Abstract
Probiotics have tremendous potential to develop healthy diets, treatment and prevention. Investigation of in vitro cultural properties of health-promoting microorganisms like lactic acid bacteria (LAB) and Bifidobacteria is crucial to select probiotic strains for treatments based on gut microbiota modulation to justify individualized and personalized approach for nutrition and prevention of variety of diseases.
The studied strains of LAB and bifidobacteria did not form spores, were positively stained by Gram, grow on medium in a wide range of pH (1.0–9.0, optimum pH 5.5–6.5), were sensitive to wide range of antibiotics; and showed different resistance to gastric juice, bile and pancreatic enzymes.
The most resistant to antibiotics were L. rhamnosus LB-3 VK6 and L. delbrueckii LE VK8 strains. The most susceptible to gastric juice was L. plantarum LM VK7, which stopped its growth at 8% of gastric juice; L. acidophilus IMV B-7279, B. animalis VKL and B. animalis VKB strains were resistant even in the 100% concentration.
Strains L. acidophilus IMV В-7279, L. casei IMV В-7280, B. animalis VKL, B. animalis VKB, L. rhamnosus LB-3 VK6, L. delbrueckii LE VK8 and L. delbrueckii subsp. bulgaricus IMV В-7281 were resistant to pancreatic enzymes.
Adhesive properties of the strains according to AIM index were high in L. casei IMV В-7280, B. animalis VKL and B. animalis VKB; were moderate in L. delbrueckii subsp. bulgaricus IMV В-7281; and were low in L. acidophilus IMV В-7279, L. rhamnosus LB-3 VK6, L. delbrueckii LE VK8 and L. plantarum LM VK7.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Predictive preventive personalized medicine
- Lactobacillus
- Bifidobacterium
- Probiotics
- Gut microbiota
- Antibiotics
- Gastric juice
- Bile
- Pancreatic enzymes
- Adhesive properties
- Pili
- Patient phenotype
- Individualized medicine
13.1 Relevance of In Vitro Research to Support Strains Stratification for Effective Personalized Probiotic Interventions
Intestinal microbiota (mainly represented by Firmicutes and Bacteroidetes) impact on health and homeostasis [1]. The definition of a probiotic was determined in 2001 by the World Health Organization (WHO) and Food and Agriculture Organization of the United Nations (FAO) [2] and confirmed in 2014 by the experts of International Scientific Association for Probiotics and Prebiotics (ISAPP) [3]. Probiotic microorganisms have large potential as a part of healthy diets and treatment of immunity-related disease [3,4,5,6,7,8,9,10,11,12]. They are effective actors both in distant sites and in the gut [12] and have potential for use in nutrition and personalized medicine [13, 14].
Gut microbiota modification in metabolic syndrome and other chronic diseases [14, 15] is a leading task of microbiome studying and for clinical use of probiotics [16]. Excluding few aspects, evidence-supported knowledge on the role of probiotics in applicability to clinical care and disease pathophysiology is not yet sufficient [17]. The effects of probiotic microorganisms are considered evidence-based in cases of diarrhea associated with Clostridium difficile and respiratory tract infections [18]. In vitro and in vivo studies, including ecology of gut microbiota, mechanism of probiotics functioning, and metabolomic researches for strains screening are needed to implement personalized probiotic treatment [8, 19, 20]. Taking into account that clinical studies are complicated to conduct and design, in vitro studies [21] and the use of animal models [22] can still provide a high quality data. For now original postulates of Koch were adapted and requirements for different microorganism to be considered a probiotic were formulated:
-
Commensal microorganism’ strain is associated with the health of the host, which is regularly manifested in healthy hosts, but less common in patients with disease.
-
Commensal microorganism’ strain can be identified as pure culture and cultivated in the laboratory.
-
Commensal microorganism’ strain improves or alleviate the disease when introduced into a new host organism.
-
Commensal microorganism’ strain can be detected after its introduction into a host [23].
The principle mechanisms of the role of probiotics in several diseases pathogenesis is the ability of microorganisms to change the immune response, matabolize nutrients, regulate balance of energy and prevent pathogens colonization [1, 2]. Probiotics have antagonistic relations with pathogens due to the producing of a number of organic acids, lysozyme, bacteriocins, and hydrogen peroxide [2, 3, 24]. Probiotic microorganisms are also able to synthesize digestive enzymes and vitamins C, K, A, PP, E, B and others [25, 26], produce metabolites such as histamine and short-chain fatty acids [27]. Due to the presence of these substances, probiotic microorganisms are able to change microbiota and participate in the metabolism of bile, fatty acids, cholesterol, glucose, bilirubin, and choline; as well as influence on the metabolism of calcium and iron and have antitoxic and immunomodulatory properties [28].
Each strain of probiotic microorganism may have specific phenotype/genotype and have multiple mechanisms of action [10]. In vitro tests have demonstrated the strain-dependent immunomodulation properties of bifidobacteria [29, 30]. In vitro studies have several limitations but they should obviously be used for preliminary screening of bacterial cells’ effects [31, 32], such as in vitro models using macrophage-like cells and cells isolated from the gut-associated lymphoid tissues (GALT) [33]. Biological properties such as parameters of the cell wall [30, 31] play an essential role in different aspects of immune response modulating and EPS-producing phenotype [32]. Such crosslinks between genotype and phenotype can warrant to stratify strains on their modulatory activity on innate immunity to justify an individual approach to prevention and nutrition.
13.2 Cultural-Morphological and Tinctorial Properties
To find the potential of lactic acid bacteria (LAB) and bifidobacteria as probiotics we determined their morphological and tinctorial properties by conventional research methods. A total of eight isolates were researched, as L. casei IMV B-7280 (B-7280 strain), L. acidophilus IMV B-7279 (B-7279 strain), L. rhamnosus LB-3 VK6 (VK6 strain), L. delbruecki subsp. bulgaricus IMV B-7281 (B-7281 strain), L. delbrueckii LE VK8 (VK8 strain), L. plantarum LM VK7 (VK7 strain), B. animalis VKB (VKB strain) and B. animalis VKL (VKL strain), which we obtained from the intestines of clinically healthy people. Freeze-dried LAB and bifidobacteria were used in our studies, therefore, we tested the viability of these bacteria by monitoring their growth on Man-Rogoza-Sharp (MRS) agar (MRSA) or bifidum agar (BA), respectively, for 24-48 h at 37 °C. Electron microscopy of bacterial samples was performed according to the generally accepted method [34] on an electron microscope JEM-1400 (Zabolotny Institute of Microbiology and Virology of the National Academy of Sciences of Ukraine), at 80 kV (Fig. 13.1).
These tested LAB and bifidobacteria strains were Gram-positive and grew on nutrient media in the pH range 1.0–9.0 (optimal pH 5.5–6.5). They did not form a dispute, they were motionless. Bifidobacteria cells were polymorphic. Instead, LAB strains were typically rod-shaped. Figure 13.1 illustrates that the shape of bifidobacterial cells dependent on the stage of culture development. Thus, it ranged from rod-shaped to spindle-shaped, hairpin, amorphous, Y- or X-shaped cells.
The dynamics of growth of these LAB and bifidobacteria strains during incubation in MRS nutrient medium at 37 °C was also investigated. We determined the beginning of the stationary phase of their growth. The beginning of the stationary phase of their growth that was 7–12 h after, t did not depend on the species or genus of these probiotic cultures. The stationary growth phase of VKB, VKL and B-7279 strains was approximately 8 h after their inoculation into the MRS nutrient medium. The stationary growth phase of VK6 and B-7280 strains began at 10 h. However, the stationary growth phase of B-7281, VK8 and VK7 strains occurred only after 11–12 h, i.e. it turned out to be the longest.
13.3 Tolerance of Probiotic Bacteria to Gastric Juice, Bile Salts and Proteolytic Enzymes (Pancreatin)
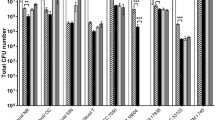
The tested LAB and bifidobacteria strains were examined for gastric juice (of 1, 2, 5, 8, 10, 20, 30, 50, 75, and 100% in medium; for 2.5 h), bile (of 0.1, 0.25, 0.5, 0.75, 1, 2, 3, 4, and 5%; for 5 h) and proteolytic enzymes tolerance (of 0.1, 0.25, 0.5, 0.75, 1, 2, 3, 4, and 5%; for 15 h) in this research (Fig. 13.2). Daily probiotic cultures strains were grown in liquid media at 37 °C with these aggressive factors, and after this, bacteria were sown to MRSА and BA, respectively. The number of colony-forming units (CFU) was counted after cultivation at 37 °C for 24–48 h. Complex effect of these aggressive factors (gastric juice 2%, for 2.5 h; of bile salts 1%; for 5 h; proteolytic enzymes 1%; for 15 h) on the LAB and bifidobacteria was also evaluated. Thus, tested LAB and bifidobacteria strains were grown with the gradual addition [35, 36].
It was found that these probiotic cultures had different tolerance to these aggressive factors. The resistance to gastric juice even at 100% concentration were for three strains, as B-7279, VKL and VKB. The growth of VK6 strains was inhibited at a concentration of 50% gastric juice in MRS medium, and VK6 and B-7280 strains were characterized by moderate resistance to its effect. The gastric juice of 10% completely inhibited the growth of VK8 strain. Note that the growth of these bacteria was significantly inhibited by gastric juice at 1% concentration. In Fig. 13.2 illustrated that complete inhibition of growth of B-7281 strain was observed at 10% concentration of gastric juice in nutrient medium. The most sensitive was the VK7 strain, the growth of which was inhibited at a concentration of gastric juice of 8%.
The bile salts at a concentration of 1–40% had also different degrees of inhibition on the tested LAB and bifidobacteria strains in our study; their tolerances to bile salts are also shown in Fig. 13.2. The B-7279, B-7280, VK6, VKB, VKL strains were the most resistant to bile salts; their viability was not completely inhibited in the presence of this aggressive factor in concentrations up to 40%. According to our unpublished data, the growth of VKB and VKL strains began to be inhibited after the introduction of bile salts into the nutrient medium at concentrations of 50% and 75%, respectively. The B-7281 and VK7 strains resisted 40 or 10% bile salts, respectively. Medium bile salt resistance was found for B-7281 and VK7 strains. At the same time, complete inhibition of the VK8 strain occurred at a concentration of bile salts of only 8%. That is, this strain of LAB was the most sensitive to them. The viability of tested LAB and bifidobacteria with different proteolytic enzymes tolerance is also strain-dependent. Similar tolerance to proteolytic enzymes was observed among the seven strains. Thus, B-7279, B-7280, VKL, VKB, VK6, VK8, B-7281 strains had the highest survival in the MRS nutrient medium with proteolytic enzymes in concentrations up to 5% (inclusive). The growth of the VKB strain was inhibited at 5% concentration of proteolytic enzymes. But, VK7 and VKB strains were the most susceptible to them. Proteolytic enzymes at a concentration of 4% completely inhibited their growth.
Subsequent studies have investigated the complex effects of gastric juice, bile and proteolytic enzymes on tested LAB and bifidobacteria strains, as live probiotic bacteria must gradually resist these aggressive factors of the digestive human or animal tract (Fig. 13.3). Thus, only four tested strains were the most tolerant. In particular, the survival of B-7279, B-7280, VKB and VKL strains was over 90% (96.7; 95.7; 90.2 and 91.9%, respectively). The survival of VK6, VK8 and B-7281 strains was lower (50.0; 86.0; 65.6 and 79.2%, respectively). That is, they were moderately sensitive to the complex action of these aggressive factors. But the survival rate of VK7 strain was only 49.3%.
Probably more critical for the complete survival of live probiotic bacteria in the digestive tract is their resistance to bile salts and pancreatic enzymes, as B-7280 strain has moderate resistance to gastric juice (alone). But more research is needed to confirm this.
13.4 Adhesive Properties of Probiotic Bacteria
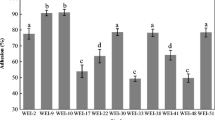
For analysis of some cell surface characteristics, the adhesive properties of the tested LAB and bifidobacteria strains were measured (Table 13.1). Adhesive properties of these probiotic culture were assessed, as AAR (average adhesion rate; this is an average number of bacteria, which attached to the one epitheliocyte); AIM (index of adhesion of microorganisms; this is the average number of bacteria attached to one epitheliocyte); and PRE (the participation rate of epithelial cells; this is a percent of epithelial cells, having on its surface-adhered bacteria), AIM is calculated by the formula [37]. It was considered that bacteria had high adhesive activity at AIM >4.0; average adhesive activity at AIM 2.51–4.04; at AIM 1.75–2.5 were low adhesive activity, and at AIM index ≤1.75 there was no adhesive activity [29]. As shown in Table 13.2, the adhesive properties of these strains were significantly different (p < 0.05), and AIM index ranged from 2.14 to 7.81. Our data showed that the tested bacteria were distributed in this way by adhesive properties: В-7280 strain ≥ VKВ strain ≥ VKL strain ≥ В-7281 strain ≥ В-7279 strain ≥ VK8 strain ≥ VK6 strain ≥ VK7 strain. Consequently, the adhesive properties of В-7280, VKВ and VKL strains are higher than that of other tested probiotic cultures.
13.5 Antibiotic Resistant of Probiotic Bacteria
The tested LAB and bifidobacteria strains resistance to antibiotics (by disc-diffusion method [19, 22]), that are inhibitors of protein, or peptidoglycan, or nucleic acid biosynthesis, was studied to determine the possibility of their use together with antimicrobial drugs (Table 13.2). In our studies the standardized discs with antibiotics were used. According to the size of the zone of growth inhibition, we determined the degree of sensitivity of bacteria to antibiotics: over 20 mm as sensitive (S); 10–20 mm as medium resistant (M); and less than 10 mm as resistant (R) [31].
In particular, VK7, B-7279 and B-7280 strains were highly sensitive to antibiotics that inhibit protein biosynthesis. The VKB strain was resistant to chlortetracycline, clarithromycin and roxithromycin. To aminoglycosides was highly sensitive only to the B-7279 strain. To nitrofurans VKB or B-7279 strains were moderately resistant or highly sensitive, respectively. But nitrofurans did not inhibit the growth of B-7280 and VK6 strains. The B-7281 strain was sensitive to furadonin, and VK8 and VK7 strains were moderately resistant to furadonin and resistant to fucidin and furazolidone. To most antibiotics, that blocking the peptidoglycan biosynthesis, were sensitive B-7279 (except oxacillin), B-7280 and VKB (except oxacillin, penicillin, imipenem, meropenem) strains. VK8 strain was also predominantly sensitive to these antibiotics. But strain VK6 3 was resistant to most of them.
Importantly, vancomycin, oxacillin, and teicoplanin did not inhibit the growth of LAB and bifidobacteria strains we studied. As shown in Table 13.1, most of them were moderately resistant or resistant to cephalosporins, except for B-7279 and B-7281 strains. The tested probiotic cultures also had different sensitivity to antibiotics, inhibitors of nucleic acid synthesis. Only the VK6 strain was resistant to these antibiotics. At the same time, they moderately inhibited the growth of VK8, VK7 and B-7280 strains. Levofloxacin and sparfloxacin inhibited the growth of VKL strain, and sparfloxacin and ciprofloxacin inhibited the growth of B-7279 strain. However, the VKB strain was sensitive only to sparfloxacin. All of these tested probiotic cultures were resistant to nalidixic acid. Our results showed that the antibiotic resistance of LAB strains and bifidobacteria dependent on the strain and not on the genus and species of microorganisms.
Therefore, based on the results of our study, the strains with the best probiotic properties were selected. These LAB and bifidobacteria strains, as B-7279, B-7280, VKB and VKL, with the highest survival (over 90%) at complex effects of gastric juice, bile salts and proteolytic enzymes, proved their tolerance to digestive tract conditions. But the adhesive properties were high only of B-7280, VKB and VKL strains (their AIM index ranged from 4.72 to 7.81). Average adhesive activity was in the В-7281 strain with moderately tolerance (79.2%) to the complex of these aggressive factors. B-7279 strain and other tested probiotic cultures had low adhesive activity (their AIM index ranged from 2.14 to 2.45) and different tolerance to gastric juice, bile salts and proteolytic enzymes. Thus, strains B-7280, B-7281, VKB and VKL, isolated from the intestines of clinically healthy people, have some probiotic properties and are promising for use in probiotic preparations. But other comprehensive preclinical studies of their safety and probiotic properties using different models, as well as clinical trials, are needed.
13.6 Data Interpretation
Obtained data partially coincide with the data of other researchers [38,39,40,41,42,43,44]. Antibiotic resistant bacteria could be transmitted from animals to humans via food chain [41]. Almost all LAB isolated from farm animals and humans are susceptible to ampicillin, amikacin, different cephalosporins, gentamicin, imipenem, erythromycin, oxacillin, and penicillin.
Teuber et al. [45] shoved that bifidobacteria and LAB have sensitivity to ampicillin, penicillin, cephalosporin, erythromycin, and tetracycline and resistance to vancomycin, gentamicin, and streptomycin. A matter of concern to use strains resistant to antibiotics is that antibiotic resistance is not a safety parameter due to the risk of plasmid transfer to pathogenic strains [35, 36, 46].
The strains studied by us meet such important criteria of selection as resistance to antibiotics according to probiotics guidelines like European Food Safety Authority (EFSA) [37].
As far as metabolism of cholesterol has strong crosslinks with circulation of bile acids and the potential associations between modulatory of immune response and hypocholesterolemic activity of probiotics [8], search for bile resistant strains might have a strong input on treatment patients with atherosclerosis, cholestasis, and other conditions. Some LAB and bifidobacteria strains had high hypocholesterolemic activity in different in vivo models of metabolic disorders and associated in vitro with degradation of bile acids [47].
Gram-positive LAB strains have bacterial pili for adhesion strengthening and protection against environmental stresses. However, their role in immune interaction is still largely unknown [48,49,50]. Pilus-associated SpaC pilin and gene cluster spaCBA makes possible exertion of both intimate and long distance contact with host tissue and provides mucus-binding [50]. Glycosylation as a sortase-dependent pili modification was shown to play a great role in the immunomodulation [50]. These findings altered the assumption about glycoconjugates’s underappreciated role in interplay between bacteria and host. Studying of cell wall of probiotic bacteria using imaging and molecular methods is important to predict or evaluate adhesion and immunomodulatory properties of the strain [31, 51,52,53].
Gut microbiota is a key player in host energy homoeostasis’ regulation and in the pathogenesis of metabolic syndrome and obesity for now is properly studied in clinical set due to number of unpredictable and uncontrollable factors in humans [54,55,56] such as genetic and environmental factors (drug assumption, diet, life style) [8, 56]. The essential task for probiotic and microbiome research in clinico is the search for reproducible in large population and reliable microbiome phenotype markers for longitudinal observation.
Immunomodulatory properties and anti-inflammatory properties [5, 7, 31] has been hypothesized to be likely correlate with clinically relevant effects, in particular the ability to demonstrate liver protective and hypocholesterolemic properties and modulate metabolic conditions [8]. All probiotic strains have primary native antiviral [57, 58], antibacterial [59] and antifungal properties [60,61,62], and have a high perspective to be alternative for antibiotics during treatment of various infections. Oxygen tolerance of probiotic microorganisms is another important parameter that should be studied in the near future, as far as for now there are only few studies in the field [63, 64]. LAB strains can potentiate hypoxia-inducible factor in the gut [65]. These results open new perspectives to propose individualized treatment to patients with Flammer syndrome [66]; manage hypoxia-associated conditions and stress [65]; use patient stratification on important gut hypoxia signaling marker, mesenteric ischemia in patients with atherosclerosis [67]. Potential of probiotic microorganisms for enhancement of safety and regenerative therapy efficacy [68], stem cells transplantation is difficult challenge aimed to develop biological preparations of a new generation. Finally, many of the mentioned properties of probiotic strains should be used in cancer case management as a part of supportive therapy [18] to facilitate, treatment-associated symptoms [69, 70].
This chapter includes concepts to the topic developed by the authors within previously performed studies [71,72,73].
References
Parekh PJ, Balart LA, Johnson DA (2015 Jun 18) The influence of the gut microbiome on obesity, metabolic syndrome and gastrointestinal disease. Clin Transl Gastroenterol 6:e91. https://doi.org/10.1038/ctg.2015.16
WHO/FAO scientific document. http://who.int/foodsafety/fs_management/en/probiotic_guidelines.pdf. Accessed 11 Feb 2018
Hill C, Guarner F, Reid G, Gibson GR, Merenstein DJ, Pot B et al (2014) Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 8:506–514. https://doi.org/10.1038/nrgastro.2014.66
Gibson GR, Hutkins R, Sanders ME, Prescott SL, Reimer RA, Salminen SJ et al (2017) Expert consensus document: the International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat Rev Gastroenterol Hepatol 14(8):491–502. https://doi.org/10.1038/nrgastro.2017.75
Bubnov RV, Spivak MY, Lazarenko LM, Bomba A, Boyko NV (2015) Probiotics and immunity: provisional role for personalized diets and disease prevention. EPMA J 6:14
Aron-Wisnewsky J, Clément K (2016 Mar) The gut microbiome, diet, and links to cardiometabolic and chronic disorders. Nat Rev Nephrol 12(3):169–181. https://doi.org/10.1038/nrneph.2015.191
Lazarenko LM, Babenko LP, Bubnov RV, Demchenko OM, Zotsenko VM, Boyko NV et al (2017) Imunobiotics are the novel biotech drugs with antibacterial and immunomodulatory properties. Mikrobiol Z 79(1):66–75
Bubnov RV, Babenko LP, Lazarenko LM, Mokrozub VV, Demchenko OA, Nechypurenko OV et al (2017) Comparative study of probiotic effects of Lactobacillus and bifidobacteria strains on cholesterol levels, liver morphology and the gut microbiota in obese mice. EPMA J 8(4):357–376. https://doi.org/10.1007/s13167-017-0117-3
Jobin C (2018 Jan 5) Precision medicine using microbiota. Science 359(6371):32–34. https://doi.org/10.1126/science.aar2946
Lebeer S, Bron PA, Marco ML, Van Pijkeren JP, O’Connell Motherway M, Hill C et al (2017) Identification of probiotic effector molecules: present state and future perspectives. Curr Opin Biotechnol 49:217–223. https://doi.org/10.1016/j.copbio.2017.10.007
Marchesi JR, Adams DH, Fava F, Hermes GD, Hirschfield GM, Hold G et al (2016) The gut microbiota and host health: a new clinical frontier. Gut 65(2):330–339. https://doi.org/10.1136/gutjnl2015-309990
Reid G, Abrahamsson T, Bailey M, Bindels LB, Bubnov R, Ganguli K et al (2017 Aug 24) How do probiotics and prebiotics function at distant sites? Benef Microbes 8(4):521–533. https://doi.org/10.3920/BM2016.0222
Golubnitschaja O, Baban B, Boniolo G, Wang W, Bubnov R, Kapalla M et al (2016) Medicine in the early twenty-first century: paradigm and anticipation—EPMA position paper 2016. EPMA J 7:23
Shapiro H, Suez J, Elinav E (2017) Personalized microbiome-based approaches to metabolic syndrome management and prevention. J Diabetes 9(3):226–236. https://doi.org/10.1111/1753-0407.12501.Review
Dao MC, Clément K (2018) Gut microbiota and obesity: concepts relevant to clinical care. Eur J Intern Med 48:18–24. https://doi.org/10.1016/j.ejim.2017.10.005
van den Nieuwboer M, Browne PD, Claassen E (2016) Patient needs and research priorities in probiotics: a quantitative KOL prioritization analysis with emphasis on infants and children. PharmaNutrition 4(1):19–28
Park S, Bae JH (2015) Probiotics for weight loss: a systematic review and meta-analysis. Nutr Res 35:566–575
Rondanelli M, Faliva MA, Perna S, Giacosa A, Peroni G, Castellazzi AM (2017) Using probiotics in clinical practice: where are we now? A review of existing meta-analyses. Gut Microbes 8(6):521–543. https://doi.org/10.1080/19490976.2017.1345414
Papadimitriou K, Zoumpopoulou G, Foligné B et al (2015) Discovering probiotic microorganisms: in vitro, in vivo, genetic and omics approaches. Front Microbiol 6:58. https://doi.org/10.3389/fmicb.2015.00058
Fijan S (2014) Microorganisms with claimed probiotic properties: an overview of recent literature. Int J Environ Res Public Health 11(5):4745–4767
Shah P, Fritz JV, Glaab E, Desai MS, Greenhalgh K, Frachet A et al (2016) A microfluidics-based in vitro model of the gastrointestinal human-microbe interface. Nat Commun 7:11535. https://doi.org/10.1038/ncomms11535
Nguyen TL, Vieira-Silva S, Liston A, Raes J (2015) How informative is the mouse for human gut microbiota research? Dis Model Mech 8(1):1–16. https://doi.org/10.1242/dmm.017400
Neville BA, Forster SC, Lawley TD (2017) Commensal Koch’s postulates: establishing causation in human microbiota research. Curr Opin Microbiol 42:47–52. https://doi.org/10.1016/j.mib.2017.10.001
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C et al (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65
D’Aimmo MR, Mattarelli P, Biavati B, Carlsson NG, Andlid T (2012) The potential of bifidobacteria as a source of natural folate. J Appl Microbiol 112(5):975–984. https://doi.org/10.1111/j.1365-2672.2012.05261.x
Lilly DM, Stillwell RH (1965) Growth promoting factors produced by probiotics. Science 147:747–748
Gao C, Ganesh BP, Shi Z, Shah RR, Fultz R, Major A et al (2017 Oct) Gut microbe-mediated suppression of inflammation-associated colon carcinogenesis by luminal histamine production. Am J Pathol 187(10):2323–2336. https://doi.org/10.1016/j.ajpath.2017.06.011
Bermudez-Brito M, Plaza-Díaz J, Muñoz-Quezada S, Gómez-Llorente C, Gil A (2012) Probiotic mechanisms of action. Ann Nutr Metab 61(2):160–174. https://doi.org/10.1159/000342079
Ruiz L, Delgado S, Ruas-Madiedo P, Sánchez B, Margolles A (2017) Bifidobacteria and their molecular communication with the immune system. Front Microbiol 8:2345. https://doi.org/10.3389/fmicb.2017.02345
Kobayashi H, Kanmani P, Ishizuka T, Miyazaki A, Soma J, Albarracin L et al (2017) Development of an in vitro immunobiotic evaluation system against rotavirus infection in bovine intestinal epitheliocytes. Benef Microbes 8:309–321. https://doi.org/10.3920/BM2016.0155
Mokrozub VV, Lazarenko LM, Sichel LM, Bubnov RV, Spivak MY (2015) The role of beneficial bacteria wall elasticity in regulating innate immune response. EPMA J 6:13
Hidalgo-Cantabrana C, Sánchez B, Milani C, Ventura M, Margolles A, Ruas-Madiedo P (2014) Genomic overview and biological functions of exopolysaccharide biosynthesis in Bifidobacterium spp. Appl Environ Microbiol 80(1):9–18. https://doi.org/10.1128/AEM.02977-13
Hidalgo-Cantabrana C, Sánchez B, Álvarez-Martín P, López P, Martínez-Álvarez N, Delley M et al (2015) A single mutation in the gene responsible for the mucoid phenotype of Bifidobacterium animalis subsp. lactis confers surface and functional characteristics. Appl Environ Microbiol 81(23):7960–7968. https://doi.org/10.1128/AEM.02095-15
Wang T, Cai G, Qiu Y, Fei N, Zhang M, Pang X et al (2012) Structural segregation of gut microbiota between colorectal cancer patients and healthy volunteers. ISME J 6:320–329
European Food Safety Authority (EFSA) (2008) Technical guidance—update of the criteria used in the assessment of bacterial resistance to antibiotics of human or veterinary importance. EFSA J 732:1–15. https://doi.org/10.2903/j.efsa.2008.732
Ruiz L, Margolles A, Sánchez B (2013) Bile resistance mechanisms in Lactobacillus and Bifidobacterium. Front Microbiol 4:396. https://doi.org/10.3389/fmicb.2013.00396
Tanaka H, Doesburg K, Iwasaki T, Mierau I (1999) Screening of lactic acid bacteria for bile salt hydrolase activity. J Dairy Sci 82:2530–2535
Penders J, Stobberingh EE, Savelkoul PHM, Wolffs PFG (2013) The human microbiome as a reservoir of antimicrobial resistance. Front Microbiol 4:87. https://doi.org/10.3389/fmicb.2013.00087
D’Aimmo MR, Modesto M, Biavati B (2007) Antibiotic resistance of lactic acid bacteria and Bifidobacterium spp. isolated from dairy and pharmaceutical products. Int J Food Microbiol 115(1):35–42
Teuber M, Meile L, Schwarz F (1999) Acquired antibiotic resistance in lactic acid bacteria from food. Antonie Van Leeuwenhoek 76(1–4):115–137. Review
Singer RS, Finch R, Wegener HC, Bywater R, Walters J, Lipsitch M (2003 Jan) Antibiotic resistance—the interplay between antibiotic use in animals and human beings. Lancet Infect Dis 3(1):47–51
Gueimonde M, Sánchez B, de los Reyes-Gavilán CG, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:202. https://doi.org/10.3389/fmicb.2013.00202
Zheng M, Zhang R, Tian X, Zhou X, Pan X, Wong A (2017) Assessing the risk of probiotic dietary supplements in the context of antibiotic resistance. Front Microbiol 8:908
Sharma P, Tomar SK, Goswami P, Sangwan V, Singh R (2014) Antibiotic resistance among commercially available probiotics. Food Res Int 57:176–195
Tannock GW, Luchansky JB, Miller L, Connell H, Thode-Andersen S, Mercer AA et al (1994) Molecular characterization of a plasmid-borne (pGT633) erythromycin resistance determinant (ermGT) from Lactobacillus reuteri 100-63. Plasmid 31(1):60–71
EFSA NDA Panel (EFSA Panel on Dietetic Products, Nutrition and Allergies) (2016) General scientific guidance for stakeholders on health claim applications. EFSA J 14(1):4367 [38 pp.] https://doi.org/10.2903/j.efsa.2016.4367
Lebeer S, Vanderleyden J, De Keersmaecker SC (2008) Genes and molecules of lactobacilli supporting probiotic action. Mucosal adhesion properties of the probiotic Lactobacillus rhamnosus GG SpaCBA and SpaFED pilin subunits. Microbiol Mol Biol Rev 72(4):728–764
Lebeer S, Claes I, Tytgat HL, Verhoeven TL, Marien E, von Ossowski I et al (2012) Functional analysis of Lactobacillus rhamnosus GG pili in relation to adhesion and immunomodulatory interactions with intestinal epithelial cells. Appl Environ Microbiol 78(1):185–193. https://doi.org/10.1128/AEM.06192-11
von Ossowski I, Reunanen J, Satokari R, Vesterlund S, Kankainen M, Huhtinen H et al (2010) Mucosal adhesion properties of the probiotic Lactobacillus rhamnosus GG SpaCBA and SpaFED pilin subunits. Appl Environ Microbiol 76(7):2049–2057. https://doi.org/10.1128/AEM.01958-09
Tytgat HL, van Teijlingen NH, Sullan RM, Douillard FP, Rasinkangas P, Messing M et al (2016) Probiotic gut microbiota isolate interacts with dendritic cells via glycosylated heterotrimeric pili. PLoS One 11(3):e0151824. https://doi.org/10.1371/journal.pone.0151824.eCollection.2016
Burgain J, Gaiani C, Francius G, Revol-Junelles AM, Cailliez-Grimal C, Lebeer S et al (2013) In vitro interactions between probiotic bacteria and milk proteins probed by atomic force microscopy. Colloids Surf B Biointerfaces 104:153–162. https://doi.org/10.1016/j.colsurfb.2012.11.032
Tytgat HL, Schoofs G, Vanderleyden J, Van Damme EJ, Wattiez R, Lebeer S et al (2016) Systematic exploration of the glycoproteome of the beneficial gut isolate Lactobacillus rhamnosus GG. J Mol Microbiol Biotechnol 26(5):345–358. https://doi.org/10.1159/000447091
Guerin J, Burgain J, Borges F, Bhandari B, Desobry S, Scher J et al (2017) Use of imaging techniques to identify efficient controlled release systems of Lactobacillus rhamnosus GG during in vitro digestion. Food Funct 8(4):1587–1598. https://doi.org/10.1039/c6fo01737a
Garcia SL, Buck M, McMahon KD, Grossart HP, Eiler A, Auxotrophy WF (2015) Intrapopulation complementary in the ‘interactome’ of a cultivated freshwater model community. Mol Ecol 24(17):4449–4459. https://doi.org/10.1111/mec.13319
Garcia SL, Stevens SLR, Crary B, Martinez-Garcia M, Stepanauskas R, Woyke T et al (2018) Contrasting patterns of genome-level diversity across distinct co-occurring bacterial populations. ISME J 12(3):745–755. https://doi.org/10.1038/s41396-017-0001-0
Compare D, Rocco A, Zamparelli MS, Nardone G (2016) The gut bacteria-driven obesity development. Dig Dis 34(3):221–229
Al Kassaa I, Hober D, Hamze M, Chihib NE, Drider D (2014) Antiviral potential of lactic acid bacteria and their bacteriocins. Probiotics Antimicrob Proteins 6(3–4):177–185. https://doi.org/10.1007/s12602-014-9162-6
Dillon SM, Frank DN, Wilson CC (2016) The gut microbiome and HIV1 pathogenesis: a two-way street. AIDS 30(18):2737–2751
Babenko LP, Lazarenko LM, Shynkarenko LM, Mokrozub VV, Pidgorskyi VS, Spivak MY (2012 Nov–Dec) The effect of lacto- and bifidobacteria compositions on the vaginal microflora in cases of intravaginal staphylococcosis. Mikrobiol Z 74(6):80–89
Jiang TT, Shao TY, Ang WXG, Kinder JM, Turner LH, Pham G et al (2017) Commensal fungi recapitulate the protective benefits of intestinal bacteria. Cell Host Microbe 22(6):809–816.e4. https://doi.org/10.1016/j.chom.2017.10.013
Ilavenil S, Park HS, Vijayakumar M, Arasu MV, Kim DH, Ravikumar S et al (2015) Probiotic potential of Lactobacillus strains with antifungal activity isolated from animal manure. Sci World J 2015:802570–802510. https://doi.org/10.1155/2015/802570
Hager CL, Ghannoum MA (2017) The mycobiome: role in health and disease, and as a potential probiotic target in gastrointestinal disease. Dig Liver Dis 49(11):1171–1176. https://doi.org/10.1016/j.dld.2017.08.025
Talwalkar A, Kailasapathy K (2003) Metabolic and biochemical responses of probiotic bacteria to oxygen. J Dairy Sci 86(8):2537–2546
Talwalkar A, Kailasapathy K (2004 Mar) The role of oxygen in the viability of probiotic bacteria with reference to L. acidophilus and Bifidobacterium spp. Curr Issues Intest Microbiol 5(1):1–8
Wang Y, Kirpich I, Liu Y, Ma Z, Barve S, McClain CJ et al (2011) Lactobacillus rhamnosus GG treatment potentiates intestinal hypoxia-inducible factor, promotes intestinal integrity and ameliorates alcohol-induced liver injury. Am J Pathol 179(6):2866–2875. https://doi.org/10.1016/j.ajpath.2011.08.039
Bubnov R, Polivka J Jr, Zubor P, Koniczka K, Golubnitschaja O (2017) Premetastatic niches in breast cancer: are they created by or prior to the tumour onset? “Flammer syndrome” relevance to address the question. EPMA J 8:141–157. https://doi.org/10.1007/s13167-017-0092-8
Bubnov RV (2011) Ultrasonography diagnostic capability for mesenteric vascular disorders. Gut 60(Suppl 3):A104
Taur Y, Jenq RR, Perales MA, Littmann ER, Morjaria S, Ling L et al (2014) The effects of intestinal tract bacterial diversity on mortality following allogeneic hematopoietic stem cell transplantation. Blood 124(7):1174–1182
Grech G, Zhan X, Yoo BC, Bubnov R, Hagan S, Danesi R et al (2015) Position paper in cancer: current overview and future perspectives. EPMA J 6(1):9. https://doi.org/10.1186/s13167-015-0030-6
York A (2018) Microbiome: gut microbiota sways response to cancer immunotherapy. Nat Rev Microbiol 16(3):121. https://doi.org/10.1038/nrmicro.2018.12
Bubnov RV, Babenko LP, Lazarenko LM, Mokrozub VV, Spivak MY (2018) Specific properties of probiotic strains: relevance and benefits for the host. EPMA J 9(2):205–223. https://doi.org/10.1007/s13167-018-0132-z
Bubnov R, Babenko L, Lazarenko L, Kryvtsova M, Shcherbakov O, Zholobak N, Golubnitschaja O, Spivak M (2019) Can tailored nanoceria act as a prebiotic? Report on improved lipid profile and gut microbiota in obese mice. EPMA J 10(4):317–335. https://doi.org/10.1007/s13167-019-00190-1
Golubnitschaja O (ed) (2019) Flammer syndrome – from phenotype to associated pathologies, prediction, prevention and personalisation, vol 11. isbn 978-3-030-13549-2 isbn 978-3-030-13550-8 (eBook). https://doi.org/10.1007/978-3-030-13550-8
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bubnov, R.V., Babenko, L.P., Lazarenko, L.M., Mokrozub, V.V., Spivak, M. (2023). In Vitro Study of Specific Properties of Probiotic Strains for Effective and Personalized Probiotic Therapy. In: Boyko, N., Golubnitschaja, O. (eds) Microbiome in 3P Medicine Strategies. Advances in Predictive, Preventive and Personalised Medicine, vol 16. Springer, Cham. https://doi.org/10.1007/978-3-031-19564-8_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-19564-8_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-19563-1
Online ISBN: 978-3-031-19564-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)