Abstract
The detection of Caenorhabditis elegans (C. elegans) is a complex problem due to the variety of poses they can adopt, the aggregations and the problems of dirt accumulation on the plates. Artificial neural networks have achieved great results in detection tasks in recent years.
In this article, two detection network architectures have been compared: YOLOv5s and Faster R-CNN. Accuracy and computational cost have been used as comparison criteria. The results show that both models perform similar in terms of accuracy but the smaller version of the YOLOv5 network (YOLOv5s) is able to obtain better results in computational cost.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The nematode Caenorhabditis elegans (C. elegans) has been used as an important animal model for many years [2]. In advanced ages, just like humans, they present some pathologies, physical deterioration, mobility loss, etc. It has a short life (approximately 3 weeks) and a length of 1 mm, which makes it easy to handle and grow. They can be stored in large quantities and small spaces (Petri dishes) [15]. These characteristics allow large-scale assays to be performed economically and make this nematode an ideal model for research into neurodegenerative diseases and the development of new drugs.
Despite these advantages, handling and observation of these nematodes are costly and require qualified technicians. For this reason, computer vision techniques allow these processes to be automated, increasing productivity and precision in the development of assays.
The detection of C. elegans is the first task in automatic image processing applications such as monitoring, tracking [9,10,11,12], lifespan [6, 18] and healthspan [4, 7, 13]. To date, many methods perform segmentation pre-processing and search for worm characteristics in order to filter worms (object of interest) from non-worms (noise), which takes an additional computational cost.
In recent years, image processing methods based on artificial neural networks have gained much popularity in these C. elegans applications, as they have great advantages compared to traditional computer vision methods based on feature extraction (using segmentation, edge detection, textures). Feature extraction is a costly process as it requires experience and trial and error to select the optimal ones. In addition, it is difficult to adjust the parameters of the image processing algorithms to work on different images.
Detection methods based on artificial neural networks have achieved great results, outperforming classical methods. The main detection methods based on deep learning can be classified into two groups: one-stage methods (YOLO, SSD, RetinaNet) and two-stage methods (Faster R-CNN, RFCN, Mask R-CNN).
These are some of the latest studies that show the advantages of artificial neural networks in the detection of C. elegans:
-
[1] implement Faster R-CNN object detection models for detection and location of C. elegans in different conditions. This study demonstrates some of the advantages of Faster R-CNN algorithm to identify, count and track behaviors in comparison to traditional methods in terms of speed and flexibility. These traditional image processing methods, such as background subtraction or using morphological features, are more sensitive to changes in the environmental conditions due to complex experimental setups, resulting in low accuracy to detect objects of interest.
-
[5] present a low-cost system including a DIY microscope to capture low-resolution images. They use the framework Mask R-CNN to detect, locate and classify C. elegans with high accuracy, rather than traditional methods such as human counting and analyzing high-resolution microscope images, which result in longer times and lower efficiency. They also generate a great number of masks by a preprocessing method that includes visual inspection, since the Mask R-CNN algorithm needs them as an input.
In summary, the main goals of this study are:
2 Materials and Method
2.1 C. elegans Strains and Culture Conditions
The C. elegans that were used to perform detection assays had the following features: strains N2, Bristol (wild-type) and CB1370, daf-2 (e1370). They were provided by the Caenorhabditis Genetics Centre at the University of Minnesota. All worms were age-synchronized and pipetted onto Nematode Growth Medium (NGM) in 55 mm Petri plates, where the temperature was kept at 20 \(^\circ \)C.
The worms were fed with strain OP50 of Escherichia coli, seeded in the middle of the plate to avoid occluded wall zones, as C. elegans tend to stay on the lawn. To reduce fungal contamination, fungizone was added to the plates [23], and FUdR (0.2 mM) was added to the plates to decrease the probability of reproduction.
In the laboratory, the operator had to follow a specific methodology to obtain the images: (1) Extract the plates from the incubator to put them in the acquisition system; (2) Examine the lid, before initiating the capture, to check there is no condensation and wipe it if detected; (3) Capture a sequence of 30 images per plate at 1 fps and return the plates to the incubator. This method was designed to prevent condensation in the cover.
2.2 Image Capture Method
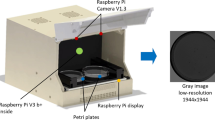
Images were captured using the monitoring system developed in [19], that uses the active backlight illumination method proposed in [20]. Active backlighting technique has been shown to be effective for low-resolution C. elegans applications for both the mentioned [20] and [17] capture systems. The monitoring system is based on an RGB Raspberry Pi camera v1.3 (OmniVision OV5647, with a resolution of 2592 \(\times \) 1944 pixels, a pixel size of 1.4\(\,\times \,\)1.4 \(\upmu \)m, a view field of 53.50\(^{\underline{\textrm{o}}} \times 41.41^\circ \), optical size of 1/4", and focal ratio of 2.9) placed in front of the lighting system (a 7" Raspberry Pi display 800 \(\times \) 480 at a resolution at 60 fps, 24 bit RGB colour) and the inspected plate in between both devices. A Raspberry Pi 3 was used as a processor to control lighting. The distance between the camera and the Petri plate was sufficient to enable a complete picture of the Petri plate, and the camera lens was focused at this distance (about 77 mm). The system captured images of 1944\(\,\times \,\)1944 pixels at a frequency 1 Hz. A graphic of the capture system is shown in Fig. 1.
The code, components and the guidelines to build the monitoring system and the assembly description can be found in the repository https://github.com/JCPuchalt/SiViS.
2.3 Dataset Generation and Labeling
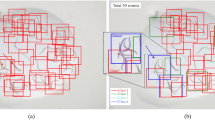
The dataset is composed of 1901 images captured from 106 test plates with the SiViS system [20]. Each plate contains between 10 and 15 nematodes. These images were manually labeled by marking the bounding boxes of each nematode.
The dataset was divided into three subsets using the following proportions: 70% for training, 10% for validation and 20% for test. Number of images and labels in each subset are shown Table 1.
2.4 Detection Method
In this article it was decided to test with the best-known detection methods based on artificial neural networks: YOLOv5 and Faster R-CNN.
Faster RCNN is an architecture proposed in [22]. It consists of three parts: (1) a convolutional neural network that extracts the image features; (2) a Region Proposal Network (RPN) in charge of obtaining the candidate bounding boxes and (3) a network in charge of performing the classification and regression of the bounding box coordinates.
YOLOv5 is an object detection algorithm created by Glenn Jocher in 2020. Considering previous models of the YOLO series, YOLOv5 offers higher accuracy, faster calculated speed and smaller size. Among the YOLOv5 models, YOLOv5s works the fastest, although the average precision is the lowest. The YOLOv5 network consists in three main parts: Backbone, Neck and Head. Backbone gathers and produces picture characteristics on various image granularities when the image is supplied. The picture characteristics are then stitched and transmitted to the prediction layer by Neck, where the image features are predicted by Head to generate bounding boxes and predicted categories [3].
2.5 Training and Evaluation Method
The networks were implemented and trained using the Pytorch deep learning framework [16] on a computer Gigabyte Technology Z390 AORUS PRO machine, Intel(R) Core (TM) i9-9900KF CPU @ 3.60 GHz x16 with 32 GB of RAM, and NVIDIA GeForce RTX 2080 Ti graphics card with 4352 Cuda cores, Ubuntu 19.04 64bits operating system.
Faster R-CNN. In this work the Faster R-CNN model architecture was used with a ResNet-50-FPN backbone pre-trained with COCO dataset [14] (Fig. 2).
The network was trained for 150 epochs with a batch size of 6 samples using the implementation of https://pytorch.org/tutorials/intermediate/torchvision_tutorial.html. A Reduce Learning Rate on Plateau scheduler with a patience of 3 epochs and a factor of 0.1 was used. The optimizer used was SGD with momentum 0.9 and weight decay 0.0005. Vertical flip (0.5), HSV augmentation (hue = 0.015, saturation = 0.7, value = 0.4), image scale (0.5) and translation (0.1) were used as data augmentation techniques.
Image pipeline through Fast R-CNN architecture. The blocks of the Fast R-CNN architecture were made using the PlotNeuralNet tool [8].
YOLOv5. To train YOLOv5 we used the implementation of (https://github.com/ultralytics/yolov5) in its YOLOv5s version, which is the smallest and fastest (Fig. 3).
Image pipeline through YOLOv5 architecture. The blocks of the YOLOv5 architecture were made using the PlotNeuralNet tool [8].
We used the weights pretrained on the COCO dataset. It was trained with a batch size of 6 samples for 150 epochs using the SGD optimizer with momentum 0.937 and weight decay 0.0005. LambdaLR Learning Rate scheduler was used with the configuration proposed in the original implementation and other data augmentations, such as Vertical flip (0.5), HSV augmentation (hue = 0.015, saturation = 0.7, value = 0.4), image scale (0.5) and translation (0.1) were used.
The YOLOv5 model needed an image size multiple of 32 and, consequently, this model resized automatically the images to 1952 pixels. To make both algorithms work in the same image conditions, the images were resized with the same scale factor for Faster R-CNN model. Before this, in this model’s dataloader, a function to crop and resize the images was implemented, since Faster R-CNN model needed an image size between 800 and 1333 pixels.
2.6 Evaluation Methods (Metrics)
To compare YOLOv5 and Faster R-CNN it is necessary to use some metrics that can be applied to both models, so the comparison is fair and adequate. The main metrics that will be used are computational cost and mean Average Precision (mAP), which is obtained from other metrics, precision and recall.
To decide whether a prediction is correct, the IoU is used, which measures the degree of overlap between the real box delimiting the object and the one predicted by the network. If the IoU value exceeds a threshold, it is considered True Positive (TP), otherwise it is considered False Positive (FP). If the network does not find an object, it is considered False Negative (FN).
Precision: It represents the relation between the correct detections and all the detections made by the model (Eq 1).
Recall: It shows the ability of the model to make correct detections out of all ground true bounding boxes (Eq 2).
Average Precision (AP): It is calculated as the area under the precision-recall curve, thus summarizing the information of the curve in one parameter (Eq 3).
Mean Average Precision (mAP): This is obtained calculating the mean of APs over all IoU tresholds. In this case, the range of IoU tresholds used is [0.5; 0.95] (Eq. 4).
where N is the total number of IoU thresholds and AP(n) is the average precision for each IoU threshold value.
Computational Cost: It measures the amount of resources a neural network uses in a process, which determines the amount of time the process lasts. In this case, time per prediction will be measured to compare the computational costs in inference, which allows to determine how both models would work for possible real-time applications. For this reason, this metric will be measured in seconds.
3 Experimental Results
This section presents the results obtained after training with the YOLOv5s and Faster R-CNN networks. The results were obtained with the best trained models in the three subsets (train, validation, test, Table 2, 3, 4 respectively).
In Figs. 4, 5, 6 we can find examples of true positives, false positives and false negatives respectively obtained when predicting with the models.
4 Discussion
The experiments carried out in this study have shown that, as expected, due to the greater number of parameters of the Faster R-CNN model used (backbone resnet50) with respect to YOLOv5s, the computational cost is higher, as reflected in the inference times of both models. Generally, in C. elegans image acquisition, the images are captured at a frequency 1 Hz, maximum 15 Hz, due to the low speed of movement of the nematodes. It has thus been shown that both detection techniques could be used for real-time monitoring of C. elegans. This would be useful for healthspan assays in which the movements of the nematodes are analysed.
In terms of network performance, the results of the experiments show that they perform similarly, achieving a mAP value of around 93–94% for an IoU treshold of 0.5. As explained in section 2.6 Evaluation methods (Metrics), this metric is quite representative, as it considers precision and recall values. The YOLOv5s model obtains a better mAP result for IoU treshold [0.5–0.95].
5 Conclusion and Future Work
Finally, it can be concluded that the YOLOv5s model performs better in this application of C. elegans detection, as it achieves similar accuracy results to those obtained with Faster R-CNN (accuracy values of 93.2% and 94.4% respectively for our dataset) but in less time and, consequently, with a lower computational cost.
As future work, the available dataset could be labelled by obtaining the nematode masks. With this new dataset, a Mask R-CNN network could be trained, which would have the advantage over YOLO of obtaining a segmentation. Another option to avoid the cost of labelling would be to use synthetic images to train the network.
References
Bates, K., Le, K.N., Lu, H.: Deep learning for robust and flexible tracking in behavioral studies for C. elegans. PLOS Comput. Biol. 18(4), e1009942 (2022)
Biron, D., Haspel, G. (eds.): C. elegans. MMB, vol. 1327. Humana Press, Totowa (2015). https://doi.org/10.1007/978-1-4939-2842-2
Chen, Z., et al.: Plant disease recognition model based on improved YOLOv5. Agronomy 12(2), 365 (2022)
Di Rosa, G., et al.: Healthspan enhancement by olive polyphenols in C. elegans wild type and Parkinson’s models. Int. J. Mol. Sci. 21(11) (2020). https://doi.org/10.3390/ijms21113893
Fudickar, S., Nustede, E.J., Dreyer, E., Bornhorst, J.: Mask R-CNN based C. elegans detection with a DIY microscope. Biosensors 11(8), 257 (2021)
García Garví, A., Puchalt, J.C., Layana Castro, P.E., Navarro Moya, F., Sánchez-Salmerón, A.J.: Towards lifespan automation for Caenorhabditis elegans based on deep learning: analysing convolutional and recurrent neural networks for dead or live classification. Sensors 21(14) (2021). https://doi.org/10.3390/s21144943
Hahm, J.H., et al.: C. elegans maximum velocity correlates with healthspan and is maintained in worms with an insulin receptor mutation. Nat. Commun. 6(1), 1–7 (2015). https://doi.org/10.1038/ncomms9919
Iqbal, H.: Harisiqbal88/plotneuralnet v1.0.0 (2018). code https://github.com/HarisIqbal88/PlotNeuralNet
Javer, A., et al.: An open-source platform for analyzing and sharing worm-behavior data. Nat. Methods 15 (2018). https://doi.org/10.1038/s41592-018-0112-1
Koopman, M., et al.: Assessing motor-related phenotypes of Caenorhabditis elegans with the wide field-of-view nematode tracking platform. Nat. Protoc. 15, 1–36 (2020). https://doi.org/10.1038/s41596-020-0321-9
Layana Castro, P.E., Puchalt, J.C., García Garví, A., Sánchez-Salmerón, A.J.: Caenorhabditis elegans multi-tracker based on a modified skeleton algorithm. Sensors 21(16) (2021). https://doi.org/10.3390/s21165622
Layana Castro, P.E., Puchalt, J.C., Sánchez-Salmerón, A.J.: Improving skeleton algorithm for helping Caenorhabditis elegans trackers. Sci. Rep. 10(1), 22247 (2020). https://doi.org/10.1038/s41598-020-79430-8
Le, K.N., Zhan, M., Cho, Y., Wan, J., Patel, D.S., Lu, H.: An automated platform to monitor long-term behavior and healthspan in Caenorhabditis elegans under precise environmental control. Commun. Biol. 3(1), 1–13 (2020). https://doi.org/10.1038/s42003-020-1013-2
Lin, T.-Y., et al.: Microsoft COCO: common objects in context. In: Fleet, D., Pajdla, T., Schiele, B., Tuytelaars, T. (eds.) ECCV 2014. LNCS, vol. 8693, pp. 740–755. Springer, Cham (2014). https://doi.org/10.1007/978-3-319-10602-1_48
Olsen, A., Gill, M.S. (eds.): Ageing: Lessons from C. elegans. HAL. Springer, Cham (2017). https://doi.org/10.1007/978-3-319-44703-2
Paszke, A., et al.: PyTorch: an imperative style, high-performance deep learning library. In: Advances in Neural Information Processing Systems 32 (2019)
Puchalt, J.C., Gonzalez-Rojo, J.F., Gómez-Escribano, A.P., Vázquez-Manrique, R.P., Sánchez-Salmerón, A.J.: Multiview motion tracking based on a cartesian robot to monitor Caenorhabditis elegans in standard petri dishes. Sci. Rep. 12(1), 1–11 (2022). https://doi.org/10.1038/s41598-022-05823-6
Puchalt, J.C., Layana Castro, P.E., Sánchez-Salmerón, A.J.: Reducing results variance in lifespan machines: an analysis of the influence of vibrotaxis on wild-type Caenorhabditis elegans for the death criterion. Sensors 20(21) (2020). https://doi.org/10.3390/s20215981
Puchalt, J.C., Sánchez-Salmerón, A.J., Eugenio, I., Llopis, S., Martínez, R., Martorell, P.: Small flexible automated system for monitoring Caenorhabditis elegans lifespan based on active vision and image processing techniques. Sci. Rep. 11 (2021). https://doi.org/10.1038/s41598-021-91898-6
Puchalt, J.C., Sánchez-Salmerón, A.J., Martorell Guerola, P., Genovés Martínez, S.: Active backlight for automating visual monitoring: an analysis of a lighting control technique for Caenorhabditis elegans cultured on standard petri plates. PLoS ONE 14(4), 1–18 (2019). https://doi.org/10.1371/journal.pone.0215548
Redmon, J., Divvala, S., Girshick, R., Farhadi, A.: You only look once: unified, real-time object detection. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 779–788 (2016)
Ren, S., He, K., Girshick, R., Sun, J.: Faster R-CNN: towards real-time object detection with region proposal networks. In: Advances in Neural Information Processing Systems 28 (2015)
Stiernagle, T.: Maintenance of C. elegans. WormBook. The C. elegans research community. WormBook (2006)
Acknowledgment
This study was supported by the Plan Nacional de I+D with Project RTI2018-094312-B-I00, FPI Predoctoral contract PRE2019-088214, Ministerio de Universidades (Spain) under grantFPU20/02639 and by European FEDER funds. ADM Nutrition, Biopolis SL, and Archer Daniels Midland provided support in the supply of C. elegans.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Rico-Guardiola, E.J., Layana-Castro, P.E., García-Garví, A., Sánchez-Salmerón, AJ. (2022). Caenorhabditis Elegans Detection Using YOLOv5 and Faster R-CNN Networks. In: Pereira, A.I., Košir, A., Fernandes, F.P., Pacheco, M.F., Teixeira, J.P., Lopes, R.P. (eds) Optimization, Learning Algorithms and Applications. OL2A 2022. Communications in Computer and Information Science, vol 1754. Springer, Cham. https://doi.org/10.1007/978-3-031-23236-7_53
Download citation
DOI: https://doi.org/10.1007/978-3-031-23236-7_53
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-23235-0
Online ISBN: 978-3-031-23236-7
eBook Packages: Computer ScienceComputer Science (R0)