Abstract
Uniting three self-supervised learning (SSL) ingredients (discriminative, restorative, and adversarial learning) enables collaborative representation learning and yields three transferable components: a discriminative encoder, a restorative decoder, and an adversary encoder. To leverage this advantage, we have redesigned five prominent SSL methods, including Rotation, Jigsaw, Rubik’s Cube, Deep Clustering, and TransVW, and formulated each in a United framework for 3D medical imaging. However, such a United framework increases model complexity and pretraining difficulty. To overcome this difficulty, we develop a stepwise incremental pretraining strategy, in which a discriminative encoder is first trained via discriminative learning, the pretrained discriminative encoder is then attached to a restorative decoder, forming a skip-connected encoder-decoder, for further joint discriminative and restorative learning, and finally, the pretrained encoder-decoder is associated with an adversarial encoder for final full discriminative, restorative, and adversarial learning. Our extensive experiments demonstrate that the stepwise incremental pretraining stabilizes United models training, resulting in significant performance gains and annotation cost reduction via transfer learning for five target tasks, encompassing both classification and segmentation, across diseases, organs, datasets, and modalities. This performance is attributed to the synergy of the three SSL ingredients in our United framework unleashed via stepwise incremental pretraining. All codes and pretrained models are available at GitHub.com/JLiangLab/StepwisePretraining.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Self-supervised learning
- Discriminative learning
- Restorative learning
- Adversarial learning
- United framework
- Stepwise pretraining
1 Introduction
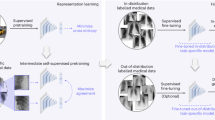
Self-supervised learning (SSL) [11] pretrains generic source models [20] without using expert annotation, allowing the pretrained generic source models to be quickly fine-tuned into high-performance application-specific target models with minimal annotation cost [18]. The existing SSL methods may employ one or a combination of the following three learning ingredients [9]: (1) discriminative learning, which pretrains an encoder by distinguishing images associated with (computer-generated) pseudo labels; (2) restorative learning, which pretrains an encoder-decoder by reconstructing original images from their distorted versions; and (3) adversarial learning, which pretrains an additional adversary encoder to enhance restorative learning. Haghighi et al. articulated a vision and insights for integrating three learning ingredients in one single framework for collaborative learning [9], yielding three learned components: a discriminative encoder, a restorative decoder, and an adversary encoder (Fig. 1). However, such integration would inevitably increase model complexity and pretraining difficulty, raising these two questions: (a) how to optimally pretrain such complex generic models and (b) how to effectively utilize pretrained components for target tasks?
Our United model consists of three components: a discriminative encoder, a restorative decoder, and an adversary encoder, where the discriminative encoder and the restorative decoder are skip connected, forming an encoder-decoder. To overcome the United model complexity and pretraining difficulty, we develop a strategy to incrementally train the three components in a stepwise fashion: (1) Step D trains the discriminative encoder via discriminative learning; (2) Step (D)+R attaches the pretrained discriminative encoder to the restorative decoder for further joint discriminative and restorative learning; (3) Step ((D)+R)+A associates the pretrained encoder-decoder with the adversarial encoder for final full discriminative, restorative, and adversarial learning. This stepwise incremental pretraining has proven to be reliable across multiple SSL methods (Fig. 2) for a variety of target tasks across diseases, organs, datasets, and modalities.
To answer these two questions, we have redesigned five prominent SSL methods for 3D imaging, including Rotation [7], Jigsaw [13], Rubik’s Cube [21], Deep Clustering [4], and TransVW [8], and formulated each in a single framework called “United” (Fig. 2), as it unites discriminative, restorative, and adversarial learning. Pretraining United models, i.e., all three components together, directly from scratch is unstable; therefore, we have investigated various training strategies and discovered a stable solution: stepwise incremental pretraining. An example of such pretraining follows: first training a discriminative encoder via discriminative learning (called Step D), then attaching the pretrained discriminative encoder to a restorative decoder (i.e., forming an encoder-decoder) for further combined discriminative and restorative learning (called Step (D)+R), and finally associating the pretrained autoencoder with an adversarial-encoder for the final full discriminative, restorative, and adversarial training (called Step ((D)+R)+A). This stepwise pretraining strategy provides the most reliable performance across most target tasks evaluated in this work encompassing both classification and segmentation (see Table 2 and 3 as well as Table 4 in the Supplementary Material).
Redesigning five prominent SSL methods: (a) Jigsaw, (b) Rubik’s Cube, (c) Deep Clustering, (d) Rotation, and (e) TransVW in a United framework. The original Jigsaw [13], Deep Clustering [4], and Rotation [7] were proposed for 2D image analysis employing discriminative learning alone and provided only pretrained encoders; therefore, in our United framework (a, c, d), these methods have been augmented with two new components (in light blue) for restorative learning and adversarial learning and re-implemented in 3D. The code for the original Rubik’s Cube [21] is not released and thus reimplemented and augmented with new learning ingredients in light blue (b). The original TransVW [8] is supplemented with adversarial learning (c). Following our redesign, all five methods provide all three learned components: discriminative encoders, restorative decoders, and adversary encoders, which are transferable to target classification and segmentation tasks. (Color figure online)
Through our extensive experiments, we have observed that (1) discriminative learning alone (i.e., Step D) significantly enhances discriminative encoders on target classification tasks (e.g., +3% and +4% AUC improvement for lung nodule and pulmonary embolism false positive reduction as shown in Table 2) relative to training from scratch; (2) in comparison with (sole) discriminative learning, incremental restorative pretraining combined with continual discriminative learning (i.e., Step (D)+R) enhances discriminative encoders further for target classification tasks (e.g., +2% and +4% AUC improvement for lung nodule and pulmonary embolism false positive reduction as shown in Table 2) and boosts encoder-decoder models for target segmentation tasks (e.g., +3%, +7%, and +5% IoU improvement for lung nodule, liver, and brain tumor segmentation as shown in Table 3); and (3) compared with Step (D)+R, the final stepwise incremental pretraining (i.e., Step: ((D)+R)+A) generates sharper and more realistic medical images (e.g., FID decreases from 427.6 to 251.3 as shown in Table 5 in the Supplementary Material) and further strengthens each component for representation learning, leading to considerable performance gains (see Fig. 3) and annotation cost reduction (e.g., 28%, 43%, and 26% faster for lung nodule false positive reduction, lung nodule tumor segmentation, and pulmonary embolism false positive reduction as shown in Fig. 4) for five target tasks across diseases, organs, datasets, and modalities.
We should note that recently Haghighi et al. [9] also combined discriminative, restorative, and adversarial learning, but our findings complement theirs, and more importantly, our method significantly differs from theirs, because they were more concerned with contrastive learning (e.g., MoCo-v2 [5], Barlow Twins [19], and SimSiam [6]) and focused on 2D medical image analysis. By contrast, we are focusing on 3D medical imaging by redesigning five popular SSL methods beyond contrastive learning. As they acknowledged [9], their results on TransVW [8] augmented with an adversarial encoder were based on the experiments presented in this paper. Furthermore, this paper focuses on a stepwise incremental pretraining to stabilize United model training, revealing new insights into synergistic effects and contributions among the three learning ingredients.
In summary, we make the following three main contributions:
-
1.
A stepwise incremental pretraining strategy that stabilizes United models’ pretraining and unleashes the synergistic effects of the three SSL ingredients;
-
2.
A collection of pretrained United models that integrate discriminative, restorative, and adversarial learning in a single framework for 3D medical imaging, encompassing both classification and segmentation tasks;
-
3.
A set of extensive experiments that demonstrate how various pretraining strategies benefit target tasks across diseases, organs, datasets, and modalities.
2 Stepwise Incremental Pretraining
We have redesigned five prominent SSL methods, including Rotation, Jigsaw, Rubik’s Cube, Deep Clustering, and TransVW, and augmented each with the missing components under our United framework (Fig. 2). A United model (Fig. 1) is a skip-connected encoder-decoder associated with an adversary encoder. With our redesign, for the first time, all five methods have all three SSL components. We incrementally train United models component by component in a stepwise manner, yielding three learned transferable components: discriminative encoders, restorative decoders, and adversarial encoders. The pretrained discriminative encoder can be fine-tuned for target classification tasks; the pretrained discriminative encoder and restorative decoder, forming a skip-connected encoder-decoder network (i.e., U-Net [14, 16]), can be fine-tuned for target segmentation tasks.
Discriminative learning trains a discriminative encoder \(D_\theta \), where \(\theta \) represents the model parameters, to predict target label \(y \in Y\) from input \(x \in X\) by minimizing a loss function for \(\forall x \in X\) defined as
where N is the number of samples, K is the number of classes, and \(p_{nk}\) is the probability predicted by \(D_\theta \) for \(x_{n}\) belonging to Class k; that is, \(p_{n}=D_\theta (x_{n})\) is the probability distribution predicted by \(D_\theta \) for \(x_{n}\) over all classes. In SSL, the labels are automatically obtained based on the properties of the input data, involving no manual annotation. All five SSL methods in this work have a discriminative component formulated as a classification task, while other discriminative losses can be used, such as contrastive losses in MoCo-v2 [5], Barlow Twins [19], and SimSiam [6].
Restorative learning trains an encoder-decoder \((D_\theta ,R_{\theta '})\) to reconstruct an original image x from its distorted version \(\mathcal {T}(x)\), where \(\mathcal {T}\) is a distortion function, by minimizing pixel-level reconstruction error:
where \(L_2(u,v)\) is the sum of squared pixel-by-pixel differences between u and v.
Adversarial learning trains an additional adversary encoder, \(A_{\theta ''}\), to help the encoder-decoder \((D_\theta ,R_{\theta '})\) reconstruct more realistic medical images and in turn strengthen representation learning. The adversary encoder learns to distinguish fake image pair \((R_{\theta '}(D_\theta (\mathcal {T}(x))), \mathcal {T}(x))\) from real pair \((x, \mathcal {T}(x))\) via an adversarial loss:
The final objective combines all losses:
where \(\lambda _d\), \(\lambda _r\), and \(\lambda _a\) controls the importance of each learning ingredients.
Stepwise incremental pretraining trains our United models continually component-by-component because training a whole United model in an end-to-end fashion (i.e., all three components together directly from scratch)—a strategy called (D+R+A)—is unstable. For example, as shown in Table 1, Strategy ((D)+R)+A) (see Fig. 1) always outperforms Strategy (D+R+A) and provides the most reliable performance across most target tasks evaluated in this work.
 and
and
 denote a non-pretrained decoder and with pretrained decoders but not used for target tasks. With incremental restorative learning, the performance gains were consistent for both target tasks.
denote a non-pretrained decoder and with pretrained decoders but not used for target tasks. With incremental restorative learning, the performance gains were consistent for both target tasks.3 Experiments and Results
Datasets and Metrics. To pretrain all five United models, we used 623 CT scans from the LUNA16 [15] dataset. We adopted the same strategy as [20], and cropped sub-volumes with a pixel size of \(64\times 64\times 64\). To evaluate the effectiveness of pretraining the five methods, we tested their performance on five 3D medical imaging tasks (See §B) including BraTS [2, 12], LUNA16 [15], LiTS [3], PE-CAD [17], and LIDC-IDRI [1]. The acronyms BMS, LCS, and NCS denote the tasks of segmenting a brain tumor, liver, and lung nodules; NCC and ECC denote the tasks of reducing lung nodule and pulmonary embolism false positives results, respectively. We measured the performances of the pretrained models on five target tasks and reported the AUC (Area Under the ROC Curve) for classification tasks and IoU (Intersection over Union) for segmentation tasks. All target tasks ran at least 10 times and statistical analysis was performed using independent two-sample t-test.
 ,
,
 , and ✓ denote a non-pretrained decoder, not using pretrained decoders, and using pretrained decoders, respectively. Statistic analysis (*\(p<\)0\(\cdot \)5, **\(p<\)0\(\cdot \)1, ***\(p<\)0\(\cdot \)05) was conducted between
, and ✓ denote a non-pretrained decoder, not using pretrained decoders, and using pretrained decoders, respectively. Statistic analysis (*\(p<\)0\(\cdot \)5, **\(p<\)0\(\cdot \)1, ***\(p<\)0\(\cdot \)05) was conducted between
 and ✓.
and ✓.(1) Incremental restorative learning ((D)+R) enhances discriminative encoders further for classification tasks. After pretraining discriminative encoders, we append restorative decoders to the end of the encoders and continue to pretrain discriminative encoder and restorative decoder together. The incremental restorative learning significantly enhances encoders in classification tasks, as shown in Table 2. Specifically, compared with the original methods, the incremental restorative learning improves Jigsaw by AUC scores of 1.9% and 2.6% in NCC and ECC; similarly, it improves Rubik’s Cube by 1.9% and 2.4%, Deep Clustering by 0.9% and 0.3%, TransVW by 1.0% and 2.9%, and Rotation by 1.0% and 1.2%. The discriminative encoders are enhanced because they not only learn global features for discriminative tasks but also learns fine-grained features through incremental restorative learning.
(2) Incremental restorative learning ((D)+R) directly boost target segmentation tasks. Most state-of-the-art segmentation methods do not pretrain their decoders but instead initialize them at random [5, 10]. We argue that the random decoders are suboptimal, evidenced by the data in Table 3, and we demonstrate that the incremental pretrained restorative decoders can directly boost target segmentation tasks. In particular, compared with the original methods, the incremental pretrained restorative decoder improves Jigsaw by 1.2%, 2.1% and 2.0% IoU improvement in NCS, LCS and BMS; similarly, it improves Rubik’s Cube by 2.8%, 7.6%, and 3.1%, Deep Clustering by 1.1%, 2.0%, and 0.9%, TransVW by 0.4%, 1.4%, and 4.8% and Rotation by 0.6%, 2.2% and 1.5%. The consistent performance gain suggests that a wide variety of target segmentation tasks can benefit from our incremental pretrained restorative decoders.
Adversarial training strengthens learned representation. Target tasks performance are generally increased (red) following the adversarial training. Although some target tasks show a decrease (pink), these reductions do not reach statistical significance according to the t-test. (Color figure online)
Adversarial training reduces annotation costs. TransVW combined with adversarial learning reduces the annotation cost by 28%, 43%, and 26% for target tasks of NCC, NCS, and ECC, respectively, compared with the original TransVW. It also reduces the annotation cost by 57%, 61%, and 66% for target tasks of NCC, NCS, and ECC, respectively, compared with initializing the network from random.
(3) Adversarial training strengthens representation and reduces annotation costs. Quantitative measurements shown in Table 5 reveal that adversarial training can generate sharper and more realistic images in the restoration proxy task. More importantly, we found that adversarial training also makes a significant contribution to pretraining. First, as shown in Fig. 3, adding adversarial training can benefit most target tasks, particularly segmentation tasks. The incremental adversarial pretraining improves Jigsaw by AUC scores of 0.3%, 0.7%, and 0.7% in NCS, LCS, and BMS; similarly, it improves Rubik’s Cube by 0.4%, 1.0%, and 1.0%, Deep Clustering by 0.5%, 0.5%, and 0.5%, TransVW by 0.2%, 0.3%, and 0.8% and Rotation by 0.1%, 0.1%, and 0.7%. Additionally, incremental adversarial pretraining improves performance on small data regimes. Figure 4 shows that incremental adversarial pretrained TransVW [8] can reduce the annotation cost by 28%, 43%, and 26% on NCC, NCS, and ECC, respectively, compared with TransVW [8].
4 Conclusion
We have developed a United framework that integrates discriminative SSL methods with restorative and adversarial learning. Our extensive experiments demonstrate that our pretrained United models consistently outperform the SoTA baselines. This performance improvement is attributed to our stepwise pertaining scheme, which not only stabilizes the pretraining but also unleashes the synergy of discriminative, restorative, and adversarial learning. We expect that our pretrained United models will exert an important impact on medical image analysis across diseases, organs, modalities, and specialties.
References
Armato III, S.G., et al.: The lung image database consortium (LIDC) and image database resource initiative (IDRI): a completed reference database of lung nodules on CT scans. Med. Phys. 38(2), 915–931 (2011)
Bakas, S., et al.: Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the brats challenge. arXiv preprint arXiv:1811.02629 (2018)
Bilic, P., et al.: The liver tumor segmentation benchmark (LiTS). arXiv preprint arXiv:1901.04056 (2019)
Caron, M., Bojanowski, P., Joulin, A., Douze, M.: Deep clustering for unsupervised learning of visual features. In: European Conference on Computer Vision (2018)
Chen, X., Fan, H., Girshick, R., He, K.: Improved baselines with momentum contrastive learning (2020)
Chen, X., He, K.: Exploring simple Siamese representation learning. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 15750–15758, June 2021
Gidaris, S., Singh, P., Komodakis, N.: Unsupervised representation learning by predicting image rotations (2018)
Haghighi, F., Taher, M.R.H., Zhou, Z., Gotway, M.B., Liang, J.: Transferable visual words: exploiting the semantics of anatomical patterns for self-supervised learning. IEEE Trans. Med. Imaging, 1 (2021). https://doi.org/10.1109/TMI.2021.3060634
Haghighi, F., Taher, M.R.H., Gotway, M.B., Liang, J.: DiRA: discriminative, restorative, and adversarial learning for self-supervised medical image analysis. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 20824–20834 (2022)
He, K., Fan, H., Wu, Y., Xie, S., Girshick, R.: Momentum contrast for unsupervised visual representation learning (2020)
Jing, L., Tian, Y.: Self-supervised visual feature learning with deep neural networks: a survey. IEEE Trans. Pattern Anal. Mach. Intell. 43(11), 4037–4058 (2020)
Menze, B.H., et al.: The multimodal brain tumor image segmentation benchmark (BRATS). IEEE Trans. Med. Imaging 34(10), 1993–2024 (2014)
Noroozi, M., Favaro, P.: Unsupervised learning of visual representations by solving jigsaw puzzles. In: Leibe, B., Matas, J., Sebe, N., Welling, M. (eds.) ECCV 2016. LNCS, vol. 9910, pp. 69–84. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46466-4_5
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Setio, A.A.A., et al.: Validation, comparison, and combination of algorithms for automatic detection of pulmonary nodules in computed tomography images: the LUNA16 challenge. Med. Image Anal. 42, 1–13 (2017)
Siddique, N., Sidike, P., Elkin, C., Devabhaktuni, V.: U-Net and its variants for medical image segmentation: theory and applications (2020). http://arxiv.org/abs/2011.01118
Tajbakhsh, N., Gotway, M.B., Liang, J.: Computer-aided pulmonary embolism detection using a novel vessel-aligned multi-planar image representation and convolutional neural networks. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9350, pp. 62–69. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24571-3_8
Tajbakhsh, N., Roth, H., Terzopoulos, D., Liang, J.: Guest editorial annotation-efficient deep learning: the holy grail of medical imaging. IEEE Trans. Med. Imaging 40(10), 2526–2533 (2021)
Zbontar, J., Jing, L., Misra, I., LeCun, Y., Deny, S.: Barlow twins: self-supervised learning via redundancy reduction. arXiv:2103.03230 (2021)
Zhou, Z., Sodha, V., Pang, J., Gotway, M.B., Liang, J.: Models genesis. Med. Image Anal. 67, 101840 (2021). https://doi.org/10.1016/j.media.2020.101840
Zhuang, X., Li, Y., Hu, Y., Ma, K., Yang, Y., Zheng, Y.: Self-supervised feature learning for 3D medical images by playing a Rubik’s cube. In: Shen, D., et al. (eds.) MICCAI 2019. LNCS, vol. 11767, pp. 420–428. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-32251-9_46
Acknowledgments
We thank F. Haghighi, M. R. Hosseinzadeh Taher, and Z. Zhou for their discussions, debates, and supports in implementing the earlier ideas behind “United & Unified” and in drafting earlier versions. This research has been supported in part by ASU and Mayo Clinic through a Seed Grant and an Innovation Grant, and in part by the NIH under Award Number R01HL128785. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH. This work has utilized the GPUs provided in part by the ASU Research Computing and in part by the Extreme Science and Engineering Discovery Environment (XSEDE) funded by the National Science Foundation (NSF) under grant numbers: ACI-1548562, ACI-1928147, and ACI-2005632. The content of this paper is covered by patents pending.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Guo, Z., Islam, N.U., Gotway, M.B., Liang, J. (2022). Discriminative, Restorative, and Adversarial Learning: Stepwise Incremental Pretraining. In: Kamnitsas, K., et al. Domain Adaptation and Representation Transfer. DART 2022. Lecture Notes in Computer Science, vol 13542. Springer, Cham. https://doi.org/10.1007/978-3-031-16852-9_7
Download citation
DOI: https://doi.org/10.1007/978-3-031-16852-9_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-16851-2
Online ISBN: 978-3-031-16852-9
eBook Packages: Computer ScienceComputer Science (R0)