA, Colacino JM, Babiak J, Ju WD, Hirawat S, Northcutt VJ, Miller LL, Spatrick P, He F, Kawana M, Feng H, Jacobson A, Peltz SW, Sweeney HL (2007) PTC124 targets genetic disorders caused by nonsense mutations. Nature 447(7140):87–91
Welch EM, Jacobson A (1999) An internal open reading frame triggers nonsense-mediated decay of the yeast SPT10 mRNA. EMBO J 18(21):6134–6145
Weng Y, Czaplinski K, Peltz SW (1996) Identification and characterization of mutations in the UPF1 gene that affect nonsense suppression and the formation of the Upf protein complex but not mRNA turnover. Mol Cell Biol 16(10):5491–5506
Wilschanski M, Yahav Y, Yaacov Y, Blau H, Bentur L, Rivlin J, Aviram M, Bdolah-Abram T, Bebok Z, Shushi L, Kerem B, Kerem E (2003) Gentamicin-induced correction of CFTR function in patients with cystic fibrosis and CFTR stop mutations. N Engl J Med 349(15):1433–1441
Wilson MA, Meaux S, van Hoof A (2007) A genomic screen in yeast reveals novel aspects of nonstop mRNA metabolism. Genetics 177(2):773–784
Wilusz CJ, Wilusz J (2005) Eukaryotic Lsm proteins: lessons from bacteria. Nat Struct Mol Biol 12(12):1031–1036
Wittmann J, Hol EM, Jack HM (2006) hUPF2 silencing identifies physiologic substrates of mammalian nonsense-mediated mRNA decay. Mol Cell Biol 26(4):1272–1287
Wurtmann EJ, Wolin SL (2009) RNA under attack: cellular handling of RNA damage. Crit Rev Biochem Mol Biol 44(1):34–49
Zhang S, Ruiz-Echevarria MJ, Quan Y, Peltz SW (1995) Identification and characterization of a sequence motif involved in nonsense-mediated mRNA decay. Mol Cell Biol 15(4):2231–2244
Zund D, Gruber AR, Zavolan M, Muhlemann O (2013) Translation-dependent displacement of UPF1 from coding sequences causes its enrichment in 3′ UTRs. Nat Struct Mol Biol 20(8):936–943
Acknowledgments
MG acknowledges funding from the Centre National pour la Recherche Scientifique ATIP-AVENIR program, the Agence Nationale pour la Recherche (grants ANR-06-BLAN-0075-02 and ANR-11-BSV800902), the Association Française contre les Myopathies, and the Human Frontiers Science Program (grant RGP0018). ZF acknowledges the financial support from the Fondation pour la Recherche Médicale (FRM). The authors apologise for the many studies that could not be cited due to space constraints.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Fourati, Z., Graille, M. (2014). Cytoplasmic mRNA Surveillance Pathways. In: Sesma, A., von der Haar, T. (eds) Fungal RNA Biology. Springer, Cham. https://doi.org/10.1007/978-3-319-05687-6_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-05687-6_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-05686-9
Online ISBN: 978-3-319-05687-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)