Abstract
Arbuscular mycorrhizal fungi (AMF) of the Glomeromycotina subphylum are one of the oldest fungal lineages for which the mechanistic underpinning of genetic diversity is unknown. They are present in all terrestrial ecosystems and interact with the majority of land plants, significantly impacting global nutrient cycling. The study of genomes of AMF is of fundamental importance for understanding their evolutionary history and the molecular bases of symbiosis. Here we summarize the current knowledge of AMF genome organization, regulation, and transmission. We discuss the implications of recent findings in our understanding of AMF biodiversity, adaptation, and evolution.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Fossil evidence and molecular phylogenies suggest that mycorrhizal symbioses were established with the earliest plants to colonize the earth land surface >450 Mya, initiating long-term co-evolution (Morris et al. 2018; Rich et al. 2021; Selosse and Le Tacon 1998; Strullu-Derrien et al. 2018). While plants and other fungal groups have experienced extinction, radiation events, and diversification, fungi from the Glomeromycotina appear to have poorly diversified morphologically (Kruger et al. 2012). Yet, their ecological success is undeniable since today arbuscular mycorrhizal fungi (AMF) are present on all continents, in environments ranging from tropical forests to Antarctica. Despite their ecological importance (Smith and Read 2008), the molecular mechanisms underlying AMF adaptation and evolution are still elusive. This is partly because AMF defy many of the cellular and molecular approaches that are possible for other eukaryotes.
The definition of the cell in AMF is peculiar. Hyphae form a non-septate mycelium (syncytium) where nuclei and cytoplasmic contents flow bidirectionally. AMF spores contain hundreds of nuclei, and the transition from one ‘generation’ to the next involves carrying over of multiple nuclear genomes (Ehinger et al. 2012; Jany and Pawlowska 2010; Marleau et al. 2011). The concept of the individual is also blurry: compatible mycelial networks can fuse (a process called anastomosis) (Bago et al. 1999; Cardenas-Flores et al. 2011; Purin and Morton 2013; Sbrana et al. 2018), allowing for horizontal exchange of genetic material which can give rise to a network bearing more than one nucleotype (heterokaryon) (Croll et al. 2009; Hijri and Sanders 2005; Ropars et al. 2016; Serghi et al. 2021). At least three AMF species can anastomose (Rhizophagus irregularis, Rhizophagus clarus, and Funneliformis mosseae) although heterokaryosis has been reported so far only in R. irregularis. It is unclear whether these processes occur in other fungi of the Glomeromycotina. As obligate biotrophs, AMF are not amenable to genetic transformation approaches that require cultivation outside the host. The AMF lifecycle is dependent on interaction with plants and the provision of fatty acids (Bravo et al. 2017; Jiang et al. 2017; Keymer et al. 2017; Luginbuehl et al. 2017). Axenic culture is therefore limited to in vitro spore germination experiments (Dallaire et al. 2021; Kamel et al. 2017; Nadal et al. 2017), which can be extended up to a single next generation of spores by the addition of specific fatty acids and plant-derived hormones (Kameoka et al. 2019; Sugiura et al. 2020). AMF can be grown in co-culture with transformed host hairy roots, a system that is useful for symbiosis-related research, but which likely does not fully recapitulate the metabolic and developmental complexity of symbiotic relationships occurring in nature.
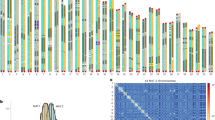
These features make AMF a unique biological mystery, with the unfortunate drawback of being intractable to molecular genetics approaches. Transcriptomic, proteomic, and metabolomic analyses have therefore proven most useful to reveal molecular components involved in AMF biology. The recent accessibility of genome sequencing technologies that can resolve complex repeats and haplotype heterozygosity promises to enable genome-scale analyses of the Glomeromycotina phylum and accelerate our understanding of AMF genetic diversity and evolution. Comparative and population genomics are therefore the next frontier to bridge molecular biology and evolutionary genetics of AMF. In this chapter, we discuss key observations of genome organization, regulation, and transmission in AMF and identify gaps in our understanding of symbiotic genome evolution and lifestyles (Fig. 4.1).
2 Organization of the Genome
To date, few AMF species have been isolated and sustainably cultivated. While over 200 AMF species have been defined so far (Davison et al. 2015; Öpik and Davison 2016), full genome sequences are available for only nine AMF species—Diversispora epigaea, R. irregularis, R. clarus, R. cerebriforme, R. diaphanus, R. proliferus, Gigaspora margarita, Gigaspora rosea, and Geosiphon pyriformis (Chen et al. 2018b; Kobayashi et al. 2018; Lin et al. 2014; Malar et al. 2021; Morin et al. 2019; Prasad Singh et al. 2019; Sun et al. 2019; Tisserant et al. 2013; Venice et al. 2020). AMF genomes have been difficult to assemble because of their high content of repetitive sequences. Thanks to long-read sequencing and chromatin proximity-guided scaffolding, this problem is now solved and five strains of the model species R. irregularis have been assembled to chromosome-scale (Yildirir et al. 2022). Given their widespread distribution across the globe, considerable genetic diversity remains to be explored, and the community will greatly benefit from large-scale isolation and (re)-sequencing projects.
Fungi from the Glomeromycotina have some of the largest genome sizes in the fungal kingdom (Stajich 2017). The model AMF, R. irregularis, has telomeres with the classical (TTAGGG)n sequence (Yildirir et al. 2022), but centromeric structures are still undefined (Friedman and Freitag 2017). In the early-diverging fungal phyla Chytridiomycota and Mucoromycota, the coding space is characterized by the retention of ancestral introns and high intron densities (Lim et al. 2021). Intron retention is proposed to correlate with reduced rates of evolutionary change, whereas fast-evolving lineages experienced more intron loss (Lim et al. 2021). Supporting the notion that Glomeromycotina fungi are slow evolving, recent analyses of small subunit rRNA gene diversity showed that their speciation rates are one order of magnitude lower than those of other eukaryotes (Perez-Lamarque et al. 2022).
Comparative analyses revealed that AMF have lost genes required for plant cell wall degradation, thiamine biosynthesis, secondary metabolism, and lipid production (Lin et al. 2014; Tisserant et al. 2013; Wewer et al. 2014). These results proved to be physiologically relevant, since it was later discovered that the obligate biotrophy of AMF relies on the provision of lipids by plants (Bravo et al. 2017; Jiang et al. 2017; Keymer et al. 2017; Luginbuehl et al. 2017). AMF genomes also contain gene family expansions with predicted protein functions in signalling, protein–protein interaction, and RNA interference (RNAi) (Chen et al. 2018b; Maeda et al. 2018; Morin et al. 2019; Tisserant et al. 2013).
Evidence of within-strain phenotypic and genetic diversity has been accumulating for decades (Angelard and Sanders 2011; Angelard et al. 2014; Boon et al. 2010; Corradi et al. 2007; Corradi and Sanders 2006; Croll et al. 2009; Koch et al. 2004; Mathieu et al. 2018; Ropars et al. 2016; Savary et al. 2018; Wyss et al. 2016). Recently, methods of prokaryotic pangenome analysis have been applied to R. irregularis and other fungi to investigate the functional outcome of intraspecific genetic variation. The pangenome is the distinction between a set of “core” genes found among all strains or individuals of a species, and a set of “accessory” genes, found in a subset of, but not all strains of a species. Accessory genes might encode functions that are not essential, but which confer selective advantages, such as adaptation to different niches, resistance to pathogens, or host range expansion (Medini et al. 2005). Knowledge of the content and the dynamics of a pangenome informs on the selective pressures experienced by a population. Evidence for pangenomic structure was found in five species of fungi. The accessory gene sets of the non-pathogenic fungi Saccharomyces cerevisiae, Candida albicans, Cryptococcus neoformans, and Aspergillus fumigatus species range between 10% and 20% (McCarthy and Fitzpatrick 2019), while the major wheat pathogen Zymoseptoria tritici contains up to 40% accessory genes (Bergstrom et al. 2014; Dunn et al. 2012; Peter et al. 2018; Plissonneau et al. 2018; Song et al. 2015). In Z. tritici and Fusarium species, entire chromosomes have been deemed accessory because they show extensive presence/absence variation and rearrangements among strains, and encode accessory virulence factors (Ma et al. 2010; Moller et al. 2018; Plissonneau et al. 2018). A pangenomic analysis of five R. irregularis strains revealed up to 50% of accessory coding sequence (Chen et al. 2018b). These accessory genes are part of large family expansions with predicted functions in signalling and signal transduction, and small secreted proteins (Chen et al. 2018b). Since their expression tends to be induced in planta, these genes were proposed to play adaptive roles in communication and exchange with host plants (Chen et al. 2018b; Mathieu et al. 2018; Reinhardt et al. 2021). Contrary to Z. tritici, the number of R. irregularis chromosomes appears stable in strains studied so far (Yildirir et al. 2022), lessening the possibility of extreme genome instability in AMF. Nevertheless, chromosomal rearrangements have been observed in high-quality long-read-based assemblies (Yildirir et al. 2022), and further analyses of strain-specific structural variation may explain presence/absence variation of accessory genes.
In R. irregularis, core and accessory genes have different distributions relative to repetitive elements: accessory genes tend to be sparsely distributed and located next to specific transposable elements (TEs), while core genes form denser clusters in less repetitive regions (Dallaire et al. 2021; Yildirir et al. 2022). In filamentous/eukaryotic plant pathogens, virulence effectors are sometimes embedded into TE-rich regions, and this effectively increases the likelihood of their sequences being reshuffled (Dong et al. 2015; Faino et al. 2016). The increased mutation rate observed in TE-rich regions was therefore proposed to support an evolutionary arms race with correspondingly highly variable immune associated gene sets in the host plant. In AMF, the molecular rate of evolution of TE-linked accessory genes remains to be investigated and may reflect the selective pressures associated with an obligate mutualist lifestyle.
3 Regulation of the Genome(s)
Eukaryotic genomic DNA is folded and compacted to form chromatin, which regulates gene expression, as well as DNA replication and repair. Chromatin is divided into euchromatin and heterochromatin, where euchromatin is generally gene-rich, permissive to transcription, and localized towards the periphery of the nucleolus, whereas heterochromatin is typically gene-poor, repeat-rich, and refractory to the transcription machinery. Historically, TEs were the first genetic elements shown to be enriched in heterochromatin (McClintock 1951), a location that restrains their activity and maintains genomic stability. Since then, the presence or absence of specific histone modifications, DNA methylation, pericentromeric regions, and targeting by small RNAs have also been used to distinguish heterochromatin from euchromatin in various organisms, including fungi (Tamaru 2010). In the model AMF R. irregularis, DNA folding was assessed using chromosome conformation capture (Hi-C), which revealed a checkerboard pattern consistent with intra- and inter-chromosomal contacts (Yildirir et al. 2022). Two compartments named A and B were detected, which reflect the folding of the chromosomes into euchromatin and heterochromatin, respectively.
Consistent with the fact that the location of DNA sequences within chromatin coincides with particular transcriptional states, genes and TEs of R. irregularis were shown to be differentially partitioned into A and B compartments (Dallaire et al. 2021; Yildirir et al. 2022). The euchromatic A compartment preferentially comprises genes with higher expression levels, lower DNA methylation frequencies, and fewer repeats. Strikingly, accessory genes and genes up-regulated during symbiosis were found to be enriched in heterochromatic B compartments, suggesting that their repression by epigenetic mechanisms and chromosome topology might be environmentally-responsive, even perhaps host-responsive. Although the formation of heterochromatin is essential for regulating gene expression and silencing mobile genetic elements, some heterochromatic regions can retain the potential to switch into a euchromatic state following certain cues. Genes that are regulated developmentally or in a cell-type-specific manner are often found in such facultative heterochromatin (Trojer and Reinberg 2007). The regulation of facultative heterochromatin in AMF may involve histone modification, DNA methylation, and small and long non-coding RNAs that modulate boundary formation between different chromatin domains (Cohen and Jia 2014; Freitag et al. 2004; Klocko et al. 2016; Saksouk et al. 2015; Smith et al. 2011).
AMF genomes encode DNA cytosine methyltransferases and an expanded repertoire of RNAi pathway genes (Dallaire et al. 2021; Silvestri et al. 2019, 2020; Tisserant et al. 2013). Whole-genome epigenomic profiling of R. irregularis provided direct evidence of 5-methylcytosine (5mC) DNA methylation and small RNA production which occurs mostly at TE loci, suggesting ongoing epigenetic regulation (Dallaire et al. 2021; Silvestri et al. 2019; Yildirir et al. 2022). In the AMF G. margarita, most small RNA-generating loci are intergenic and show similarity to fungal repetitive elements (Silvestri et al. 2020), indicating a conserved contribution of RNAi in suppressing TE activity. In both G. margarita and R. irregularis, few genes were found to be highly methylated or to be targeted by small RNAs (Dallaire et al. 2021; Silvestri et al. 2020), and in R. irregularis these genes did not encode proteins with a clear enrichment for specific domains or functions (Dallaire et al. 2021). The biological relevance of DNA methylation and RNAi in regulating AMF gene expression is therefore still elusive. DNA N6-methyldeoxyadenine (6 mA), a modification with very low abundance and an unclear role in eukaryotes (Zhao et al. 2020), was also investigated as a potential epigenetic mark in R. irregularis and other early-diverging fungi and Dikarya (Chaturvedi et al. 2021; Mondo et al. 2017). In R. irregularis, the presence of 6 mA on DNA is associated with a subset of transcriptionally active genes with predicted functions in phosphate regulation (Chaturvedi et al. 2021). It is important to consider that in mammalian cells, genomic m6A was shown to originate from the misincorporation of ribo-N6-methyldeoxyadenine from degraded, modified RNA (Musheev et al. 2020). So far, none of the proposed m6A methyltransferase enzymes have been biochemically shown to act as a DNA methyltransferase. Therefore, the detection of genomic m6A on specific sets of genes may be a consequence of their active transcription, which can cause DNA damage and repair with low-level misincorporation of modified nucleotides (Sebastian and Oberdoerffer 2017). While the presence of 6 mA on DNA may have biological consequences, its causative regulatory effect should be carefully interpreted.
In AMF, genome expression and regulation need to be considered in the context of a multinucleate, syncytial state, and occasional heterokaryosis. Throughout the different life stages ranging from spore germination and hyphal growth, to symbiotically engaged mycelial networks, AMF nuclei experience different environments and stresses. Nuclei that are localized at the root interface would likely transcribe different gene sets than those at a foraging hyphal growth tip. However, since AMF nuclei travel freely and coexist in one large cytoplasm, it is puzzling to imagine how gene expression is effectively regulated in space and time. To compartmentalize functions, transcriptionally regulated responses of fungi to either their environment or their host likely need to be coupled with transport of messenger ribonucleoprotein complexes, localized translation, post-transcriptional and post-translational regulation. In filamentous fungi, the subcellular compartmentalization of functions depends on microtubule-dependent mRNP transport for the fast polar growth of hyphae (Becht et al. 2005, Becht et al. 2006; Feldbrugge et al. 2008; Konig et al. 2009), on vesicles and endosomes for RNA transport (Bethune et al. 2019; Haag et al. 2015), and on RNA-binding proteins for post-transcriptional regulation (Hall and Wallace 2022). In other words, the position of a nucleus within a mycelium might dictate the stability and fate of its transcriptional output (Schuurs et al. 1998). In AMF, single-molecule techniques will help identify the subcellular locations where RNAs are transcribed and proteins are synthesized, and will pinpoint the specialized cellular tasks that are required in different fungal structures.
In dikaryotic AMF strains, genome expression and regulation bear an additional layer of complexity since different alleles can be transcribed from different nuclear genomes. Allelic expression can therefore depend on the ratio of nuclei present in the mycelium, as well as differential transcriptional activity from either nucleus. The transcriptional output of both nuclear genomes would be subject to regulation by common components of cytoplasmic regulatory mechanisms (e.g. translation, RNA decay), which can theoretically buffer transcriptional imbalances. In R. irregularis dikaryons, the expression ratios of bi-allelic genes largely correspond to the ratios of nuclear genomes present (Robbins et al. 2021). Interestingly, the expression of a small number of bi-allelic genes deviated from nuclear genotype ratios, suggesting that transcriptional or post-transcriptional mechanisms can differentially impact the output of either nuclear genome.
So far, nuclear ratios of heterokaryotic strains have proven stable over time, indicating compatibility between coexisting genotypes, rather than genomic conflict. In addition, the dikaryotic state was associated with higher growth rates, which might provide a fitness advantage (Serghi et al. 2021). However, while the dikaryotic state and its associated genetic diversity intuitively translate to AMF having greater resilience and ecological success, dikaryotic strains are relatively rare in AMF culture collections (four out of 114 R. irregularis strains; (Kokkoris et al. 2021)), and the real prevalence of dikaryons versus monokaryons still needs to be assessed at the population level, in natural systems. Although cooperation might be the most apparent state of so far described dikaryons, it is possible that genomic conflict arising from incompatible genotypes leads to strong selective pressure, and will simply never be observed. Experiments that aim to recreate existing dikaryotic strains from homokaryotic parents, or to create new dikaryotic strains, will inform on the nuclear dynamics that occur at the onset of dikaryosis. Additionally, investigating the existence of dikaryosis in species other than R. irregularis will indicate whether this evolutionary strategy is widely used in AMF. Although nuclear ratios of AMF dikaryons are stable under standard laboratory conditions, they can be affected by host shifts (Kokkoris et al. 2020, 2021) and abiotic conditions such as pH, phosphorus concentration and temperature (Cornell et al. 2022). Mechanisms that regulate the relative number of coexisting nuclei are unknown, but could include asynchronous replication and mitosis, and nuclear degradation (Kokkoris et al. 2020). The ability to acquire and maintain more than one genotype may provide functional and adaptive benefits to AMF, but doing so may also create great opportunity for selection. It will be relevant to assess how stable the dikaryotic state is in planta and in the wild, as opposed to in vitro culture conditions, in order to understand the parameters acting on AMF adaptation and evolution. Since variations in nuclear ratios can be experimentally triggered, it will be interesting to test whether they are accompanied by corresponding variation in allelic gene expression, and whether these processes are reversible. Such a system could provide an elegant demonstration as to how dikaryotic AMF can make use of available genotypes to adapt to new environmental conditions (Fig. 4.2).
AMF genome compartmentalization and epigenetic regulation. Genes and transposable elements (TEs) are partitioned into euchromatin (A compartments, open) and heterochromatin (B compartments, closed), respectively. Contrary to core genes, accessory genes tend to be located in B compartments, where their expression can be affected by the spreading of heterochromatin, in cis or trans. Genes in B compartments may experience more sequence disturbance caused by TE activity and proximity, and therefore have higher rates of evolutionary change. Figure inspired by Liang and Fu (2021). mCG, methylated CG dinucleotides
4 Transmission of the Genome
In the fungal kingdom, the existence of diverse nucleotypes within one mycelium creates genotypic diversity and possibly phenotypic plasticity (Croll et al. 2009; Maheshwari 2005; Rayner 1991; Roper et al. 2011). The potential for genetic variation to be acquired within the lifetime of a single fungus was proposed to play a role in adaptation to the environment. The coexistence of multiple genotypes within one mycelium can cause competitive and cooperative genome dynamics, and differential segregation and selection of genotypes may also be adaptive (James et al. 2008; Jinks 1952; Roper et al. 2011; Samils et al. 2014). A possible outcome of nuclei coexisting as a heterokaryon is that they can fuse, undergo karyogamy, mitotic recombination, and ploidy reduction, resulting in haploid and aneuploid genomes that are different from the parental ones (Anderson et al. 2019; Forche et al. 2008; Strom and Bushley 2016). This process called parasexuality is independent of sexual reproduction, but retains hallmarks of meiosis and typically yields transiently aneuploid cells which then undergo ploidy reduction, creating de novo genetic diversity (Hickman et al. 2015; Hirakawa et al. 2017; Mishra et al. 2021). In most cases, the heterokaryon stage is expected to precede nuclear fusion, meiosis, and completion of a sexual life cycle. AMF can experience a dikaryotic life stage, express meiosis-related genes, and can recombine genetic material (Chen et al. 2018a; Corradi and Brachmann 2017; Dallaire et al. 2021; Halary et al. 2011; Hofstatter and Lahr 2019; Mateus et al. 2020; Ropars et al. 2016; Yildirir et al. 2020). However, since direct evidence of nuclear fusion, karyogamy, meiosis, or a sexual structure is lacking, the current assumption is that AMF are at least partially clonal and may undergo either parasexual or sexual reproduction at a low frequency and under unknown developmental or environmental conditions. Experimental crosses of homokaryotic AMF may reveal signs of genomic cooperation or conflict between nuclear and cytoplasmic (e.g. mitochondria) elements, and if recombination occurs, whether it is accompanied by transient aneuploidy or not (suggesting parasexual or sexual mechanisms, respectively). Limited recombination over long evolutionary timescales is problematic because each generation may generate too little genetic variation to adapt to environmental change, possibly leading to extinction. It is therefore reasonable to expect that low-frequency recombination occurs in AMF, and that alternative mechanisms may also generate the plasticity and genomic heterogeneity required for these fungi to adapt to incredibly varied environments and hosts (Fig. 4.3).
Potential mechanisms contributing to genome evolution in AMF. (a) Mobile transposable elements (TEs) can induce duplications, deletions, and chromosome rearrangements. (b) Mutagenic mechanisms (shown in red) include errors in DNA replication, errors in DNA repair, but as well as (not shown here) base deamination, oxidative DNA damage, and base methylation. (c) Sexual or parasexual recombination can generate genetic diversity, and the latter may be accompanied by asymmetric segregation of chromosomes and transient aneuploidy. Different coloured nuclei illustrate dikaryon formation, which may be associated with TE activity and recombination
5 Perspectives on Adaptation and Evolution
Knowledge of the mechanisms, rates, and consequences of AMF evolution is crucial to understand which functional traits are under selection, and for predicting the capacity of AMF to support ecosystems in the face of rapid environmental change. While AMF display remarkable persistence over long time periods, the mechanisms underlying short-term evolutionary dynamics are still elusive. Are extant AMF in an evolutionary stasis, or diversifying? Is coevolution with plants constraining or promoting evolutionary divergence of AMF, and how can we use this information to predict and support ecosystem functioning? At the molecular level, evolution manifests itself in a variety of changes in DNA sequence, ranging from point mutations to chromosomal rearrangements. Understanding how different mechanisms contribute to genome evolution and quantifying genomic diversity and evolutionary rates will help explain AMF genome organization in adaptive terms.
AMF genes and TEs tend to be partitioned in different regions of the genome. At present, epigenetic landscapes globally correlate with transcriptional activity within these compartments and may play direct roles in regulating core and accessory gene expression (Kumar et al. 2021). While accessory genes are enriched in heterochromatic compartments and display presence/absence variation, it is still unclear whether they have a higher molecular rate of evolution compared to core genes, which would imply ongoing selection pressure. In parallel, since TE expression was detected in R. irregularis spores, suggesting ongoing or recent TE mobility (Dallaire et al. 2021; Maeda et al. 2018), quantifying new TE insertions in R. irregularis strains would indicate which genomic regions were most recently shaped by TEs and are therefore potentially still dynamically evolving. This is important because in the absence of recombination, innovation could still be driven by TE activity or local mutagenesis, rather than sex-dependent recombination. In other fungi, for example, TEs tend to accumulate in accessory chromosomes or accessory compartments in core chromosomes (Croll and McDonald 2012; Sanchez-Vallet et al. 2018), possibly identifying candidate genes or genetic regions with adaptive potential and evolutionary plasticity, and pointing to regions where TEs are selected against. Another mutagenic mechanism to be considered is deamination of methylated cytosines. In analyses of human somatic mutations, methylated cytosines spontaneously deaminate at a higher rate than non-methylated cytosines, and, when not correctly repaired, result in mutations (Alexandrov et al. 2015; Kong et al. 2012). Excess mutagenicity can therefore be observed at methylated CG dinucleotides. In Ascomycota and certain Basidiomycota fungi, methylation of transposons during meiosis is associated with extremely high rates of C-T transitions and rapid mutagenesis (Gladyshev 2017; Hood et al. 2005; Horns et al. 2012). This process called repeat-induced point (RIP) mutation has however not been detected in the AMF G. margarita or in species of the Mucoromycotina (Venice et al. 2020). Nevertheless, the endogenous rate of spontaneous deamination can account for some mutagenesis in AMF, and likely influences point mutation rates.
Quantifying the rate of de novo mutation occurring during AMF vegetative growth would help estimate how much genetic diversity can be expected to arise in the absence of sex, in comparison to rates of molecular evolution in fungi and other eukaryotes (Bezmenova et al. 2020; Hiltunen et al. 2019; Kasuga et al. 2002; Obbard et al. 2012; Wolfe et al. 1987). The combination of mutation rate analysis with intraspecific divergence would help calibrate evolutionary models of species divergence in AMF and provide valuable tools for the reconstruction of their natural history. Since the rate of recombination of a species is critical for estimates of mutation and genomic change, further insights into nuclear dynamics will also be important. Investigating which of the dikaryotic or monokaryotic states is prevalent in nature and across AMF species, the extent to which inter-nuclear recombination occurs, and whether AMF have an aneuploid stage may require large numbers of isolates and species to be analysed, but will provide invaluable insights into the frequency of recombination among lineages, and whether clonal lineages arise by recombination.
Until the advent of a stable transformation protocol for AMF, increasing genomic sampling, both in numbers and in geographical diversity, is arguably the best way to accelerate our understanding of how AMF genomes work, how genetic variation shapes their phenotypes, how they evolve, and how systems-level patterns in ecological diversity arise. Scaling up AMF biodiversity genomics will improve taxonomic delineation of AMF, which will allow us to tackle major questions about evolutionary selection pressures that shape AMF biodiversity, adaptation, and evolution. Reports on host plants affecting the distribution of AMF communities are accumulating (Alguacil et al. 2019; Croll et al. 2008; Koch et al. 2006; Munkvold et al. 2004; Sanders 2003; Van Der Heijden and Scheublin 2007). Factors selecting AMF and driving their diversification may therefore include a combination of components such as host plant identity, environmental conditions and resources, interaction with microbes (including other AMF), and us (host breeding, agricultural systems). Identifying which functional traits of AMF can be selected and what pressures promote or diminish their diversity in natural communities hold the potential we need for protecting these fungal networks that sustain life on planet Earth.
References
Alexandrov LB, Jones PH, Wedge DC, Sale JE, Campbell PJ, Nik-Zainal S, Stratton MR (2015) Clock-like mutational processes in human somatic cells. Nat Genet 47(12):1402–1407. https://doi.org/10.1038/ng.3441
Alguacil MM, Diaz G, Torres P, Rodriguez-Caballero G, Roldan A (2019) Host identity and functional traits determine the community composition of the arbuscular mycorrhizal fungi in facultative epiphytic plant species. Fungal Ecol 39:307–315. https://doi.org/10.1016/j.funeco.2019.02.002
Anderson MZ, Thomson GJ, Hirakawa MP, Bennett RJ (2019) A ‘parameiosis’ drives depolyploidization and homologous recombination in Candida albicans. Nat Commun 10(1):4388. https://doi.org/10.1038/s41467-019-12376-2
Angelard C, Sanders IR (2011) Effect of segregation and genetic exchange on arbuscular mycorrhizal fungi in colonization of roots. New Phytol 189(3):652–657. https://doi.org/10.1111/j.1469-8137.2010.03602.x
Angelard C, Tanner CJ, Fontanillas P, Niculita-Hirzel H, Masclaux F, Sanders IR (2014) Rapid genotypic change and plasticity in arbuscular mycorrhizal fungi is caused by a host shift and enhanced by segregation. ISME J 8(2):284–294. https://doi.org/10.1038/ismej.2013.154
Bago B, Zipfel W, Williams RM, Piche Y (1999) Nuclei of symbiotic arbuscular mycorrhizal fungi as revealed by in vivo two-photon microscopy. Protoplasma 209(1–2):77–89. https://doi.org/10.1007/BF01415703
Becht P, Vollmeister E, Feldbrugge M (2005) Role for RNA-binding proteins implicated in pathogenic development of Ustilago maydis. Eukaryot Cell 4(1):121–133. https://doi.org/10.1128/EC.4.1.121-133.2005
Becht P, Konig J, Feldbrugge M (2006) The RNA-binding protein Rrm4 is essential for polarity in Ustilago maydis and shuttles along microtubules. J Cell Sci 119(Pt 23):4964–4973. https://doi.org/10.1242/jcs.03287
Bergstrom A, Simpson JT, Salinas F, Barre B, Parts L, Zia A, Nguyen Ba AN, Moses AM, Louis EJ, Mustonen V, Warringer J, Durbin R, Liti G (2014) A high-definition view of functional genetic variation from natural yeast genomes. Mol Biol Evol 31(4):872–888. https://doi.org/10.1093/molbev/msu037
Bethune J, Jansen RP, Feldbrugge M, Zarnack K (2019) Membrane-associated RNA-binding proteins orchestrate organelle-coupled translation. Trends Cell Biol 29(2):178–188. https://doi.org/10.1016/j.tcb.2018.10.005
Bezmenova AV, Zvyagina EA, Fedotova AV, Kasianov AS, Neretina TV, Penin AA, Bazykin GA, Kondrashov AS (2020) Rapid accumulation of mutations in growing mycelia of a hypervariable fungus Schizophyllum commune. Mol Biol Evol 37(8):2279–2286. https://doi.org/10.1093/molbev/msaa083
Boon E, Zimmerman E, Lang BF, Hijri M (2010) Intra-isolate genome variation in arbuscular mycorrhizal fungi persists in the transcriptome. J Evol Biol 23(7):1519–1527. https://doi.org/10.1111/j.1420-9101.2010.02019.x
Bravo A, Brands M, Wewer V, Dormann P, Harrison MJ (2017) Arbuscular mycorrhiza-specific enzymes FatM and RAM2 fine-tune lipid biosynthesis to promote development of arbuscular mycorrhiza. New Phytol 214(4):1631–1645. https://doi.org/10.1111/nph.14533
Cardenas-Flores A, Cranenbrouck S, Draye X, Guillet A, Govaerts B, Declerck S (2011) The sterol biosynthesis inhibitor molecule fenhexamid impacts the vegetative compatibility of Glomus clarum. Mycorrhiza 21(5):443–449. https://doi.org/10.1007/s00572-011-0385-z
Chaturvedi A, Cruz Corella J, Robbins C, Loha A, Menin L, Gasilova N, Masclaux FG, Lee SJ, Sanders IR (2021) The methylome of the model arbuscular mycorrhizal fungus, Rhizophagus irregularis, shares characteristics with early diverging fungi and Dikarya. Commun Biol 4(1):901. https://doi.org/10.1038/s42003-021-02414-5
Chen E, Mathieu S, Hoffrichter A, Sedzielewska-Toro K, Peart M, Pelin A, Ndikumana S, Ropars J, Dreissig S, Fuchs J, Brachmann A, Corradi N (2018a) Single nucleus sequencing reveals evidence of inter-nucleus recombination in arbuscular mycorrhizal fungi. elife 7. https://doi.org/10.7554/eLife.39813
Chen E, Morin E, Beaudet D, Noel J, Yildirir G, Ndikumana S, Charron P, St-Onge C, Giorgi J, Kruger M, Marton T, Ropars J, Grigoriev IV, Hainaut M, Henrissat B, Roux C, Martin F, Corradi N (2018b) High intraspecific genome diversity in the model arbuscular mycorrhizal symbiont Rhizophagus irregularis. New Phytol 220(4):1161–1171. https://doi.org/10.1111/nph.14989
Cohen AL, Jia S (2014) Noncoding RNAs and the borders of heterochromatin. Wiley Interdiscip Rev RNA 5(6):835–847. https://doi.org/10.1002/wrna.1249
Cornell C, Kokkoris V, Turcu B, Dettman J, Stefani F, Corradi N (2022) The arbuscular mycorrhizal fungus Rhizophagus irregularis harmonizes nuclear dynamics in the presence of distinct abiotic factors. Fungal Genet Biol 158:103639. https://doi.org/10.1016/j.fgb.2021.103639
Corradi N, Brachmann A (2017) Fungal mating in the most widespread plant symbionts? Trends Plant Sci 22(2):175–183. https://doi.org/10.1016/j.tplants.2016.10.010
Corradi N, Sanders IR (2006) Evolution of the P-type II ATPase gene family in the fungi and presence of structural genomic changes among isolates of Glomus intraradices. BMC Evol Biol 6:21. https://doi.org/10.1186/1471-2148-6-21
Corradi N, Croll D, Colard A, Kuhn G, Ehinger M, Sanders IR (2007) Gene copy number polymorphisms in an arbuscular mycorrhizal fungal population. Appl Environ Microbiol 73(1):366–369. https://doi.org/10.1128/AEM.01574-06
Croll D, McDonald BA (2012) The accessory genome as a cradle for adaptive evolution in pathogens. PLoS Pathog 8(4):e1002608. https://doi.org/10.1371/journal.ppat.1002608
Croll D, Wille L, Gamper HA, Mathimaran N, Lammers PJ, Corradi N, Sanders IR (2008) Genetic diversity and host plant preferences revealed by simple sequence repeat and mitochondrial markers in a population of the arbuscular mycorrhizal fungus Glomus intraradices. New Phytol 178(3):672–687. https://doi.org/10.1111/j.1469-8137.2008.02381.x
Croll D, Giovannetti M, Koch AM, Sbrana C, Ehinger M, Lammers PJ, Sanders IR (2009) Nonself vegetative fusion and genetic exchange in the arbuscular mycorrhizal fungus Glomus intraradices. New Phytol 181(4):924–937. https://doi.org/10.1111/j.1469-8137.2008.02726.x
Dallaire A, Manley BF, Wilkens M, Bista I, Quan C, Evangelisti E, Bradshaw CR, Ramakrishna NB, Schornack S, Butter F, Paszkowski U, Miska EA (2021) Transcriptional activity and epigenetic regulation of transposable elements in the symbiotic fungus Rhizophagus irregularis. Genome Res 31:2290. https://doi.org/10.1101/gr.275752.121
Davison J, Moora M, Opik M, Adholeya A, Ainsaar L, Ba A, Burla S, Diedhiou AG, Hiiesalu I, Jairus T, Johnson NC, Kane A, Koorem K, Kochar M, Ndiaye C, Partel M, Reier U, Saks U, Singh R et al (2015) Global assessment of arbuscular mycorrhizal fungus diversity reveals very low endemism. Science 349(6251):970–973. https://doi.org/10.1126/science.aab1161
Dong S, Raffaele S, Kamoun S (2015) The two-speed genomes of filamentous pathogens: waltz with plants. Curr Opin Genet Dev 35:57–65. https://doi.org/10.1016/j.gde.2015.09.001
Dunn B, Richter C, Kvitek DJ, Pugh T, Sherlock G (2012) Analysis of the Saccharomyces cerevisiae pan-genome reveals a pool of copy number variants distributed in diverse yeast strains from differing industrial environments. Genome Res 22(5):908–924. https://doi.org/10.1101/gr.130310.111
Ehinger MO, Croll D, Koch AM, Sanders IR (2012) Significant genetic and phenotypic changes arising from clonal growth of a single spore of an arbuscular mycorrhizal fungus over multiple generations. New Phytol 196(3):853–861. https://doi.org/10.1111/j.1469-8137.2012.04278.x
Faino L, Seidl MF, Shi-Kunne X, Pauper M, van den Berg GC, Wittenberg AH, Thomma BP (2016) Transposons passively and actively contribute to evolution of the two-speed genome of a fungal pathogen. Genome Res 26(8):1091–1100. https://doi.org/10.1101/gr.204974.116
Feldbrugge M, Zarnack K, Vollmeister E, Baumann S, Koepke J, Konig J, Munsterkotter M, Mannhaupt G (2008) The posttranscriptional machinery of Ustilago maydis. Fungal Genet Biol 45(Suppl 1):S40–S46. https://doi.org/10.1016/j.fgb.2008.03.013
Forche A, Alby K, Schaefer D, Johnson AD, Berman J, Bennett RJ (2008) The parasexual cycle in Candida albicans provides an alternative pathway to meiosis for the formation of recombinant strains. PLoS Biol 6(5):e110. https://doi.org/10.1371/journal.pbio.0060110
Freitag M, Hickey PC, Khlafallah TK, Read ND, Selker EU (2004) HP1 is essential for DNA methylation in neurospora. Mol Cell 13(3):427–434. https://doi.org/10.1016/s1097-2765(04)00024-3
Friedman S, Freitag M (2017) Centrochromatin of fungi. Prog Mol Subcell Biol 56:85–109. https://doi.org/10.1007/978-3-319-58592-5_4
Gladyshev E (2017) Repeat-induced point mutation and other genome defense mechanisms in fungi. Microbiol Spectr 5(4). https://doi.org/10.1128/microbiolspec.FUNK-0042-2017
Haag C, Steuten B, Feldbrugge M (2015) Membrane-coupled mRNA trafficking in fungi. Annu Rev Microbiol 69:265–281. https://doi.org/10.1146/annurev-micro-091014-104242
Halary S, Malik SB, Lildhar L, Slamovits CH, Hijri M, Corradi N (2011) Conserved meiotic machinery in Glomus spp., a putatively ancient asexual fungal lineage. Genome Biol Evol 3:950–958. https://doi.org/10.1093/gbe/evr089
Hall RA, Wallace EWJ (2022) Post-transcriptional control of fungal cell wall synthesis. Cell Surf 8:100074. https://doi.org/10.1016/j.tcsw.2022.100074
Hickman MA, Paulson C, Dudley A, Berman J (2015) Parasexual ploidy reduction drives population heterogeneity through random and transient aneuploidy in Candida albicans. Genetics 200(3):781–794. https://doi.org/10.1534/genetics.115.178020
Hijri M, Sanders IR (2005) Low gene copy number shows that arbuscular mycorrhizal fungi inherit genetically different nuclei. Nature 433(7022):160–163. https://doi.org/10.1038/nature03069
Hiltunen M, Grudzinska-Sterno M, Wallerman O, Ryberg M, Johannesson H (2019) Maintenance of high genome integrity over vegetative growth in the fairy-ring mushroom Marasmius oreades. Curr Biol 29(16):2758–2765 e2756. https://doi.org/10.1016/j.cub.2019.07.025
Hirakawa MP, Chyou DE, Huang D, Slan AR, Bennett RJ (2017) Parasex generates phenotypic diversity de novo and impacts drug resistance and virulence in Candida albicans. Genetics 207(3):1195–1211. https://doi.org/10.1534/genetics.117.300295
Hofstatter PG, Lahr DJG (2019) All eukaryotes are sexual, unless proven otherwise: many so-called asexuals present meiotic machinery and might be able to have sex. BioEssays 41(6):e1800246. https://doi.org/10.1002/bies.201800246
Hood ME, Katawczik M, Giraud T (2005) Repeat-induced point mutation and the population structure of transposable elements in Microbotryum violaceum. Genetics 170(3):1081–1089. https://doi.org/10.1534/genetics.105.042564
Horns F, Petit E, Yockteng R, Hood ME (2012) Patterns of repeat-induced point mutation in transposable elements of basidiomycete fungi. Genome Biol Evol 4(3):240–247. https://doi.org/10.1093/gbe/evs005
James TY, Stenlid J, Olson A, Johannesson H (2008) Evolutionary significance of imbalanced nuclear ratios within heterokaryons of the basidiomycete fungus Heterobasidion parviporum. Evolution 62(9):2279–2296. https://doi.org/10.1111/j.1558-5646.2008.00462.x
Jany JL, Pawlowska TE (2010) Multinucleate spores contribute to evolutionary longevity of asexual glomeromycota. Am Nat 175(4):424–435. https://doi.org/10.1086/650725
Jiang Y, Wang W, Xie Q, Liu N, Liu L, Wang D, Zhang X, Yang C, Chen X, Tang D, Wang E (2017) Plants transfer lipids to sustain colonization by mutualistic mycorrhizal and parasitic fungi. Science 356(6343):1172–1175. https://doi.org/10.1126/science.aam9970
Jinks JL (1952) Heterokaryosis; a system of adaption in wild fungi. Proc R Soc Lond B Biol Sci 140(898):83–99. https://doi.org/10.1098/rspb.1952.0046
Kamel L, Tang N, Malbreil M, San Clemente H, Le Marquer M, Roux C, Frey FD, N. (2017) The comparison of expressed candidate secreted proteins from two arbuscular mycorrhizal fungi unravels common and specific molecular tools to invade different host plants. Front Plant Sci 8:124. https://doi.org/10.3389/fpls.2017.00124
Kameoka H, Tsutsui I, Saito K, Kikuchi Y, Handa Y, Ezawa T, Hayashi H, Kawaguchi M, Akiyama K (2019) Stimulation of asymbiotic sporulation in arbuscular mycorrhizal fungi by fatty acids. Nat Microbiol 4(10):1654–1660. https://doi.org/10.1038/s41564-019-0485-7
Kasuga T, White TJ, Taylor JW (2002) Estimation of nucleotide substitution rates in Eurotiomycete fungi. Mol Biol Evol 19(12):2318–2324. https://doi.org/10.1093/oxfordjournals.molbev.a004056
Keymer A, Pimprikar P, Wewer V, Huber C, Brands M, Bucerius SL, Delaux PM, Klingl V, Ropenack-Lahaye EV, Wang TL, Eisenreich W, Dormann P, Parniske M, Gutjahr C (2017) Lipid transfer from plants to arbuscular mycorrhiza fungi. elife 6. https://doi.org/10.7554/eLife.29107
Klocko AD, Ormsby T, Galazka JM, Leggett NA, Uesaka M, Honda S, Freitag M, Selker EU (2016) Normal chromosome conformation depends on subtelomeric facultative heterochromatin in Neurospora crassa. Proc Natl Acad Sci U S A 113(52):15048–15053. https://doi.org/10.1073/pnas.1615546113
Kobayashi Y, Maeda T, Yamaguchi K, Kameoka H, Tanaka S, Ezawa T, Shigenobu S, Kawaguchi M (2018) The genome of Rhizophagus clarus HR1 reveals a common genetic basis for auxotrophy among arbuscular mycorrhizal fungi. BMC Genomics 19(1):465. https://doi.org/10.1186/s12864-018-4853-0
Koch AM, Kuhn G, Fontanillas P, Fumagalli L, Goudet J, Sanders IR (2004) High genetic variability and low local diversity in a population of arbuscular mycorrhizal fungi. Proc Natl Acad Sci U S A 101(8):2369–2374. https://doi.org/10.1073/pnas.0306441101
Koch AM, Croll D, Sanders IR (2006) Genetic variability in a population of arbuscular mycorrhizal fungi causes variation in plant growth. Ecol Lett 9(2):103–110. https://doi.org/10.1111/j.1461-0248.2005.00853.x
Kokkoris V, Stefani F, Dalpe Y, Dettman J, Corradi N (2020) Nuclear dynamics in the arbuscular mycorrhizal fungi. Trends Plant Sci 25(8):765–778. https://doi.org/10.1016/j.tplants.2020.05.002
Kokkoris V, Chagnon PL, Yildirir G, Clarke K, Goh D, MacLean AM, Dettman J, Stefani F, Corradi N (2021) Host identity influences nuclear dynamics in arbuscular mycorrhizal fungi. Curr Biol 31(7):1531–1538 e1536. https://doi.org/10.1016/j.cub.2021.01.035
Kong A, Frigge ML, Masson G, Besenbacher S, Sulem P, Magnusson G, Gudjonsson SA, Sigurdsson A, Jonasdottir A, Jonasdottir A, Wong WS, Sigurdsson G, Walters GB, Steinberg S, Helgason H, Thorleifsson G, Gudbjartsson DF, Helgason A, Magnusson OT et al (2012) Rate of de novo mutations and the importance of father’s age to disease risk. Nature 488(7412):471–475. https://doi.org/10.1038/nature11396
Konig J, Baumann S, Koepke J, Pohlmann T, Zarnack K, Feldbrugge M (2009) The fungal RNA-binding protein Rrm4 mediates long-distance transport of ubi1 and rho3 mRNAs. EMBO J 28(13):1855–1866. https://doi.org/10.1038/emboj.2009.145
Kruger M, Kruger C, Walker C, Stockinger H, Schussler A (2012) Phylogenetic reference data for systematics and phylotaxonomy of arbuscular mycorrhizal fungi from phylum to species level. New Phytol 193(4):970–984. https://doi.org/10.1111/j.1469-8137.2011.03962.x
Kumar S, Kaur S, Seem K, Kumar S, Mohapatra T (2021) Understanding 3D genome organization and its effect on transcriptional gene regulation under environmental stress in plant: a chromatin perspective. Front Cell Dev Biol 9:774719. https://doi.org/10.3389/fcell.2021.774719
Liang Z, Fu XD (2021) 3D genome encoded by LINE and SINE repeats. Cell Res 31(6):603–604. https://doi.org/10.1038/s41422-021-00485-x
Lim CS, Weinstein BN, Roy SW, Brown CM (2021) Analysis of fungal genomes reveals commonalities of intron gain or loss and functions in intron-poor species. Mol Biol Evol 38(10):4166–4186. https://doi.org/10.1093/molbev/msab094
Lin K, Limpens E, Zhang Z, Ivanov S, Saunders DG, Mu D, Pang E, Cao H, Cha H, Lin T, Zhou Q, Shang Y, Li Y, Sharma T, van Velzen R, de Ruijter N, Aanen DK, Win J, Kamoun S et al (2014) Single nucleus genome sequencing reveals high similarity among nuclei of an endomycorrhizal fungus. PLoS Genet 10(1):e1004078. https://doi.org/10.1371/journal.pgen.1004078
Luginbuehl LH, Menard GN, Kurup S, Van Erp H, Radhakrishnan GV, Breakspear A, Oldroyd GED, Eastmond PJ (2017) Fatty acids in arbuscular mycorrhizal fungi are synthesized by the host plant. Science 356(6343):1175–1178. https://doi.org/10.1126/science.aan0081
Ma LJ, van der Does HC, Borkovich KA, Coleman JJ, Daboussi MJ, Di Pietro A, Dufresne M, Freitag M, Grabherr M, Henrissat B, Houterman PM, Kang S, Shim WB, Woloshuk C, Xie X, Xu JR, Antoniw J, Baker SE, Bluhm BH et al (2010) Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464(7287):367–373. https://doi.org/10.1038/nature08850
Maeda T, Kobayashi Y, Kameoka H, Okuma N, Takeda N, Yamaguchi K, Bino T, Shigenobu S, Kawaguchi M (2018) Evidence of non-tandemly repeated rDNAs and their intragenomic heterogeneity in Rhizophagus irregularis. Commun Biol 1:87. https://doi.org/10.1038/s42003-018-0094-7
Maheshwari R (2005) Nuclear behavior in fungal hyphae. FEMS Microbiol Lett 249(1):7–14. https://doi.org/10.1016/j.femsle.2005.06.031
Malar CM, Kruger M, Kruger C, Wang Y, Stajich JE, Keller J, Chen ECH, Yildirir G, Villeneuve-Laroche M, Roux C, Delaux PM, Corradi N (2021) The genome of Geosiphon pyriformis reveals ancestral traits linked to the emergence of the arbuscular mycorrhizal symbiosis. Curr Biol 31(7):1570–1577 e1574. https://doi.org/10.1016/j.cub.2021.01.058
Marleau J, Dalpe Y, St-Arnaud M, Hijri M (2011) Spore development and nuclear inheritance in arbuscular mycorrhizal fungi. BMC Evol Biol 11:51. https://doi.org/10.1186/1471-2148-11-51
Mateus ID, Rojas EC, Savary R, Dupuis C, Masclaux FG, Aletti C, Sanders IR (2020) Coexistence of genetically different Rhizophagus irregularis isolates induces genes involved in a putative fungal mating response. ISME J 14(10):2381–2394. https://doi.org/10.1038/s41396-020-0694-3
Mathieu S, Cusant L, Roux C, Corradi N (2018) Arbuscular mycorrhizal fungi: intraspecific diversity and pangenomes. New Phytol 220(4):1129–1134. https://doi.org/10.1111/nph.15275
McCarthy CGP, Fitzpatrick DA (2019) Pan-genome analyses of model fungal species. Microb Genom 5(2). https://doi.org/10.1099/mgen.0.000243
McClintock B (1951) Chromosome organization and genic expression. Cold Spring Harb Symp Quant Biol 16:13–47. https://doi.org/10.1101/sqb.1951.016.01.004
Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R (2005) The microbial pan-genome. Curr Opin Genet Dev 15(6):589–594. https://doi.org/10.1016/j.gde.2005.09.006
Mishra A, Forche A, Anderson MZ (2021) Parasexuality of Candida species. Front Cell Infect Microbiol 11:796929. https://doi.org/10.3389/fcimb.2021.796929
Moller M, Habig M, Freitag M, Stukenbrock EH (2018) Extraordinary genome instability and widespread chromosome rearrangements during vegetative growth. Genetics 210(2):517–529. https://doi.org/10.1534/genetics.118.301050
Mondo SJ, Dannebaum RO, Kuo RC, Louie KB, Bewick AJ, LaButti K, Haridas S, Kuo A, Salamov A, Ahrendt SR, Lau R, Bowen BP, Lipzen A, Sullivan W, Andreopoulos BB, Clum A, Lindquist E, Daum C, Northen TR et al (2017) Widespread adenine N6-methylation of active genes in fungi. Nat Genet 49(6):964–968. https://doi.org/10.1038/ng.3859
Morin E, Miyauchi S, San Clemente H, Chen ECH, Pelin A, de la Providencia I, Ndikumana S, Beaudet D, Hainaut M, Drula E, Kuo A, Tang N, Roy S, Viala J, Henrissat B, Grigoriev IV, Corradi N, Roux C, Martin FM (2019) Comparative genomics of Rhizophagus irregularis, R. cerebriforme, R. diaphanus and Gigaspora rosea highlights specific genetic features in Glomeromycotina. New Phytol 222(3):1584–1598. https://doi.org/10.1111/nph.15687
Morris JL, Puttick MN, Clark JW, Edwards D, Kenrick P, Pressel S, Wellman CH, Yang Z, Schneider H, Donoghue PCJ (2018) The timescale of early land plant evolution. Proc Natl Acad Sci U S A 115(10):E2274–E2283. https://doi.org/10.1073/pnas.1719588115
Munkvold L, Kjoller R, Vestberg M, Rosendahl S, Jakobsen I (2004) High functional diversity within species of arbuscular mycorrhizal fungi. New Phytol 164(2):357–364. https://doi.org/10.1111/j.1469-8137.2004.01169.x
Musheev MU, Baumgartner A, Krebs L, Niehrs C (2020) The origin of genomic N(6)-methyl-deoxyadenosine in mammalian cells. Nat Chem Biol 16(6):630–634. https://doi.org/10.1038/s41589-020-0504-2
Nadal M, Sawers R, Naseem S, Bassin B, Kulicke C, Sharman A, An G, An K, Ahern KR, Romag A, Brutnell TP, Gutjahr C, Geldner N, Roux C, Martinoia E, Konopka JB, Paszkowski U (2017) An N-acetylglucosamine transporter required for arbuscular mycorrhizal symbioses in rice and maize. Nat Plants 3:17073. https://doi.org/10.1038/nplants.2017.73
Obbard DJ, Maclennan J, Kim KW, Rambaut A, O'Grady PM, Jiggins FM (2012) Estimating divergence dates and substitution rates in the Drosophila phylogeny. Mol Biol Evol 29(11):3459–3473. https://doi.org/10.1093/molbev/mss150
Öpik M, Davison J (2016) Uniting species- and community-oriented approaches to understand arbuscular mycorrhizal fungal diversity. Fungal Ecol 24:106–113. https://doi.org/10.1016/j.funeco.2016.07.005
Perez-Lamarque B, Öpik M, Maliet O, Afonso Silva AC, Selosse M-A, Martos F, Morlon H (2022) Global drivers of obligate mycorrhizal symbionts diversification. bioRXiv. https://doi.org/10.1101/2020.07.28.224790
Peter J, De Chiara M, Friedrich A, Yue JX, Pflieger D, Bergstrom A, Sigwalt A, Barre B, Freel K, Llored A, Cruaud C, Labadie K, Aury JM, Istace B, Lebrigand K, Barbry P, Engelen S, Lemainque A, Wincker P et al (2018) Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556(7701):339–344. https://doi.org/10.1038/s41586-018-0030-5
Plissonneau C, Hartmann FE, Croll D (2018) Pangenome analyses of the wheat pathogen Zymoseptoria tritici reveal the structural basis of a highly plastic eukaryotic genome. BMC Biol 16(1):5. https://doi.org/10.1186/s12915-017-0457-4
Prasad Singh P, Srivastava D, Jaiswar A, Adholeya A (2019) Effector proteins of Rhizophagus proliferus: conserved protein domains may play a role in host-specific interaction with different plant species. Braz J Microbiol 50(3):593–601. https://doi.org/10.1007/s42770-019-00099-x
Purin S, Morton JB (2013) Anastomosis behavior differs between asymbiotic and symbiotic hyphae of Rhizophagus clarus. Mycologia 105(3):589–602. https://doi.org/10.3852/12-135
Rayner A (1991) The challenge of the individualistic mycelium. Mycologia 83:48–71. https://doi.org/10.1080/00275514.1991.12025978
Reinhardt D, Roux C, Corradi N, Di Pietro A (2021) Lineage-specific genes and cryptic sex: parallels and differences between arbuscular mycorrhizal fungi and fungal pathogens. Trends Plant Sci 26(2):111–123. https://doi.org/10.1016/j.tplants.2020.09.006
Rich MK, Vigneron N, Libourel C, Keller J, Xue L, Hajheidari M, Radhakrishnan GV, Le Ru A, Diop SI, Potente G, Conti E, Duijsings D, Batut A, Le Faouder P, Kodama K, Kyozuka J, Sallet E, Becard G, Rodriguez-Franco M et al (2021) Lipid exchanges drove the evolution of mutualism during plant terrestrialization. Science 372(6544):864–868. https://doi.org/10.1126/science.abg0929
Robbins C, Cruz Corella J, Aletti C, Seiler R, Mateus ID, Lee SJ, Masclaux FG, Sanders IR (2021) Generation of unequal nuclear genotype proportions in Rhizophagus irregularis progeny causes allelic imbalance in gene transcription. New Phytol 231(5):1984–2001. https://doi.org/10.1111/nph.17530
Ropars J, Toro KS, Noel J, Pelin A, Charron P, Farinelli L, Marton T, Kruger M, Fuchs J, Brachmann A, Corradi N (2016) Evidence for the sexual origin of heterokaryosis in arbuscular mycorrhizal fungi. Nat Microbiol 1(6):16033. https://doi.org/10.1038/nmicrobiol.2016.33
Roper M, Ellison C, Taylor JW, Glass NL (2011) Nuclear and genome dynamics in multinucleate ascomycete fungi. Curr Biol 21(18):R786–R793. https://doi.org/10.1016/j.cub.2011.06.042
Saksouk N, Simboeck E, Dejardin J (2015) Constitutive heterochromatin formation and transcription in mammals. Epigenetics Chromatin 8:3. https://doi.org/10.1186/1756-8935-8-3
Samils N, Oliva J, Johannesson H (2014) Nuclear interactions in a heterokaryon: insight from the model Neurospora tetrasperma. Proc Biol Sci 281(1786). https://doi.org/10.1098/rspb.2014.0084
Sanchez-Vallet A, Fouche S, Fudal I, Hartmann FE, Soyer JL, Tellier A, Croll D (2018) The genome biology of effector gene evolution in filamentous plant pathogens. Annu Rev Phytopathol 56:21–40. https://doi.org/10.1146/annurev-phyto-080516-035303
Sanders IR (2003) Preference, specificity and cheating in the arbuscular mycorrhizal symbiosis. Trends Plant Sci 8(4):143–145. https://doi.org/10.1016/S1360-1385(03)00012-8
Savary R, Masclaux FG, Wyss T, Droh G, Cruz Corella J, Machado AP, Morton JB, Sanders IR (2018) A population genomics approach shows widespread geographical distribution of cryptic genomic forms of the symbiotic fungus Rhizophagus irregularis. ISME J 12(1):17–30. https://doi.org/10.1038/ismej.2017.153
Sbrana C, Strani P, Pepe A, de Novais CB, Giovannetti M (2018) Divergence of Funneliformis mosseae populations over 20 years of laboratory cultivation, as revealed by vegetative incompatibility and molecular analysis. Mycorrhiza 28(4):329–341. https://doi.org/10.1007/s00572-018-0830-3
Schuurs TA, Dalstra HJ, Scheer JM, Wessels JG (1998) Positioning of nuclei in the secondary mycelium of Schizophyllum commune in relation to differential gene expression. Fungal Genet Biol 23(2):150–161. https://doi.org/10.1006/fgbi.1997.1028
Sebastian R, Oberdoerffer P (2017) Transcription-associated events affecting genomic integrity. Philos Trans R Soc Lond Ser B Biol Sci 372(1731):20160288. https://doi.org/10.1098/rstb.2016.0288
Selosse MA, Le Tacon F (1998) The land flora: a phototroph-fungus partnership? Trends Ecol Evol 13(1):15–20. https://doi.org/10.1016/s0169-5347(97)01230-5
Serghi EU, Kokkoris V, Cornell C, Dettman J, Stefani F, Corradi N (2021) Homo- and dikaryons of the arbuscular mycorrhizal fungus Rhizophagus irregularis differ in life history strategy. Front Plant Sci 12:715377. https://doi.org/10.3389/fpls.2021.715377
Silvestri A, Fiorilli V, Miozzi L, Accotto GP, Turina M, Lanfranco L (2019) In silico analysis of fungal small RNA accumulation reveals putative plant mRNA targets in the symbiosis between an arbuscular mycorrhizal fungus and its host plant. BMC Genomics 20(1):169. https://doi.org/10.1186/s12864-019-5561-0
Silvestri A, Turina M, Fiorilli V, Miozzi L, Venice F, Bonfante P, Lanfranco L (2020) Different genetic sources contribute to the small RNA population in the arbuscular mycorrhizal fungus Gigaspora margarita. Front Microbiol 11:395. https://doi.org/10.3389/fmicb.2020.00395
Smith SE, Read DJ (2008) Mycorrhizal symbiosis, 3rd edn. Academic Press, London, pp 1–787
Smith KM, Phatale PA, Sullivan CM, Pomraning KR, Freitag M (2011) Heterochromatin is required for normal distribution of Neurospora crassa CenH3. Mol Cell Biol 31(12):2528–2542. https://doi.org/10.1128/MCB.01285-10
Song G, Dickins BJ, Demeter J, Engel S, Gallagher J, Choe K, Dunn B, Snyder M, Cherry JM (2015) AGAPE (automated genome analysis PipelinE) for pan-genome analysis of Saccharomyces cerevisiae. PLoS One 10(3):e0120671. https://doi.org/10.1371/journal.pone.0120671
Stajich JE (2017) Fungal genomes and insights into the evolution of the kingdom. Microbiol Spectr 5(4). https://doi.org/10.1128/microbiolspec.FUNK-0055-2016
Strom NB, Bushley KE (2016) Two genomes are better than one: history, genetics, and biotechnological applications of fungal heterokaryons. Fungal Biol Biotechnol 3:4. https://doi.org/10.1186/s40694-016-0022-x
Strullu-Derrien C, Selosse MA, Kenrick P, Martin FM (2018) The origin and evolution of mycorrhizal symbioses: from palaeomycology to phylogenomics. New Phytol 220(4):1012–1030. https://doi.org/10.1111/nph.15076
Sugiura Y, Akiyama R, Tanaka S, Yano K, Kameoka H, Marui S, Saito M, Kawaguchi M, Akiyama K, Saito K (2020) Myristate can be used as a carbon and energy source for the asymbiotic growth of arbuscular mycorrhizal fungi. Proc Natl Acad Sci U S A 117(41):25779–25788. https://doi.org/10.1073/pnas.2006948117
Sun X, Chen W, Ivanov S, MacLean AM, Wight H, Ramaraj T, Mudge J, Harrison MJ, Fei Z (2019) Genome and evolution of the arbuscular mycorrhizal fungus Diversispora epigaea (formerly Glomus versiforme) and its bacterial endosymbionts. New Phytol 221(3):1556–1573. https://doi.org/10.1111/nph.15472
Tamaru H (2010) Confining euchromatin/heterochromatin territory: jumonji crosses the line. Genes Dev 24(14):1465–1478. https://doi.org/10.1101/gad.1941010
Tisserant E, Malbreil M, Kuo A, Kohler A, Symeonidi A, Balestrini R, Charron P, Duensing N, Frei dit Frey N, Gianinazzi-Pearson V, Gilbert LB, Handa Y, Herr JR, Hijri M, Koul R, Kawaguchi M, Krajinski F, Lammers PJ, Masclaux FG et al (2013) Genome of an arbuscular mycorrhizal fungus provides insight into the oldest plant symbiosis. Proc Natl Acad Sci U S A 110(50):20117–20122. https://doi.org/10.1073/pnas.1313452110
Trojer P, Reinberg D (2007) Facultative heterochromatin: is there a distinctive molecular signature? Mol Cell 28(1):1–13. https://doi.org/10.1016/j.molcel.2007.09.011
Van Der Heijden MGA, Scheublin TR (2007) Functional traits in mycorrhizal ecology: their use for predicting the impact of arbuscular mycorrhizal fungal communities on plant growth and ecosystem functioning. New Phytol 174(2):244–250. https://doi.org/10.1111/j.1469-8137.2007.02041.x
Venice F, Ghignone S, Salvioli di Fossalunga A, Amselem J, Novero M, Xianan X, Sedzielewska Toro K, Morin E, Lipzen A, Grigoriev IV, Henrissat B, Martin FM, Bonfante P (2020) At the nexus of three kingdoms: the genome of the mycorrhizal fungus Gigaspora margarita provides insights into plant, endobacterial and fungal interactions. Environ Microbiol 22(1):122–141. https://doi.org/10.1111/1462-2920.14827
Wewer V, Brands M, Dormann P (2014) Fatty acid synthesis and lipid metabolism in the obligate biotrophic fungus Rhizophagus irregularis during mycorrhization of Lotus japonicus. Plant J 79(3):398–412. https://doi.org/10.1111/tpj.12566
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci U S A 84(24):9054–9058. https://doi.org/10.1073/pnas.84.24.9054
Wyss T, Masclaux FG, Rosikiewicz P, Pagni M, Sanders IR (2016) Population genomics reveals that within-fungus polymorphism is common and maintained in populations of the mycorrhizal fungus Rhizophagus irregularis. ISME J 10(10):2514–2526. https://doi.org/10.1038/ismej.2016.29
Yildirir G, Malar CM, Kokkoris V, Corradi N (2020) Parasexual and sexual reproduction in arbuscular mycorrhizal fungi: room for both. Trends Microbiol 28(7):517–519. https://doi.org/10.1016/j.tim.2020.03.013
Yildirir G, Sperschneider J, Malar CM, Chen ECH, Iwasaki W, Cornell C, Corradi N (2022) Long reads and Hi-C sequencing illuminate the two-compartment genome of the model arbuscular mycorrhizal symbiont Rhizophagus irregularis. New Phytol 233(3):1097–1107. https://doi.org/10.1111/nph.17842
Zhao LY, Song J, Liu Y, Song CX, Yi C (2020) Mapping the epigenetic modifications of DNA and RNA. Protein Cell 11(11):792–808. https://doi.org/10.1007/s13238-020-00733-7
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Dallaire, A., Paszkowski, U. (2023). Genomes of Arbuscular Mycorrhizal Fungi. In: Scott, B., Mesarich, C. (eds) Plant Relationships. The Mycota, vol 5. Springer, Cham. https://doi.org/10.1007/978-3-031-16503-0_4
Download citation
DOI: https://doi.org/10.1007/978-3-031-16503-0_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-16502-3
Online ISBN: 978-3-031-16503-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)