Abstract
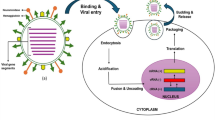
The frequent emergence of pathogenic viruses with pandemic potential has posed a significant threat to human health and economy, despite enormous advances in our understanding of infection mechanisms and devising countermeasures through developing various prophylactic and therapeutic strategies. The recent coronavirus disease (COVID-19) pandemic has re-emphasised the importance of rigorous research on virus infection mechanisms and highlighted the need for our preparedness for potential pandemics. Although viruses cannot self-replicate, they tap into host cell factors and processes for their entry, propagation and dissemination. Upon entering the host cells, viruses ingeniously utilise the innate biological functions of the host cell to replicate themselves and maintain their existence in the hosts. Influenza A virus (IAV), which has a negative-sense, single-stranded RNA as its genome, is no exception. IAVs are enveloped viruses with a lipid bilayer derived from the host cell membrane and have a surface covered with the spike glycoprotein haemagglutinin (HA) and neuraminidase (NA). Viral genome is surrounded by an M1 shell, forming a “capsid” in the virus particle. IAV particles use HA to recognise sialic acids on the cell surface of lung epithelial cells for their attachment. After attachment to the cell surface, IAV particles are endocytosed and sorted into the early endosomes. Subsequently, as the early endosomes mature into late endosomes, the endosomal lumen becomes acidified, and the low pH of the late endosomes induces conformational reaggangements in the HA to initiate fusion between the endosomal and viral membranes. Upon fusion, the viral capsid disintegrates and the viral ribonucleoprotein (vRNP) complexes containing the viral genome are released into the cytosol. The process of viral capsid disintegration is called “uncoating”. After successful uncoating, the vRNPs are imported into the nucleus by importin α/β (IMP α/β), where viral replication and transcription take place and the new vRNPs are assembled. Recently, we have biochemically elucidated the molecular mechanisms of the processes of viral capsid uncoating subsequent viral genome dissociation. In this chapter, we present the molecular details of the viral uncoating process.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Molecular Mechanisms of Influenza A Virus Entry
To establish successful infection in the target cell, virus particles need to deliver their genome and the accessory proteins for replication and transcription. Due to small viral particle sizes, viruses can only accommodate a limited number of protein-producing genes to help with their multiplication and spread. Therefore, they largely rely on the host proteins and processes to carry out essential functions. Virus particles produced from infected cells need to be stable enough to endure the challenges in the hostile extracellular environment for their survival. The stability of the virion is imparted by strong macromolecular interactions among the viral genome and accessory proteins during viral packaging, ensuring highly ordered condensation of the viral components into an extremely compact structure. However, when a highly stable virus in the extracellular environment finds its target cell, the macromolecular interactions that lead to its assembly need to be reversed to release its genome into the cell for replication and viral protein synthesis. Growing evidence suggests that viruses alone are unable to undergo self-disassembly in the host cell, therefore, they exploit various cellular factors to accomplish uncoating (Edinger et al. 2015; He et al. 2013; Huotari et al. 2012; Konig et al. 2010; Pohl et al. 2014; Su et al. 2013; Yanguez et al. 2018; Banerjee et al. 2014; Gschweitl et al. 2016).
Upon binding to the cell surface, IAV enters the target cell via either receptor-mediated endocytosis (Eierhoff et al. 2010; Fujioka et al. 2018) or macropinocytosis (de Vries et al. 2011). Receptor-mediated endocytosis is a process by which cells internalise specific proteins, metabolites, hormones, viruses, etc., by inwardly budding the plasma membrane and forming endocytic vesicles, whereas in macropinocytosis, cells engulf non-specific soluble materials in bulk into large uncoated vesicles called macropinosomes (Mercer et al. 2010). After entry into the host cells, IAV is sorted into the mildly acidic, Rab5-positive early endosomes (Chavrier et al. 1990; Gorvel et al. 1991). The activity of vacuolar ATPase (vATPase) present on the early endosomes causes a gradual drop in the luminal pH, and as the early endosomes mature into Rab7-positive late endosomes, the pH drops to 5.0 (Liang et al. 2016). The low pH in the late endosomes induces rapid structural rearrangements in the HA, exposing the fusion peptide which mediates the fusion between the viral membrane and the limiting membrane of the late endosome (Harrison 2008, 2015; Chlanda et al. 2016; Kanaseki et al. 1997; Lee 2010).
Although some viruses can enter the host cells by penetrating at the plasma membrane, the vast majority of enveloped viruses depend on the host cell endocytic pathways for entry and subsequent penetration at endosomes for transmission of their genome into the host cells (Yamauchi and Greber 2016; Mercer et al. 2010). Upon fusion at the endosomes, the uncoating process may differ from various viruses and cell types; however, the general processes of capsid disassembly and genome release are still not well understood. Previously it was believed that the viral fusion process is coupled to the capsid disassembly process, but recent studies indicate that host molecules play a critical role in capsid disassembly after fusion (Hao et al. 2013; Banerjee et al. 2014; Miyake et al. 2019; Kawaguchi et al. 2003). Uncoating is a highly regulated, step-wise process, which is dependent on host cell cues; dysregulation of the endocytic pathways or absence of critical host factors may lead to abortive uncoating.
After initial attachment to the plasma membrane, IAV, is internalised by either clathrin-mediated endocytosis or macropinocytosis, and the virus is sorted into early endosomes. The vATPase present in the early endosome causes an accumulation of protons (H+) in the lumen of the early endosome, as a result of which the luminal pH gradually drops. As the early endosome matures to late endosome while travelling toward the microtubule organising centre (MTOC) from the cell periphery, the pH drops to 5.0–5.5, which serves as the cue for HA conformational change to make the virus fusion-competent (Alvarado-Facundo et al. 2015; Grambas and Hay 1992). The low pH also opens up the viral M2 channel through which, H+ and potassium ions (K+) enter the viral core, loosening up the macromolecular interactions and priming the virus for uncoating (Stauffer et al. 2014). The pH-induced fusogenic HA brings the viral and endosomal membranes in close proximity, which is followed by the merging of the two membranes (Harrison 2015). Fusion of the virus in the late endosome is followed by uncoating, which ensures the escape of the virus from the endosome and the release of the vRNPs into the cytosol (Yamauchi and Helenius 2013; Yamauchi and Greber 2016; Staring et al. 2018; Yamauchi 2020). For example, IAV uncoating requires free ubiquitin chains in the virus and the host histone deacetylase 6 (HDAC6) (Banerjee et al. 2014).
In influenza virions, the vRNPs are attached to the viral envelope by nucleoprotein (NP) and M1 interactions (Martin and Helenius 1991). M1 also interacts with the cytoplasmic tails of HA and NA (Sempere Borau and Stertz 2021). The influx of H+ and K+ triggers conformational rearrangements in M1, leading to the disruption in the M1-NP interactions. Although priming of the virus in the late endosome is a prerequisite for uncoating, it is not sufficient for efficient capsid disassembly; the virus needs assistance from the host to complete the process of uncoating to release the vRNPs into the cytosol. One of the host proteins found to play a critical role in IAV uncoating is HDAC6, which deacetylases tubulin, Hsp90, cortactin and DDX3X (Zhang et al. 2006; Zhang et al. 2008; Hubbert et al. 2002; Kovacs et al. 2005; Zhang et al. 2007; Saito et al. 2019). HDAC6 plays a role in the transport of ubiquitinated protein aggregates to the aggresome for clearance via recruitment of the motor protein dynein (Fig. 14.1) (Hao et al. 2013; Kawaguchi et al. 2003; Banerjee et al. 2014). HDAC6 has two catalytic domains whose functions have been reported to involve tubulin deacetylation and the regulation of liquid phase separation of target proteins (Miyake et al. 2016; Saito et al. 2019). On the other hand, the ZnF domain of HDAC6 at the C-terminus recognises and strongly binds to the—LRGG sequence at the free C-terminus of the ubiquitin chain (Hao et al. 2013; Banerjee et al. 2014). The ubiquitin-binding activity of HDAC6 has been shown to be important for IAV uncoating. Although IAV could successfully internalise and fuse at the late endosome in the HDAC6-knockout (KO) mouse embryonic fibroblasts (MEFs), it was unable to undergo uncoating, and the virus particles remained trapped in the endosomal compartments. A similar phenotype was observed in the lung alveolar cell line (A549) deficient in HDAC6. The block in uncoating resulted in the inhibition of viral replication. Further, it was found that uncoating was independent of the mutations abrogating the enzymatic functions of HDAC6 but required the ubiquitin-binding activity; a point mutation in the ubiquitin-binding ZnF motif suppressed uncoating to the level observed in the HDAC6 KO cells. Biochemical investigations and super-resolution microscopy of purified IAVs revealed the presence of host-derived free ubiquitin chains within the virus particles. After confirming the presence of free ubiquitin chains within virions, the localisation of the ubiquitin chains was examined; if ubiquitin molecules were present on the viral surface, they should have been degraded by protease K in the absence of detergent (Triton). But no degradation of the ubiquitin molecules was observed upon protease K treatment. On the other hand, viral particles wrapped in the lipid bilayers showed membrane breakdown in the presence of Triton, and the ubiquitins present inside the particles were degraded by proteases. This suggests that ubiquitin molecules are present beneath the viral membrane and inside the core (Fig. 14.1). An in vitro assay with purified HDAC6 and IAV revealed an interaction of the viral ubiquitin with the ZnF domain of HDAC6. HDAC6 is a factor that responds to various cellular stresses and is known to transport untranslated mRNAs to stress granules (SGs). Misfolded and ubiquitinated proteins that have not moved to microtubule organising centres (MTOCs) along microtubules form aggregates corresponding to membraneless organelles called aggresomes. HDAC6 has been reported to interact with dynein motor proteins that move on microtubules in the retrograde direction (Kawaguchi et al. 2003), and HDAC6 mutants lacking the dynein binding motif also showed reduced viral infection. In addition to dynein, HDAC6 was found to interact with the actin-based motor protein myosin 10 (MYO10). We found that after fusion at the late endosome, the free ubiquitin chains present in the viral particles are detected by HDAC6, which, after binding to the ubiquitin molecules, recruit dynein and MYO10 to the fusion site. The activity of the motor proteins generates a pulling force on the softened virus capsid, which further helps in the dissociation of the capsid and release of the vRNPs into the cytosol. This suggests that IAV, with its packaged ubiquitin chains, make clever use of the HDAC6-dependent ubiquitin-sensing pathways lined to aggresome processing for its entry and establishment of a successful infection (Rudnicka and Yamauchi 2016). Biochemical fractionation of endosomal fractions of IAV-infected cells by ultracentrifugation revealed the presence of HDAC6 in one fraction containing viral component M1 and a late endosome marker Rab7 in the soluble endosomal fraction (Fig. 14.2; see fraction (7)). As a fraction containing Rab7 is also found in the pellet fraction, the endosomes that are undergoing uncoating may be precipitated in complex with various other proteins (Fig. 14.2). A recent study on Zika virus has shown how a similar HDAC6-dependent uncoating mechanism is exploited by the virus for uncoating. Designed ankyrin repeat proteins (DARPins) that bind to the ZnF motif of HDAC6 disrupt ubiquitin binding to HDAC6 and prevent Zika virus infection by suppressing viral uncoating (Wang et al. 2022).
Schematic of influenza A virus entry. Viral particles are taken up into the cell by endocytosis and become acidified in the endosomes as they mature (Priming). Within matured endosomes, HA undergoes structural conformation change and fuses to the endosomal membrane. Then, host histone deacetylase HDAC6 is recruited by the ubiquitin chains derived from the virus. The ubiquitin-binding domain of HDAC6 tightly binds to unanchored ubiquitin chains. By protein quality control mechanisms, unfold or misfold proteins that are no longer required are ubiquitinated for its degradation and transport to the aggresome by binding to HDAC6. As a stress response, HDAC6 bound to ubiquitin chains in the virus interacts with microtubules and actin fibres to promote uncoating response. After uncoating, the viral nucleoprotein complexes (vRNPs) are released into the cytoplasm. Host factor TNPO1 pulls off the M1 protein on the vRNPs bundle, and the eight bundles of vRNPs are separated into each vRNP. Dissociated vRNPs bind to IMPα/β and migrate into the nucleus for replication and transcription
Biochemical analysis of the endosomal fraction of virus-infected cells. (a) After collection of the virus-infected cells, the supernatant of the cell lysate was fractionated by Optiprep-density gradient centrifugation. NPTs; Nuclear pellet fractions. (b) Analysis of each fraction after density gradient centrifugation by Western blotting. Fraction (2) up to fraction (7) is positive for the late endosomal marker Rab7. The M1 matrix protein, used as a marker for IAV, is also co-localised in the endosomal fraction (right panel). Fraction (7) is considered to be the endosomal fraction from which HDAC6 is recruited immediately after uncoating
In addition to HDAC6, several other host factors have been found to promote IAV uncoating. Larson and colleagues reported that the epidermal growth factor receptor pathway substrate 8 (EPS8) physically interacts with IAV NP and facilitates viral uncoating at a post-fusion step. IAV uncoating and subsequent infection processes were significantly compromised in the EPS8-deficient cells, and complementing EPS8 in the cells restored infection (Larson et al. 2019). IAV uncoating is also blocked in cells with endosomal maturation defect. Houtari and colleagues found that Cullin-3, which serves as a scaffolding subunit in a large family of E3 ubiquitin ligases, plays a critical role in endosome maturation. Upon Cullin-3 depletion, although IAV can enter the cell and the HA can assume an acid-induced conformation in the acidic lumen of the late endosome, the virus fails to uncoat due to impaired endosomal maturation (Huotari et al. 2012). Another study found that the lymphocyte antigen 6 family member E (LY6E) promotes IAV uncoating; ectopic expression of LY6E increases capsid disassembly, suggesting the supporting role of the protein in uncoating at a post-fusion step (Mar et al. 2018). A genome-wide pooled shRNA screen identified an E3 ubiquitin ligase ITCH, which participates in the release of IAV from late endosomes. In the early stage of IAV infection, ITCH is activated by phosphorylation and is recruited to endosomes. In ITCH-depleted cells, the amount of mono- and poly-ubiquitinated M1 was reduced and the M1 co-immunoprecipitated with ITCH. Together, the results suggested that ITCH ubiquitinates M1 in the endosomes and facilitates IAV uncoating after fusion (Su et al. 2013). It is clear from the above studies that IAV uncoating is not a spontaneous event that follows fusion at the late endosome, but requires the participation of the host molecules. Additional uncoating factors could be involved, and future studies may illuminate new molecules that promote IAV uncoating.
After uncoating, the IAV genome is transferred into the nucleus, where the genome is transcribed and replicated. The IAV genome is divided into eight vRNPs segments, which encode 11 structural and non-structural proteins. These eight vRNPs are linked to each other by RNA-RNA interactions or protein-mediated interactions (Hutchinson et al. 2010; Noda et al. 2012; Dadonaite et al. 2019; Noda 2021). When the vRNPs are released into the cytosol following uncoating, they are bundled together and, therefore, cannot travel into the nucleus due to size limitation for the passage of the vRNPs through the nuclear pores. To identify the host factors promoting vRNP nuclear import, we conducted an RNAi screen targeting 70 potential genes involved in nucleocytoplasmic transport and identified transportin 1 (TNPO1) as one of the top hits (Miyake et al. 2019). RNAi-mediated knockdown of TNPO1 significantly arrested vRNP nuclear import. In the TNPO1-deficient cells, both viral capsid uncoating and vRNP nuclear import were impeded, and strong NP and M1 signals were observed in the cytoplasm. We reasoned that reduced uncoating and the block in vRNP nuclear import was possibly due to clustered vRNPs that needed to be debundled into individual segments in order to pass through the nuclear pores. Close examination of the amino acid sequence of the M1 protein revealed the presence of a sequence similar to the existing Proline-Tyrosine nuclear localisation signal (PY-NLS) at the N-terminus of the protein. TNPO1 was previously documented to interact with PY-NLS. Hydrophobic-rich amino acid sequences, including proline, tyrosine and glycine and their neighbouring basic sequences are widely conserved in the sequences of proteins interacting with TNPO1. This TNPO1-binding sequence is also conserved in the capsid protein CA of HIV-1, and it was reported that TNPO1 is an essential factor that facilitates HIV-1 uncoating (Fernandez et al. 2019). IAVs are exposed to acidic conditions in late endosomes for fusion, but upon penetration, the matrix protein M1, vRNPs and the accessory proteins are released into the cytosol at an almost neutral pH. Mimicking both endosomal and cytosolic pH conditions, we examined in vitro whether TNPO1 interacts with M1. Extracts were prepared after treatment of the virus in a buffer solution of pH 5.6 containing potassium salts mimicking the late endosome luminal environment, and immunoprecipitation with purified TNPO1 protein was attempted. Interestingly, TNPO1 interacted with M1 exclusively at pH 5.6 but failed to bind to the protein at pH 7.0, indicating the interaction of TNPO1 with M1 was dependent on the acidic pH environment found in the lumen of late endosomes. On the other hand, in a mutant (G18A) virus in which the glycine at position 18 of M1, the most conserved protein in the TNPO1-binding sequence, was replaced by alanine, the interaction with TNPO1 was abrogated, and the infectivity of the virus was also greatly reduced. X-ray crystallography of the M1 protein with mutation at glycine 18 revealed a significant structural change around the mutated amino acid. This conformational change is expected to have a significant impact on the interaction with TNPO1, but molecular interaction details are still unclear and are currently being investigated. Taken together, these results suggest that TNPO1 acts on the eight-bundle vRNPs complex of IAVs and binds to the M1 protein, dissociating them into individual vRNPs to be subsequently imported into the nucleus for replication and transcription (vRNPs uncoating, Fig. 14.1).
Uncoating Reaction In Vitro
To further understand the mechanism of IAV uncoating, we performed an in vitro assay, exposing purified viruses to buffer solutions at neutral or acidic pH conditions to mimic reactions within late endosomes, and analysed the properties of ubiquitin molecules interacting with HDAC6 in the viral extracts. Virus particles exposed to pH 7.0 or pH 5.4 were disrupted by lysis buffer. Viral lysates were reacted with ZnF, the ubiquitin-binding domain of HDAC6, and purification was attempted by histidine tagging of the ZnF domain. Samples that reacted with viral extracts and were exposed to acidic conditions resulted in smears on the gel, corresponding to high molecular weight ubiquitin chains (Fig. 14.3; lane (12)). This suggests that upon acidification, the ubiquitin chains in the virus can increase to higher molecular weight. A detailed molecular mechanism of viral uncoating is currently being investigated.
Several attempts have been made to reproduce this uncoating reaction in vitro using biochemical experimental methods so far (Zhirnov 1990; Stauffer et al. 2014; Stauffer et al. 2016). To mimic the maturation process in endosomes, viral particles were exposed to different pH conditions. The lipid bilayer containing surface glycoproteins was biochemically removed by treatment with non-ionic detergents such as NP-40. In a follow-up study, we attempted to extract vRNP complexes from viral particles using this method. The surface glycoproteins of IAV particles that passed through a buffer containing NP-40 at pH 7.4 (near neutral) were removed, and only the viral core was purified in the pellet fraction (Fig. 14.4a, b). In contrast, under acidic conditions (pH 5.4) mimicking late endosomes, the shell consisting of a lipid bilayer and M1 was removed and the eight segments of vRNPs, were observed in the lower fractions by glycerol density gradient centrifugation (Fig. 14.4c). Thus, this process allowed the extraction of vRNPs with rod-like octameric segments of different lengths from the viral particles without shells, as previously demonstrated by Noda et al. (2006) in ultrathin sections. Negative staining electron microscopy of the vRNPs fraction containing NPs extracted from this intermediate layer revealed individually dispersed vRNPs as well as multiple segments of vRNPs. Since many octameric vRNPs have been observed, it is likely that IAV vRNPs are organised by RNA-RNA interactions and/or RNA-protein interactions (Fig. 14.4d). In future, such purified vRNPs could be used as substrates to reveal the molecular mechanisms of dispersion and assembly of vRNPs.
In vitro uncoating of IAV. (a) Two-layer glycerol ultracentrifugation with NP-40 at different pH precipitated viral cores under neutral conditions and vRNPs genomes under acidic conditions, respectively. (b) IAV core without HA and NA and lipid bilayer (left); vRNPs without M1 shell (right). (c) Viral fractionation by glycerol density gradient centrifugation. IAV particles uncoated in buffer solution with acidic conditions pH 5.4 containing NP-40 were layered on top of buffer containing 0–70% glycerol and fractionated by ultracentrifugation. The fractions containing NPs were collected and observed under an electron microscope. (d) Electron microscopic image of the vRNPs fraction. In addition to individually isolated rod-shaped vRNPs (arrow), bundled vRNPs complexes (arrowhead) were also observed. Enlarged images are shown on the right panels
Application to Coronavirus Endosomal Uncoating
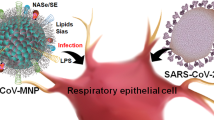
The in vitro uncoating approach to extracting vRNPs of IAVs can be applied to other viruses. For example, coronaviruses, such as SARS-CoV-2, are also RNA viruses covered with a lipid bilayer. As with SARS-CoV-2, common cold coronaviruses have spike proteins on their viral surface and a 30 kb positive-sense single-stranded RNA genome inside the virus (V’Kovski et al. 2021; Evans and Liu 2021). Coronaviruses have been reported to have an envelope protein (E protein) on the lipid bilayer membrane, which functions as an ion channel (Ruch and Machamer 2012). Since the 229E and OC43 strains have been reported to be taken up into cells in a Rab7-dependent manner (Schneider et al. 2021), the seasonal coronaviruses also internalise via endosocytosis and uncoat after fusion at the late endosome. However, the detailed mechanisms of coronavirus uncoating are still unclear. Similar to IAV, intact viral particles were precipitated by in vitro uncoating by ultracentrifugation under neutral conditions without NP-40 (Fig. 14.5; left panel). On the other hand, when ultracentrifugation was performed at neutral and/or acidic pH conditions in a buffer containing NP-40, cleaved vRNP fragments were observed under neutral conditions at pH 7.4. Interestingly, individual vRNPs form condensates at pH 5.4, mimicking the late endosomal luminal environment, and some condensates were also observed with the neighbouring virus-derived vRNPs (Fig. 14.5). The aggregation of these N-protein-wrapped RNA complexes under acidic conditions may favour the efficient release of the genome into the cytoplasm during uncoating from the endosome. It has also been suggested that coronavirus RNA-N-protein complexes may undergo phase separation (Carlson et al. 2020; Iserman et al. 2020; Jack et al. 2021), and there may be a mechanism by which, the phenomenon of phase separation can be successfully exploited to facilitate genome packaging and uncoating reactions (Etibor et al. 2021).
In vitro uncoating of coronavirus 229E strain. Purified coronavirus strain 229E was ultracentrifuged in two-layer glycerol buffer as in IAV. Virus particles precipitate in the buffer without NP-40 (left panel). Viral genome vRNPs are purified when ultracentrifuged in the buffer containing NP-40. Under neutral conditions, the structure was dissociated (centre panel). Under acidic conditions, vRNPs from each virus particle condensed (right panel). Viral genomes incorporated within endosomes in the cell may form condensates under acidic conditions as the endosomes mature. Scale bar; 200 nm
Conclusions
As viral entry represents the first step in the life cycle of a virus, inhibition of these early infection events can block the downstream infection processes. Therefore, a mechanistic understanding of virus entry has important implications for vaccine development and drug discovery research. Moreover, by targeting the entry pathways, it should be possible to inhibit different viruses that exploit the same entry route(s). In the future, the design of host-targeted molecular compounds that can block the intracellular entry of emerging and re-emerging viruses is expected to lead to the development of prophylactic and therapeutic agents common to different viruses. Alternatively, if efficient uncoating can be inhibited by drugs, viral infection can also be attenuated.
References
Alvarado-Facundo E, Gao Y, Ribas-Aparicio RM, Jimenez-Alberto A, Weiss CD, Wang W (2015) Influenza virus M2 protein ion channel activity helps to maintain pandemic 2009 H1N1 virus hemagglutinin fusion competence during transport to the cell surface. J Virol 89(4):1975–1985. https://doi.org/10.1128/JVI.03253-14
Banerjee I, Miyake Y, Nobs SP, Schneider C, Horvath P, Kopf M, Matthias P, Helenius A, Yamauchi Y (2014) Influenza A virus uses the aggresome processing machinery for host cell entry. Science 346(6208):473–477. https://doi.org/10.1126/science.1257037
Carlson CR, Asfaha JB, Ghent CM, Howard CJ, Hartooni N, Safari M, Frankel AD, Morgan DO (2020) Phosphoregulation of phase separation by the SARS-CoV-2 N protein suggests a biophysical basis for its dual functions. Mol Cell 80(6):1092–1103. e1094. https://doi.org/10.1016/j.molcel.2020.11.025
Chavrier P, Parton RG, Hauri HP, Simons K, Zerial M (1990) Localization of low molecular weight GTP binding proteins to exocytic and endocytic compartments. Cell 62(2):317–329. https://doi.org/10.1016/0092-8674(90)90369-p
Chlanda P, Mekhedov E, Waters H, Schwartz CL, Fischer ER, Ryham RJ, Cohen FS, Blank PS, Zimmerberg J (2016) The hemifusion structure induced by influenza virus haemagglutinin is determined by physical properties of the target membranes. Nat Microbiol 1(6):16050. https://doi.org/10.1038/nmicrobiol.2016.50
Dadonaite B, Gilbertson B, Knight ML, Trifkovic S, Rockman S, Laederach A, Brown LE, Fodor E, Bauer DLV (2019) The structure of the Influenza A virus genome. Nat Microbiol 4(11):1781–1789. https://doi.org/10.1038/s41564-019-0513-7
de Vries E, Tscherne DM, Wienholts MJ, Cobos-Jimenez V, Scholte F, Garcia-Sastre A, Rottier PJ, de Haan CA (2011) Dissection of the Influenza A virus endocytic routes reveals macropinocytosis as an alternative entry pathway. PLoS Pathog 7(3):e1001329. https://doi.org/10.1371/journal.ppat.1001329
Edinger TO, Pohl MO, Yanguez E, Stertz S (2015) Cathepsin W is required for escape of Influenza A virus from late endosomes. mBio 6(3):e00297. https://doi.org/10.1128/mBio.00297-15
Eierhoff T, Hrincius ER, Rescher U, Ludwig S, Ehrhardt C (2010) The epidermal growth factor receptor (EGFR) promotes uptake of Influenza A viruses (IAV) into host cells. PLoS Pathog 6(9):e1001099. https://doi.org/10.1371/journal.ppat.1001099
Etibor TA, Yamauchi Y, Amorim MJ (2021) Liquid biomolecular condensates and viral lifecycles: review and perspectives. Viruses 13(3):366. https://doi.org/10.3390/v13030366
Evans JP, Liu SL (2021) Role of host factors in SARS-CoV-2 entry. J Biol Chem 297(1):100847. https://doi.org/10.1016/j.jbc.2021.100847
Fernandez J, Machado AK, Lyonnais S, Chamontin C, Gartner K, Leger T, Henriquet C, Garcia C, Portilho DM, Pugniere M, Chaloin L, Muriaux D, Yamauchi Y, Blaise M, Nisole S, Arhel NJ (2019) Transportin-1 binds to the HIV-1 capsid via a nuclear localization signal and triggers uncoating. Nat Microbiol 4(11):1840–1850. https://doi.org/10.1038/s41564-019-0575-6
Fujioka Y, Nishide S, Ose T, Suzuki T, Kato I, Fukuhara H, Fujioka M, Horiuchi K, Satoh AO, Nepal P, Kashiwagi S, Wang J, Horiguchi M, Sato Y, Paudel S, Nanbo A, Miyazaki T, Hasegawa H, Maenaka K, Ohba Y (2018) A Sialylated voltage-dependent Ca(2+) channel binds hemagglutinin and mediates Influenza A virus entry into mammalian cells. Cell Host Microbe 23(6):809–818. e805. https://doi.org/10.1016/j.chom.2018.04.015
Gorvel JP, Chavrier P, Zerial M, Gruenberg J (1991) rab5 controls early endosome fusion in vitro. Cell 64(5):915–925. https://doi.org/10.1016/0092-8674(91)90316-q
Grambas S, Hay AJ (1992) Maturation of Influenza A virus hemagglutinin--estimates of the pH encountered during transport and its regulation by the M2 protein. Virology 190(1):11–18. https://doi.org/10.1016/0042-6822(92)91187-y
Gschweitl M, Ulbricht A, Barnes CA, Enchev RI, Stoffel-Studer I, Meyer-Schaller N, Huotari J, Yamauchi Y, Greber UF, Helenius A, Peter M (2016) A SPOPL/Cullin-3 ubiquitin ligase complex regulates endocytic trafficking by targeting EPS15 at endosomes. elife 5:e13841. https://doi.org/10.7554/eLife.13841
Hao R, Nanduri P, Rao Y, Panichelli RS, Ito A, Yoshida M, Yao TP (2013) Proteasomes activate aggresome disassembly and clearance by producing unanchored ubiquitin chains. Mol Cell 51(6):819–828. https://doi.org/10.1016/j.molcel.2013.08.016
Harrison SC (2008) Viral membrane fusion. Nat Struct Mol Biol 15(7):690–698. https://doi.org/10.1038/nsmb.1456
Harrison SC (2015) Viral membrane fusion. Virology 479-480:498–507. https://doi.org/10.1016/j.virol.2015.03.043
He J, Sun E, Bujny MV, Kim D, Davidson MW, Zhuang X (2013) Dual function of CD81 in influenza virus uncoating and budding. PLoS Pathog 9(10):e1003701. https://doi.org/10.1371/journal.ppat.1003701
Hubbert C, Guardiola A, Shao R, Kawaguchi Y, Ito A, Nixon A, Yoshida M, Wang XF, Yao TP (2002) HDAC6 is a microtubule-associated deacetylase. Nature 417(6887):455–458. https://doi.org/10.1038/417455a
Huotari J, Meyer-Schaller N, Hubner M, Stauffer S, Katheder N, Horvath P, Mancini R, Helenius A, Peter M (2012) Cullin-3 regulates late endosome maturation. Proc Natl Acad Sci U S A 109(3):823–828. https://doi.org/10.1073/pnas.1118744109
Hutchinson EC, von Kirchbach JC, Gog JR, Digard P (2010) Genome packaging in Influenza A virus. J Gen Virol 91(Pt 2):313–328. https://doi.org/10.1099/vir.0.017608-0
Iserman C, Roden CA, Boerneke MA, Sealfon RSG, McLaughlin GA, Jungreis I, Fritch EJ, Hou YJ, Ekena J, Weidmann CA, Theesfeld CL, Kellis M, Troyanskaya OG, Baric RS, Sheahan TP, Weeks KM, Gladfelter AS (2020) Genomic RNA elements drive phase separation of the SARS-CoV-2 nucleocapsid. Mol Cell 80(6):1078–1091. e1076. https://doi.org/10.1016/j.molcel.2020.11.041
Jack A, Ferro LS, Trnka MJ, Wehri E, Nadgir A, Nguyenla X, Fox D, Costa K, Stanley S, Schaletzky J, Yildiz A (2021) SARS-CoV-2 nucleocapsid protein forms condensates with viral genomic RNA. PLoS Biol 19(10):e3001425. https://doi.org/10.1371/journal.pbio.3001425
Kanaseki T, Kawasaki K, Murata M, Ikeuchi Y, Ohnishi S (1997) Structural features of membrane fusion between influenza virus and liposome as revealed by quick-freezing electron microscopy. J Cell Biol 137(5):1041–1056. https://doi.org/10.1083/jcb.137.5.1041
Kawaguchi Y, Kovacs JJ, McLaurin A, Vance JM, Ito A, Yao TP (2003) The deacetylase HDAC6 regulates aggresome formation and cell viability in response to misfolded protein stress. Cell 115(6):727–738. https://doi.org/10.1016/s0092-8674(03)00939-5
Konig R, Stertz S, Zhou Y, Inoue A, Hoffmann HH, Bhattacharyya S, Alamares JG, Tscherne DM, Ortigoza MB, Liang Y, Gao Q, Andrews SE, Bandyopadhyay S, De Jesus P, Tu BP, Pache L, Shih C, Orth A, Bonamy G, Miraglia L, Ideker T, Garcia-Sastre A, Young JA, Palese P, Shaw ML, Chanda SK (2010) Human host factors required for influenza virus replication. Nature 463(7282):813–817. https://doi.org/10.1038/nature08699
Kovacs JJ, Murphy PJ, Gaillard S, Zhao X, Wu JT, Nicchitta CV, Yoshida M, Toft DO, Pratt WB, Yao TP (2005) HDAC6 regulates Hsp90 acetylation and chaperone-dependent activation of glucocorticoid receptor. Mol Cell 18(5):601–607. https://doi.org/10.1016/j.molcel.2005.04.021
Larson GP, Tran V, Yu S, Cai Y, Higgins CA, Smith DM, Baker SF, Radoshitzky SR, Kuhn JH, Mehle A (2019) EPS8 facilitates uncoating of Influenza A virus. Cell Rep 29(8):2175–2183. e2174. https://doi.org/10.1016/j.celrep.2019.10.064
Lee KK (2010) Architecture of a nascent viral fusion pore. EMBO J 29(7):1299–1311. https://doi.org/10.1038/emboj.2010.13
Liang R, Swanson JMJ, Madsen JJ, Hong M, DeGrado WF, Voth GA (2016) Acid activation mechanism of the Influenza A M2 proton channel. Proc Natl Acad Sci U S A 113(45):E6955–E6964. https://doi.org/10.1073/pnas.1615471113
Mar KB, Rinkenberger NR, Boys IN, Eitson JL, McDougal MB, Richardson RB, Schoggins JW (2018) LY6E mediates an evolutionarily conserved enhancement of virus infection by targeting a late entry step. Nat Commun 9(1):3603. https://doi.org/10.1038/s41467-018-06000-y
Martin K, Helenius A (1991) Nuclear transport of influenza virus ribonucleoproteins: the viral matrix protein (M1) promotes export and inhibits import. Cell 67(1):117–130. https://doi.org/10.1016/0092-8674(91)90576-k
Mercer J, Schelhaas M, Helenius A (2010) Virus entry by endocytosis. Annu Rev Biochem 79:803–833. https://doi.org/10.1146/annurev-biochem-060208-104626
Miyake Y, Keusch JJ, Wang L, Saito M, Hess D, Wang X, Melancon BJ, Helquist P, Gut H, Matthias P (2016) Structural insights into HDAC6 tubulin deacetylation and its selective inhibition. Nat Chem Biol 12(9):748–754. https://doi.org/10.1038/nchembio.2140
Miyake Y, Keusch JJ, Decamps L, Ho-Xuan H, Iketani S, Gut H, Kutay U, Helenius A, Yamauchi Y (2019) Influenza virus uses transportin 1 for vRNP debundling during cell entry. Nat Microbiol 4(4):578–586. https://doi.org/10.1038/s41564-018-0332-2
Noda T (2021) Selective genome packaging mechanisms of Influenza A viruses. Cold Spring Harb Perspect Med 11(7):a038497. https://doi.org/10.1101/cshperspect.a038497
Noda T, Sagara H, Yen A, Takada A, Kida H, Cheng RH, Kawaoka Y (2006) Architecture of ribonucleoprotein complexes in Influenza A virus particles. Nature 439(7075):490–492. https://doi.org/10.1038/nature04378
Noda T, Sugita Y, Aoyama K, Hirase A, Kawakami E, Miyazawa A, Sagara H, Kawaoka Y (2012) Three-dimensional analysis of ribonucleoprotein complexes in Influenza A virus. Nat Commun 3:639. https://doi.org/10.1038/ncomms1647
Pohl MO, Edinger TO, Stertz S (2014) Prolidase is required for early trafficking events during Influenza A virus entry. J Virol 88(19):11271–11283. https://doi.org/10.1128/JVI.00800-14
Ruch TR, Machamer CE (2012) The coronavirus E protein: assembly and beyond. Viruses 4(3):363–382. https://doi.org/10.3390/v4030363
Rudnicka A, Yamauchi Y (2016) Ubiquitin in influenza virus entry and innate immunity. Viruses 8(10):293. https://doi.org/10.3390/v8100293
Saito M, Hess D, Eglinger J, Fritsch AW, Kreysing M, Weinert BT, Choudhary C, Matthias P (2019) Acetylation of intrinsically disordered regions regulates phase separation. Nat Chem Biol 15(1):51–61. https://doi.org/10.1038/s41589-018-0180-7
Schneider WM, Luna JM, Hoffmann HH, Sanchez-Rivera FJ, Leal AA, Ashbrook AW, Le Pen J, Ricardo-Lax I, Michailidis E, Peace A, Stenzel AF, Lowe SW, MacDonald MR, Rice CM, Poirier JT (2021) Genome-scale identification of SARS-CoV-2 and pan-coronavirus host factor networks. Cell 184(1):120–132. e114. https://doi.org/10.1016/j.cell.2020.12.006
Sempere Borau M, Stertz S (2021) Entry of Influenza A virus into host cells - recent progress and remaining challenges. Curr Opin Virol 48:23–29. https://doi.org/10.1016/j.coviro.2021.03.001
Staring J, Raaben M, Brummelkamp TR (2018) Viral escape from endosomes and host detection at a glance. J Cell Sci 131(15):jcs216259. https://doi.org/10.1242/jcs.216259
Stauffer S, Feng Y, Nebioglu F, Heilig R, Picotti P, Helenius A (2014) Stepwise priming by acidic pH and a high K+ concentration is required for efficient uncoating of Influenza A virus cores after penetration. J Virol 88(22):13029–13046. https://doi.org/10.1128/JVI.01430-14
Stauffer S, Nebioglu F, Helenius A (2016) In vitro disassembly of Influenza A virus capsids by gradient centrifugation. J Vis Exp 109:e53909. https://doi.org/10.3791/53909
Su WC, Chen YC, Tseng CH, Hsu PW, Tung KF, Jeng KS, Lai MM (2013) Pooled RNAi screen identifies ubiquitin ligase Itch as crucial for Influenza A virus release from the endosome during virus entry. Proc Natl Acad Sci U S A 110(43):17516–17521. https://doi.org/10.1073/pnas.1312374110
V’Kovski P, Kratzel A, Steiner S, Stalder H, Thiel V (2021) Coronavirus biology and replication: implications for SARS-CoV-2. Nat Rev Microbiol 19(3):155–170. https://doi.org/10.1038/s41579-020-00468-6
Wang L, Moreira EA, Kempf G, Miyake Y, Oliveira Esteves BI, Fahmi A, Schaefer JV, Dreier B, Yamauchi Y, Alves MP, Pluckthun A, Matthias P (2022) Disrupting the HDAC6-ubiquitin interaction impairs infection by influenza and Zika virus and cellular stress pathways. Cell Rep 39(4):110736. https://doi.org/10.1016/j.celrep.2022.110736
Yamauchi Y (2020) Influenza A virus uncoating. Adv Virus Res 106:1–38. https://doi.org/10.1016/bs.aivir.2020.01.001
Yamauchi Y, Greber UF (2016) Principles of virus uncoating: cues and the snooker ball. Traffic 17(6):569–592. https://doi.org/10.1111/tra.12387
Yamauchi Y, Helenius A (2013) Virus entry at a glance. J Cell Sci 126(Pt 6):1289–1295. https://doi.org/10.1242/jcs.119685
Yanguez E, Hunziker A, Dobay MP, Yildiz S, Schading S, Elshina E, Karakus U, Gehrig P, Grossmann J, Dijkman R, Schmolke M, Stertz S (2018) Phosphoproteomic-based kinase profiling early in influenza virus infection identifies GRK2 as antiviral drug target. Nat Commun 9(1):3679. https://doi.org/10.1038/s41467-018-06119-y
Zhang Y, Gilquin B, Khochbin S, Matthias P (2006) Two catalytic domains are required for protein deacetylation. J Biol Chem 281(5):2401–2404. https://doi.org/10.1074/jbc.C500241200
Zhang X, Yuan Z, Zhang Y, Yong S, Salas-Burgos A, Koomen J, Olashaw N, Parsons JT, Yang XJ, Dent SR, Yao TP, Lane WS, Seto E (2007) HDAC6 modulates cell motility by altering the acetylation level of cortactin. Mol Cell 27(2):197–213. https://doi.org/10.1016/j.molcel.2007.05.033
Zhang Y, Kwon S, Yamaguchi T, Cubizolles F, Rousseaux S, Kneissel M, Cao C, Li N, Cheng HL, Chua K, Lombard D, Mizeracki A, Matthias G, Alt FW, Khochbin S, Matthias P (2008) Mice lacking histone deacetylase 6 have hyperacetylated tubulin but are viable and develop normally. Mol Cell Biol 28(5):1688–1701. https://doi.org/10.1128/MCB.01154-06
Zhirnov OP (1990) Solubilization of matrix protein M1/M from virions occurs at different pH for orthomyxo- and paramyxoviruses. Virology 176(1):274–279. https://doi.org/10.1016/0042-6822(90)90253-n
Acknowledgements
The authors wish to acknowledge Division for Medical Research Engineering, Nagoya University Graduate School of Medicine, for technical support of electron microscopy; Dr. Koji Itakura, analysis with Mass spec facility; Dr. Kentaro Taki, technical support for microscopy; Dr. Eri Yorifuji. We also thank Prof. Yohei Yamauchi (ETH Zürich), Prof. Hiroshi Kimura and Mr. Atzin Bolanos-Ceron for comments on the manuscript, Prof. Jiro Usukura (Nagoya University Graduate School of Science) and Ms. Tomoko Kunogi for technical assistance.
Funding
This work was supported by JSPS RPD research fellowship (Grant Number 18J40214), JSPS KAKENHI (Grant-in-Aid for Scientific Research (C)) Grant Number JP20K07513, JST FOREST Program (Grant Number JPMJFR206B, Japan), and MRC-AMED SICORP (Grant Number JP22jm0210070, Japan) to Y.M.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Miyake, Y., Hara, Y., Umeda, M., Banerjee, I. (2023). Influenza A Virus: Cellular Entry. In: Vijayakrishnan, S., Jiu, Y., Harris, J.R. (eds) Virus Infected Cells. Subcellular Biochemistry, vol 106. Springer, Cham. https://doi.org/10.1007/978-3-031-40086-5_14
Download citation
DOI: https://doi.org/10.1007/978-3-031-40086-5_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-40085-8
Online ISBN: 978-3-031-40086-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)