Abstract
Cardiac optogenetics offers unprecedented opportunities to study the role of different cell populations in complex biological tissues, such as the heart. To this end, light-emitting sensor proteins or light-inducible effector proteins are expressed in the target cells to either observe or steer their activity with high spatiotemporal resolution. Optogenetic tools to monitor cardiac activity include genetically encoded Ca2+ and voltage indicators. In addition, photo-activated ion pumps and channels are used to modulate transmembrane ionic flow and membrane voltage. In cardiac research, optogenetic approaches have been applied successfully for optical pacing, resynchronization, and defibrillation, and they have offered novel insight into cell-specific contributions to arrhythmia formation and termination, as well as simplified automatization of cardiac toxicity screening. Combining optogenetic experiments on intact myocardium with computational modelling allows one to quantitatively assess hypotheses on arrhythmia mechanisms, with the vision of developing novel, targeted approaches to prevent or terminate cardiac arrhythmias. In the following chapter, we summarize principles of optogenetic interrogations of cardiac tissue and present key experiments towards optical control of heart rhythm.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Optogenetics
- Channelrhodopsin
- Optical pacing
- Optical defibrillation
- Cardiotoxicity screening
- Computational modelling
1 Introduction
Successful optogenetic experiments require suitable probes, cell-population targeted gene delivery, and optical technologies for spatiotemporally defined, yet minimally invasive light application or collection. Despite the idea of selectively manipulating cellular activity in brain tissue being proposed as early as 1979 [1], and the discovery of light-driven ion pumps in the 1970s [2, 3], only a few optogenetic experiments were reported before the millennium. These include the pioneering work of H. Gobind Khorana expressing bovine rhodopsin in Xenopus oocytes [4] and early studies using fluorescent sensor proteins to image vesicular pH changes, intracellular Ca2+ concentrations, and membrane voltage dynamics [5,6,7]. First optogenetic manipulation of excitable cells was realized by Boris Zemelman and colleagues, co-expressing arrestin-2, rhodopsin, and the α-subunit of the corresponding heterotrimeric G protein in hippocampal neurons to increase action potential firing rate during photostimulation [8]. The breakthrough of optogenetics was the characterization of light-gated ion channels, called channelrhodopsins (ChR), at a time when methods for efficient gene transfer were available. In 2005/2006, several groups used channelrhodopsin-2 (ChR2) to drive depolarizing ion currents in neurons, thereby eliciting action potentials (AP) in vitro and in vivo [7,8,9,10,11,12]. At the same time, Tallini et al. employed the genetically encoded Ca2+ sensor (GCamP2) for measuring Ca2+ transients in vivo and for imaging Ca2+ signaling in the developing heart [13]. However, the use of ChR2-based tools for cardiac applications was only implemented about five years later [14, 15].
In the following, we introduce commonly applied optogenetic probes (see Figs. 17.1 and 17.2) and summarize strategies for gene targeting and optical measurements, before presenting key studies using optogenetic approaches to monitor and steer cardiac rhythm in vertebrate cells, tissues, and hearts. Furthermore, we highlight how optogenetics will foster our understanding of cardiac arrhythmias, and how we can use it for facilitated cardiotoxicity screening and for dissecting non-myocyte contributions to arrhythmia inducibility and termination.
Optogenetic actuators for membrane potential modulation and their effects in cardiomyocytes. Comparison of common optogenetic actuators and their respective effects on cardiomyocyte resting membrane potential (RMP). Wavelengths of maximal activation are indicated. Abbreviations: ChR2 channelrhodopsin-2 from Chlamydomonas reinhardtii, ReachR red-activatable channelrhodopsin, iChloC improved chloride-conducting channelrhodopsin, iC++ improved chloride-conducting chimeric channelrhodopsin, GtACR1 anion channelrhodopsin-1 from Guillardia theta, ArchT proton pump from Halorubrum sp. TP009, NpHR halorhodopsin from Natronomonas pharaonis, AP action potential, bPAC photoactivated cyclase from Beggiatoa, SthK cyclic-nucleotide-gated K+ channel from Spirochaeta thermophile
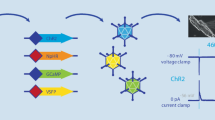
Additional optogenetic tools: Light-activated G protein coupled-receptors (GPCR) and fluorescent reporters. (a) GPCR used to modulate cardiac activity include JellyOP from Carybdea rastonii and murine melanopsin. Of note, by binding a different Gα protein, melanopsin can also activate the Gi/o signaling pathway, similar to short- and long-wavelength cone opsins (SWO and LWO). (b) Fluorescent voltage reporters comprise the Förster-resonance energy transfer (FRET)-based indicators mermaid and VSFP2.3, and rhodopsin-based, single-wavelength indicators such as QuasAr2. In FRET-based reporters, voltage changes are transmitted from the voltage-sensitive domain (VSD) of Ciona intestinalis voltage-sensing phosphatase to a FRET pair, thereby increasing FRET efficiency upon membrane depolarization. Mermaid employs mUKG and mKOκ, VSFP2.3 uses cyan and yellow fluorescent protein (CFP and YFP) as FRET donors and acceptors, respectively. (c) Genetically encoded Ca2+ sensors of the GCamP family are single-wavelength indicators, wherein Ca2+-induced changes in the interaction of the myosin light chain kinase fragment (M13) and calmodulin (CaM) change fluorescence intensity of a circular permutated green fluorescent protein (cpGFP). (d) CaViar can be used for simultaneous imaging of Ca2+ and voltage dynamics
2 Optogenetic Actuators in Cardiac Optogenetics
Optogenetic tools can be divided into light-activated effector proteins, also referred to as optogenetic actuators, and light-emitting sensor proteins [16]. Most commonly used actuators are based on microbial rhodopsins, small heptahelical transmembrane proteins binding all-trans retinal as co-factor for light absorption. More specifically, light-driven proton, chloride, and sodium pumps use the energy provided by visible light for the active transport of ions against the transmembrane gradient (proton and sodium pumps drive outward transport of cations, chloride pumps drive inward transport of anions). When expressed in excitable cells, light-driven pumps will thus mediate hyperpolarizing membrane currents during illumination, which may inhibit AP both in neurons and in cardiomyocytes (Fig. 17.1c). The orange-light activated, inward-directed chloride pump NpHR from Natronomonas pharaonis was first used in zebrafish hearts to induce reversible block of contractions upon global cardiac illumination. Using spatially restricted illumination patterns, it was further utilized to identify the location of pacemaker cells in the developing zebrafish heart and to optically mimic different degrees of atrioventricular conduction block [14]. NpHR-mediated hyperpolarization was also shown to silence or shorten AP in monolayers of neonatal rat ventricular myocytes [17]. Pumping protons out of the cell, the green-light-driven proton pump ArchT shows hyperpolarizing effects on the membrane potential that are comparable to NpHR effects. ArchT-mediated hyperpolarization has been exploited to silence spontaneous excitation of fibroblast–cardiomyocyte co-cultures. Interestingly, in those experiments, ArchT was expressed in fibroblasts, with the resulting hyperpolarization transmitted to cardiomyocytes by electronic coupling [18, 19]. In mouse hearts expressing ArchT in cardiomyocytes, hyperpolarizing currents upon green-light exposure were sufficient to terminate ventricular arrhythmia, albeit at low efficiency [20]. This could be explained either by the limited inhibitory effect of hyperpolarization per se or by the high light intensities required, as maximally one ion can be transported per absorbed photon by light-driven ion pumps.
In contrast to pumps, channelrhodopsins (ChR) are light-gated ion channels mediating passive ion flux along the electrochemical gradient upon light activation. ChR can be divided into cation-selective ChR, such as the frequently used channelrhodopsin-2 (ChR2) from Chlamydomonas reinhardtii, and the more recently discovered group of anion-conducting channelrhodopsins (ACR), including engineered channels (e.g. iChloC, iC++) and naturally occurring representatives from Guillardia theta (GtACR1, GtACR2) [21,22,23]. Cation-selective ChR conducts protons, sodium, potassium, calcium, and magnesium ions [24], resulting in depolarizing membrane currents at negative membrane potentials (reversal potential at around 0 mV). Their activation by short light pulses thus leads to short, reversible membrane depolarization of the target cells, sufficient to reliably trigger AP in cardiomyocytes (Fig. 17.1a). Prolonged illumination results in sustained depolarization, preventing repolarization to the resting membrane potential. This precludes recovery from the inactivation of fast sodium channels, thereby suppressing re-excitation of cardiomyocytes [15]. In cardiac optogenetic studies, the blue-light-activated ChR2 and the green-light-activated chimera ReachR [25] have been widely used for modulating cardiac electrophysiology, with applications ranging from cardiac pacing, resynchronization, arrhythmia termination, and drug screening, to studying the role of intracardiac neurons and interstitial non-excitable cells [26].
Anion-selective channelrhodopsins predominantly conduct chloride ions under physiological conditions, thus their reversal potential is determined by the transmembrane gradient for chloride. In cardiomyocytes, ACR activation leads to membrane depolarization, suitable either for optically pacing with short light flashes or for maintaining cells at constant depolarized potentials during prolonged illumination, thereby suppressing further AP (Fig. 17.1b) [27, 28]. Despite high expression levels and large photocurrents of ACR in cardiomyocytes, their use has been restricted to proof-of-principles studies so far. Holding cardiomyocytes at their diastolic membrane potential can be achieved using light-activated K+-conducting channel systems (Fig. 17.1d). However, currently available systems are either limited by insufficient expression levels in mammalian cells [29] or by slow off-kinetics [30], rendering them unsuitable for applications aiming at beat-by-beat control of cardiac excitation. These challenges might be overcome by the recently discovered class of natural occuring Kalium Rhodopsins, including the potent K+- selective channel WiChR from Wobblia lunata [31, 32].
In addition to microbial rhodopsin-based tools to control the membrane potential, there is a vast range of other optogenetic actuators, including, but not limited to, light-activated G protein-coupled receptors (GPCR, Fig. 17.2a), photo-activated enzymes, and light-controlled protein interaction systems [16]. Optical control of GPCR signaling is of special interest for understanding intracellular signaling in cardiomyocytes. Suitable GPCR comprise naturally occurring light-sensitive GPCR (visual and non-visual vertebrate rhodopsins and invertebrate rhodopsins) [33, 34] and custom-engineered rhodopsins referred to as opto-XR.
Light-induced activation of the invertebrate rhodopsin JellyOP allows mimicking β-adrenergic stimulation with unprecedented spatiotemporal precision within the intact heart [35]. Illumination of the right atrium of isolated murine hearts expressing JellyOP leads to an instantaneous increase in spontaneous beating rate. In contrast, illumination of the left posterior atrium at the sites of pulmonary vein insertions results in the generation of supraventricular extrasystoles. Thus, JellyOP activation can be used to look for pathway-specific arrhythmia hotspots within the intact heart. Furthermore, the use of short activating light pulses revealed different on-kinetics of the positive inotropic and lusitropic effects, with the latter being significantly faster. These experiments indicate that optogenetic approaches can facilitate the study of amplification mechanisms and temporal dynamics of G protein signaling cascades, as well as compartment-specific signaling behavior. Importantly, optogenetic control of GPCR activation can mimic physiological responses to pulsatile neurotransmitter release by repeated application of short light flashes.
In embryonic stem-cell-derived cardiomyocytes, heterologous expression of the vertebrate rhodopsin melanopsin enables light-induced Gq protein activation leading to inositol-1,4,5-trisphosphate production and elevation of intracellular Ca2+ concentration, thereby enhancing beating frequency of spontaneously beating embryoid bodies [36]. However, melanopsin is a promiscuous receptor also able to activate the Gi signaling cascade [37, 38]. While optogenetic GPCR for specific control of Gi pathways have been identified [39, 40], the optogenetic toolbox currently lacks probes to specifically control Gq signaling – despite the importance of this pathway for acute and chronic adaption of the heart. One possible candidate GPCR could be human Neuropsin (hOPN5), as recently proposed [41].
3 Fluorescent Reporters in Cardiac Optogenetics
Complementary to optogenetic actuators, fluorescent sensor proteins can be used for cell-type specific imaging of selected cellular parameters. With respect to cardiac arrhythmias, genetically encoded Ca2+ indicators (GECI) and genetically encoded voltage indicators (GEVI) represent the most important tools for imaging cardiac activity (Fig. 17.2b–d). GECI comprise single-fluorophore (e.g. GCaMP family) or FRET-based Ca2+ sensors (e.g. Cameleon) that change their fluorescence intensity and/or FRET efficiency upon Ca2+ binding [42]. GCaMP2 was first used to image Ca2+ dynamics in mouse hearts in vivo and to assess the effects of isoproterenol on diastolic and systolic Ca2+ levels as well as kinetics of Ca2+ transients in cardiomyocytes. In Langendorff-perfused hearts, GCaMP2 further enabled measurements of the conduction velocity of Ca2+ waves, and for combined recordings of Ca2+ transients and membrane voltage, the latter using the red-fluorescent voltage dye RH237 [13]. GCaMP2-based imaging also showed functional coupling between engrafted embryonic cardiomyocytes and native myocardium, preventing the risk of post-infarct arrhythmia [43]. To date, a variety of GECI with optimized properties are available, including variants with red-shifted fluorescence spectra [44, 45], fast response times [46, 47], and low-affinity indicators suitable for measuring Ca2+ dynamics in intracellular organelles with elevated Ca2+ levels such as the sarcoplasmic reticulum [48,49,50].
GEVI include single-wavelength and ratiometric fluorescent indicators that enable imaging cell-type specific membrane voltage dynamics in real-time [51, 52]. Being expressed under the control of the cardiomyocyte-specific myosin light chain 2 promoter, the ratiometric GEVI mermaid, for example, was used for non-invasive imaging of cardiac activity in embryonic zebrafish hearts, revealing altered cardiac excitation patterns in the presence of the hERG channel blocker astemizole [53]. Similarly, cardiomyocyte-targeted expression of the voltage-sensitive fluorescent protein VSFP2.3 was applied for measuring optical cardiograms, both during sinus rhythm and ventricular tachyarrhythmia in intact mouse hearts [54]. When targeted to cardiac non-myocytes VSFP2.3 was further used to explore electrotonic coupling from cardiomyocytes to non-myocytes in the scar border zone of murine hearts [55]. Additionally, GEVI facilitated rapid phenotyping of stem-cell-derived cardiomyocytes [52, 56]. Improved GEVI comprise variants exhibiting minimized photobleaching and optimized performance for 2-photon imaging, as well as near-IR fluorescent sensors [51, 57], but their utility for cardiac application still needs to be demonstrated.
4 Targeted Transgene Delivery
After selecting the appropriate molecular tools for an optogenetic experiment, they need to be targeted to the specific cell population of interest. The four main strategies for targeted transgene delivery to cardiac cells are (1) the generation of knock-in animals expressing the gene of interest under a cell-type specific promoter, (2) the use of recombinatorial animal models (mainly mice) where a cell-type specific driver line (e.g. expressing Cre recombinase) is cross-bred with a driver-dependent line coding for the probe (e.g. Cre-dependent line containing a floxed or flexed transgene) [58,59,60,61,62,63], (3) viral delivery of genes of interest [64,65,66], and (4) injection of cells expressing the respective optogenetic probe [67]. The technical details and challenges of the individual methods have been discussed earlier [68, 69]. We would like to strengthen the point that the model generation itself is one of the essential steps towards meaningful optogenetic experiments, which requires thorough controls to exclude off-target expression [70, 71] and side-effects such as cardiotoxicity [72, 73] and immunogenicity [74].
5 Ex Vivo Optical Stimulation and Readouts
Another challenge is suitable light delivery and collection for spatiotemporally precise photostimulation and optical readouts of cardiac activity. In principle, for ex vivo experiments, light delivery via conventional light sources for fluorescence microscopy such as shutter-controlled halogen or mercury lamps with suitable bandpass filters, or standard LEDs in combination with microscope/macroscope optics for spatial focusing provide sufficiently high light intensities for optical probe activation. Patterned illumination can be achieved with different optical approaches, e.g. using digital micromirror devices (DMD) from projectors [14, 75], or by rapid scanning of focused excitation light with acousto-optical deflectors [76]. Combining optical stimulation with simultaneous readouts of electrical activity, e.g. via optical mapping, allows one to establish closed-loop systems for real-time adaptation of optical stimulation patterns to observed electrical activation maps. Potential applications for all-optical systems include fast light-controlled restoration of normal electrical activity in hearts showing AV block, and optical termination of ventricular arrhythmias [77, 78].
6 Optogenetic Approaches for Controlling Heart Rate and Rhythm
Commonly used devices for heart rhythm control are artificial electrical pacemakers (atrial, ventricular, or dual-chamber pacemakers) and implantable cardioverter defibrillators (ICD), which rely on the application of electrical shocks by electrodes for triggering cardiac excitation and defibrillation, respectively. Pacemakers, on the one hand, have proven extremely useful for long-term maintenance of cardiac activity and come with the advantage that electrical pulses usually remain unnoticeable to patients. ICD shocks, on the other hand, use approximately one thousand times more energy, leading to non-specific tissue excitation of myocardium, nerves, and skeletal muscle. This is associated with adverse effects, including severe pain, chronic anxiety, and structural tissue remodeling [79, 80]. In contrast, optogenetic approaches enable depolarization or hyperpolarization of spatially defined subsets of cardiomyocytes, with minimal effects on other cardiac cell populations. Optogenetic-based systems for external heart rhythm control may thus, in the long term, provide more specific, pain-free, and effective alternatives for heart rhythm management in patients.
6.1 Optical Pacing
Depolarizing ChR2-expressing myocytes by short blue-light pulses reliably triggers AP, allowing atrial and ventricular optical pacing in intact hearts from transgenic or virally transduced animals [14, 15, 64, 66]. Furthermore, dual- and multi-site optical stimulation effectively synchronizes ventricular activation, indicating the feasibility of light-driven cardiac resynchronization therapy [66]. Cell-type-specific optogenetic depolarization was also used to assess the number of activated cardiomyocytes or Purkinje cells required for inducing focal ectopic beats, finding that simultaneous depolarization of at least 1300–1800 working cardiomyocytes or 90–160 Purkinje fibers was necessary to trigger extrabeats in murine hearts [58]. Finally, ChR2 activation in human induced pluripotent stem cells in 3D-engineered heart tissue allowed for intermittent tachypacing over the duration of several weeks, providing novel insights into electrical and mechanical remodeling in ventricular and atrial tachycardia, as well as into long-term effects of optogenetic interventions [81, 82].
6.2 Optical Defibrillation
Using optogenetic approaches for terminating either atrial or ventricular tachyarrhythmias does not only improve our mechanistic understanding of cardioversion but may also facilitate the development of strategies towards optimized defibrillation therapy, e.g. by testing different locations, geometries, and light levels in a reversible and minimally invasive manner. So far, this relied on ChR2- or ReachR-based depolarization of cardiomyocytes by longer light pulses (hundreds of milliseconds to seconds), thereby suppressing effective repolarization, extending the period of Na+ channel inactivation and thus inhibiting re-excitation. Bruegmann et al. showed effective termination of ventricular tachycardia by ChR2 activation via blue light application to the anterior ventricular epicardium of Langendorff-perfused mouse hearts, both for healthy hearts and following myocardial infarction [79]. Crocini et al. [76] used a similar experimental approach to terminate induced ventricular tachycardia in ChR2-expressing mouse hearts and showed that a three-barrier pattern of ventricular illumination—based on prior knowledge of the location and geometry of the underlying re-entry pathway—was equally potent to block VT as unfocussed light application on the left ventricular surface, albeit at much lower total irradiation energy (~ 4% of energy). In contrast, single-barrier or point stimulation was insufficient to effectively block arrhythmias in their study [76]. Successful optical defibrillation was also demonstrated on rat hearts expressing the green-light-activated ChR chimera ReaChR [65]. Follow-up studies have since demonstrated the feasibility of optical defibrillation also for terminating atrial fibrillation [83, 84].
Based on experimental studies varying the timing, location and/or intensity of optical stimulation in combination with in silico models, several mechanisms have been suggested for effective optogenetic cardioversion. These include ChR-mediated depolarization and conduction block into illuminated tissue volumes [79]. In this case, transmural depolarization to keep Na+ channels refractory seems to be mandatory [79, 85]. Alternatively, filling the “excitable gap” can be achieved by pacing the excitable region between wavefronts, promoting the extinction of self-sustained re-entry. While the energy requirements are much lower for this mechanism, filling the excitable gap either relies on spatiotemporal information on rotating wavefronts or requires global illumination of the entire ventricular surface (which obliterates the energetic advantage) [86]. Optogenetic tissue depolarization is also associated with prolonged action potential duration (APD), which has been proposed as a distinct mechanism for terminating VT [65]. Finally, a recent study suggested that low-intensity illumination, leading to sub-threshold depolarization, may be effective for steering rotors towards more depolarized tissue areas, which could be used to terminate cardiac arrhythmias [87]. Interestingly, optogenetic termination of VT is also possible using the light-driven proton pump ArchT, although at lower efficacy compared to depolarizing ChR, potentially via an increased electrical sink mechanism [20]. Light-gated K+ channels [29, 30] could serve as potent tools for arrhythmia termination if current amplitudes were sufficiently large to counteract the large Na+ inward currents associated with the propagating wave of excitation, thereby preventing electrical activation and conduction in illuminated tissue areas.
Combining monitoring of heart rhythm and cardiac conduction with optical arrhythmia termination offers the opportunity for automated, real-time correction of rhythm disturbances. Following this idea, Burton et al. implemented light-controlled reversal of spiral wave chirality based on dye-free imaging of contractions in cardiomyocyte monolayers [78]. Scardigli et al. developed a closed-loop system comprising a high-speed sCMOS camera for optical mapping of membrane voltage and a DMD-based projector for optical stimulation, which allowed them to correct for atrioventricular node delay and to optically simulate re-entrant tachycardia [77]. A hybrid bioelectric system with automated ECG-based optogenetic stimulation of the right-atrial epicardial surface was used by Nyns et al. and was shown to be effective in terminating atrial tachyarrhythmias in rats in vivo [84].
The studies described here demonstrate the feasibility of optical rhythm control in select animal models. Further advances in optogenetic proteins (e.g. red-light activated channels with high light sensitivity), methods for safe and targeted gene delivery (ensuring long-lasting, stable protein expression), and software/hardware for optical stimulation (including biocompatible miniature LEDs for photostimulation) will be needed before these concepts may be transferrable to patients in the future [73].
7 Using Optogenetics for Cardiotoxicity Screening
Cardiac toxicity and, in particular, drug-induced long QT syndrome are major reasons why newly developed drugs fail to enter or are withdrawn from the market. To avoid life-threatening side effects, as well as the related financial consequences for the pharmaceutical industry, new approaches providing more accuracy and predictive power are required during preclinical drug development [88]. Drug-induced cardiotoxicity is often related to inhibiting Kv11.1 channels, reducing hERG currents, and prolonging APD. Moreover, drugs may also modulate Nav1.5 channels. Inhibition of Nav1.5 channels can induce Brugada-like syndromes, whereas activation of late sodium currents may cause AP prolongation similar to LQT3 syndrome [89,90,91].
Classically, cardiotoxicity screening of cardiac syncytia relies on spontaneous beating (limited to slow beating rates) or electrical field stimulation (associated with potential stimulation artefacts and/or side effects on stimulated cells) for rate control. In contrast, all-optical approaches provide the unique opportunity for precise external pacing, simultaneous recordings of membrane potential or Ca2+ levels, and measurements of myocyte contractility [17, 92, 93]. This further comes with the advantage of potential automatization [94], even if spectral separation can be difficult (i.e. one needs to separate light used for ChR2 and dye excitation from emission light by fluorescent proteins and dyes) [95]. OptoDyce is a platform combining ChR2-mediated stimulation and synthetic dyes for voltage and Ca2+ imaging with dye-free video tracking of contractions of cardiomyocyte monolayers. OptoDyce uses short light flashes for optical pacing and temporally separated (intermitted) optical recordings to avoid cross-contamination of the different light signals [92]. In subsequent work, the same group described a quantitative all-optical assay of inter-cellular connectivity (i.e., OptoGap) [96]; this study also used computational modelling, as discussed in the next section, to show that it may be feasible to scale up this approach to intact human hearts, even after accounting for light attenuation effects. Dempsey et al. used a co-culture of two different, spatially separated transgenic cardiomyocyte lines differentiated from human-induced pluripotent stem cells (a ChR line for optical pacing and a reporter line expressing CaViar; Fig. 17.2d) [93]. Spectral separation was also achieved by employing infrared-compatible dyes for voltage and Ca2+ imaging [97]. Alternatively, optical pacing was combined with the recording of electrical field potentials [98]. Global illumination results in near-synchronous activation of all cardiomyocytes, allowing spatial averaging of measured field potentials, as shown with averaging from 60 electrodes within an array [99] and later using one larger electrode [100], thereby simplifying high-throughput screening. Importantly, studies employing optogenetics for cardiotoxicity screening neither reported side effects from overexpression of ChR2, nor acute effects on myocyte electrophysiology. Furthermore, no drug effect on ChR photoactivation has been reported so far. However, chronic optogenetic tachypacing has been shown to be arrhythmogenic by itself [81, 82]. Taken together, combined optogenetic rate control and imaging of cardiomyocyte behaviour provide a unique opportunity for high-throughput cardiotoxicity screening.
8 Computational Modelling and Simulation of Cardiac Optogenetics
Multi-scale simulations of cardiac electrophysiology have emerged as a means for investigating mechanistic underpinnings of arrhythmia perpetuation, and for exploring new therapeutic approaches in realistic image-based human atrial or ventricular models [101]. Thanks to the development of robust mathematical representations of light-, voltage-, and time-dependent responses of microbial rhodopsins (e.g., ChR2-H134R) to light stimulation [102], it is possible to simulate optogenetic manipulation of the heart [103, 104]. This involves modifications of existing computational approaches at the cell, tissue, and organ scale to account for the expression of photo-sensitive channels and pumps, spatial distribution of light-sensitized cells resulting from viral gene delivery, and attenuation of light applied to the myocardial surface due to photon scattering and absorption.
Simulations have been used to explore the theoretical possibility of deploying optogenetics in the hearts of larger mammals (e.g., rabbits, dogs, and humans [103]), which thus far cannot be tested in an experimental setting. This work provided proof-of-concept evidence for the feasibility of light-based arrhythmia termination in human atrial [105, 106] and ventricular [79, 107] models. A prominent take-home message has been that the significant attenuation of blue light in the thicker-walled chambers of larger species is expected to be a major hindrance to the translation of approaches from smaller animals, leading to the prediction that optogenetic actuators with red-shifted action spectra would be necessary. Another important finding is that spatially targeted illumination of critical areas to disrupt re-entrant circuits could dramatically reduce the energy needs for optogenetic defibrillation [79, 107], confirming experimental findings in smaller hearts [83]. More recent work has assessed the use of ACR to disrupt re-entrant arrhythmia in human hearts, showing that utilizing these high-efficiency chloride conducting channels may lower the energy required for defibrillation by 2–3 orders of magnitude [108].
A second application of computational cardiac optogenetics research is the use of simulations to enhance the understanding of data from wet lab preparations like tandem cell units [67, 103] or monolayers of cardiomyocytes co-cultured with light-sensitized donor cells [109], or to validate and explore the expansion of methodologies like the OptoGap approach described in the prior section [96]. Likewise, this type of modelling work has been employed to explore completely new applications of optogenetics like the use of subthreshold light-induced depolarization to steer re-entrant drivers towards non-conductive boundaries, ultimately resulting in arrhythmia termination [87]. Notably, while much of the recent interest in computational modelling of cardiac optogenetics has focused on actuator opsins (e.g., ChR2, GtACR1), similar tools have also been developed to realistically simulate light emission from dyes used extensively in optical mapping experiments [110,111,112].
9 Cardiac Optogenetics beyond Cardiomyocytes
Taking advantage of the possibility to selectively target individual cell populations of interest, optogenetic approaches have been extended to cardiac non-myocytes, including cardiac interstitial cells, and intracardiac sympathetic and parasympathetic neurons. While interstitial cells (e.g. fibroblasts and resident immune cells) have traditionally been viewed as electrical insulators, there has been long-standing evidence that cardiac fibroblasts can be electrotonically coupled to cardiomyocytes in vitro, acting as an electrical load and/or interlinking otherwise unconnected cardiomyocytes [113]. Based on fibroblast-specific activation of either ChR2, ArchT, or eNpHR, several optogenetic studies have confirmed electrotonic coupling between (myo)fibroblasts and cardiomyocytes in diverse in vitro systems, with direct effects on cardiomyocyte excitability, AP properties, and conduction velocity [18, 67, 114,115,116]. In 2016, Quinn et al. showed that fibroblasts expressing VSFP2.3 follow rhythmic depolarizations of cardiomyocytes in scar border zone tissue following ventricular cryoinjury in Langendorff-perfused whole hearts [55]. Similar coupling was subsequently observed by Rubart et al. studying myofibroblasts after myocardial infarction [117]. In 2017, Hulsmans et al. described Cx43-based gap junctions between cardiomyocytes and tissue-resident macrophages and demonstrated that ChR2-mediated macrophage depolarization could facilitate AV node conduction at high atrial stimulation rates [63]. These studies exemplify how cell-type-specific optogenetic manipulation and observation can be applied for unravelling cellular functions that are difficult to assess with classic electrophysiological techniques such as patch-clamp measurements or dye-based optical mapping of membrane voltage.
Another exciting avenue of cardiac optogenetics is the selective photostimulation of intracardiac neurons. Wengrowski et al. established a mouse model expressing ChR2 in murine catecholaminergic sympathetic neurons. In Langendorff-perfused hearts, photostimulation increased both heart rate and developed force of contraction and led to a significant shortening of the AP plateau phase. Moreover, the optogenetic release of norepinephrine increases both the incidence and severity of ventricular arrhythmias following burst pacing [59]. Based on experiments using optogenetic stimulation of sympathetic neurons and FRET-based imaging of cAMP levels, Prando et al. concluded that neurotransmission from sympathetic neurons to cardiomyocytes occurs within spatially (diffusion)-restricted intermembrane domains [60]. In line with the observed chronotropic effects of sympathetic nerve activation, reversible neuronal hyperpolarization by ArchT activation in the left stellate ganglion decreases the amplitude and frequency of nerve activity. This resulted in reduced systolic blood pressure increase upon neuronal stimulation, longer APD and effective refractory periods, decreased heart rate variability, and suppression of ischemia-induced ventricular arrhythmias in an in vivo study using virally transduced beagles [118].
Moreno et al. targeted ChR2 to choline acetyltransferase-expressing neurons; hence, blue-light mediated ChR2 activation induced acetylcholine release from cardiac parasympathetic neurons. Accordingly, illumination slowed the sinus rate and delayed atrioventricular node conduction [119]. When using ChR2-mediated activation of cholinergic neurons of the inferior pulmonary vein ganglionated plexus, Rajendran et al. also observed a light-dependent decrease in the heart rate; however, in their experimental setting, no change in AV node conduction was seen, suggesting that neuronal fibers originating from this plexus may pass the AV node without forming synapses. Photostimulation at higher frequencies further induced ectopic atrial activation and asystole. Finally, the study showed differential effects of optogenetic vs. electrical stimulation of the vagus nerve, enabling authors to dissect the effects of efferent and afferent vagal nerve activation [120]. Taken together, optogenetic interrogation of intracardiac neurons thus represents an elegant method to elucidate the output of defined neuronal subsets on myocyte activity from single-cell to whole-heart level, as selective photostimulation enables superior specificity compared to electrical activation that cannot discriminate well between different neuronal classes, and that may also directly stimulate neighboring myocytes.
10 Conclusions
We have summarized the experimental approaches in cardiac optogenetics and how they can be used for studying cardiac electrophysiology and intercellular communication. Moreover, we have highlighted key applications of optical heart rhythm control and cardiotoxicity screening. Computational modelling facilitates the interpretation of experimental data from laboratory-based optogenetic studies and data integration across scales and species.
As cardiac optogenetics opens up a host of new opportunities to tackle basic research questions at present, it may pave the way for cell-type specific anti-arrhythmic therapies in the future. However, proof-of-concept studies are still limited to small animals and major hurdles have to be taken before transfer from bench to patient is realistic, with efficient and safe gene transfer, development of implantable light sources, and prevention of an immune response representing some of the most important challenges. The intrinsic technical advantages of optogenetics compared to pharmacological and unspecific electrical stimulation have already provided important insights into cardiac arrhythmia mechanisms—and we envision further exciting insights to come.
References
Crick FHC. Thinking about the brain. Sci Am. 1979;241:219–32.
Oesterhelt D, Stoeckenius W. Rhodopsin-like protein from the purple membrane of Halobacterium halobium. Nature. 1971;233:149–52.
Matsuno-Yagi A, Mukohata Y. Two possible roles of bacteriorhodopsin; a comparative study of strains of Halobacterium halobium differing in pigmentation. Biochem Biophys Res Commun. 1977;78:237–43.
Khorana HG, Knox BE, Nasi E, Swanson R, Thompson DA. Expression of a bovine rhodopsin gene in Xenopus oocytes: demonstration of light-dependent ionic currents. Proc Natl Acad Sci U S A. 1988;85:7917–21.
Miesenböck G, de Angelis DA, Rothman JE. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature. 1998;394:192–5.
Miyawaki A, Llopis J, Heim R, McCaffery JM, Adams JA, et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature. 1997;388:882–7.
Siegel MS, Isacoff EY. A genetically encoded optical probe of membrane voltage. Neuron. 1997;19:735–41.
Zemelman BV, Lee GA, Ng M, Miesenböck G. Selective photostimulation of genetically chARGed neurons. Neuron. 2002;33:15–22.
Ishizuka T, Kakuda M, Araki R, Yawo H. Kinetic evaluation of photosensitivity in genetically engineered neurons expressing green algae light-gated channels. Neurosci Res. 2006;54:85–94.
Boyden ES, Zhang F, Bamberg E, Nagel G, Deisseroth K. Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci. 2005;8:1263–8.
Li X, Gutierrez DV, Hanson MG, Han J, Mark MD, et al. Fast noninvasive activation and inhibition of neural and network activity by vertebrate rhodopsin and green algae channelrhodopsin. Proc Natl Acad Sci U S A. 2005;102:17816–21.
Nagel G, Brauner M, Liewald JF, Adeishvili N, Bamberg E, et al. Light activation of channelrhodopsin-2 in excitable cells of Caenorhabditis elegans triggers rapid behavioral responses. Curr Biol. 2005;15:2279–84.
Tallini YN, Ohkura M, Choi B-R, Ji G, Imoto K, et al. Imaging cellular signals in the heart in vivo: cardiac expression of the high-signal Ca2+ indicator GCaMP2. Proc Natl Acad Sci U S A. 2006;103:4753–8.
Arrenberg AB, Stainier DYR, Baier H, Huisken J. Optogenetic control of cardiac function. Science. 2010;330:971–4.
Bruegmann T, Malan D, Hesse M, Beiert T, Fuegemann CJ, et al. Optogenetic control of heart muscle in vitro and in vivo. Nat Methods. 2010;7:897–900.
Rost BR, Schneider-Warme F, Schmitz D, Hegemann P. Optogenetic tools for subcellular applications in neuroscience. Neuron. 2017;96:572–603.
Park SA, Lee S-R, Tung L, Yue DT. Optical mapping of optogenetically shaped cardiac action potentials. Sci Rep. 2014;4:6125.
Nussinovitch U, Shinnawi R, Gepstein L. Modulation of cardiac tissue electrophysiological properties with light-sensitive proteins. Cardiovasc Res. 2014;102:176–87.
Nussinovitch U, Gepstein L. Optogenetics for suppression of cardiac electrical activity in human and rat cardiomyocyte cultures. Neurophotonics. 2015;2:031204.
Funken M, Malan D, Sasse P, Bruegmann T. Optogenetic hyperpolarization of cardiomyocytes terminates ventricular arrhythmia. Front Physiol. 2019;10:1–7.
Wietek J, Beltramo R, Scanziani M, Hegemann P, Oertner TG, et al. An improved chloride-conducting channelrhodopsin for light-induced inhibition of neuronal activity in vivo. Sci Rep. 2015;5:14807.
Berndt A, Lee SY, Wietek J, Ramakrishnan C, Steinberg EE, et al. Structural foundations of optogenetics: determinants of channelrhodopsin ion selectivity. Proc Natl Acad Sci U S A. 2016;113:822–9.
Govorunova EG, Sineshchekov OA, Janz R, Liu X, Spudich JL. Natural light-gated anion channels: a family of microbial rhodopsins for advanced optogenetics. Science. 2015;349:647–50.
Schneider F, Gradmann D, Hegemann P. Ion selectivity and competition in channelrhodopsins. Biophys J. 2013;105:91–100.
Lin JY, Knutsen PM, Muller A, Kleinfeld D, Tsien RY. ReaChR: a red-shifted variant of channelrhodopsin enables deep transcranial optogenetic excitation. Nat Neurosci. 2013;16:1499–508.
Entcheva E, Kay MW. Cardiac optogenetics : a decade of enlightenment. Nat Rev Cardiol. 2021;18:349–67.
Govorunova EG, Cunha SR, Sineshchekov OA, Spudich JL. Anion channelrhodopsins for inhibitory cardiac optogenetics. Sci Rep. 2016;6:1–7.
Kopton RA, Baillie JS, Rafferty SA, Moss R, Zgierski-Johnston CM, et al. Cardiac electrophysiological effects of light-activated chloride channels. Front Physiol. 2018;9:1806.
Alberio L, Locarno A, Saponaro A, Romano E, Bercier V, et al. A light-gated potassium channel for sustained neuronal inhibition. Nat Methods. 2018;15:969–76.
Bernal Sierra YA, Rost BR, Pofahl M, Fernandes AM, Kopton RA, et al. Potassium channel-based optogenetic silencing. Nat Commun. 2018;9:4611.
Govorunova EG, Gou Y, Sineshchekov OA, Li H, Lu X et al. Kalium channelrhodopsins are natural light-gated potassium channels that mediate optogenetic inhibition. Nat Neurosci 2022; 25: 967–974.
Vierock J, Peter E, Grimm C, Rozenberg A, Chen IW et al. WiChR, a highly potassium-selective channelrhodopsin for lowlight one- and two-photon inhibition of excitable cells. Sci Adv. 2022; 8: eadd7729.
Ernst OP, Lodowski DT, Elstner M, Hegemann P, Brown LS, et al. Microbial and animal rhodopsins: structures, functions, and molecular mechanisms. Chem Rev. 2014;114:126–63.
Koyanagi M, Terakita A. Diversity of animal opsin-based pigments and their optogenetic potential. Biochim Biophys Acta Bioenerg. 2014;1837:710–6.
Makowka P, Bruegmann T, Dusend V, Malan D, Beiert T, et al. Optogenetic stimulation of Gs-signaling in the heart with high spatio-temporal precision. Nat Commun. 2019;10:1281.
Beiert T, Bruegmann T, Sasse P. Optogenetic activation of Gq signalling modulates pacemaker activity of cardiomyocytes. Cardiovasc Res. 2014;102:507–16.
Tennigkeit SA, Karapinar R, Rudack T, Dreier MA, Althoff P, et al. Design of an ultrafast G protein switch based on a mouse melanopsin variant. Chembiochem. 2019;20:1766–71.
Spoida K, Eickelbeck D, Karapinar R, Eckhardt T, Mark MD, et al. Melanopsin variants as intrinsic optogenetic on and off switches for transient versus sustained activation of G protein pathways. Curr Biol. 2016;26:1206–12.
Gutierrez DV, Mark MD, Masseck O, Maejima T, Kuckelsberg D, et al. Optogenetic control of motor coordination by Gi/o protein-coupled vertebrate rhodopsin in cerebellar Purkinje cells. J Biol Chem. 2011;286:25848–58.
Masseck OA, Spoida K, Dalkara D, Maejima T, Rubelowski JM, et al. Vertebrate cone opsins enable sustained and highly sensitive rapid control of Gi/o signaling in anxiety circuitry. Neuron. 2014;81:1263–73.
Wagdi A, Malan D, Sathyanarayanan U, Beauchamp JS, Vogt M et al. Selective optogenetic control of Gq signaling using human Neuropsin. Nat Commun. 2022; 13: 1765
Kaestner L, Scholz A, Tian Q, Ruppenthal S, Tabellion W, et al. Genetically encoded Ca2+ indicators in cardiac myocytes. Circ Res. 2014;114:1623–39.
Roell W, Lewalter T, Sasse P, Tallini YN, Choi BR, et al. Engraftment of connexin 43-expressing cells prevents post-infarct arrhythmia. Nature. 2007;450:819–24.
Dana H, Mohar B, Sun Y, Narayan S, Gordus A, et al. Sensitive red protein calcium indicators for imaging neural activity. elife. 2016;5:e12727.
Qian Y, Cosio DMO, Piatkevich KD, Aufmkolk S, Su WC, et al. Improved genetically encoded near-infrared fluorescent calcium ion indicators for in vivo imaging. PLoS Biol. 2020;18:e3000965.
Chen T-W, Wardill TJ, Sun Y, Pulver SR, Renninger SL, et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature. 2013;499:295–300.
Dana H, Sun Y, Mohar B, Hulse BK, Kerlin AM, et al. High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nat Methods. 2019;16:649–57.
Suzuki J, Kanemaru K, Ishii K, Ohkura M, Okubo Y, et al. Imaging intraorganellar Ca2+ at subcellular resolution using CEPIA. Nat Commun. 2014;5:1–13.
Henderson MJ, Baldwin HA, Werley CA, Boccardo S, Whitaker LR, et al. A low affinity GCaMP3 variant (GCaMPer) for imaging the endoplasmic reticulum calcium store. PLoS One. 2015;10:1–17.
de Juan-Sanz J, Holt GT, Schreiter ER, de Juan F, Kim DS, et al. Axonal endoplasmic reticulum Ca2+ content controls release probability in CNS nerve terminals. Neuron. 2017;93:867–881.e6.
Bando Y, Grimm C, Cornejo VH, Yuste R. Genetic voltage indicators. BMC Biol. 2019;17:71.
Kaestner L, Tian Q, Kaiser E, Xian W, Müller A, et al. Genetically encoded voltage indicators in circulation research. Int J Mol Sci. 2015;16:21626–42.
Tsutsui H, Higashijima S, Miyawaki A, Okamura Y. Visualizing voltage dynamics in zebrafish heart. J Physiol. 2010;588:2017–21.
Chang LML, de Boer TP, Mutoh H, Raad N, Richter C, et al. Sensing cardiac electrical activity with a cardiac myocyte-targeted optogenetic voltage indicator. Circ Res. 2015;117:401–12.
Quinn TA, Camelliti P, Rog-Zielinska EA, Siedlecka U, Poggioli T, et al. Electrotonic coupling of excitable and nonexcitable cells in the heart revealed by optogenetics. Proc Natl Acad Sci U S A. 2016;113:14852–7.
Leyton-Mange JS, Mills RW, Macri VS, Jang MY, Butte FN, et al. Rapid cellular phenotyping of human pluripotent stem cell-derived cardiomyocytes using a genetically encoded fluorescent voltage sensor. Stem Cell Reports. 2014;2:163–70.
Monakhov MV, Matlashov ME, Colavita M, Song C, Shcherbakova DM, et al. Screening and cellular characterization of genetically encoded voltage indicators based on near-infrared fluorescent proteins. ACS Chem Neurosci. 2020;11:3523–31.
Zaglia T, Pianca N, Borile G, Da Broi F, Richter C, et al. Optogenetic determination of the myocardial requirements for extrasystoles by cell type-specific targeting of ChannelRhodopsin-2. Proc Natl Acad Sci U S A. 2015;112:E4495–504.
Wengrowski AM, Wang X, Tapa S, Posnack NG, Mendelowitz D, et al. Optogenetic release of norepinephrine from cardiac sympathetic neurons alters mechanical and electrical function. Cardiovasc Res. 2015;105:143–50.
Prando V, Da Broi F, Franzoso M, Plazzo AP, Pianca N, et al. Dynamics of neuro-effector coupling at ‘cardiac sympathetic’ synapses. J Physiol. 2018;596:2055–75.
Wang Y, Lin WK, Crawford W, Ni H, Bolton EL, et al. Optogenetic control of heart rhythm by selective timulation of cardiomyocytes derived from Pnmt+ cells in murine heart. Sci Rep. 2017;7:40687.
Fernández MC, Kopton RA, Simon-Chica A, Madl J, Hilgendorf I, et al. Channelrhodopsins for cell-type specific illumination of cardiac electrophysiology. Methods Mol Biol. 2021;2191:287–307.
Hulsmans M, Clauss S, Xiao L, Aguirre AD, King KR, et al. Macrophages facilitate electrical conduction in the heart. Cell. 2017;169:510–522.e20.
Vogt CC, Bruegmann T, Malan D, Ottersbach A, Roell W, et al. Systemic gene transfer enables optogenetic pacing of mouse hearts. Cardiovasc Res. 2015;106:338–43.
Nyns ECA, Kip A, Bart CI, Plomp JJ, Zeppenfeld K, et al. Optogenetic termination of ventricular arrhythmias in the whole heart: towards biological cardiac rhythm management. Eur Heart J. 2017;38:2132–6.
Nussinovitch U, Gepstein L. Optogenetics for in vivo cardiac pacing and resynchronization therapies. Nat Biotechnol. 2015;33:750–4.
Jia Z, Valiunas V, Lu Z, Bien H, Liu H, Wang HZ, et al. Stimulating cardiac muscle by light: cardiac optogenetics by cell delivery. Circ Arrhythm Electrophysiol. 2001;4:753–60.
Boyle PM, Karathanos TV, Trayanova NA. “Beauty is a light in the heart”: the transformative potential of optogenetics for clinical applications in cardiovascular medicine. Trends Cardiovasc Med. 2015;25:73–81.
Bruegmann T, Sasse P. Optogenetic cardiac pacemakers: science or fiction? Trends Cardiovasc Med. 2015;25:82–3.
Johnston CM, Rog-Zielinska EA, Wülfers EM, Houwaart T, Siedlecka U, et al. Optogenetic targeting of cardiac myocytes and non-myocytes: tools, challenges and utility. Prog Biophys Mol Biol. 2017;130:140–9.
Prabhakar A, Vujovic D, Cui L, Olson W, Luo W. Leaky expression of channelrhodopsin-2 (ChR2) in Ai32 mouse lines. PLoS One. 2019;14:e0213326.
Li Q, Ni RR, Hong H, Gho KY, Rossi M, et al. Electrophysiological properties and viability of neonatal rat ventricular myocyte cultures with inducible ChR2 expression. Sci Rep. 2017;7:1531.
Richter C, Bruegmann T. No light without the dark: perspectives and hindrances for translation of cardiac optogenetics. Prog Biophys Mol Biol. 2020;154:39–50.
Maimon BE, Diaz M, Revol ECM, Schneider AM, Leaker B, et al. Optogenetic peripheral nerve immunogenicity Sci Rep. 2018;8:14076.
Richter C, Christoph J, Lehnart SE, Luther S. Optogenetic light crafting tools for the control of cardiac arrhythmias. Methods Mol Biol. 2016;1408:293–302.
Crocini C, Ferrantini C, Coppini R, Scardigli M, Yan P, et al. Optogenetics design of mechanistically-based stimulation patterns for cardiac defibrillation. Sci Rep. 2016;6:35628.
Scardigli M, Müllenbroich C, Margoni E, Cannazzaro S, Crocini C, et al. Real-time optical manipulation of cardiac conduction in intact hearts. J Physiol. 2018;596:3841–58.
Burton RAB, Klimas A, Ambrosi CM, Tomek J, Corbett A, et al. Optical control of excitation waves in cardiac tissue. Nat Photonics. 2015;9:813–6.
Bruegmann T, Boyle PM, Vogt CC, Karathanos TV, Arevalo HJ, et al. Optogenetic defibrillation terminates ventricular arrhythmia in mouse hearts and human simulations. J Clin Invest. 2016;126:3894–904.
Pedersen SS, van den Broek KC, Theuns DA, Erdman RA, Alings M, et al. Risk of chronic anxiety in implantable defibrillator patients: a multi-center study. Int J Cardiol. 2011;147:420–3.
Lemme M, Braren I, Prondzynski M, Aksehirlioglu B, Ulmer BM, et al. Chronic intermittent tachypacing by an optogenetic approach induces arrhythmia vulnerability in human engineered heart tissue. Cardiovasc Res. 2020;116:1487–99.
Lemoine MD, Lemme M, Ulmer BM, Braren I, Krasemann S, et al. Intermittent optogenetic tachypacing of atrial engineered heart tissue induces only limited electrical remodelling. J Cardiovasc Pharmacol. 2021;77:291–9.
Bruegmann T, Beiert T, Vogt CC, Schrickel JW, Sasse P. Optogenetic termination of atrial fibrillation in mice. Cardiovasc Res. 2017;114:713–23.
Nyns ECA, Poelma RH, Volkers L, Plomp JJ, Bart CI, et al. An automated hybrid bioelectronic system for autogenous restoration of sinus rhythm in atrial fibrillation. Sci Transl Med. 2019;11(481):eaau6447.
Watanabe M, Feola I, Majumder R, Angsangthong W, Teplenin AS, et al. Optogenetic manipulation of anatomical re-entry by light-guided generation of a reversible local conduction block. Cardiovasc Res. 2017;113:354–66.
Quiñonez Uribe RA, Luther S, Diaz-Maue L, Richter C. Energy-reduced arrhythmia termination using global photostimulation in optogenetic murine hearts. Front Physiol. 2018;9:1651.
Hussaini S, Venkatesan V, Biasci V, Romero Sepúlveda JM, Quiñonez Uribe RA, et al. Drift and termination of spiral waves in optogenetically modified cardiac tissue at sub-threshold illumination. elife. 2021;10:e59954. https://doi.org/10.7554/eLife.59954.
Li X, Zhang R, Zhao B, Lossin C, Cao Z. Cardiotoxicity screening: a review of rapid-throughput in vitro approaches. Arch Toxicol. 2016;90:1803–16.
Kannankeril P, Roden DM, Darbar D. Drug-induced long QT syndrome. Pharmacol Rev. 2010;62:760–81.
Konstantopoulou A, Tsikrikas S, Asvestas D, Korantzopoulos P, Letsas KP. Mechanisms of drug-induced proarrhythmia in clinical practice. World J Cardiol. 2013;5:175–85.
Meyer T, Sartipy P, Blind F, Leisgen C, Guenther E. New cell models and assays in cardiac safety profiling. Expert Opin Drug Metab Toxicol. 2007;3:507–17.
Klimas A, Ambrosi CM, Yu J, Williams JC, Bien H, et al. OptoDyCE as an automated system for high-throughput all-optical dynamic cardiac electrophysiology. Nat Commun. 2016;7:11542.
Dempsey GT, Chaudhary KW, Atwater N, Nguyen C, Brown BS, et al. Cardiotoxicity screening with simultaneous optogenetic pacing, voltage imaging and calcium imaging. J Pharmacol Toxicol Methods. 2016;81:240–50.
Entcheva E, Bub G. All-optical control of cardiac excitation: combined high-resolution optogenetic actuation and optical mapping. J Physiol. 2016;594:2503–10.
O’Shea C, Holmes AP, Winter J, Correia J, Ou X, et al. Cardiac optogenetics and optical mapping – overcoming spectral congestion in all-optical cardiac electrophysiology. Front Physiol. 2019;10:182.
Boyle PM, Yu J, Klimas A, Williams JC, Trayanova NA, et al. OptoGap is an optogenetics-enabled assay for quantification of cell-cell coupling in multicellular cardiac tissue. Sci Rep. 2021;11:9310.
Klimas A, Ortiz G, Boggess SC, Miller EW, Entcheva E. Multimodal on-axis platform for all-optical electrophysiology with near-infrared probes in human stem-cell-derived cardiomyocytes. Prog Biophys Mol Biol. 2020;154:62–70.
Yakushenko A, Gong Z, Maybeck V, Hofmann B, Gu E, et al. On-chip optical stimulation and electrical recording from cells. J Biomed Opt. 2013;18:111402.
Lapp H, Bruegmann T, Malan D, Friedrichs S, Kilgus C, et al. Frequency-dependent drug screening using optogenetic stimulation of human iPSC-derived cardiomyocytes. Sci Rep. 2017;7:9629.
Rehnelt S, Malan D, Juhasz K, Wolters B, Doerr L, et al. Frequency-dependent multi-well cardiotoxicity screening enabled by optogenetic stimulation. Int J Mol Sci. 2017;18:2634.
Bifulco SF, Akoum N, Boyle PM. Translational applications of computational modelling for patients with cardiac arrhythmias. Heart 2020, Dec 10:heartjnl-2020-316854. https://doi.org/10.1136/heartjnl-2020-316854.
Williams JC, Xu J, Lu Z, Klimas A, Chen X, et al. Computational optogenetics: empirically-derived voltage- and light-sensitive channelrhodopsin-2 model. PLoS Comput Biol. 2013;9(9):e1003220.
Boyle PM, Williams JC, Ambrosi CM, Entcheva E, Trayanova NA. A comprehensive multiscale framework for simulating optogenetics in the heart. Nat Commun. 2013;4:2370.
Boyle PM, Karathanos TV, Entcheva E, Trayanova NA. Computational modeling of cardiac optogenetics: methodology overview and review of findings from simulations. Comput Biol Med. 2015;65:200–8.
Karathanos TV, Boyle PM, Trayanova NA. Optogenetics-enabled dynamic modulation of action potential duration in atrial tissue: feasibility of a novel therapeutic approach. Europace. 2014;16(Suppl 4):iv69–iv76:iv69.
Boyle PM, Murphy MJ, Karathanos TV, Zahid S, Blake RC 3rd, et al. Termination of re-entrant atrial tachycardia via optogenetic stimulation with optimized spatial targeting: insights from computational models. J Physiol. 2018;596:181–96.
Karathanos TV, Bayer JD, Wang D, Boyle PM, Trayanova NA. Opsin spectral sensitivity determines the effectiveness of optogenetic termination of ventricular fibrillation in the human heart: a simulation study. J Physiol. 2016;594:6879–91.
Ochs AR, Karathanos TV, Trayanova NA, Boyle PM. Optogenetic stimulation using anion channelrhodopsin (GtACR1) facilitates termination of reentrant arrhythmias with low light energy requirements: a computational study. Front Physiol. 2021;12:718622. https://doi.org/10.3389/fphys.2021.718622.
Ambrosi CM, Boyle PM, Chen K, Trayanova NA, Entcheva E. Optogenetics-enabled assessment of viral gene and cell therapy for restoration of cardiac excitability. Sci Rep. 2015;5:17350.
Bishop MJ, Rodriguez B, Eason J, Whiteley JP, Trayanova N, et al. Synthesis of voltage-sensitive optical signals: application to panoramic optical mapping. Biophys J. 2006;90:2938–45.
Bishop MJ, Rodriguez B, Qu F, Efimov IR, Gavaghan DJ, et al. The role of photon scattering in optical signal distortion during arrhythmia and defibrillation. Biophys J. 2007;93:3714–26.
Bishop MJ, Rowley A, Rodriguez B, Plank G, Gavaghan DJ, et al. The role of photon scattering in voltage-calcium fluorescent recordings of ventricular fibrillation. Biophys J. 2011;101:307–18.
Kohl P, Gourdie RG. Fibroblast-myocyte electrotonic coupling: does it occur in native cardiac tissue? J Mol Cell Cardiol. 2014;70:37–46.
Funken M, Bruegmann T, Sasse P. Selective optogenetic stimulation of fibroblasts enables quantification of hetero-cellular coupling to cardiomyocytes in a three-dimensional model of heart tissue. Europace. 2020;22:1590–9.
De SSA, Moyle S, Buccarello A, Dellenbach C, Kucera JP, et al. The role of membrane capacitance in cardiac impulse conduction: an optogenetic study with non-excitable cells coupled to cardiomyocytes. Front Physiol. 2020;11:194.
Kostecki GM, Shi Y, Chen CS, Reich DH, Entcheva E, et al. Optogenetic current in myofibroblasts acutely alters electrophysiology and conduction of co-cultured cardiomyocytes. Sci Rep. 2021;11:4430.
Rubart M, Tao W, Lu X-L, Conway SJ, Reuter SP, et al. Electrical coupling between ventricular myocytes and myofibroblasts in the infarcted mouse heart. Cardiovasc Res. 2018;114:389–400.
Yu L, Zhou L, Cao G, Po SS, Huang B, et al. Optogenetic modulation of cardiac sympathetic nerve activity to prevent ventricular arrhythmias. J Am Coll Cardiol. 2017;70:2778–90.
Moreno A, Endicott K, Skancke M, Dwyer MK, Brennan J, et al. Sudden heart rate reduction upon optogenetic release of acetylcholine from cardiac parasympathetic neurons in perfused hearts. Front Physiol. 2019;10:16.
Rajendran PS, Challis RC, Fowlkes CC, Hanna P, Tompkins JD, et al. Identification of peripheral neural circuits that regulate heart rate using optogenetic and viral vector strategies. Nat Commun. 2019;10:1944.
Acknowledgements
We thank all members of the Institute for Experimental Cardiovascular Medicine (IEKM) for the critical discussion of the manuscript. This work was supported by the German Centre for Cardiovascular Research (DZHK) and the German Research Foundation (DFG: Priority Program 1926 [#315212873;#315193289], CRC1002 [#193793266], CRC1425 [#4226818459], and the Emmy-Noether Programme [#412853334]). FSW is an associate member of the excellence cluster Centre for Integrated Biological Signalling Studies (CIBSS).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bruegmann, T., Boyle, P.M., Schneider-Warme, F. (2023). Enlightening Cardiac Arrhythmia with Optogenetics. In: Tripathi, O.N., Quinn, T.A., Ravens, U. (eds) Heart Rate and Rhythm. Springer, Cham. https://doi.org/10.1007/978-3-031-33588-4_17
Download citation
DOI: https://doi.org/10.1007/978-3-031-33588-4_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-33587-7
Online ISBN: 978-3-031-33588-4
eBook Packages: MedicineMedicine (R0)