Abstract
Functional brain dynamics is supported by parallel and overlapping functional network modes that are associated with specific neural circuits. Decomposing these network modes from fMRI data and finding their temporal characteristics is challenging due to their time-varying nature and the non-linearity of the functional dynamics. Dynamic Mode Decomposition (DMD) algorithms have been quite popular for solving this decomposition problem in recent years. In this work, we apply GraphDMD—an extension of the DMD for network data—to extract the dynamic network modes and their temporal characteristics from the fMRI time series in an interpretable manner. GraphDMD, however, regards the underlying system as a linear dynamical system that is sub-optimal for extracting the network modes from non-linear functional data. In this work, we develop a generalized version of the GraphDMD algorithm—DeepGraphDMD—applicable to arbitrary non-linear graph dynamical systems. DeepGraphDMD is an autoencoder-based deep learning model that learns Koopman eigenfunctions for graph data and embeds the non-linear graph dynamics into a latent linear space. We show the effectiveness of our method in both simulated data and the HCP resting-state fMRI data. In the HCP data, DeepGraphDMD provides novel insights into cognitive brain functions by discovering two major network modes related to fluid and crystallized intelligence.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Component Analysis
- Dynamic Mode Decomposition
- Koopman Operator
- Non-linear Graph Dynamical Systems
- Functional Network Dynamics
- State-space models
- Graph Representation Learning
1 Introduction
The human brain has evolved to support a set of complementary and temporally varying brain network organizations enabling parallel and higher-order information processing [16, 20, 24]. Decoupling these networks from a non-linear mixture of signals (such as functional MRI) and extracting their temporal characteristics in an interpretable manner has been a long-standing challenge in the neuroscience community.
Conventional mode/component decomposition methods such as Principal Component Analysis (PCA) or Independent Component Analysis (ICA) assume the modes to be static [7, 15, 26] and thus sub-optimal for the functional networks generated by time-varying modes. Dynamic Mode Decomposition (DMD) can be treated as a dynamic extension of such component analysis methods since it allows its modes to oscillate over time with a fixed frequency [18]. This assumption is appropriate for the human brain as the functional brain organizations are supported by oscillatory network modes [2, 8, 11, 23, 27]. An extension of DMD for network data called GraphDMD [4] preserves the graph structure of the networks during the decomposition. In our work, we extend GraphDMD to a sequence of sliding window based dynamic functional connectivity (dNFC) networks to extract independent and oscillatory functional network modes.
Under the hood, GraphDMD regards the network sequence as a linear dynamical system (LDS) where a linear operator shifts the current network state one time-point in the future. The LDS assumption, however, is not optimal for modeling functional brain networks that exhibit complex non-linearity such as rapid synchronization and desynchronization as well as transient events [6]. Articles [3, 13] propose switching linear dynamical system (SLDS) to tackle the nonlinearity of spatiotemporal data by a piecewise linear approximation. While these models offer interpretability, their shallow architecture limits their generalizability to arbitrary nonlinear systems. On the other hand, the methods in [5, 10, 21] model the non-linearity with a deep neural network. While these models have more representation capabilities compared to SLDS, the latent states are not interpretable. More importantly, all of these methods consider the node-level dynamics instead of the network dynamics.
Here, we propose a novel Deep Graph Dynamic Mode Decomposition (DeepGraphDMD) algorithm that applies to arbitrary non-linear network dynamics while maintaining interpretability in the latent space. Our method uses Koopman operator theory to lift a non-linear dynamical system into a linear space through a set of Koopman eigenfunctions (Fig. 1a). There has been a growing line of work that learns these measurement functions using deep autoencoder architectures [14, 22]. Training these autoencoders for network data, however, has two unique challenges – 1. preserving the edge identity in the latent space so that the network modes are interpretable, 2. enforcing linearity in the latent space for the high dimensional network data. In DeepGraphDMD, we tackle the first challenge by indirectly computing the network embeddings by a novel node embedding scheme. For the second challenge, we introduce a sparse Koopman operator to reduce the complexity of the learning problem. We evaluate the effectiveness of our novel method in both simulated data and resting-state fMRI (rs-fMRI) data from Human Connectome Project.
2 Methodology
Let’s assume \(X \in \mathbb {R}^{n \times t}\) is a matrix containing the BOLD (blood-oxygen-level-dependent) signal of n brain regions (ROIs) in its rows at t time frames sampled at every \(k\varDelta t\) time points, where \(\varDelta t\) is the temporal resolution. To compute the dynamic connectivity matrix at time point \(k \varDelta t\), a snapshot \(X_k = X_{:,k:k+s}\) is taken in a sliding window of s time frames. A correlation matrix \(G_k \in \mathbb {R}^{n \times n}\) is then computed from \(X_k\) by taking the pearson correlation between the rows of \(X_k\), i.e., \(G_k^{ij} = \texttt {pearson}(x_k^i, x_k^j)\) where \(x_k^i\), \(x_k^j\) are the \(i^{th}\) and \(j^{th}\) row of \(X_k\) respectively. This yields a sequence of graphs \(G = [G_1, G_2, \cdots , G_{t-s+1}]\). Let’s also assume that \(g_k \in \mathcal {R}^{n^2}\) is a vectorized version of \(G_k\), i.e. \(g_k = vec(G_k)\) and \(g \in \mathcal {R}^{n^2 \times (t-s+1)}\) is a matrix containing \(g_k\) in its columns. The goal is to decouple the overlapping spatiotemporal modes from the network sequence G using – 1. Graph Dynamic Mode Decomposition algorithm, and 2. a novel Deep Learning-based Graph Dynamic Mode Decomposition algorithm.
2.1 Graph Dynamic Mode Decomposition
GraphDMD [4] assumes that \(g_k\) follows an LDS:
where \(A \in \mathbb {R}^{n^2 \times n^2}\) is a linear operator that shifts the current state \(g_k\) to the state at the next time frame \(g_{k+1}\). To extract the low dimensional global network dynamics, GraphDMD projects A into a lower dimensional space \(\hat{A}\) using tensor-train decomposition, applies eigendecomposition of \(\hat{A}\), and projects the eigenvectors back to the original space which we refer to as dynamic modes (DMs). GraphDMD uses tensor-train decomposition to maintain the network structure of \(g_k\) and thus, the DMs from GraphDMD can be reshaped into \(n\times n\) adjacency matrix forms. Let’s assume these DMs are \(\varPhi _1, \varPhi _2, \cdots , \varPhi _r\) where \(\varPhi _p \in \mathbb {C}^{n \times n}\) and the corresponding eigenvalues are \(\lambda _1, \lambda _2, \cdots , \lambda _r\) where \(\lambda _p \in \mathbb {C}\) (Fig. 1b). Here, r is the total number of DMs. \(\varPhi _p\) corresponds to the coherent spatial mode and \(\lambda _p\) defines its temporal characteristics (growth/decay rate and frequencies). We can see this by unrolling Eq. 1 in time:
where \(\lambda _p = a_p \exp (\omega _p \varDelta t)\), \(\varPhi ^\dag \) is the conjugate transpose of \(\varPhi \), \(b_p = vec(\varPhi _p^\dag ) g_1\) is the projection of the initial value onto the DMD modes, \(a_p = ||\lambda _p||\) is the growth/decay rate and \(\omega _p = {\text {Im}}(\ln \lambda _p) / \varDelta t\) is the angular frequency of \(\varPhi _p\).
2.2 Adaptation of Graph-DMD for Nonlinear Graph Dynamics
Since the dynamics of the functional networks are often non-linear, the linearity assumption of Eq. 1 is sub-optimal. In this regard, we resort to Koopman operator theory to transform the non-linear system into an LDS using a set of Koopman eigenfunctions \(\psi \), i.e., \(\psi (g_{k+1}) = A \psi (g_k)\) [9]. We learn \(\psi \) using a deep autoencoder-based architecture—DeepGraphDMD—where the encoder and the decoder are trained to approximate \(\psi \) and \(\psi ^{-1}\), respectively. We enforce \(\psi (g_k)\) to follow an LDS by applying Latent Koopman Invariant Loss [22] in the form:
where  are two matrices with columns stacked with \(\psi (g_k)\) and \(Y^\dag \) is the right inverse of Y. After training, we reshape \(\psi (g_k)\) into a \(n\times n\) network \(\psi (G_k)\) and generate the latent network sequence \(\psi (G_1), \cdots , \psi (G_{t-s+1})\). We then apply GraphDMD (described in Sect. 2.1) on this latent and linearized network sequence to extract the DMs \(\varPhi _p\) and their corresponding \(\lambda _p\).
are two matrices with columns stacked with \(\psi (g_k)\) and \(Y^\dag \) is the right inverse of Y. After training, we reshape \(\psi (g_k)\) into a \(n\times n\) network \(\psi (G_k)\) and generate the latent network sequence \(\psi (G_1), \cdots , \psi (G_{t-s+1})\). We then apply GraphDMD (described in Sect. 2.1) on this latent and linearized network sequence to extract the DMs \(\varPhi _p\) and their corresponding \(\lambda _p\).
However, there are two unique challenges of learning network embeddings using the DeepGraphDMD model: 1. the edge identity and, thus, the interpretability will be lost in the latent space if we directly embed \(g_k\) using \(\psi \), and 2. \(Y^\dag \) doesn’t exist, and thus \(\mathcal {L}_{lkis}\) can’t be computed because Y is low rank with the number of rows \(\frac{n (n - 1)}{2} >>\) the number of columns \(t - s + 1\).
To solve the first problem, instead of learning \(\psi (g_k)\) directly, we embed the BOLD signal \(x_k^i\) of each ROI independently using the encoder to learn the latent embeddings \(z_k^i\) (Fig. 1a). We then compute the pearson correlation between the latent embeddings of the ROIs to get the Koopman eigenfunctions of \(g_k\) i.e., \(\psi (g_{k}^{ij}) = \texttt {pearson}(z_k^i, z_k^j)\). The weights of the encoder and decoder are shared across the ROIs.
The second problem arises because the Koopman operator A regresses the value of an edge at the next time-point as a linear combination of all the other edges at the current time-point, i.e., \(g^{ij}_{k+1} = \sum _{p,q=1} ^ N w_{pq} g^{pq}_{k}\). This results in \(\mathcal {O}(n^2)\) covariates with \(t - s + 1 << \mathcal {O}(n^2)\) samples making the regression ill-posed. We propose a sparse Koopman operator where each edge \(g^{ij}_k\) is regressed using only the edges that share a common end-point with it, i.e., \(g^{ij}_{k+1} = \sum _{p=1, p \ne i,j}^n w_{ip}g^{ip}_{k} + \sum _{q=1, q \ne i,j}^n w_{qj}g^{qj}_k + w_{ij}g^{ij}_k\) (Supplementary Fig. 1). Since there are only \(\mathcal {O}(n)\) such edges, it solves the ill-posedness of the regression.
Other than \(\mathcal {L}_{lkis}\), we also train the autoencoder with a reconstruction loss \(\mathcal {L}_{recon}\) which is the mean-squared error (MSE) between \(x_k^i\) and the reconstructed output from the decoder \(\hat{x}_k^i\). Moreover, a regularizer \(\mathcal {L}_{reg}\) in the form of an MSE loss between \(g_k\) and the latent \(\psi (g_k)\) is also added. The final loss is the following:
where \(\alpha \) and \(\beta \) are hyper-parameters. We choose \(\alpha \), \(\beta \), and other hyperparameters using grid search on the validation set. The network architecture and the values of the hyper-parameters of DeepGraphDMD training are shown in Supplementary Fig. 1. The code is available inFootnote 1.
2.3 Window-Based GraphDMD
We apply GraphDMD in a short window of size 64 time frames with a step size of 4 time frames instead of the whole sequence G because, in real-world fMRI data, both the frequency and the structure of the DMs can change over time. We then combine the DMs across different sliding windows using the following post-processing steps:
Post-processing of the DMs: We first group the DMs within the frequency bins: 0–0.01 Hz, 0.01–0.04 Hz, 0.04–0.08 Hz, 0.08–0.12 Hz, and 0.12–0.16 Hz. We then cluster the DMs within each frequency bin using a clustering algorithm and select the cluster centroids as the representative DMs (except for the first bin where we average the DMs). We chose the optimal clustering algorithm to be Spherical KMeans [1] (among Gaussian Mixture Model, KMeans, Spherical KMeans, DBSCAN, and, KMedoids) and the optimal number of clusters to be 3 for every frequency bin based on silhouette analysis [17] (Supplementary Fig. 2). We use this frequency binning technique to allow for slight variations of \(\omega \) of a DM over the scanning session. To align these representative DMs across subjects, we apply another round of spherical clustering on the DMs from all subjects and align them based on their cluster memberships.
3 Experiments
3.1 Dataset
We use rs-fMRI for 840 subjects from the HCP Dense Connectome datasetFootnote 2 [25]. Each fMRI image was acquired with a temporal resolution (\(\varDelta t\)) of 0.72 s and a 2 mm isotropic spatial resolution using a 3T Siemens Skyra scanner. Individual subjects underwent four rs-fMRI runs of 14.4 min each (1200 frames per run). Group-ICA using FSL’s MELODIC tool [7] was applied to parcellate the brain into 50 functional regions (ROIs). To find the correlation between cognition with the rs-fMRI data, we select two behavioral measures related to fluid intelligence: CogFluidComp, PMAT24_A_CR and one measure related to crystallized intelligence: ReadEng, and, the normalized scores of the fluid and crystallized cognition measures: CogTotalComp. We regress out the confounding factors: age, gender, and head motion from these behavioral measures using ordinary least squares [12].
3.2 Baseline Methods
We compare GraphDMD and DeepGraphDMD against three decomposition methods: Principal Component Analysis (PCA), Independent Component Analysis (ICA), and standard Dynamic Mode Decomposition (DMD) [18]. We use the sklearn decomposition library for PCAFootnote 3 and ICAFootnote 4 and the pyDMDFootnote 5 library for standard DMD. We apply PCA and ICA on g, and DMD directly on the bold signal X instead of g (for reasons described in Sect. 4.2). We choose the number of components (n_components) to be three for these decomposition methods, (Results for other n_components values are shown in Supplementary Table 1). The components are aligned across subjects using spherical clustering similar to the GraphDMD modes (Sect. 2.3). We also compare with static functional connectivity (sFC), which is the pairwise pearson correlation between brain regions across all time frames.
3.3 Simulation Study
We generate a sequence of dynamic adjacency matrices G using Eq. 2 from three time-varying modes \(\varPhi _1, \varPhi _2, \varPhi _3\) with corresponding frequencies \(\omega _1 \sim \mathcal {N}(0.1, 0.05)\), \(\omega _2 \sim \mathcal {N}(1, 0.1)\), \(\omega _3 \sim \mathcal {N}(2.5, 0.1)\) (Hz). Each \(\varPhi _p\) is a \(32 \times 32\) block diagonal matrices with block sizes 16, 8, and 4. We choose \(a_1=1.01\), \(a_2=0.9\), \(a_3=1.05\) and \(b_1=b_2=b_3=1\). We simulate the process for \(k=1, \cdots , 29\) time-points yielding a sequence of 30 matrices of shape \(32 \times 32\). We repeat the process ten times with different \(\omega _1, \omega _2, \omega _3\) and generate ten matrix sequences. We apply PCA, ICA, and GraphDMD (Sect. 2.1) on G to extract three components and compare them against the ground truth modes using pearson correlation.
3.4 Application of GraphDMD and DeepGraphDMD in HCP Data
Comparison of DMs with sFC: The ground truth DMs are unknown for the HCP dataset; however, we can use the sFC as a substitute for the ground truth DM with \(\omega = 0\) (static DM). sFC offsets the DMs with \(\omega > 0\) as they have both positive and negative cycles, and thus only retain the static DM. For comparison, we compute the pearson correlation between the DM within the frequency bin 0–0.01 Hz and sFC for both GraphDMD and DeepGraphDMD. For PCA and ICA, we take the maximum value of the correlation between sFC and the (PCA or ICA) components.
Regression Analysis of Behavioral Measures from HCP: In this experiment, we regress the behavioral measures with the DMs within each frequency bin (Sect. 2.3) using Elastic-net. As an input to the Elastic-net, we take the real part of the upper diagonal part of the DM and flatten it into a vector. We then train the Elastic-net in two ways—1. single-band: where we train the Elastic-net independently with the DMs in each frequency bin, and 2. multi-band: we concatenate two DMs in the frequency bins: 0–0.01 Hz and 0.08–0.12 Hz and regress using the concatenated vector. For evaluation, we compute the correlation coefficient r between the predicted and the true values of the measures.
4 Results
4.1 Simulation Study
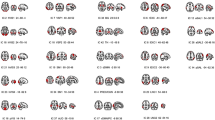
In Fig. 2a, we show the results after applying PCA, ICA, and, GraphDMD on the simulated data described in Sect. 3.3. Since the DMs in this data are oscillating, the data generated from this process are more likely to be overlapping compared to when the modes are static. As a result, methods that assume static modes, such as PCA and ICA, struggle to decouple the DMs and discover modes in overlapping high-density regions. For example, in Mode 2 of ICA, we can see the remnants of Mode 3 in the blue boxes and Mode 1 (negative cycle) in the orange boxes. We observe similar scenarios in the Mode 3 of ICA and Mode 1, Mode 2, and Mode 3 of PCA (the red boxes). On the other hand, the DMs from GraphDMD have fewer remnants from other modes and closely resemble the ground truth. To empirically compare, the mean (± std) pearson correlation for PCA, ICA, and, GraphDMD are \(0.81(\pm 0.04)\), \(0.88(\pm 0.03)\), and \(0.98(\pm 0.01)\).
(a) Ground truth network modes from simulated data (column 1) and extracted network modes from PCA (2nd column), ICA (3rd column), and, GraphDMD (4th column), (b) Circle plot of the average DMs with \(\omega \approx 0\), (c) \(\omega \in [0.08-0.12]\) from DeepGraphDMD organized based on common resting-state networks [19].
4.2 Application of GraphDMD and DeepGraphDMD in HCP Data
Comparison of DMs with sFC: The average pearson correlations with sFC across all the subjects are \(0.6 (\pm 0.09)\), \(0.6 (\pm 0.09)\), \(0.84 (\pm 0.09)\), and \(0.86 (\pm 0.05)\) for PCA, ICA, GraphDMD, and, DeepGraphDMD (Fig. 2b) respectively. This shows that the DMD-based methods can robustly decouple the static DM from time-varying DMs. In comparison, the corresponding PCA and ICA component has significantly lower correlation due to the overlap from the higher frequency components.
Regression Analysis of Behavioral Measures from HCP: We show the values of r across different methods in Table 1. We only show the results for two frequency bins 0–0.01 Hz and 0.08–0.12 Hz, as the DMs in the other bins are not significantly correlated (\(r < 0.2\)) with the behavioral measures (Supplementary Table 2). The table shows that multi-band training with the DMs from the DMD-based methods significantly improves the regression performance over the baseline methods. Compared to sFC, GraphDMD improves r by 22%, 6%, 0.7%, and, 3% for CogFluidComp, PMAT24_A_CR, ReadEng, CogTotalComp, respectively and DeepGraphDMD further improves the performance by 5%, 2.2%, 0.7%, and, 1.5%, respectively. Significant performance improvement for CogFluidComp can be explained by the DM within the bin 0.08–0.12 Hz. This DM provides additional information related to fluid intelligence (\(r = 0.227\) for GraphDMD) to which the sFC doesn’t have access. By considering non-linearity, DeepGraphDMD extracts more robust and less noisy DMs (Fig. 2b–c), and hence, it improves the regression performance by \(8\%\) compared to GraphDMD in this frequency bin. By contrast, the standard DMD algorithm yields unstable modes with \(a_p << 1\) when applied to the network sequence G. These modes have no correspondence across subjects and thus can’t be used for regression. We instead apply DMD on the BOLD signal X, but the DMD modes show little correlation with the behavioral measures. PCA and ICA perform significantly worse than the baseline sFC method for all behavioral measures.
Traditional dynamical functional connectivity analysis methods (such as sliding window-based techniques) consider a sequence of network states. However, our results show that these states can be further decomposed into more atomic network modes. The importance of decoupling these network modes from nonlinearly mixed fMRI signals using DeepGraphDMD has been shown in regressing behavioral measures from HCP data.
5 Conclusion
In this paper, we proposed a novel algorithm—DeepGraphDMD—to decouple spatiotemporal network modes in dynamic functional brain networks. Unlike other decomposition methods, DeepGraphDMD accounts for both the non-linear and the time-varying nature of the functional modes. As a result, these functional modes from DeepGraphDMD are more robust compared to their linear counterpart in GraphDMD and are shown to be correlated with fluid and crystallized intelligence measures.
Notes
- 1.
- 2.
- 3.
sklearn.decomposition.PCA.
- 4.
sklearn.decomposition.FastICA.
- 5.
References
Banerjee, A., Dhillon, I.S., Ghosh, J., Sra, S.: Clustering on the unit hypersphere using von Mises-Fisher distributions. J. Mach. Learn. Res. 6(46), 1345–1382 (2005). http://jmlr.org/papers/v6/banerjee05a.html
Casorso, J., Kong, X., Chi, W., Van De Ville, D., Yeo, B.T., Liégeois, R.: Dynamic mode decomposition of resting-state and task fMRI. NeuroImage 194, 42–54 (2019). https://doi.org/10.1016/j.neuroimage.2019.03.019. https://www.sciencedirect.com/science/article/pii/S1053811919301922
Fox, E., Sudderth, E., Jordan, M., Willsky, A.: Nonparametric Bayesian learning of switching linear dynamical systems. In: Koller, D., Schuurmans, D., Bengio, Y., Bottou, L. (eds.) Advances in Neural Information Processing Systems, vol. 21. Curran Associates, Inc. (2008). https://proceedings.neurips.cc/paper_files/paper/2008/file/950a4152c2b4aa3ad78bdd6b366cc179-Paper.pdf
Fujii, K., Takeishi, N., Hojo, M., Inaba, Y., Kawahara, Y.: Physically-interpretable classification of biological network dynamics for complex collective motions. Sci. Rep. 10 (2020). https://doi.org/10.1038/s41598-020-58064-w
Gao, Y., Archer, E.W., Paninski, L., Cunningham, J.P.: Linear dynamical neural population models through nonlinear embeddings. In: Lee, D., Sugiyama, M., Luxburg, U., Guyon, I., Garnett, R. (eds.) Advances in Neural Information Processing Systems, vol. 29. Curran Associates, Inc. (2016). https://proceedings.neurips.cc/paper_files/paper/2016/file/76dc611d6ebaafc66cc0879c71b5db5c-Paper.pdf
He, B.J.: Robust, transient neural dynamics during conscious perception. Trends Cogn. Sci. 22(7), 563–565 (2018). https://doi.org/10.1016/J.TICS.2018.04.005. https://pubmed.ncbi.nlm.nih.gov/29764721/
Hyvarinen, A.: Fast and robust fixed-point algorithms for independent component analysis. IEEE Trans. Neural Netw. 10(3), 626–634 (1999)
Ikeda, S., Kawano, K., Watanabe, S., Yamashita, O., Kawahara, Y.: Predicting behavior through dynamic modes in resting-state fMRI data. NeuroImage 247, 118801 (2022). https://doi.org/10.1016/j.neuroimage.2021.118801. https://www.sciencedirect.com/science/article/pii/S1053811921010727
Koopman, B.O.: Hamiltonian systems and transformation in Hilbert space. Proc. Natl. Acad. Sci. U.S.A. 17(5), 315 (1931). https://doi.org/10.1073/PNAS.17.5.315. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1076052/
Krishnan, R.G., Shalit, U., Sontag, D.A.: Structured inference networks for nonlinear state space models. In: AAAI Conference on Artificial Intelligence (2016)
Kunert-Graf, J.M., Eschenburg, K.M., Galas, D.J., Kutz, J.N., Rane, S.D., Brunton, B.W.: Extracting reproducible time-resolved resting state networks using dynamic mode decomposition. Front. Comput. Neurosci. 13 (2019). https://doi.org/10.3389/fncom.2019.00075. https://www.frontiersin.org/articles/10.3389/fncom.2019.00075
Liégeois, R., et al.: Resting brain dynamics at different timescales capture distinct aspects of human behavior. Nat. Commun. 10(1), 2317 (2019)
Linderman, S., Johnson, M., Miller, A., Adams, R., Blei, D., Paninski, L.: Bayesian learning and inference in recurrent switching linear dynamical systems. In: Singh, A., Zhu, J. (eds.) Proceedings of the 20th International Conference on Artificial Intelligence and Statistics. Proceedings of Machine Learning Research, vol. 54, pp. 914–922. PMLR (2017). https://proceedings.mlr.press/v54/linderman17a.html
Lusch, B., Nathan Kutz, J., Brunton, S.L.: Deep learning for universal linear embeddings of nonlinear dynamics. Nat. Commun. 9, 4950 (2018)
McKeown, M.J., et al.: Analysis of fMRI data by blind separation into independent spatial components. Hum. Brain Mapp. 6(3), 160–188 (1998)
Osada, T., et al.: Parallel cognitive processing streams in human prefrontal cortex: parsing areal-level brain network for response inhibition. Cell Rep. 36(12), 109732 (2021). https://doi.org/10.1016/j.celrep.2021.109732. https://www.sciencedirect.com/science/article/pii/S2211124721011815
Rousseeuw, P.J.: Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987). https://doi.org/10.1016/0377-0427(87)90125-7. https://www.sciencedirect.com/science/article/pii/0377042787901257
Schmid, P.J.: Dynamic mode decomposition of numerical and experimental data. J. Fluid Mech. 656, 5–28 (2010)
Seitzman, B.A., et al.: A set of functionally-defined brain regions with improved representation of the subcortex and cerebellum. NeuroImage 206, 116290 (2020). https://doi.org/10.1016/j.neuroimage.2019.116290. https://www.sciencedirect.com/science/article/pii/S105381191930881X
Sigman, M., Dehaene, S.: Brain mechanisms of serial and parallel processing during dual-task performance. J. Neurosci. 28(30), 7585–7598 (2008)
Sussillo, D., Jozefowicz, R., Abbott, L., Pandarinath, C.: LFADS - latent factor analysis via dynamical systems (2016)
Takeishi, N., Kawahara, Y., Yairi, T.: Learning Koopman invariant subspaces for dynamic mode decomposition. In: Proceedings of the 31st International Conference on Neural Information Processing Systems, NIPS 2017, pp. 1130–1140. Curran Associates Inc., Red Hook (2017)
Turja, M.A., Wu, G., Yang, D., Styner, M.A.: Learning the latent heat diffusion process through structural brain network from longitudinal \(\beta \)-amyloid data. In: Heinrich, M., Dou, Q., de Bruijne, M., Lellmann, J., Schläfer, A., Ernst, F. (eds.) Proceedings of the Fourth Conference on Medical Imaging with Deep Learning. Proceedings of Machine Learning Research, vol. 143, pp. 761–773. PMLR (2021). https://proceedings.mlr.press/v143/turja21a.html
Turja, M.A., Zsembik, L.C.P., Wu, G., Styner, M.: Constructing consistent longitudinal brain networks by group-wise graph learning. In: Shen, D., et al. (eds.) MICCAI 2019. LNCS, vol. 11766, pp. 654–662. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-32248-9_73
Van Essen, D.C., et al.: The human connectome project: a data acquisition perspective. Neuroimage 62(4), 2222–2231 (2012)
Viviani, R., Grön, G., Spitzer, M.: Functional principal component analysis of fMRI data. Hum. Brain Mapp. 24(2), 109–129 (2005)
Xiao, J., et al.: A spatio-temporal decomposition framework for dynamic functional connectivity in the human brain. NeuroImage 263, 119618 (2022). https://doi.org/10.1016/j.neuroimage.2022.119618. https://www.sciencedirect.com/science/article/pii/S1053811922007339
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Turja, M.A., Styner, M., Wu, G. (2023). DeepGraphDMD: Interpretable Spatio-Temporal Decomposition of Non-linear Functional Brain Network Dynamics. In: Greenspan, H., et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2023. MICCAI 2023. Lecture Notes in Computer Science, vol 14227. Springer, Cham. https://doi.org/10.1007/978-3-031-43993-3_35
Download citation
DOI: https://doi.org/10.1007/978-3-031-43993-3_35
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-43992-6
Online ISBN: 978-3-031-43993-3
eBook Packages: Computer ScienceComputer Science (R0)