Abstract
MicroRNAs (miRNAs) constitute an evolutionarily conserved class of small, noncoding RNA molecules that regulate gene expression by targeting specific mRNAs for degradation and/or translational repression. MiRNAs have been widely investigated due to their potential role in regulating a variety of cellular processes, including proliferation, differentiation, and apoptosis. Many miRNAs are implicated in various human cancers. Functional analysis of cancer-related miRNAs has proposed that they might act as either oncogenes or tumor suppressors. In fact, the link between aberrant miRNA expression and cancer development and progression can be observed either through the loss of tumor suppressor miRNAs or the over-expression of oncogenic miRNAs. This chapter aims to provide a succinct framework to gain insight into miRNA function in cancer.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 MicroRNAs: Biogenesis, Processing and Mode of Action
MicroRNAs (miRNAs or miRs) are a class of non-coding small RNAs of ~22 nucleotides that regulate gene expression by targeting specific mRNAs bearing partially complementary target sequences for degradation and/or translational repression (Liu et al. 2008; Babashah and Soleimani 2011). The first discovery of a small non-coding RNA dates back to 1993, when Victor Ambros and collaborators identified lin-4 in Caenorhabditis elegans (Lee et al. 1993). Lin-4 was believed to be a unique species until year 2000 when another small non-coding RNA, let-7, was reported in C. elegans (Reinhart et al. 2000) and in a variety of other organisms (Pasquinelli et al. 2000). Since then, hundreds of small non-coding RNA sequences (now known to be miRNAs) have been identified in a wide range of organisms from nematodes to vertebrates, plants and human. Currently, the official miRNA database miRBase lists 1,872 human miRNA gene loci, generating 2,578 mature miRNA sequences (http://www.mirbase.org, Release 20.0, June 2013). Precise attribution of miRNA effects on gene expression can be complicated by the fact that often each miRNA may control several hundred target genes directly or indirectly, whereas a single protein coding gene target could be regulated by more than one miRNA. In fact, miRNAs are predicted to target up to one-third of human transcripts (Zhong et al. 2012; Friedman et al. 2009).
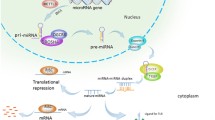
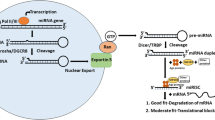
The biogenesis of miRNAs begins in the nucleus with the synthesis of a relatively long double-stranded RNA molecule, known as primary (pri)-miRNA, by RNA polymerase II or III. The resultant pri-miRNA transcript is often more longer than 1 kb in length and includes a stable stem-loop hairpin structure that contains the sequence for the mature miRNA. The hairpin structure is excised in the nucleus from pri-miRNA as a ~70-nucleotide long precursor (pre)-miRNA by the nuclear RNase III endonuclease Drosha and DGCR8 (the “microprocessor complex”) (Lee et al. 2003; Denli et al. 2004; Gregory et al. 2004). DGCR8 is essential as a molecular anchor for Drosha’s activity on pri-mRNAs, as it recognizes the pri-miRNA at double-stranded RNA – single-stranded RNA junction and directs Drosha to cleave approximately 11 nucleotides from the base of the stem to free the hairpin from the primary transcript (Han et al. 2006). Members of the microprocessor complex have additional cellular functions, as Drosha is also involved in the processing of ribosomal RNA (Wu et al. 2000) and DGCR8 also acts as a heme-binding protein (Faller et al. 2007). The resultant pre-miRNA contains a 5′ phosphate and a distinctive 3′ two-nucleotide overhang which is signal to transport into the cytoplasm by a protein complex consisting of Exportin-5 and Ran-GTPase (Yi et al. 2003; Lund et al. 2004; Bohnsack et al. 2004) (Fig. 1.1). In cytoplasm, further processing facilitated by the second RNase III endonuclease Dicer, cuts off the terminal loop and generates an imperfect double-stranded RNA with about 17-26-nucleotide in length. This duplex molecule contains the mature miRNA (often designated miR) and its complementary miRNA*. The duplex binds to one of four proteins of the Argonaute (Ago) family, which are part of the RNA-induced silencing complex (RISC). After unwinding the double-stranded RNA and discarding and degrading the passenger strand (miRNA*), the mature miRNA is loaded onto the RISC and interacts with the complementary sequences that are mostly located in the 3′ untranslated region (3′ UTR) of the targeted mRNAs (Cullen 2004; Liu et al. 2008; Ikeda et al. 2007). Subsequent mechanisms by which miRNAs regulate gene expression depend on the degree of complementarity between mRNA target sites and the nucleotide sequence from position 2–8 at the 5′ end of miRNAs (the seed region). The rare occasion of perfect (or near perfect) Watson-Crick complementarity leads to Ago-catalysed cleavage of the targeted mRNA. More commonly, imperfect complementarity leads to translational inhibition, although the precise mechanisms and the players involved are still under debate (reviewed in Fabian and Sonenberg (2012); Pasquinelli (2012)) (Fig. 1.1).
Schematic representation of biogenesis, processing and function of microRNA. The biogenesis of miRNAs begins in the nucleus and is completed in the cytoplasm. For more details, see the text. Pri-miRNA primary miRNA, Pre-miRNA precursor miRNA, Drosha RNase III endonuclease, DGCR8 DiGeorge syndrome critical region 8, Dicer RNase III endonuclease, RISC RNA-induced silencing complex
Owing to the imperfect complementarity between miRNAs and their target mRNAs almost observed in mammals, direct prediction of relevant downstream targets of a miRNA is particularly difficult. Several bioinformatic approaches and various algorithms have been developed to predict miRNA-controlled target mRNAs in silico (Lewis et al. 2003; Krek et al. 2005; Paraskevopoulou et al. 2013). A list of computational tools for miRNA target prediction is summarized in Table 1.1. However, as the bioinformatic approach focuses on identifying conserved targets in the 3′-UTR of an mRNA, many non-conserved targets are missed. In addition, there are several lines of evidence indicating that miRNAs can also regulate gene expression through binding to “seedless” 3′-UTR miRNA recognition elements (Lal et al. 2009) or to sites located within the coding regions of transcript (Lee et al. 2009). Therefore, the efficacy of such a bioinformatic approach needs to be validated by in vitro or in vivo experiments.
MiRNAs are involved in the control of a variety of biological processes, including cellular proliferation, tissue differentiation, organ development, maintenance of stem cell potency and apoptosis (Babashah and Soleimani 2011; Cheng et al. 2005; Chen et al. 2004; Ambros 2004). Given this wide variety of functions, it is not surprising that miRNAs are affected in many diseases such as cancer. In fact, dysregulation of miRNAs has been widely observed in different types and stages of cancer and mounting evidence points to their important roles in the development of a variety of human cancers (Bandyopadhyay et al. 2010; Esquela-Kerscher and Slack 2006; Lu et al. 2005; Volinia et al. 2006).
2 The Oncogenic and Tumor Suppressive Roles of MicroRNAs in Cancer
Aberrant expression of miRNAs has been frequently noted in almost all types of cancer (Croce 2009; Farazi et al. 2011). Functional analysis of these aberrantly expressed miRNAs indicates that they might function as either oncogenes or tumor suppressors. The oncogenic miRNAs, called as “oncomiRs”, are up-regulated in cancer and usually promote tumor development by inhibiting tumor suppressor genes and/or genes that control cell differentiation or apoptosis. On the contrary, there are many down-regulated miRNAs which may be considered as tumor suppressors in cancer. These miRNAs are called as “TSmiRs” and may function by inhibiting oncogenes and/or genes that inhibit cell differentiation or apoptosis (Bandyopadhyay et al. 2010; Esquela-Kerscher and Slack 2006; Lu et al. 2005; Babashah and Soleimani 2011). Deregulation of miRNA expression frequently results from genetic mutations and/or epigenetic alterations, represented by deletions, amplifications, point mutations and aberrant DNA methylation events. Indeed, about half of the cancer-related miRNA genes are located at fragile sites of the genome as well as in minimal regions with loss of heterozygosity, minimal regions of amplification or common breakpoint regions (Calin et al. 2002, 2004b).
The first evidence for the involvement of miRNAs in tumorigenesis was reported by Calin et al. (2002) in describing a chromosome region containing the miR-15a/miR-16-1 cluster, which is frequently lost or down-regulated in B-cell chronic lymphocytic leukemia (B-CLL). Down-regulation of the miR-15a/miR-16-1 cluster in CLL and several solid tumors raised the question whether they might function as tumor suppressors (Calin et al. 2002). Cimmino et al. (2005) demonstrated that both miR-15a and miR-16-1 promote the normal apoptotic response by directly targeting the anti-apoptotic gene BCL-2, indicating the possible tumor suppressive role of these two miRNAs in tumorigenesis.
A common tumor suppressive role for the let-7 family of miRNAs has been described in different types of human tissues, particularly in lung. It has been shown that let-7 is able to negatively regulate the expression of various oncogenes such as RAS and MYC as well as other cell cycle progression genes (Johnson et al. 2005; Bhat-Nakshatri et al. 2009). Reduced expression of Let-7 has been observed in different types of cancers, including lung, breast and prostate cancers. It has been shown that down-regulation of let-7 correlates with increased lymph node metastasis and proliferation capacity, suggesting a potential tumor suppressive role for this family of miRNAs in cancer progression (Lynam-Lennon et al. 2009; Liu et al. 2012). Although it has been demonstrated that induction of let-7 reduces tumor growth in a murine model of lung cancer (Esquela-Kerscher et al. 2008; Kumar et al. 2008), the regulation of individual let-7 targets on tumorigenesis needs to be further investigated in more in vivo models of human cancers.
The miR-17-92 cluster (containing seven homologous miRNAs: miR-17-3p, miR-17-5p, miR-18a, miR-20a, miR-19a, miR-19b-1 and miR-92a-1; with genomic positions on chromosomes X, 7 and 13) is the first and well-studied miRNA cluster with oncogenic activity. He et al. (2005) investigated the potential oncogenic role of the miR-17-92 cluster. They demonstrated that over-expression of the miR-17-92 cluster in the hematopoietic system acted with c-myc expression to accelerate tumor development and progression in a transgenic mouse model of B-cell lymphoma. Importantly, tumors resulting from combined c-Myc and miR-17-92 expression were able to evade from normal apoptotic responses that were otherwise prevalent in tumors lacking the cluster. O’Donnell et al. (2005) found that c-Myc activates expression of a set of six miRNAs on human chromosome 13 that was tied to the development of human lymphoma. They also found that expression of E2F1 was negatively regulated by two miRNAs in this cluster, miR-17-5p and miR-20a. These findings reveal a mechanism through which the c-Myc simultaneously promotes E2F1 transcription and represses following translation, indicating a tightly controlled proliferative signal. Woods et al. (2007) proposed a model in which the miR-17-92 cluster promotes cell proliferation by shifting the E2F transcriptional balance away from the pro-apoptotic E2F1 and toward the proliferative E2F3 transcriptional network. The miR-17-92 cluster might also inhibit apoptosis by negatively regulating the tumor suppressor PTEN and the pro-apoptotic protein Bim (Xiao et al. 2008; Mendell 2008). Bim is induced by Myc in B-cells and is able to antagonize anti-apoptotic proteins such as Bcl-2. Therefore, down-regulation of Bim by the miR-17-92 cluster may contribute to the ability of these miRNAs to exacerbate disease progression in a mouse model of B-cell leukemia (Egle et al. 2004).
As stated above, miRNAs can function either as oncogenes or tumor suppressors. However, it has been demonstrated that a miRNA can exploit both functions according to the cellular context of their target genes. For instance, there is a body of evidence pointing to the tumor suppressive activity of the miR-17-92 cluster, which contrasts with the hypothesized oncogenic role observed in other cancers (Yu et al. 2008). This implies that the tissue- and developmental-stage-specific expression decisively controls appropriate function of a miRNA.
3 MicroRNAs and Tumor Metastasis
Tumor invasion and metastasis are major characteristics of aggressive phenotypes observed in human cancers (Steeg 2003). During the “invasion-metastasis cascade”, cancer cells (a) are detached and migrate out of the primary tumor site; (b) invade the basement membrane to enter the circulatory system (intravasation); (c) are translocated through the vasculature; (d) exit circulatory vessels at the metastatic site (extravasation); (e) survive within the foreign microenvironment; and finally (f) re-initiate their proliferative machinery to establish macroscopic secondary tumors (colonization) (Fig. 1.2) (Harquail et al. 2012; Fidler 2003). Despite the clinical significance of metastasis for determining disease outcome in human cancers, our current understanding on how cancer cells actually migrate out of primary tumors, adapt to distant tissues and organs, and form a secondary tumor are still not completely understood (Gupta and Massague 2006).
MiRNAs have recently been more widely investigated due to their potential role as critical regulators of tumor metastasis in cancer development. The link between altered expression levels of miRNAs and cancer development and metastasis can be observed either through the loss of tumor suppressor miRNAs or the over-expression of oncogenic miRNAs in different cancer cells. Some miRNAs involved in metastasis are summarized in Table 1.2, most of which will be discussed in more detail in the sections below.
3.1 Pro-metastatic miRNAs
Multiple lines of evidence highlight the contribution of certain miRNAs to promoting tumor metastasis. MiR-10b is the first miRNA identified to positively regulate the metastatic potential of human cancer cell. Ma et al (2007) showed that miR-10b over-expression endowed otherwise non-metastatic breast cancer cells with the capacity to acquire invasive and metastatic behavior. MiR-10b is able to induce migration and invasion capacities in breast cancer cells through direct targeting of homeobox D10 (HOXD10), a receptor of genes involved in cell migration and extracellular matrix remodeling. Notably, systemic treatment of breast tumor-bearing mice with miR-10b antagomirs decreased the metastatic tumor burden, providing promising evidence that antagomirs can be efficiently delivered to rapidly growing tumor cells in vivo, preventing metastasis (Ma et al. 2010). To identify miRNAs that have the capacity to promote metastasis, Huang et al. (2008) set up a genetic screen involving over-expression of approximately 450 miRNAs in non-metastatic, human breast tumor cell line. They found that miR-373 and miR-520c (both belonging to a miRNA family that shares similar seed sequence) can induce tumor cell migration and invasion in vitro and in vivo, and that the migratory phenotype of certain cancer cell lines depends on endogenous miR-373 expression. They proposed that suppression of cell migration by an anti-miR-373 oligonucleotide may be a potential strategy for developing efficient therapies against tumor metastasis. After that, two independent studies indicated that apart from the oncogenic role of miR-21 in tumorigenesis, this miRNA also plays a critical role in invasion and metastasis of human breast and colorectal carcinoma cells (Asangani et al. 2008; Zhu et al. 2008). These studies suggest that suppression of miR-21 might offer another promising therapeutic approach against advanced cancers (Table 1.2).
3.2 Anti-metastatic miRNAs
Multiple lines of evidence highlight the contribution of certain miRNAs to suppressing tumor metastasis. MiR-31 expression levels correlate inversely with metastasis in human breast cancer patients. By deploying gain- and loss-of-function strategies, Valastyan et al. (2009) demonstrated that miR-31 is capable of suppressing the metastatic potential of human breast tumor cells. They also successfully showed that miR-31 is involved during the multiple step metastatic process in vivo, including local invasion, extravasation or initial survival at a distant site, and metastatic colonization. MiR-126 and miR-335 have been identified as human breast cancer metastasis suppressor miRNAs that exert their unique effects on distinct steps of the invasion-metastasis cascade. By performing array-based miRNA profiling, Tavazoie et al. (2008) revealed that the expression of both miRNAs is lost in the majority of primary breast tumors with metastatic relapse, and the loss of expression of either miRNA is associated with poor distal metastasis-free survival. Importantly, in vivo experiments showed that miR-126 restoration reduced overall tumor growth and proliferation (at both primary site and distant organs), whereas miR-335 caused a significant reduction in cell motility and invasive capacity. The strong association of the loss of miR-335 and miR-126 expression with clinical metastatic relapse suggests the potential for the use of these miRNAs in prognostic assessment of breast cancer patients in addition to conventional clinical and pathological staging markers. Moreover, another study identified that miR-193b significantly inhibited the growth and dissemination of xenograft breast tumors in an immunodeficient mouse model. This study showed that the loss of miR-193b confers the metastatic colonization ability to the cells. As the loss of miR-193b expression is strongly correlated with metastasis, the use of this miRNA in addition to conventional clinical and pathological staging markers could be an attractive option for the prognostic stratification of patients with breast cancer (Li et al. 2009) (Table 1.2).
3.3 MiRNAs and Epithelial to Mesenchymal Transition
Epithelial to mesenchymal transition (EMT), in which polarized epithelial cells are converted into motile cells, plays an important role in tumor invasion and metastasis (Thiery 2002; Yang and Weinberg 2008; Togawa et al. 2011). The effect of miR-125b on metastatic activities of breast cancer cells was studied by Tang et al. They reported that miR-125b significantly up-regulates the expression of two EMT markers (i.e. vimentin and α-SMA expression) but another EMT marker (E-Cadherin) shows no significant change. Elevating vimentin and α-SMA expression results in a high metastasis potentiality and some mesenchymal cell characteristics in breast cancer cells (Tang et al. 2012). A large body of evidence indicates that the miR-200 family inhibits EMT and cancer cell migration by enhancing E-cadherin expression through direct targeting of the EMT-promoting transcription factors Zeb1 and Zeb2 (Korpal et al. 2008; Gregory et al. 2008; Park et al. 2008; Burk et al. 2008; Bracken et al. 2008). However, a study reported that over-expressing miR-200 in Murine breast cancer cell line 4TO7 enhances the ability of these cells to metastasize to lung and liver. This study reported that miR-200 expression leads to promote a mesenchymal to epithelial cell transition (MET) by suppressing Zeb2 expression. This finding contrasts with the EMT hypothesis of cancer metastasis that implies that the induction of epithelial characteristics would inhibit the formation of metastasis. This apparent contradiction could be explained on the basis that for some tumors, a reversion of the mesenchymal phenotype of malignant cells may facilitate tumor colonization at metastatic sites. This suggests that the epithelial nature of a tumor does not predict metastatic outcome. Moreover, these results imply that the cellular context of miRNA expression decisively controls the function of a miRNA (Dykxhoorn et al. 2009).
4 MicroRNAs and Tumor Angiogenesis
Angiogenesis is characterized by growth of new blood vessels from pre-existing vasculature in response to physiological or pathophysiological stimuli. This process, which involves proliferation, migration, and maturation of endothelial cells, plays an important role during tumor growth and metastasis (Urbich et al. 2008; Chung et al. 2010).
Evidence for the significance of miRNAs as regulators of angiogenesis comes from observations that Dicer is a critical component for embryonic angiogenesis. It has been shown that blood vessel formation/maintenance in Dicer-deficient mice embryos and their yolk sacs was severely compromised, suggesting a possible role for Dicer in angiogenesis through its function in the processing of miRNAs (Yang et al. 2005). Consistent with this observation, another studies showed that genetic silencing of Dicer in endothelial cells leads to down-regulation of several key positive regulators of the angiogenic phenotype and impairs tube formation activity in vitro and in vivo (Suarez et al. 2007; Kuehbacher et al. 2007). Mounting studies suggest that a number of angiogenesis-related miRNAs affect cancerous phenotype of malignant cells. MiRNAs can modulate angiogenesis by targeting positive or negative regulators in angiogenic signaling pathways (Hong et al. 2013; Landskroner-Eiger et al. 2013). Some miRNAs involved in tumor angiogenesis are summarized in Table 1.3, most of which will be discussed in more detail in the sections below.
4.1 Pro-angiogenic miRNAs
Up-regulation of pro-angiogenic growth factor receptors (such as platelet-derived growth factor receptor, “PDGFR” and vascular endothelial growth factor receptor, “VEGFR”) on endothelial cells is a common feature of angiogenesis (Batchelor et al. 2007; Shih and Holland 2006). Wurdinger et al. (2008) showed that glioma- or growth factor-mediated induction of miR-296 in endothelial cells leads to increased levels of pro-angiogenic growth factor receptors VEGFR2 and PDGFR-β. Possible role of miR-296 in promoting angiogenesis in tumor was further supported when inhibition of miR-296 with antagomirs reduced angiogenesis in tumor xenografts in vivo.
Some other miRNAs, such as miR-378 and miR-17-92 cluster, have been also implicated in tumor angiogenesis. MiR-378 functions as an oncogene by enhancing tumor cell survival, blood vessel expansion, and tumor growth by targeting two tumor suppressors, SuFu (suppressor of fused) and Fus-1 (Lee et al. 2007a). The miR-17-92 cluster not only augments angiogenesis in endothelial cells during normal development (Suarez et al. 2008), but also its upregulation in cancer cells can serve to promote angiogenesis during tumor growth in a xenograft model (Dews et al. 2006). Importantly, this angiogenic effect is exerted through down-regulation of anti-angiogenic thrombospondin-1 (TSP-1) and related proteins, such as connective tissue growth factor (CTGF) (Dews et al. 2006).
One study showed that many miRNAs derived from tumor cells are packaged into microvesicles and then directly delivered to their microenvironment. These tumor-secreted microvesicles are then capable of interacting with proximal endothelial cells to transport miRNAs in endothelial cells. Among these miRNAs, it was shown that tumor-secreted miR-9 promotes endothelial cell migration and tumor angiogenesis by activating JAK-STAT pathway, one of the major oncogenic signaling pathways activated in a variety of human malignancies. Importantly, administration of miR-9 antagomiRs (anti-miR-9) or JAK inhibitors impaired microvesicles-induced cell migration in vitro and decreased tumor burden in vivo. Taken together, these observations support a novel intercellular communication in which tumor-secreted miRNAs function as pro-angiogenic mediators during tumorigenesis (Zhuang et al. 2012) (Table 1.3).
4.2 Anti-angiogenic miRNAs
The miR-34 family of miRNAs (miR-34a, b and c) as direct, conserved p53 target genes presumably induces apoptosis, cell cycle arrest and senescence (Bommer et al. 2007; Chang et al. 2007). MiR-34a functions as a tumor suppressor that is frequently down-regulated in various tumor types. Kumar et al. (2012) demonstrated that miR-34a expression is significantly down-regulated in head and neck squamous cell carcinoma tumors and cell lines. Ectopic expression of miR-34a reduced head and neck tumor cell proliferation, colony formation and migration and also significantly inhibited tumor growth and tumor angiogenesis in a SCID mouse xenograft model. This in vivo tumor growth study revealed that miR-34a inhibits tumor angiogenesis by down-regulating VEGF, a key angiogenic factor.
Siragam et al. (2012) defined a regulatory role for miR-98 in tumor angiogenesis and invasion using a highly aggressive breast cancer model in vitro. They showed that miR-98 inhibits tumor angiogenesis and invasion by repressing activin receptor-like kinase-4 (ALK4) and matrix metalloproteinase-11 (MMP11) expression.
Another study showed that transient induction of miR-125b inhibits in vitro tube formation of endothelial cells through suppression of vascular endothelial (VE)-cadherin. Importantly, induction of miR-125b induced non-functional blood vessels, resulting in inhibition of tumor growth. It seems that prolonged over-expression of miR-125b could be an option in cancer therapy by causing collapse of the lumen of endothelial cells (Muramatsu et al. 2013) (Table 1.3).
5 MicroRNA Profiling by High-Throughput Technologies
Considering the fact that current cancer detection tests have their own limitations, the use of miRNAs as promising biomarkers for diagnosis and prognosis of cancer has aroused intense research interests. Additionally, distinctive pattern of miRNA expression also serves as markers of important histopathologic features such as tumor stage, proliferative capacity and vascular invasion (Lynam-Lennon et al. 2009).
Many expression profiling studies of miRNA genes have been performed on different types of cancer. However, the results of analyses of the same type of cancer by different groups are not always consistent. The disparity in these results might attribute to the different platforms for miRNA profiling in each case and the use of different sample storage methods (Calin and Croce 2009).
Currently, the most widely used methods for miRNA profiling are based on sequencing, microarray, and real-time quantitative PCR. Microarray platforms have been used for miRNA profiling, but suffer from background and cross-hybridization problems and are generally restricted to identifying the relative abundance of previously discovered miRNAs (Calin et al. 2004a; Chen et al. 2009). Sequencing-based applications for identifying and profiling miRNAs have been hindered by laborious cloning techniques and the expense of capillary DNA sequencing (Pfeffer et al. 2005; Cummins et al. 2006). High-throughput sequencing-based approaches to generate miRNA profiles, hugely enabled by next-generation technologies, provide several advantages over probe-based methodologies, including the ability to discover novel miRNAs and the potential to detect variations in the mature miRNA length and miRNA editing (Morozova and Marra 2008). Next-generation sequencing technologies are able to identify low abundance miRNAs or those exhibiting modest expression differences among samples, which may not be detected by hybridization-based methods. Real-time quantitative PCR, another highly sensitive technique for miRNA quantification, is capable of distinguishing mature and precursor miRNA, and produces fewer false-positives and reduced bias when compared with microarray or sequencing approaches (Chen et al. 2009; Fuller et al. 2009; Petriv et al. 2010). Real-time PCR may be used to validate the expression of miRNAs discovered during high throughput arrays and study the expression of individual miRNAs. This method provides several important advantages for miRNA profiling studies including low cost, superior detection of low-abundance species and high throughput (Schmittgen et al. 2008). The emergence of novel high-throughput technologies will allow more sensitive and efficient miRNA detection in patient samples, and identification of novel miRNAs. However, standardization of these novel methods is necessary to overcome the variability observed when different miRNA-expression detection platforms are used.
6 Potential Use of MicroRNAs in Cancer Therapy
Dysregulation of miRNA has been widely observed in different types of human cancers (Table 1.4), and there is mounting evidence demonstrating their important roles during cancer development and progression. Uncovering the possible mechanisms underlying the importance of miRNAs in the pathogenesis of human cancers may lead to the development of miRNA-based therapeutic strategies or diagnostic/prognostic biomarkers.
Since cancer cells often have a distinctive expression pattern of oncogenic and tumor suppressive miRNAs (Babashah et al. 2012; Babashah and Soleimani 2011; Calin and Croce 2006), approaches that manipulate miRNA expression levels, either alone or in combination with currently used therapies, may prove to be therapeutically beneficial. Sequence-specific knockdown of oncogenic miRNAs by chemically engineered oligonucleotides termed “antagomirs” or locked nucleic acid (LNA)-modified oligonucleotides is a plausible therapeutic approach for inhibiting expression levels of oncogenic miRNAs in cancer (Orom et al. 2006; Krutzfeldt et al. 2005). In contrast, elevating the expression level of tumor suppressive miRNAs that could be achieved by viral or liposomal delivery of mimic miRNAs represents a potential therapeutic strategy against cancer (Calin and Croce 2006; Meng et al. 2006). However, many concerns need to be addressed before consideration of conducting miRNA-based therapy including dosage, safety, specificity, stability, efficacy, and problems of delivery to the target (Chen et al. 2010; Cho 2010b; Tong and Nemunaitis 2008; Wu et al. 2007).
7 Conclusions and Perspectives
-
As miRNAs can regulate various target genes, precise attribution of their functions on gene expression is very complicated. However, the critical involvement of miRNAs in many aspect of cancer biology is irrefutable.
-
Although miRNAs are postulated to function as either oncogenes or tumor suppressors in human cancers, further studies establishing such roles for miRNAs using in vivo experimental models are needed to elucidate precise mechanisms of miRNAs functions in cancer.
-
MiRNA expression profiling of human cancers has identified diagnostic and prognostic signatures. Additionally, miRNA signatures could be used for cancer classification and prediction of therapeutic efficacy.
-
The association of miRNA dysregulation with oncogenesis demonstrates the feasibility of manipulating miRNA levels as a potential strategy for therapeutic purposes.
-
Given the potential involvement of candidate miRNAs in the pathogenesis of human cancers, it seems that pharmacological modulation of miRNA expression will have a brilliant future and become a promising option in cancer therapy.
References
Akao Y, Nakagawa Y, Kitade Y, Kinoshita T, Naoe T (2007) Downregulation of microRNAs-143 and −145 in B-cell malignancies. Cancer Sci 98(12):1914–1920
Ambros V (2004) The functions of animal microRNAs. Nature 431(7006):350–355
Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S et al (2008) MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27(15):2128–2136
Babashah S, Soleimani M (2011) The oncogenic and tumour suppressive roles of microRNAs in cancer and apoptosis. Eur J Cancer 47(8):1127–1137
Babashah S, Sadeghizadeh M, Tavirani MR, Farivar S, Soleimani M (2012) Aberrant microRNA expression and its implications in the pathogenesis of leukemias. Cell Oncol (Dordr) 35(5):317–334
Bandi N, Zbinden S, Gugger M, Arnold M, Kocher V, Hasan L et al (2009) miR-15a and miR-16 are implicated in cell cycle regulation in a Rb-dependent manner and are frequently deleted or down-regulated in non-small cell lung cancer. Cancer Res 69(13):5553–5559
Bandyopadhyay S, Mitra R, Maulik U, Zhang MQ (2010) Development of the human cancer microRNA network. Silence 1(1):6
Barbarotto E, Schmittgen TD, Calin GA (2008) MicroRNAs and cancer: profile, profile, profile. Int J Cancer 122(5):969–977
Batchelor TT, Sorensen AG, di Tomaso E, Zhang WT, Duda DG, Cohen KS et al (2007) AZD2171, a pan-VEGF receptor tyrosine kinase inhibitor, normalizes tumor vasculature and alleviates edema in glioblastoma patients. Cancer Cell 11(1):83–95
Bhat-Nakshatri P, Wang G, Collins NR, Thomson MJ, Geistlinger TR, Carroll JS et al (2009) Estradiol-regulated microRNAs control estradiol response in breast cancer cells. Nucleic Acids Res 37(14):4850–4861
Bhaumik D, Scott GK, Schokrpur S, Patil CK, Campisi J, Benz CC (2008) Expression of microRNA-146 suppresses NF-kappaB activity with reduction of metastatic potential in breast cancer cells. Oncogene 27(42):5643–5647
Bohnsack MT, Czaplinski K, Gorlich D (2004) Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10(2):185–191
Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE et al (2007) p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 17(15):1298–1307
Bonci D, Coppola V, Musumeci M, Addario A, Giuffrida R, Memeo L et al (2008) The miR-15a-miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat Med 14(11):1271–1277
Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF et al (2008) A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res 68(19):7846–7854
Burk U, Schubert J, Wellner U, Schmalhofer O, Vincan E, Spaderna S et al (2008) A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep 9(6):582–589
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6(11):857–866
Calin GA, Croce CM (2009) Chronic lymphocytic leukemia: interplay between noncoding RNAs and protein-coding genes. Blood 114(23):4761–4770
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al (2002) Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 99(24):15524–15529
Calin GA, Liu CG, Sevignani C, Ferracin M, Felli N, Dumitru CD et al (2004a) MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc Natl Acad Sci U S A 101(32):11755–11760
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al (2004b) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A 101(9):2999–3004
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE et al (2005) A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353(17):1793–1801
Camps C, Buffa FM, Colella S, Moore J, Sotiriou C, Sheldon H et al (2008) hsa-miR-210 Is induced by hypoxia and is an independent prognostic factor in breast cancer. Clin Cancer Res 14(5):1340–1348
Chan JA, Krichevsky AM, Kosik KS (2005) MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65(14):6029–6033
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH et al (2007) Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 26(5):745–752
Chang SS, Jiang WW, Smith I, Poeta LM, Begum S, Glazer C et al (2008) MicroRNA alterations in head and neck squamous cell carcinoma. Int J Cancer 123(12):2791–2797
Chen Y, Gorski DH (2008) Regulation of angiogenesis through a microRNA (miR-130a) that down-regulates antiangiogenic homeobox genes GAX and HOXA5. Blood 111(3):1217–1226
Chen CZ, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303(5654):83–86
Chen Y, Gelfond JA, McManus LM, Shireman PK (2009) Reproducibility of quantitative RT-PCR array in miRNA expression profiling and comparison with microarray analysis. BMC Genomics 10:407
Chen J, Odenike O, Rowley JD (2010) Leukaemogenesis: more than mutant genes. Nat Rev Cancer 10(1):23–36
Cheng AM, Byrom MW, Shelton J, Ford LP (2005) Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res 33(4):1290–1297
Childs G, Fazzari M, Kung G, Kawachi N, Brandwein-Gensler M, McLemore M et al (2009) Low-level expression of microRNAs let-7d and miR-205 are prognostic markers of head and neck squamous cell carcinoma. Am J Pathol 174(3):736–745
Cho WC (2010a) MicroRNAs in cancer – from research to therapy. Biochim Biophys Acta 1805(2):209–217
Cho WC (2010b) MicroRNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol 42(8):1273–1281
Chung AS, Lee J, Ferrara N (2010) Targeting the tumour vasculature: insights from physiological angiogenesis. Nat Rev Cancer 10(7):505–514
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M et al (2005) miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci U S A 102(39):13944–13949
Connolly E, Melegari M, Landgraf P, Tchaikovskaya T, Tennant BC, Slagle BL et al (2008) Elevated expression of the miR-17-92 polycistron and miR-21 in hepadnavirus-associated hepatocellular carcinoma contributes to the malignant phenotype. Am J Pathol 173(3):856–864
Corney DC, Flesken-Nikitin A, Godwin AK, Wang W, Nikitin AY (2007) MicroRNA-34b and MicroRNA-34c are targets of p53 and cooperate in control of cell proliferation and adhesion-independent growth. Cancer Res 67(18):8433–8438
Croce CM (2009) Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet 10(10):704–714
Cullen BR (2004) Derivation and function of small interfering RNAs and microRNAs. Virus Res 102(1):3–9
Cummins JM, He Y, Leary RJ, Pagliarini R, Diaz LA Jr, Sjoblom T et al (2006) The colorectal microRNAome. Proc Natl Acad Sci U S A 103(10):3687–3692
Datta J, Kutay H, Nasser MW, Nuovo GJ, Wang B, Majumder S et al (2008) Methylation mediated silencing of MicroRNA-1 gene and its role in hepatocellular carcinogenesis. Cancer Res 68(13):5049–5058
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ (2004) Processing of primary microRNAs by the Microprocessor complex. Nature 432(7014):231–235
Dews M, Homayouni A, Yu D, Murphy D, Sevignani C, Wentzel E et al (2006) Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat Genet 38(9):1060–1065
Dutta KK, Zhong Y, Liu YT, Yamada T, Akatsuka S, Hu Q et al (2007) Association of microRNA-34a overexpression with proliferation is cell type-dependent. Cancer Sci 98(12):1845–1852
Dykxhoorn DM, Wu Y, Xie H, Yu F, Lal A, Petrocca F et al (2009) miR-200 enhances mouse breast cancer cell colonization to form distant metastases. PLoS One 4(9):e7181
Egle A, Harris AW, Bouillet P, Cory S (2004) Bim is a suppressor of Myc-induced mouse B cell leukemia. Proc Natl Acad Sci U S A 101(16):6164–6169
Esquela-Kerscher A, Slack FJ (2006) Oncomirs – microRNAs with a role in cancer. Nat Rev Cancer 6(4):259–269
Esquela-Kerscher A, Trang P, Wiggins JF, Patrawala L, Cheng A, Ford L et al (2008) The let-7 microRNA reduces tumor growth in mouse models of lung cancer. Cell Cycle 7(6):759–764
Fabbri M, Garzon R, Cimmino A, Liu Z, Zanesi N, Callegari E et al (2007) MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci U S A 104(40):15805–15810
Fabian MR, Sonenberg N (2012) The mechanics of miRNA-mediated gene silencing: a look under the hood of miRISC. Nat Struct Mol Biol 19(6):586–593
Faller M, Matsunaga M, Yin S, Loo JA, Guo F (2007) Heme is involved in microRNA processing. Nat Struct Mol Biol 14(1):23–29
Farazi TA, Horlings HM, Ten Hoeve JJ, Mihailovic A, Halfwerk H, Morozov P et al (2011) MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res 71(13):4443–4453
Felicetti F, Errico MC, Bottero L, Segnalini P, Stoppacciaro A, Biffoni M et al (2008) The promyelocytic leukemia zinc finger-microRNA-221/-222 pathway controls melanoma progression through multiple oncogenic mechanisms. Cancer Res 68(8):2745–2754
Felli N, Fontana L, Pelosi E, Botta R, Bonci D, Facchiano F et al (2005) MicroRNAs 221 and 222 inhibit normal erythropoiesis and erythroleukemic cell growth via kit receptor down-modulation. Proc Natl Acad Sci U S A 102(50):18081–18086
Fidler IJ (2003) The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer 3(6):453–458
Fish JE, Santoro MM, Morton SU, Yu S, Yeh RF, Wythe JD et al (2008) miR-126 regulates angiogenic signaling and vascular integrity. Dev Cell 15(2):272–284
Friedman RC, Farh KK, Burge CB, Bartel DP (2009) Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19(1):92–105
Fuller CW, Middendorf LR, Benner SA, Church GM, Harris T, Huang X et al (2009) The challenges of sequencing by synthesis. Nat Biotechnol 27(11):1013–1023
Galardi S, Mercatelli N, Giorda E, Massalini S, Frajese GV, Ciafre SA et al (2007) miR-221 and miR-222 expression affects the proliferation potential of human prostate carcinoma cell lines by targeting p27Kip1. J Biol Chem 282(32):23716–23724
Gartel AL, Kandel ES (2008) miRNAs: little known mediators of oncogenesis. Semin Cancer Biol 18(2):103–110
Gillies JK, Lorimer IA (2007) Regulation of p27Kip1 by miRNA 221/222 in glioblastoma. Cell Cycle 6(16):2005–2009
Gottardo F, Liu CG, Ferracin M, Calin GA, Fassan M, Bassi P et al (2007) Micro-RNA profiling in kidney and bladder cancers. Urol Oncol 25(5):387–392
Grady WM, Parkin RK, Mitchell PS, Lee JH, Kim YH, Tsuchiya KD et al (2008) Epigenetic silencing of the intronic microRNA hsa-miR-342 and its host gene EVL in colorectal cancer. Oncogene 27(27):3880–3888
Gramantieri L, Ferracin M, Fornari F, Veronese A, Sabbioni S, Liu CG et al (2007) Cyclin G1 is a target of miR-122a, a microRNA frequently down-regulated in human hepatocellular carcinoma. Cancer Res 67(13):6092–6099
Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N et al (2004) The Microprocessor complex mediates the genesis of microRNAs. Nature 432(7014):235–240
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G et al (2008) The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10(5):593–601
Gupta GP, Massague J (2006) Cancer metastasis: building a framework. Cell 127(4):679–695
Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK et al (2006) Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125(5):887–901
Harquail J, Benzina S, Robichaud GA (2012) MicroRNAs and breast cancer malignancy: an overview of miRNA-regulated cancer processes leading to metastasis. Cancer Biomark 11(6):269–280
Harris TA, Yamakuchi M, Ferlito M, Mendell JT, Lowenstein CJ (2008) MicroRNA-126 regulates endothelial expression of vascular cell adhesion molecule 1. Proc Natl Acad Sci U S A 105(5):1516–1521
Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S et al (2005) A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 65(21):9628–9632
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S et al (2005) A microRNA polycistron as a potential human oncogene. Nature 435(7043):828–833
Hong L, Han Y, Zhou Y, Nita A (2013) Angiogenesis-related microRNAs in colon cancer. Expert Opin Biol Ther 13(1):77–84
Hu X, Macdonald DM, Huettner PC, Feng Z, El Naqa IM, Schwarz JK et al (2009) A miR-200 microRNA cluster as prognostic marker in advanced ovarian cancer. Gynecol Oncol 114(3):457–464
Huang Q, Gumireddy K, Schrier M, le Sage C, Nagel R, Nair S et al (2008) The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat Cell Biol 10(2):202–210
Huang L, Dai T, Lin X, Zhao X, Chen X, Wang C et al (2012) MicroRNA-224 targets RKIP to control cell invasion and expression of metastasis genes in human breast cancer cells. Biochem Biophys Res Commun 425(2):127–133
Ibrahim SA, Yip GW, Stock C, Pan JW, Neubauer C, Poeter M et al (2012) Targeting of syndecan-1 by microRNA miR-10b promotes breast cancer cell motility and invasiveness via a Rho-GTPase- and E-cadherin-dependent mechanism. Int J Cancer 131(6):E884–E896
Ichimi T, Enokida H, Okuno Y, Kunimoto R, Chiyomaru T, Kawamoto K et al (2009) Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int J Cancer 125(2):345–352
Ikeda S, Kong SW, Lu J, Bisping E, Zhang H, Allen PD et al (2007) Altered microRNA expression in human heart disease. Physiol Genomics 31(3):367–373
Iorio MV, Visone R, Di Leva G, Donati V, Petrocca F, Casalini P et al (2007) MicroRNA signatures in human ovarian cancer. Cancer Res 67(18):8699–8707
Jiang J, Gusev Y, Aderca I, Mettler TA, Nagorney DM, Brackett DJ et al (2008) Association of microRNA expression in hepatocellular carcinomas with hepatitis infection, cirrhosis, and patient survival. Clin Cancer Res 14(2):419–427
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A et al (2005) RAS is regulated by the let-7 microRNA family. Cell 120(5):635–647
Kluiver J, Poppema S, de Jong D, Blokzijl T, Harms G, Jacobs S et al (2005) BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J Pathol 207(2):243–249
Kondo N, Toyama T, Sugiura H, Fujii Y, Yamashita H (2008) miR-206 expression is down-regulated in estrogen receptor alpha-positive human breast cancer. Cancer Res 68(13):5004–5008
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS et al (2008) MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol 28(22):6773–6784
Korpal M, Lee ES, Hu G, Kang Y (2008) The miR-200 family inhibits epithelial-mesenchymal transition and cancer cell migration by direct targeting of E-cadherin transcriptional repressors ZEB1 and ZEB2. J Biol Chem 283(22):14910–14914
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ et al (2005) Combinatorial microRNA target predictions. Nat Genet 37(5):495–500
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M et al (2005) Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438(7068):685–689
Kuehbacher A, Urbich C, Zeiher AM, Dimmeler S (2007) Role of Dicer and Drosha for endothelial microRNA expression and angiogenesis. Circ Res 101(1):59–68
Kumar MS, Erkeland SJ, Pester RE, Chen CY, Ebert MS, Sharp PA et al (2008) Suppression of non-small cell lung tumor development by the let-7 microRNA family. Proc Natl Acad Sci U S A 105(10):3903–3908
Kumar B, Yadav A, Lang J, Teknos TN, Kumar P (2012) Dysregulation of microRNA-34a expression in head and neck squamous cell carcinoma promotes tumor growth and tumor angiogenesis. PLoS One 7(5):e37601
Lal A, Navarro F, Maher CA, Maliszewski LE, Yan N, O’Day E et al (2009) miR-24 inhibits cell proliferation by targeting E2F2, MYC, and other cell-cycle genes via binding to “seedless” 3′UTR microRNA recognition elements. Mol Cell 35(5):610–625
Landskroner-Eiger S, Moneke I, Sessa WC (2013) miRNAs as modulators of angiogenesis. Cold Spring Harb Perspect Med 3(2):a006643
Le XF, Almeida MI, Mao W, Spizzo R, Rossi S, Nicoloso MS et al (2012) Modulation of microRNA-194 and cell migration by HER2-targeting trastuzumab in breast cancer. PLoS One 7(7):e41170
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J et al (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425(6956):415–419
Lee DY, Deng Z, Wang CH, Yang BB (2007a) MicroRNA-378 promotes cell survival, tumor growth, and angiogenesis by targeting SuFu and Fus-1 expression. Proc Natl Acad Sci U S A 104(51):20350–20355
Lee EJ, Gusev Y, Jiang J, Nuovo GJ, Lerner MR, Frankel WL et al (2007b) Expression profiling identifies microRNA signature in pancreatic cancer. Int J Cancer 120(5):1046–1054
Lee I, Ajay SS, Yook JI, Kim HS, Hong SH, Kim NH et al (2009) New class of microRNA targets containing simultaneous 5′-UTR and 3′-UTR interaction sites. Genome Res 19(7):1175–1183
Lehmann U, Hasemeier B, Christgen M, Muller M, Romermann D, Langer F et al (2008) Epigenetic inactivation of microRNA gene hsa-mir-9-1 in human breast cancer. J Pathol 214(1):17–24
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115(7):787–798
Li XF, Yan PJ, Shao ZM (2009) Downregulation of miR-193b contributes to enhance urokinase-type plasminogen activator (uPA) expression and tumor progression and invasion in human breast cancer. Oncogene 28(44):3937–3948
Liu X, Fortin K, Mourelatos Z (2008) MicroRNAs: biogenesis and molecular functions. Brain Pathol 18(1):113–121
Liu X, Jiang L, Wang A, Yu J, Shi F, Zhou X (2009) MicroRNA-138 suppresses invasion and promotes apoptosis in head and neck squamous cell carcinoma cell lines. Cancer Lett 286(2):217–222
Liu Q, Lv GD, Qin X, Gen YH, Zheng ST, Liu T et al (2012) Role of microRNA let-7 and effect to HMGA2 in esophageal squamous cell carcinoma. Mol Biol Rep 39(2):1239–1246
Lopez JI, Camenisch TD, Stevens MV, Sands BJ, McDonald J, Schroeder JA (2005) CD44 attenuates metastatic invasion during breast cancer progression. Cancer Res 65(15):6755–6763
Lowery AJ, Miller N, McNeill RE, Kerin MJ (2008) MicroRNAs as prognostic indicators and therapeutic targets: potential effect on breast cancer management. Clin Cancer Res 14(2):360–365
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al (2005) MicroRNA expression profiles classify human cancers. Nature 435(7043):834–838
Lujambio A, Calin GA, Villanueva A, Ropero S, Sanchez-Cespedes M, Blanco D et al (2008) A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci U S A 105(36):13556–13561
Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U (2004) Nuclear export of microRNA precursors. Science 303(5654):95–98
Lynam-Lennon N, Maher SG, Reynolds JV (2009) The roles of microRNA in cancer and apoptosis. Biol Rev Camb Philos Soc 84(1):55–71
Ma L, Teruya-Feldstein J, Weinberg RA (2007) Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449(7163):682–688
Ma L, Reinhardt F, Pan E, Soutschek J, Bhat B, Marcusson EG et al (2010) Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat Biotechnol 28(4):341–347
Marton S, Garcia MR, Robello C, Persson H, Trajtenberg F, Pritsch O et al (2008) Small RNAs analysis in CLL reveals a deregulation of miRNA expression and novel miRNA candidates of putative relevance in CLL pathogenesis. Leukemia 22(2):330–338
Mendell JT (2008) miRiad roles for the miR-17-92 cluster in development and disease. Cell 133(2):217–222
Meng F, Henson R, Lang M, Wehbe H, Maheshwari S, Mendell JT et al (2006) Involvement of human micro-RNA in growth and response to chemotherapy in human cholangiocarcinoma cell lines. Gastroenterology 130(7):2113–2129
Mertens-Talcott SU, Chintharlapalli S, Li X, Safe S (2007) The oncogenic microRNA-27a targets genes that regulate specificity protein transcription factors and the G2-M checkpoint in MDA-MB-231 breast cancer cells. Cancer Res 67(22):11001–11011
Miller TE, Ghoshal K, Ramaswamy B, Roy S, Datta J, Shapiro CL et al (2008) MicroRNA-221/222 confers tamoxifen resistance in breast cancer by targeting p27Kip1. J Biol Chem 283(44):29897–29903
Morozova O, Marra MA (2008) Applications of next-generation sequencing technologies in functional genomics. Genomics 92(5):255–264
Muller DW, Bosserhoff AK (2008) Integrin beta 3 expression is regulated by let-7a miRNA in malignant melanoma. Oncogene 27(52):6698–6706
Muramatsu F, Kidoya H, Naito H, Sakimoto S, Takakura N (2013) microRNA-125b inhibits tube formation of blood vessels through translational suppression of VE-cadherin. Oncogene 32(4):414–421
Nam EJ, Yoon H, Kim SW, Kim H, Kim YT, Kim JH et al (2008) MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 14(9):2690–2695
Nasser MW, Datta J, Nuovo G, Kutay H, Motiwala T, Majumder S et al (2008) Down-regulation of micro-RNA-1 (miR-1) in lung cancer. Suppression of tumorigenic property of lung cancer cells and their sensitization to doxorubicin-induced apoptosis by miR-1. J Biol Chem 283(48):33394–33405
Nicoloso MS, Spizzo R, Shimizu M, Rossi S, Calin GA (2009) MicroRNAs–the micro steering wheel of tumour metastases. Nat Rev Cancer 9(4):293–302
Nikiforova MN, Tseng GC, Steward D, Diorio D, Nikiforov YE (2008) MicroRNA expression profiling of thyroid tumors: biological significance and diagnostic utility. J Clin Endocrinol Metab 93(5):1600–1608
Noonan EJ, Place RF, Pookot D, Basak S, Whitson JM, Hirata H et al (2009) miR-449a targets HDAC-1 and induces growth arrest in prostate cancer. Oncogene 28(14):1714–1724
O’Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT (2005) c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435(7043):839–843
Orom UA, Kauppinen S, Lund AH (2006) LNA-modified oligonucleotides mediate specific inhibition of microRNA function. Gene 372:137–141
Paraskevopoulou MD, Georgakilas G, Kostoulas N, Vlachos IS, Vergoulis T, Reczko M et al (2013) DIANA-microT web server v5.0: service integration into miRNA functional analysis workflows. Nucleic Acids Res 41(Web Server issue):W169–W173
Park SM, Gaur AB, Lengyel E, Peter ME (2008) The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev 22(7):894–907
Pasquinelli AE (2012) MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nat Rev Genet 13(4):271–282
Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI, Maller B et al (2000) Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408(6808):86–89
Pekarsky Y, Santanam U, Cimmino A, Palamarchuk A, Efanov A, Maximov V et al (2006) Tcl1 expression in chronic lymphocytic leukemia is regulated by miR-29 and miR-181. Cancer Res 66(24):11590–11593
Petriv OI, Kuchenbauer F, Delaney AD, Lecault V, White A, Kent D et al (2010) Comprehensive microRNA expression profiling of the hematopoietic hierarchy. Proc Natl Acad Sci U S A 107(35):15443–15448
Pfeffer S, Sewer A, Lagos-Quintana M, Sheridan R, Sander C, Grasser FA et al (2005) Identification of microRNAs of the herpesvirus family. Nat Methods 2(4):269–276
Poliseno L, Tuccoli A, Mariani L, Evangelista M, Citti L, Woods K et al (2006) MicroRNAs modulate the angiogenic properties of HUVECs. Blood 108(9):3068–3071
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE et al (2000) The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403(6772):901–906
Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA et al (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9(6):435–443
Schmittgen TD, Lee EJ, Jiang J, Sarkar A, Yang L, Elton TS et al (2008) Real-time PCR quantification of precursor and mature microRNA. Methods 44(1):31–38
Scott GK, Goga A, Bhaumik D, Berger CE, Sullivan CS, Benz CC (2007) Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b. J Biol Chem 282(2):1479–1486
Shih AH, Holland EC (2006) Platelet-derived growth factor (PDGF) and glial tumorigenesis. Cancer Lett 232(2):139–147
Siragam V, Rutnam ZJ, Yang W, Fang L, Luo L, Yang X et al (2012) MicroRNA miR-98 inhibits tumor angiogenesis and invasion by targeting activin receptor-like kinase-4 and matrix metalloproteinase-11. Oncotarget 3(11):1370–1385
Slaby O, Svoboda M, Fabian P, Smerdova T, Knoflickova D, Bednarikova M et al (2007) Altered expression of miR-21, miR-31, miR-143 and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology 72(5–6):397–402
Spizzo R, Nicoloso MS, Croce CM, Calin GA (2009) SnapShot: microRNAs in cancer. Cell 137(3):586–586 e581
Stamatopoulos B, Meuleman N, Haibe-Kains B, Saussoy P, Van Den Neste E, Michaux L et al (2009) microRNA-29c and microRNA-223 down-regulation has in vivo significance in chronic lymphocytic leukemia and improves disease risk stratification. Blood 113(21):5237–5245
Steeg PS (2003) Metastasis suppressors alter the signal transduction of cancer cells. Nat Rev Cancer 3(1):55–63
Su H, Yang JR, Xu T, Huang J, Xu L, Yuan Y et al (2009) MicroRNA-101, down-regulated in hepatocellular carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer Res 69(3):1135–1142
Suarez Y, Fernandez-Hernando C, Pober JS, Sessa WC (2007) Dicer dependent microRNAs regulate gene expression and functions in human endothelial cells. Circ Res 100(8):1164–1173
Suarez Y, Fernandez-Hernando C, Yu J, Gerber SA, Harrison KD, Pober JS et al (2008) Dicer-dependent endothelial microRNAs are necessary for postnatal angiogenesis. Proc Natl Acad Sci U S A 105(37):14082–14087
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H et al (2004) Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64(11):3753–3756
Tang F, Zhang R, He Y, Zou M, Guo L, Xi T (2012) MicroRNA-125b induces metastasis by targeting STARD13 in MCF-7 and MDA-MB-231 breast cancer cells. PLoS One 7(5):e35435
Tavazoie SF, Alarcon C, Oskarsson T, Padua D, Wang Q, Bos PD et al (2008) Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451(7175):147–152
Tazawa H, Tsuchiya N, Izumiya M, Nakagama H (2007) Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc Natl Acad Sci U S A 104(39):15472–15477
Thiery JP (2002) Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer 2(6):442–454
Togawa H, Nakanishi K, Mukaiyama H, Hama T, Shima Y, Sako M et al (2011) Epithelial-to-mesenchymal transition in cyst lining epithelial cells in an orthologous PCK rat model of autosomal-recessive polycystic kidney disease. Am J Physiol Renal Physiol 300(2):F511–F520
Tong AW, Nemunaitis J (2008) Modulation of miRNA activity in human cancer: a new paradigm for cancer gene therapy? Cancer Gene Ther 15(6):341–355
Urbich C, Kuehbacher A, Dimmeler S (2008) Role of microRNAs in vascular diseases, inflammation, and angiogenesis. Cardiovasc Res 79(4):581–588
Valastyan S, Reinhardt F, Benaich N, Calogrias D, Szasz AM, Wang ZC et al (2009) A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137(6):1032–1046
Varambally S, Cao Q, Mani RS, Shankar S, Wang X, Ateeq B et al (2008) Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science 322(5908):1695–1699
Visone R, Pallante P, Vecchione A, Cirombella R, Ferracin M, Ferraro A et al (2007) Specific microRNAs are downregulated in human thyroid anaplastic carcinomas. Oncogene 26(54):7590–7595
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F et al (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A 103(7):2257–2261
Wang S, Aurora AB, Johnson BA, Qi X, McAnally J, Hill JA et al (2008a) The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev Cell 15(2):261–271
Wang Y, Lee AT, Ma JZ, Wang J, Ren J, Yang Y et al (2008b) Profiling microRNA expression in hepatocellular carcinoma reveals microRNA-224 up-regulation and apoptosis inhibitor-5 as a microRNA-224-specific target. J Biol Chem 283(19):13205–13215
Webster RJ, Giles KM, Price KJ, Zhang PM, Mattick JS, Leedman PJ (2009) Regulation of epidermal growth factor receptor signaling in human cancer cells by microRNA-7. J Biol Chem 284(9):5731–5741
Woods K, Thomson JM, Hammond SM (2007) Direct regulation of an oncogenic micro-RNA cluster by E2F transcription factors. J Biol Chem 282(4):2130–2134
Wu H, Xu H, Miraglia LJ, Crooke ST (2000) Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J Biol Chem 275(47):36957–36965
Wu W, Sun M, Zou GM, Chen J (2007) MicroRNA and cancer: current status and prospective. Int J Cancer 120(5):953–960
Wurdinger T, Tannous BA, Saydam O, Skog J, Grau S, Soutschek J et al (2008) miR-296 regulates growth factor receptor overexpression in angiogenic endothelial cells. Cancer Cell 14(5):382–393
Xiao C, Srinivasan L, Calado DP, Patterson HC, Zhang B, Wang J et al (2008) Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nat Immunol 9(4):405–414
Yan D, Zhou X, Chen X, Hu DN, Dong XD, Wang J et al (2009) MicroRNA-34a inhibits uveal melanoma cell proliferation and migration through downregulation of c-Met. Invest Ophthalmol Vis Sci 50(4):1559–1565
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M et al (2006) Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9(3):189–198
Yang J, Weinberg RA (2008) Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev Cell 14(6):818–829
Yang WJ, Yang DD, Na S, Sandusky GE, Zhang Q, Zhao G (2005) Dicer is required for embryonic angiogenesis during mouse development. J Biol Chem 280(10):9330–9335
Yang H, Kong W, He L, Zhao JJ, O’Donnell JD, Wang J et al (2008) MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res 68(2):425–433
Yi R, Qin Y, Macara IG, Cullen BR (2003) Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev 17(24):3011–3016
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C et al (2007) let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131(6):1109–1123
Yu Z, Wang C, Wang M, Li Z, Casimiro MC, Liu M et al (2008) A cyclin D1/microRNA 17/20 regulatory feedback loop in control of breast cancer cell proliferation. J Cell Biol 182(3):509–517
Zhang X, Liu S, Hu T, He Y, Sun S (2009) Up-regulated microRNA-143 transcribed by nuclear factor kappa B enhances hepatocarcinoma metastasis by repressing fibronectin expression. Hepatology 50(2):490–499
Zhao H, Wang D, Du W, Gu D, Yang R (2010) MicroRNA and leukemia: tiny molecule, great function. Crit Rev Oncol Hematol 74(3):149–155
Zhong X, Coukos G, Zhang L (2012) miRNAs in human cancer. Methods Mol Biol 822:295–306
Zhu S, Si ML, Wu H, Mo YY (2007) MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J Biol Chem 282(19):14328–14336
Zhu S, Wu H, Wu F, Nie D, Sheng S, Mo YY (2008) MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res 18(3):350–359
Zhuang G, Wu X, Jiang Z, Kasman I, Yao J, Guan Y et al (2012) Tumour-secreted miR-9 promotes endothelial cell migration and angiogenesis by activating the JAK-STAT pathway. EMBO J 31(17):3513–3523
Acknowledgments
I would like to thank all authors responsible for the insights that I attempted to summarize. I apologize to the colleagues whose work was not cited because of space considerations.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Babashah, S. (2014). MicroRNAs and Cancer: An Overview. In: Babashah, S. (eds) MicroRNAs: Key Regulators of Oncogenesis. Springer, Cham. https://doi.org/10.1007/978-3-319-03725-7_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-03725-7_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-03724-0
Online ISBN: 978-3-319-03725-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)