Abstract
Autophagy is one of the intracellular machinery for maintaining organelle as well as physiological homeostasis in cells by clearance of cellular debris and recycling of essential raw materials. It is different from other cellular processes like apoptosis and necrosis in the sense that it acts as a double-edged sword that might lead either to survival or death based on the stimuli. There are broadly three different types of autophagy: macroautophagy, microautophagy, and chaperone mediated autophagy. Macroautophagy is one of the commonly understood forms of autophagy and has been discussed simply as autophagy throughout the chapter. The role of autophagy in stem cell maintenance and differentiation is essential as both the processes require intensive intracellular remodeling which involves a continuous cycle of synthesis and degradation of event-specific proteins. Several pathways are involved in the regulation of autophagy and vice versa in stem cells. Among them, there are master proteins mandatory for stem cell maintenance and/or differentiation reported to be directly regulating autophagy. The current chapter discusses the different signaling pathways in stem cells; regulating or being regulated by autophagy and its role in the maintenance and differentiation of various types of stem cells.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Autophagy, which literally means ‘Self eating’ although might seem like an intimidating devouring process, actually refers to an evolutionarily conserved intracellular cleansing process for the recycling of damaged organelles, unwanted proteins, surplus storage nutrients (glycogen and lipids), reactive oxygen species (ROS), macro and micro-molecules (nucleotides and metabolic byproducts) in the form of essential intracellular raw materials like amino acids, sugars, fatty acids, nucleosides, etc., to maintain cellular homeostasis [1,2,3,4]. Even intracellular pathogens can be taken up by the autophagy program to break them down into antigenic peptides to further incite specific immune responses [5, 6]. Autophagy was first discovered in yeast by Yoshinori Ohsumi and his colleagues at the University of Tokyo, in 1992, although the terminology was given long back by Christian de Duve in 1963 [7]. Induction of autophagy was initially believed to be only under starvation [8], but later on, it was also found to be activated by DNA damage, cellular stress, infection, and hypoxia [4]. Autophagy has been implicated in the prognosis of various human diseases, cellular, and host immunity as well as in developmental processes like stem cell renewal, maintenance, and metabolic remodeling leading to differentiation and morphogenesis, especially (but not limited to) during embryo development [5, 9,10,11,12,13,14]. The cellular machinery for autophagy basically includes the formation of a double-membraned vesicular body called autophagosome which carries the intracellular junk and wholly moves toward the lysosome and fuses with it for degradation and recycling. The fused body is called auto-phagolysosome which finally completes the process of autophagy [1, 2].
Autophagy Compared to Necrosis and Apoptosis
Autophagy is far different from apoptosis and necrosis. Although apoptosis and necrosis themselves differ from each other in terms of the process and what they do to the cell; both are bound to cause cell death unlike autophagy. Both are incited by mostly similar but somewhat different intracellular or extracellular cues. In short, apoptosis is a caspase-mediated programmed cell death that is characterized by chromosome condensation, nuclear fragmentation, and membrane blebbing intended to basically get rid of excess, undesirable, and sometimes intracellularly damaged cells, whereas necrosis is considered to be an accidental cell death caused by nonspecific or nonphysiological stimuli, mostly extracellular stress inducers, and is characterized by the expansion of cellular organelles, disintegration of cell membrane, and further inflammatory responses mediated by the release of the intracellular contents. However, autophagy is basically a cleaning and recycling process associated with the formation of the autophagosome, which is a bilayer vesicle containing damaged organelles, proteins, and other cytoplasmic components [15,16,17]. Although autophagy is intended to be more of a survival mechanism maintaining cellular homeostasis by resource recycling and consuming cellular waste; its excessive triggering by single or multiple upstream signaling might lead to unplanned cell death [4].
The Autophagic Cascade
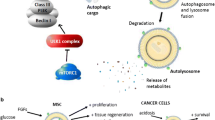
The whole process of autophagy is basically divided into 6 steps: Induction, Nucleation, Elongation, Maturation, Fusion, and ‘Degradation and recycling’. Unless there is a signal; the whole process is in check and is negatively regulated by the mTOR complex. Induction of autophagy results in the dephosphorylation and activation of ULK1 which then forms a ULK1–ATG13–FIP200–ATG101 complex. The ULK1 complex translocates to sites on the Endoplasmic Reticulum where the nucleation of autophagosome takes place leading to pre-autophagosome formation. ULK1 further activates another effector protein called Beclin1 from the phosphatidylinositol 3-kinase (PI3K) complex by phosphorylation. Beclin1 is one of the most important proteins within the PI3 Kinase complex associated with other accessory proteins like VPS34 and VPS15. Now, this PI3 Kinase converts PIP2 (Phospho-inositol diphosphate) to PIP3 (Phospho-inositol triphosphate) on the pre-autophagosome membrane which leads to the recruitment of WIPI proteins on the surface. This happens during the elongation process. WIPI proteins help in the recruitment of P62 and NBR1 sequestered LC3-I proteins to PE (Phosphatidylethanolamine) via another complex called ATG16-ATG5-ATG12 complex and facilitate the conversion of LC3-I to LC3-II by lipidation. Now, P62 and NBR1 are cargo carrier proteins that carry the debris for degradation and recycling into the fully formed autophagosomes. The presence of LC3-II generally is an indicator of a fully matured autophagosome. Once the matured autophagosome is formed, it finally fuses with the lysosome for degradation of the cargo and recycling of resources [2, 6, 7].
Types of Autophagy
There are basically three different types of autophagy: macroautophagy, microautophagy, and chaperone mediated autophagy [18].
Macroautophagy is an umbrella term for several different sub-types of autophagy and essentially refers to bulk degradation. All different types of organellophagy (bulk degradation of organelles through autophagy) like mitophagy (mitochondria), reticulophagy (Endoplasmic reticulum), nucelophagy (nucleus) including lipidophagy (lipid droplets), xenophagy (foreign micro-organisms), etc., come under macroautophagy [19,20,21].
Microautophagy by far is understood to be a simpler form of autophagy where the lysosomes directly take up cytoplasmic contents. The different forms of microautophagy are general microautophagy (cytoplasmic contents), Micro-ER-phagy (Endoplasmic reticulum), Micropexophagy (Peroxisome), Micronucleophagy (Nucleus), Micromitophagy (Mitochondria), and Microlipophagy (Lipid droplets). There is no need for ATG proteins or core autophagy machinery for general microautophagy as such though ESCRT (Endosomal Sorting Complexes Required for Transport) proteins mediate membrane budding and scission in many cellular processes and other proteins like VSP4 and lipidated LC3 (for cargo selection) might be required. However, special kinds of microautophagy like micropexophagy might require few other proteins from the core autophagy machinery as well [22].
Chaperone mediated autophagy (CMA) is a special kind of autophagy wherein a chaperone protein Hsc-70 is involved in the identification of the proteins destined for degradation. The cytosolic proteins targeted for degradation must contain a specific amino acid signature, namely, ‘KFERQ peptide sequence’. In this process, first, the target protein is identified based on the signature sequence followed by its unfolding and later taken up by the lysosomes for degradation. Regulation may depend on whether the KFERQ motif present in the target protein is accessible to Hsc-70 or not [23].
Role of Autophagy in Stem Cell Maintenance
Stem cells are unique cells of the human body that have the ability to self-renew for pool maintenance and differentiate to develop into any cell of the physiological system. They are mainly classified into the following four types based on their potency to form the various cell types—totipotent stem cells, pluripotent stem cells, multipotent stem cells, and unipotent stem cells. Stem cells are found not only in growing embryos, but also in adults where they are preserved for a long period in order to give rise to progenitor cells in the hour of need. This is to be kept in mind that with each differentiation, the stem cells’ potential to form a variety of cells becomes restricted; such as pluripotent stem cells can form any cell of the body but a multipotent stem cell can give rise to cells of a specific lineage [24].
Stem cell maintenance is one of the essential requirements both in case of embryonic, as well as adult stem cells, for replenishing the mother stem cell reserve. Maintenance of stem cells requires a lot of intracellular metabolic remodeling which involves periodic synthesis and degradation of proteins and thus requires a very active or multiple modes of protein degradation machinery [11].
The two major machineries to control the turnover of proteins in cells are autophagy and ubiquitin mediated proteasomal degradation [25]. If we consider autophagy as a tool that the cells can use as per their requirement; for each requirement, the cells might induce autophagy in a different way and the outcomes could be different depending on the upstream signaling induced by the cell responding to a particular stimulus. To understand whether autophagy plays a role in stem cell maintenance, several researchers have inhibited different key proteins in the pathway of autophagy and have searched for its effect on the renewal of various stem cells.
An insight into how autophagy helps in the maintenance of particular stem cells is discussed subsequently.
Role of Autophagy in Maintenance of Hematopoietic Stem Cells (HSCs)
The HSCs undergo hematopoiesis for generating blood cells of both myeloid and lymphoid lineages. For hematopoiesis to occur; a balance between hematopoietic stem cell (HSC) quiescence, activation, and differentiation is essential which is guided by autophagy [14].
In order to maintain the self-renewal capacity, intracellular reactive oxygen species (ROS) levels must be low, and the nuclear genome must be protected. Autophagy helps to maintain genome integrity and degrades defective mitochondria. This helps in the removal of excess ROS leading to the maintenance of HSC. There is also a close involvement of autophagy-related protein 7 (Atg7) whose deletion is coupled with the accumulation of damaged mitochondria, increased ROS levels, and DNA damage [26]. The absence of Atg 7 and FIP 200 genes inhibits regeneration activity and self-renewal capacity, and increases stress-induced apoptosis at an older age. Another autophagy gene, Atg12, deficiency also accompany impaired self-renewal and regenerative potential of HSCs [14, 27].
In order to maintain quiescence, when there is no need for multiplication or differentiation; HSCs show low levels of oxidative phosphorylation. Here in, autophagy plays a critical role in regulating intracellular oxidative metabolism by removing active healthy mitochondria which are fascinating, thus helping in energy conservation and maintenance of quiescence [7, 14].
Enduring metabolic stress is also important for the survival of any cell irrespective of whether it is a stem cell or somatic cell. HSCs maintain a very high level of FOXO3A which is a transcription factor that targets and induces pro-autophagy genes when the autophagy program has already been induced. In a way, higher levels of FOXO3A enhance autophagy rather than inducing it directly [7, 28].
Being an integral part of blood cell population maintenance, any disruption in hematopoiesis regulation leads to hematopoietic disorders like anemia and leukemia [7].
Role of Autophagy in Maintenance of Neural Stem Cells
Neural stem cells (NSCs) are the stem cells of the nervous system which are multipotent in nature and reside in discrete niches inside the subventricular zone of the lateral ventricles and subgranular zone of the hippocampal dentate gyrus of the adult brain.
Like HSCs; neural stem cells also depend on the FOXO family of proteins like FOXO1, FOXO3, and FOXO4 for enhancing autophagy to maintain cellular homeostasis. Inactivation of any of these proteins in NSCs has shown defective self-renewal and accumulation of protein aggregates [27]. In contrast, when compared with the quiescent NSCs, lysosome-related genes are surprisingly seen to be highly upregulated unlike activated NSCs; those instead harbor genes involved in proteasomal degradation pathway. The above is further supported by the fact that quiescent NSCs harbor a substantial amount of large lysosomes containing insoluble protein aggregates. The cells can reversibly get rid of these lysosomes whenever they are activated. This is probably a strange strategic way, how NSCs conserve energy during quiescence and stay in a docile state till activation.
Unlike HSCs, where autophagy is also known to degrade metabolically active mitochondria to maintain quiescence, NSCs only use autophagy to remove excess intracellular ROS via mitophagy [7].
Role of Autophagy in Maintenance of Muscle Stem Cells
Homeostasis and regeneration of skeletal muscles are maintained by muscle stem cells; also called Satellite cells. The nomenclature came up based on their position beneath the basal lamina of muscle fibers. These myoblasts are somite-derived that have not fused with other myoblasts and retained their stemness throughout adult life [29, 30].
The functioning of autophagy is quite interesting in case of muscle stem cells. A group of researchers has shown that starvation in the form of short-term caloric restriction could induce autophagy that could increase satellite cell number and muscle regeneration. Whereas, when talking about the basal level of autophagy, quiescent satellite cells from young mice display a higher level of autophagic flux than those isolated from aged mice. Intentional induction of autophagy by inhibition of mTOR in quiescent Satellite cells from aged mice shows enhanced regeneration. This suggests that the role of autophagy in muscle stem cells is majorly associated with delaying senescence.
On another note, the transition of Satellite cells from quiescence to active state also needs autophagy. In this case autophagy acts as a mechanism to recycle ATP to cater to the energy requirements of the cells during the transition [7].
Another unique phenomenon attributed to the muscle stem cells regarding autophagy is the expression of autophagic genes in an oscillatory fashion synchronizing with the circadian rhythm, i.e., more during the day compared to night. Such rhythm is absent in aged muscle stem cells [27]. Although the functional implication of this phenomenon is not yet attributed to anything, it might have something to do with the fluctuating energy requirements of the cells during day and night.
Role of Autophagy in Maintenance of Bone Marrow Derived Mesenchymal Stem Cells (BMSCs)
Bone Marrow Derived Mesenchymal stem cells are adult pluripotent stem cells that are capable of giving rise to diverse other cell types like adipocytes, endothelial cells, osteocytes, neurons, and cardiomyocytes in response to appropriate signal exposure.
The role of autophagy in the maintenance of Bone Marrow Derived Mesenchymal stem cells is controversial. A study made by Zhang et al. in 2016 interestingly suggests that hypoxia-induced autophagy can lead to apoptosis of BMSCs by activation of AMPK that inhibits mTOR and in turn activates autophagy. The group has shown that inhibition of autophagy by an autophagy inhibitor 3-MA has reduced apoptosis of BMSCs, while apoptosis is aggravated by Rapamycin: an autophagy inducer; in a time-dependent manner. The group checked for apoptosis by TUNEL assay and expression of cleaved caspase 3 in the presence of 3-MA or Rapamycin under hypoxic condition [31].
However, on a different note, another group of authors has established that hypoxia-based autophagy which is induced via Apelin/Apj signaling helps in the proliferation of BMSCs [32]. In this regard, both cases could be possible under different circumstances depending upon the severity of the hypoxia induced.
Besides, autophagy also plays an important role in precluding senescence in different types of BMSCs like mandible-derived BMSCs and tibia-derived BMSCs [13, 26, 33].
Role of Autophagy in Maintenance of Hepatic Progenitor Cells
Liver progenitor cells (LPCs) or Hepatic progenitor cells (HPCs) have the capacity to self-renew and differentiate to form hepatocytes and biliary epithelial cells [26]. The regenerative potential of hepatic cells after partial hepatectomy is immense and is known to be mediated by IL-6 and TNF-α along with their downstream targets STAT-3 and NF-κB. Very recently, a close association of autophagy has been found to play a role in hepatocyte differentiation and their maintenance [34]. On impairment of Atg 5, damaged mitochondria get accumulated along with the reduction in ATP, total protein, and albumin levels, and an increase in ALT (alanine aminotransferase) and glucose levels. Autophagy regulates their metabolic functions, and on ablation or impairment of this process, the intrinsic apoptotic pathway is activated, reducing proliferation and increasing the death rate of hepatocellular cells.
The abolition of autophagy proteins, Atg5 and Beclin 1, in liver progenitor cells reduces stemness and induces senescence. Upon aging, lipofuscin, which is an aging pigment gets accumulated in the lysosomes causing a reduction in the number and function of autophagosomes leading to decreased regenerative potential of the liver.
Thus, autophagy is required in both differentiation and maintenance of stemness of LPCs where it is protected during liver injury by this homeostatic mechanism.
Role of Autophagy in Maintenance of Intestinal Stem Cells (ISCs)
The intestinal epithelium, which acts as a physical, as well as a physiological barrier, to prevent the gut microbiota from entering the host system, confronts extensive wear and tear due to its protective and digestive role. Thus, it requires frequent repair and maintenance. This is where intestinal stem cells come into play.
Similar to HSCs; autophagy in ISCs is also implicated in preserving cellular homeostasis by removing excess mitochondria and intracellular ROS. Moreover, research has also shown that intentional deletion of ATG7 (an intermediary protein indirectly involved in autophagophore elongation) from ISCs leads to impaired antioxidant and DNA repair responses emphasizing the role of autophagy in the maintenance of intracellular homeostasis in ISCs [35].
The role of autophagy in the maintenance of all the above different types of stem cells has been schematically shown in Fig. 2.1.
Role of autophagy in stem cell maintenance. The maintenance of different stem cells is determined by diversified roles of autophagy which is solely context dependent and cell specific. Functions like clearing mitochondrial ROS and damaged DNA, circumventing through cell senescent pathways, ATP recycling, and saving energy by quiescence help in stem cell maintenance, survival, and proliferation. Sometimes, depending upon the upstream signaling, autophagy can directly encourage the process of stem cell proliferation by regulating energy metabolism, intracellular resource recycling, and cyclic synthesis and degradation of proteins involved in the cell proliferative pathways. HSC (Haematopoietic Stem cells), NSC (Neural Stem cells), SC (Satellite Cells), ISC (Intestinal Stem Cells), HPC (Hepatic Progenitor Cells), and BMSC (Bone marrow derived mesenchymal stem cells)
Autophagy and Stem Cell Differentiation
Differentiation is a very essential phenomenon for the continuous replacement of damaged cells from tissues to maintain tissue-specific homeostasis and another very important factor controlling stem cell maintenance. There are unique ways in which stem cells practice autophagy for undergoing differentiation. The role of autophagy for inducing or enhancing differentiation could vary in a context specific depending on different types of stem cells.
Role of Autophagy in the Differentiation of HSCs
Previously, we discussed that Hematopoietic stem cells (HSCs) have the ability to differentiate into different kinds of blood cells both from lymphoid and myeloid lineages. Along with the maintenance of the HSC pool, it is also important for differentiation to happen as well; for the replacement of lost or damaged blood cells.
One of the major roles of autophagy specifically during the differentiation of HSCs to final mature erythrocytes is the clearance of mitochondria [36]. In this process of differentiation, HSCs first form erythroblasts which further mature into reticulocytes after losing the nucleus. Reticulocytes then get rid of organelles including mitochondria to finally mature into erythrocytes (RBCs). Elimination of mitochondria and nucleus provides accommodation for more hemoglobin and also helps in debarring oxygen being used by RBCs allowing RBCs to store oxygen for supply and transport. Therefore, clearance of mitochondria is essential for erythrocyte formation [37]. This is evidenced by the fact that GATA1, a transcription factor and master regulator of hematopoiesis, regulates several autophagy genes [18, 38].
However, interestingly, it has been observed that even ATG 5 deficient embryos harbor functional autophagic vacuoles in their reticulocytes. This has been confirmed by ultrastructural analysis [39]; however, in another independent study, it was shown that ATG7 is still necessary for the seamless removal of mitochondria in erythroid cells [18, 40]. Such observations from researchers suggest that an alternative mode of autophagy is operative in reticulocytes. The occurrence of an alternative mode of autophagy is not new and has been vividly discussed by Shimizu (2018) from the Dept. of Pathological Cell Biology, Tokyo Medical and Dental University, Japan, in a mini-review [41].
Role of Autophagy in the Differentiation of NSCs
Neural stem cells (NSC) as discussed earlier are multipotent self-renewing stem cells in the developing and adult mammalian central nervous system (CNS), those have the potential to give rise to either neurons or glial cells in the brain. As NSCs have the unique capability of repairing neural circuits, they tend to remain highly metabolically active under circumstances where cell replacement is necessary. Previously, we saw that autophagy is essential for the maintenance of NSCs.
Several researchers have also shown that neurogenesis also requires autophagy to happen during embryogenesis and also during adult tissue repairing. This is evident from the fact that key proteins from the autophagy cascade like Atg9a, Atg5, Atg7, Beclin1, Ambra1, Eva1a, and LC3-II go up during neurogenesis in the forebrain, olfactory bulb, and cerebral cortex derived NSCs [18]. Moreover, silencing of Vps34 also affects neurogenesis in the cerebral cortex by reducing excitatory neuron migration and axonal growth without affecting the cell cycle of NSCs at the Ventricular and subventricular zone (VZ/SVZ). Further, in cortical NSCs, intended deletion of ATG5 has shown to impair neuronal differentiation while in VZ/SVZ NSCs proliferation is induced instead of differentiation [42, 43].
In another interesting study miR-34a: an miRNA known to regulate Atg9a (a key regulator protein in autophagy induction which provides membrane for autophagosome formation); also negatively regulates neuronal differentiation suggesting autophagy is essential for neuronal differentiation [44].
On another note, if discussed about signaling known to cross-talk with autophagy in NSCs: Wnt3a and Notch signaling are reported.
The role of Wnt3a in regulating autophagy and, in turn, affecting neuronal differentiation is controversially discussed. On one hand, wnt3a is shown to decrease autophagy in mature neurons after traumatic brain injury while escalating hippocampal neurogenesis; whereas on the other hand, wnt3a increases autophagy in cells under cultured conditions in hippocampal neuronal cultures from embryonic rat through AMPK activation. Thus, whether wnt3a affects neuronal differentiation by either promoting or demoting autophagy could be highly context dependent and/or situation specific.
In case of Notch signaling, autophagy itself has been observed to regulate it in NSCs. In the process, the Notch1 receptor is degraded through its uptake into pre-autophagosomal vesicles in an Atg16L1-dependent manner. Increased Notch1 due to faulty autophagic machinery hinders neuronal differentiation and inflates the NSC pool [42, 45].
Role of Autophagy in the Differentiation of Cardiac Stem Cells
Human heart replaces its cardiomyocyte population completely around 18 times throughout life unless there is some form of cardiac damage where the cardiac stem cells (CSCs) come into play. The CSCs inhabit the stem cell niche in the adult heart and divide symmetrically/asymmetrically as per the shift in homeostasis. The asymmetric division gives rise to a CSC and another committed cell like a myocyte [46].
It has been reported that fibroblast growth factor (FGF) signaling regulates cardiac development. It inhibits ‘premature differentiation’ of CSCs. FGF attaches to fibroblast growth factor receptor (FGFR) tyrosine kinases and activates its downstream targets via FRS2α (FGF receptor substrate 2α) thereby down regulating autophagy. With the inhibition of autophagy, differentiation of myocardial stem cells also gets inhibited. Thus, it can be concluded that FRS2α is essential for autophagy suppression by FGF through the mTOR pathway. In the presence of a FGFR inhibitor, there has been an increased expression of Beclin 1 and p27 which marks the initiation of autophagy leading to CSCs differentiation. Interestingly, it has also been found that autophagy is not only a regulator of differentiation, but also is positively correlated with ectopic foci formation of the heart.
On the contrary, the Wnt pathway, which is an upstream regulator of FGF, inhibits CSCs differentiation via GSK3-TIP60-ULK1 pathway. It is even reported that cholesterol metabolism showed an increase in Atg5 proving that induction of autophagy triggers differentiation of CSCs. The other signalings which get activated are GSK3β/β-catenin and JNK/STAT-3, which cause an increase in cardiac transcriptional proteins, factors, and enhancers’ expressions, mediating their movement inside the nucleus and enhancing differentiation [14].
Role of Autophagy in Adipocyte Formation from MSCs
Adipocytes are derived from mesenchymal stem cells (MSCs) through adipogenesis; those, in turn, give rise to adipose tissue, a process closely governed by autophagy. The adipose tissue is complex enough to be considered as a vital endocrine organ in mammals for storing energy and regulating inflammation, cell signaling, and metabolism by secreting endocrines. In general, white adipose tissue (WAT) is primarily known to store lipids during the fed state, whereas during starvation, it releases fatty acids into the bloodstream for muscle energy production by breaking down triglycerides (TGs). Instead, brown adipose tissue (BAT) stores a limited amount of TGs, does not secrete fatty acids and rather uses them for heat production in the body [13, 47].
Studies have shown that activation of autophagy is important to WAT differentiation; by attenuating the proteasome-dependent degradation of PPARγ2: a regulator of fatty acid storage and glucose metabolism and cutting down the number of mitochondria. This is evident from the fact that pharmacological inhibition or knockdown of ATG5 and ATG7 genes lead to browning of WAT, hence decreasing lipid accumulation. In detail, PPARγ2 under proliferating conditions is degraded by the proteasomal system unless an adipogenic differentiation signal is received. This is followed by induction of autophagy stabilizing PPARγ2 promoting differentiation into adipogenic fate. This establishes the fact that both ubiquitin-dependent proteasomal degradation system and autophagy cross talk with each other to regulate the level of PPARγ2, in turn, regulating adipogenesis [18, 48, 49].
Deficiency of autophagy in this tissue causes diminished differentiation and development of adipose stem cells and abnormal secretion of adipocytokine. It also leads to the conversion of white adipose tissue to brown adipose tissue, clearly demarcating calorie loss, storage of which is essential to provide energy according to body requirements [47, 50].
Along with differentiation, autophagy also helps in lipid droplet expansion. C/EBPβ is found to be responsible for autophagy activation via Atg 4b and it functions as an important adipogenic factor. Upon activation, p62 (an autophagy related protein) breaks down Klf 2 and Klf 3 which are the negative regulators of adipocyte differentiation so as to promote the formation of mature adipocytes [51].
Atg7 and Atg5 also play a keen role in the maintenance of differentiation, lipid accumulation, insulin sensitivity, and white adipose mass formation. The rate of energy production through beta oxidation of fatty acids is also controlled by autophagy-related proteins. They keep a check on the equilibrium of white and brown adipose tissue formation, suggesting a significant part of autophagy being involved in adipose stem cell differentiation and balancing its zeal in metabolic activities [11, 52].
Role of Autophagy in Osteoclastogenesis
Osteoclasts are cells involved in bone resorption which develop from hematopoietic myeloid progenitor cells under the influence of colony-stimulating factor 1 (CSF1) and RANKL signals. This differentiation of osteoclasts is also regulated by autophagy, the disruption of which causes changed osteoclast function leading to osteoporosis. Apart from differentiation, survival, and function of other bone cells like osteocytes, osteoblasts are also governed by autophagy [53].
Deletion of autophagy proteins like Atg5, Atg 7, Beclin 1, and LC3 in osteoclasts are associated with reduced bone cell differentiation and mineralization, impaired secretion, bone resorption, and decreased bone mass. On the other hand, ablation of autophagy increases endoplasmic reticulum stress and RANKL secretion causing osteoblast dysfunction; leading to the activation of osteoclasts and bone resorption. Beclin 1 ubiquitination is mediated by TRAF, which, in turn, is deubiquitinated by the CYLD (Cylindromatosis) gene under the influence of p62, an autophagy cargo protein, which also causes RANKL stimulated osteoclast differentiation [54].
Upon coming across a low oxygen tension, osteoclasts show increased differentiation and expression of BNIP3 via HIF-1α; stimulating Beclin 1, Atg5, and Atg 12 release and LC3 expression on autophagosomes clearly indicating increased autophagic flux. Alongside the expressions of RANKL, Cathepsin K, NFATc1, and MMPs are also increased leading to increased osteoclastogenesis. miR-155 microRNA targets TGF-β activated kinase 1 binding protein 2 (TAB2) to induce autophagy in osteoclasts and modulates their function and differentiation. Another protein, GIT1 (G-protein-coupled receptor kinase 2 interacting protein 1), triggers phosphorylation of Beclin 1 and its dissociation from Bcl2 leading to autophagy induction and osteoclastogenesis [54].
Role of Autophagy During Embryogenesis
Embryogenesis is a phenomenon post fertilization where the maternally obtained products like proteins and mRNA are degraded along with the simultaneous synthesis of new products from the ‘zygotic genome’. This is a very rapid process with ubiquitin mediated proteasomal degradation playing an important role. But when this operation is not adequate, autophagy comes into play bringing about cellular remodeling [55].
Autophagy is activated within four hours of fertilization, regulated by E2 and progesterone which inhibits mTOR activation, a step found to have a very important role in embryo development. Atg5 null zygotes are reported to have arrested growth at the four to eight cell stage, whereas Atg5 positive zygotes proceed to the blastocyst stage. In zygotes with autophagy deficiency, there is a decreased rate of protein synthesis due to defects in the acquired protein clearance rate. Also, in such embryos, there is a reduced rate of implantation [56].
Other autophagy proteins like Beclin 1, FIP200, Ambra 1 deficiency leads to embryonic death because of pro-amniotic canal closing defect, heart failure and liver degeneration during mid or late gestational phases, defect in neural tube respectively. Atg 3, Atg5, Atg 7, Atg 16L ablation causes death of neonates just after birth due to defects in milk sucking and adjustment issues with the normoxic conditions outside the mother’s womb [56].
Although there have been several publications, researchers are still trying to understand the complete involvement of autophagy in embryogenesis (Table 2.1).
The role of autophagy in all the above different types of stem cell differentiation has been schematically shown in Fig. 2.2.
Role of autophagy in stem cell differentiation. In case of stem cell differentiation, several upstream key proteins which are involved in deciding the fate and timing of stem cell differentiation regulate or dysregulate autophagy. As autophagy is basically involved in protein degradation followed by clearance; it might influence the half-life of several proteins which directly either enhance or inhibit stem cell differentiation. Thus, autophagy has an essential role to play in stem cell differentiation as well; along with stem cell maintenance and preserving cellular homeostasis. HSC (Hematopoietic stem cells), HMPC (Hematopoietic Mesenchymal Progenitor Cells), NSC (Neural stem cells), ADSC (Adipose Derived Mesenchymal Stem Cells), and CSC (Cardiac stem cells)
Conclusion
As we come to the end of this chapter, we can say that the role of autophagy in stem cell maintenance and differentiation is quite diversified and purely context specific. It's necessitated in both embryonic as well as adult stem cells. Along with maintaining cellular and tissue-specific homeostasis, it is involved in various other functions like energy metabolism, regulating and dysregulating essential downstream pathways resulting in cell death, immune response, proliferation, and asymmetric division along with transcriptomic shift, ultimately leading to survival or differentiation. This role of autophagy is slightly different than its normal function of cell clearance during stress. Different forms of autophagy can simultaneously exist and function in the same cell. Even macroautophagy can perform differently based on the upstream signal received, i.e., the decision whether to go for debris clearance or maintenance or differentiation. Other than autophagy, ubiquitin mediated proteasomal degradation pathway also plays a crucial role to maintain cellular homeostasis under regular intracellular physiological conditions.
Abbreviations
- 3-MA:
-
3-Methyl adenine
- ADSC:
-
Adipose derived stem cell
- ALT:
-
Alanine Aminotransferase
- Ambra1:
-
Activating molecule in Beclin1-regulated autophagy protein 1
- AMPK:
-
AMP-activated protein kinase
- APJ:
-
Apelin receptor
- ATG:
-
Autophagy related gene
- ATP:
-
Adenosine triphosphate
- BAT:
-
Brown adipose tissue
- BMSC:
-
Bone marrow derived mesenchymal stem cells
- BNIP3:
-
BCL2/Adenovirus E1B 19 kDa Protein Interacting Protein 3
- C/EBPβ:
-
CCAAT-enhancer binding protein
- CMA:
-
Chaperone mediated autophagy
- CNS:
-
Central nervous system
- CSC:
-
Cardiac stem cells
- CSF:
-
Colony stimulating factor
- CYLD:
-
Cylindromatosis
- DNA:
-
Deoxyribo Nucleic Acid
- ER:
-
Endoplasmic Reticulum
- ESCRT:
-
Endosomal sorting complex required for transport
- Eva1:
-
Eva-1 homolog A
- FGF:
-
Fibroblast growth factor
- FGFR:
-
Fibroblast growth factor receptor
- FIP200:
-
Focal adhesion kinase family-interacting protein of 200kD
- FOXO:
-
Forkhead box transcription factor
- FRS2α:
-
FGF receptor substrate 2α
- GATA 1:
-
GATA-binding factor 1
- GIT1:
-
G-protein-coupled receptor kinase-interacting protein 1
- GSK3:
-
Glycogen Synthase Kinase 3
- HIF-1α:
-
Hypoxia Inducible Factor 1 alpha
- HMPC:
-
Hematopoietic mesenchymal progenitor cells
- HPC:
-
Hepatic progenitor cells
- HSC:
-
Hematopoietic stem cells
- HSC-70:
-
Heat shock Protein 70 kDa
- IL-6:
-
Interleukin-6
- ISC:
-
Intestinal stem cells
- JNK:
-
C-Jun N-terminal kinase
- Klf:
-
Kruppel-like factors
- LC3:
-
Microtubule-associated protein 1 light chain 3 protein
- LPC:
-
Liver Progenitor Cell
- miRNA:
-
Micro Ribo-nucleic acid
- MMP:
-
Matrix metalloproteinases
- MSC:
-
Mesenchymal stem cells
- mTOR:
-
Mammalian target of rapamycin
- NBR1:
-
Neighbor of BRCA1 gene 1 protein
- NFATc1:
-
Nuclear factor of activated T-cells 1
- NF-κβ:
-
Nuclear Factor kappa light chain enhancer of activated B cells
- NSC:
-
Neural stem cells
- PE:
-
Phosphatidylethanolamine
- PI3K:
-
Phosphatidylinositol—3—kinase
- PIP2:
-
Phosphatidylinositol 4,5-bisphosphate
- PIP3:
-
Phosphatidylinositol 3,4,5-triphosphate
- PPARγ2:
-
Peroxisome proliferator activated receptor gamma
- RANKL:
-
Receptor activator of nuclear factor kappa-B ligand
- RBC:
-
Red Blood cells
- ROS:
-
Reactive oxygen species
- SC:
-
Satellite cells
- STAT-3:
-
Signal transducer and activator of transcription 3
- TAB2TGF-β:
-
activates kinase 1 binding protein 2
- TGs:
-
Triglycerides
- TIP60:
-
60KDa HIV-Tat interactive protein
- TNF-α:
-
Tumor Necrosis Factor alpha
- TRAF:
-
Tumor necrosis factor receptor-associated factor
- TUNEL:
-
Terminal Deoxy-nucleotidyl Transferase dUTP Nick End Labeling
- ULK1:
-
Unc-51-like kinase 1
- VPS:
-
Vacuolar protein sorting
- VZ/SVZ:
-
Ventricular and subventricular zone
- WAT:
-
White Adipose Tissue
- WIPI:
-
WD repeat protein interacting with phosphoinositide
- Wnt:
-
Wingless-related Integration site
References
Khan I, Baig MH, Mahfooz S, Rahim M, Karacam B (2021) deciphering the role of autophagy in treatment of resistance mechanisms in Glioblastoma
Chun Y, Kim J (2018) autophagy: an essential degradation program for cellular homeostasis and life.https://doi.org/10.3390/cells7120278
El-gowily AH, Abosheasha MA (2020) Differential mechanisms of autophagy in cancer stem cells: emphasizing gastrointestinal cancers, 1–12. https://doi.org/10.1002/cbf.3552
Jung S, Jeong H, Yu S-W (2020) Autophagy as a decisive process for cell death. Exp Mol Med 526(52):921–930. https://doi.org/10.1038/s12276-020-0455-4
Levine B, Mizushima N (2011). Autophagy in immunity and inflammation, 1–5. https://doi.org/10.1038/nature09782
He C, Klionsky DJ (2009) Regulation mechanisms and signaling pathways of autophagy. Annu Rev Genet 43:67–93. https://doi.org/10.1146/annurev-genet-102808-114910.Regulation
Chang NC (2020) Autophagy and stem cells: self-eating for self-renewal 8:1–11. https://doi.org/10.3389/fcell.2020.00138
Castro-Obregon S (2010) The discovery of lysosomes and autophagy. Scitable by Nat Educ 3:49. https://www.nature.com/scitable/topicpage/the-discovery-of-lysosomes-and-autophagy-14199828/. Accessed August 10, 2021
Turksen K (2018) Autophagy in health and disease: potential therapeutic approaches. https://books.google.co.in/books?hl=en&lr=&id=Q3xxDwAAQBAJ&oi=fnd&pg=PR5&dq=autophagy+in+health+and+disease+Kursad+Turksen&ots=h4P592smya&sig=Bj20UB_RJAH5b8OQWa3ouCjwhDo. Accessed August 10, 2021
Simon H (n.d) Autophagy in myocardial differentiation and cardiac development. https://doi.org/10.1161/CIRCRESAHA.112.265157
Vessoni AT, Muotri AR, Okamoto OK (2012) Autophagy in stem cell maintenance and differentiation, 21. https://doi.org/10.1089/scd.2011.0526
Yang ZJ, Chee CE, Huang S, Sinicrope FA (2011) The role of autophagy in cancer: therapeutic implications. Mol Cancer Ther 10:1533–1541. https://doi.org/10.1158/1535-7163.MCT-11-0047
Sbrana FV, Cortini M, Avnet S, Perut F, Columbaro M, De Milito A, Baldini N (2016) The role of autophagy in the maintenance of stemness and differentiation of mesenchymal stem cells. Stem Cell Rev Reports, 621–633. https://doi.org/10.1007/s12015-016-9690-4
Guan J, Simon AK, Prescott M, Menendez JA, Wang F, Wang C, Wolvetang E, Vazquez-martin A, Zhang J (2013) Autophagy in stem cells. Taylor Fr. 9:830–849. https://doi.org/10.4161/auto.24132
Chen Q, Kang J, Fu C (2018) The independence of and associations among apoptosis, autophagy, and necrosis. Signal Transduct Target Ther. https://doi.org/10.1038/s41392-018-0018-5
Chaabane W, User SD, El-Gazzah M, Jaksik R, Sajjadi E, Rzeszowska-Wolny J, Łos, Autophagy MJ (2012) Apoptosis, Mitoptosis and necrosis: interdependence between those pathways and effects on cancer. Arch Immunol Ther Exp 611(61):43–58. https://doi.org/10.1007/S00005-012-0205-Y
Coleman J, Liu R, Wang K, Kumar A (2016) Detecting apoptosis, autophagy, and necrosis, 77–92. https://doi.org/10.1007/978-1-4939-3588-8_5
Rodolfo C, Di Bartolomeo S, Cecconi F (2016) Autophagy in stem and progenitor cells. Cell Mol Life Sci 73:475–496. https://doi.org/10.1007/s00018-015-2071-3
Jin M, Liu X, Klionsky DJ (2013) SnapShot: selective autophagy. Cell 152:368. https://doi.org/10.1016/J.CELL.2013.01.004
Li W, He P, Huang Y, Li YF, Lu J, Li M, Kurihara H, Luo Z, Meng T, Onishi M, Ma C, Jiang L, Hu Y, Gong Q, Zhu D, Xu Y, Liu R, Liu L, Yi C, Zhu Y, Ma N, Okamoto K, Xie Z, Liu J, He RR, Feng D (2020) Selective autophagy of intracellular organelles: recent research advances. Theranostics. 11:222–256. https://doi.org/10.7150/THNO.49860
Reggiori F, Komatsu M, Finley K, Simonsen A (2012) Autophagy: more than a nonselective pathway. Int J Cell Biol. https://doi.org/10.1155/2012/219625
Schuck S (2020) Microautophagy—distinct molecular mechanisms handle cargoes of many sizes. https://doi.org/10.1242/jcs.246322
Kaushik S, Cuervo AM (2012) Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol 22:407–417. https://doi.org/10.1016/j.tcb.2012.05.006.Chaperone-mediated
Zakrzewski W, Dobrzyński M, Szymonowicz M, Rybak Z (2019) Stem cells: past, present, and future. Stem Cell Res Ther 10. https://doi.org/10.1186/S13287-019-1165-5
Dikic I (2017) Proteasomal and Autophagic degradation systems. Annu Rev Biochem 86:193–224. https://doi.org/10.1146/ANNUREV-BIOCHEM-061516-044908
Chen X, He Y, Lu (2018) Review article autophagy in stem cell biology : a perspective on stem cell self-renewal and differentiation
Boya P, Codogno P, Rodriguez-muela N (2018) Autophagy in stem cells: repair, remodelling and metabolic reprogramming, 1–14. https://doi.org/10.1242/dev.146506
Warr MR, Binnewies M, Flach J, Reynaud D, Garg T, Malhotra R, Debnath J, Passegué E (2013) FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494:323–327. https://doi.org/10.1038/NATURE11895
Almeida CF, Fernandes SA, Ribeiro Junior AF, Keith Okamoto O, Vainzof M (2016) Muscle satellite cells: exploring the basic biology to rule them. Stem Cells Int. https://doi.org/10.1155/2016/1078686
Yin H, Price F, Rudnicki MA (2013) Satellite cells and the muscle stem cell niche. Physiol Rev 93:23. https://doi.org/10.1152/PHYSREV.00043.2011
Zhang Z, Yang M, Wang Y, Wang L, Jin Z, Ding L, Zhang L, Zhang L, Jiang W, Gao G, Yang J, Lu B, Cao F, Hu T (2016) Autophagy regulates the apoptosis of bone marrow-derived mesenchymal stem cells under hypoxic condition via AMP-activated protein kinase/mammalian target of rapamycin pathway 40:671–685. https://doi.org/10.1002/cbin.10604
Li L, Li L, Zhang Z, Jiang Z (2015) Hypoxia promotes bone marrow-derived mesenchymal stem cell proliferation through apelin/APJ/autophagy pathway 47:362–367. https://doi.org/10.1093/abbs/gmv014
Dong W, Zhang P, Fu Y, Ge J, Cheng J, Yuan H, Jiang H (2015) Roles of SATB2 in site-specific Stemness, autophagy and senescence of bone marrow mesenchymal stem cells. J Cell Physiol 230:680–690. https://doi.org/10.1002/JCP.24792
Xu F, Hua C, Tautenhahn H, Dirsch O, Dahmen U (2020) The role of autophagy for the regeneration of the aging liver
Trentesaux C, Fraudeau M, Luana C, Lemarchand J, Jacques S (2020) Essential role for autophagy protein ATG7 in the maintenance of intestinal stem cell integrity, 117. https://doi.org/10.1073/pnas.1917174117
Mortensen M, Ferguson DJ, Edelmann M, Kessler B, Morten KJ, Komatsu M, Simon AK (2010) Loss of autophagy in erythroid cells leads to defective removal of mitochondria and severe anemia in vivo. Proc Natl Acad Sci USA 107:832–837. https://doi.org/10.1073/PNAS.0913170107
Moras M, Lefevre SD, Ostuni MA (2017) From erythroblasts to mature red blood cells: organelle clearance in mammals. Front Physiol 8:1076. https://doi.org/10.3389/FPHYS.2017.01076
Kang YA, Sanalkumar R, O'geen H, Linnemann AK, Chang CJ, Bouhassira EE, Farnham PJ, Keles S, Bresnick EH (2012) Autophagy driven by a master regulator of hematopoiesis. Mol Cell Biol 32L:226–239. https://doi.org/10.1128/MCB.06166-11
Honda S, Arakawa S, Nishida Y, Yamaguchi H, Ishii E, Shimizu S (2014) Ulk1-mediated Atg5-independent macroautophagy mediates elimination of mitochondria from embryonic reticulocytes. Nat Commun 5:1–5. https://doi.org/10.1038/ncomms5004
Zhang J, Randall MS, Loyd MR, Dorsey FC, Kundu M, Cleveland JL, Ney PA (2009) Mitochondrial clearance is regulated by Atg7-dependent and independent mechanisms during reticulocyte maturation. Blood 114:157. https://doi.org/10.1182/BLOOD-2008-04-151639
Shimizu S (2018) Biological roles of alternative autophagy. Mol Cells 41:50–54. https://doi.org/10.14348/molcells.2018.2215
Casares-crespo L, Calatayud-baselga I, García-corzo L, Mira H (2018) On the role of basal autophagy in adult neural stem cells and neurogenesis 12:1–9. https://doi.org/10.3389/fncel.2018.00339
Lv X, Jiang H, Li B, Liang Q, Wang S, Zhao Q, JJ-S reports (n.d.) Undefined 2014, the crucial role of Atg5 in cortical neurogenesis during early brain development. Nature.Com. https://sci-hub.do/https://www.nature.com/articles/srep06010. Accessed August 11, 2021
Morgado AL, Xavier JM, Dionísio PA, Ribeiro MFC, Dias RB, Sebastião AM, Solá S, Rodrigues CMP (2015) MicroRNA-34a modulates neural stem cell differentiation by regulating expression of synaptic and Autophagic Proteins, 1168–1183. https://doi.org/10.1007/s12035-014-8794-6
Wu X, Fleming A, Ricketts T, Pavel M, Virgin H, Menzies FM, Rubinsztein DC (2016) Autophagy regulates Notch degradation and modulates stem cell development and neurogenesis. Nat Commun 7. https://doi.org/10.1038/NCOMMS10533
Ze-wei TAO, Long-gui LI (2007) Cell therapy in congestive heart failure 8:647–660. https://doi.org/10.1631/jzus.2007.B0647
Ro SH, Jang Y, Bae J, Kim IM, Schaecher C, Shomo ZD (2019) Autophagy in adipocyte browning: emerging drug target for intervention in obesity. Front Physiol 10:1–11. https://doi.org/10.3389/fphys.2019.00022
Zhang C, He Y, Okutsu M, Ong LC, Jin Y, Zheng L, Chow P, Yu S, Zhang M, Yan Z (2021) Autophagy is involved in adipogenic differentiation by repressesing proteasome-dependent PPARγ2 degradation. Am Physiol Soc 4:530–539. https://doi.org/10.1152/ajpendo.00640.2012
Levine B (2011) Autophagy in mammalian development and differentiation 12:823–830. https://doi.org/10.1038/ncb0910-823.Autophagy
Pellegrini C, Columbaro M, Schena E, Prencipe S, Andrenacci D, Iozzo P, Guzzardi MA, Capanni C, Mattioli E, Loi M, Araujo- D, Squarzoni S, Cinti S, Morselli P, Giorgetti A, Zanotti L, Gambineri A, Lattanzi G (2019) Altered adipocyte differentiation and unbalanced autophagy in type 2 Familial Partial Lipodystrophy: an in vitro and in vivo study of adipose tissue browning. Exp Mol Med. https://doi.org/10.1038/s12276-019-0289-0
Guo L, Huang JX, Liu Y, Li X, Zhou SR, Qian SW, Liu Y, Zhu H, Huang HY, Dang YJ, Tang QQ (2013) Transactivation of Atg4b by C/EBPβ promotes autophagy to facilitate adipogenesis. Mol Cell Biol 33:3180–3190. https://doi.org/10.1128/MCB.00193-13
Singh R, Xiang Y, Wang Y, Baikati K, Cuervo AM, Luu YK, Tang Y, Pessin JE, Schwartz GJ, Czaja MJ (2009) Autophagy regulates adipose mass and differentiation in mice 119:3329–3339. https://doi.org/10.1172/JCI39228DS1
Yin X, Zhou C, Li J (2019) Autophagy in bone homeostasis and the onset of osteoporosis. Bone Res. https://doi.org/10.1038/s41413-019-0058-7
Montaseri A, Giampietri C, Rossi M, Riccioli A, Del Fattore A, Filippini A (2020) Biomolecules The role of autophagy in osteoclast differentiation and bone resorption function, 1–16
Tsukamoto S, Yamamoto A, Tsukamoto S, Yamamoto A (2013) The role of autophagy in early mammalian embryonic development the role of autophagy in early mammalian embryonic development 30:86–94
Nakashima A, Aoki A, Kusabiraki T, Shima T, Yoshino O, Cheng S, Sharma S, Saito S (2017) Role of autophagy in oocytogenesis, embryogenesis, implantation, and pathophysiology of pre-eclampsia 43:633–643. https://doi.org/10.1111/jog.13292
Mizushima N (2007) FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells 9:497–510. https://doi.org/10.1083/jcb.200712064
Frudd K, Burgoyne T, Burgoyne JR (n.d.) Oxidation of Atg3 and Atg7 mediates inhibition of autophagy. Nat Commun, 1–15. https://doi.org/10.1038/s41467-017-02352-z
Maruyama T, Noda NN (2018) Autophagy-regulating protease Atg4: structure, function, regulation and inhibition. J Antibiot (Tokyo) 71:72–78. https://doi.org/10.1038/ja.2017.104
Fujita N, Itoh T, Omori H, Fukuda M, Noda T, Yoshimori T (2008) The Atg16L complex specifies the site of LC3 Lipidation for membrane biogenesis in autophagy 19:2092–2100. https://doi.org/10.1091/mbc.E07
Feng Y, Klionsky DJ (2017) Autophagic membrane delivery through ATG9. Cell Res 27:161–162. https://doi.org/10.1038/cr.2017.4
Ungermann C, Reggiori F (2018) Atg9 proteins, not so different after all. Autophagy 14:1456–1459. https://doi.org/10.1080/15548627.2018.1477382
Alers S, Wesselborg S, Stork B (2014) ATG13: just a companion, or an executor of the autophagic program?. Autophagy 10(6):944–956
Popelka H, Klionsky DJ (2017) The molecular mechanism of Atg13 function in autophagy induction: what is hidden behind the data ? Autophagy 13:449–451. https://doi.org/10.1080/15548627.2016.1277312
Kang R, Zeh HJ, Lotze MT, Tang D (2011) The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ 18:571–580. https://doi.org/10.1038/cdd.2010.191
Menon MB, Dhamija S (2018) Beclin 1 phosphorylation—at the center of autophagy regulation. Front Cell. Dev Biol 6:1–9. https://doi.org/10.3389/fcell.2018.00137
Rostislavleva K, Soler N, Ohashi Y, Zhang L, Pardon E, Burke JE, Masson GR, Johnson C, Steyaert J, Ktistakis NT, Williams RL (2015) Structure and flexibility of the endosomal Vps34 complex reveals the basis of its function on membranes. Science 80(350):1–25. https://doi.org/10.1126/science.aac7365
Anding AL, Baehrecke EH (2015) Vps15 is required for stress induced and developmentally triggered autophagy and salivary gland protein secretion in Drosophila. Cell Death Differ 22:457–464. https://doi.org/10.1038/cdd.2014.174
Iershov A, Nemazanyy I, Alkhoury C, Girard M, Barth E, Cagnard N, Montagner A, Chretien D, Rugarli EI, Guillou H, Pende M, Panasyuk G (2019) The class 3 PI3K coordinates autophagy and mitochondrial lipid catabolism by controlling nuclear receptor PPARα. Nat Commun 10:1–18. https://doi.org/10.1038/s41467-019-09598-9
Grimmel M, Backhaus C, Proikas T (2015) WIPI-mediated autophagy and longevity. Cells 4:202–217. https://doi.org/10.3390/cells4020202
Proikas T, Takacs Z, Dönnes P, Kohlbacher O (2015) WIPI proteins: essential PtdIns3P effectors at the nascent autophagosome. J Cell Sci 128:207–217. https://doi.org/10.1242/jcs.146258
Liu WJ, Ye L, Huang WF, Guo LJ, Xu ZG, Wu HL, Yang C, Liu HF (2016) P62 links the autophagy pathway and the Ubiqutin-proteasome system upon Ubiquitinated protein degradation. Cell Mol Biol Lett 21:1–14. https://doi.org/10.1186/s11658-016-0031-z
Kageyama S, Gudmundsson SR, Sou YS, Ichimura Y, Tamura N, Kazuno S, Ueno T, Miura Y, Noshiro D, Abe M, Mizushima T, Miura N, Okuda S, Motohashi H, Lee JA, Sakimura K, Ohe T, Noda NN, Waguri S, Eskelinen EL, Komatsu M (2021) p62/SQSTM1-droplet serves as a platform for autophagosome formation and anti-oxidative stress response. Nat Commun 12. https://doi.org/10.1038/s41467-020-20185-1
Tanida I, Ueno T, Kominami E (2008) LC3 and autophagy. Methods Mol Biol 445:77–88. https://doi.org/10.1007/978-1-59745-157-4_4
Runwal G, Stamatakou E, Siddiqi FH, Puri C, Zhu Y, Rubinsztein DC (2019) LC3-positive structures are prominent in autophagy-deficient cells. Sci Rep 9:1–14. https://doi.org/10.1038/s41598-019-46657-z
Maria Fimia G, Stoykova A,. Romagnoli A, Giunta L, Di Bartolomeo S, Nardacci R, Corazzari M, Fuoco C, Ucar A, Schwartz P, Gruss P, Piacentini M, Chowdhury K, Cecconi F(2007) Ambra1 regulates autophagy and development of the nervous system. Nature 447:1121–1125. https://doi.org/10.1038/nature05925
Sun WL (2016) Ambra1 in autophagy and apoptosis: Implications for cell survival and chemotherapy resistance. Oncol Lett 12:367–374. https://doi.org/10.3892/ol.2016.4644
Shen X, Kan S, Liu Z, Lu G, Zhang X, Chen Y, Bai Y (2017) EVA1A inhibits GBM cell proliferation by inducing autophagy and apoptosis. Exp Cell Res 352:130–138. https://doi.org/10.1016/j.yexcr.2017.02.003
Hu J, Li G, Qu L, Li N, Liu W, Xia D, Hongdu B, Lin X, Xu C, Lou Y, He Q, Ma D, Chen Y (2016) TMEM166/EVA1A interacts with ATG16L1 and induces autophagosome formation and cell death. Cell Death Dis 7:1–13. https://doi.org/10.1038/cddis.2016.230
Acknowledgements
We would like to acknowledge the DST-SERB grant (EMR/2017/004149) provided to SM and DRDO-LSRB grant (O/o DG(TM)/81/48222/LSRB-351/PEE&BS/2019) provided to RC as a funding source for our research. The authors acknowledge BITS Pilani, Pilani campus, for providing infrastructural support. AKS and PB acknowledge DST-SERB (EMR/2017/004149) and DRDO (O/o DG(TM)/81/48222/LSRB-351/PEE&BS/2019), respectively, for providing fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
Funding Source
DST-SERB grant (EMR/2017/004149) provided to SM and DRDO-LSRB grant (O/o DG(TM)/81/48222/LSRB-351/PEE&BS/2019) provided to RC.
Disclosure of Interests
All authors declare they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Sahu, A.K., Bandyopadhyay, P., Chowdhury, R., Mukherjee, S. (2023). Autophagy in Stem Cell Maintenance and Differentiation. In: Shravage, B.V., Turksen, K. (eds) Autophagy in Stem Cell Maintenance and Differentiation. Stem Cell Biology and Regenerative Medicine, vol 73. Springer, Cham. https://doi.org/10.1007/978-3-031-17362-2_2
Download citation
DOI: https://doi.org/10.1007/978-3-031-17362-2_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-17361-5
Online ISBN: 978-3-031-17362-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)