Abstract
Artificial intelligence (AI) is a field that designs computer systems to perform tasks that mimic human intelligence. Machine learning (ML) is subtype of AI whereby learning is attained by training and makes decisions from the data. AI is rapidly transforming radiology. This chapter provides a brief overview of the basic concepts and common methods in ML, followed by a small sampling of potential applications in neuroradiology.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Artificial intelligence
- Machine learning
- Deep learning
- Neural networks
- Supervised learning
- Convolutional neural network
- Neuroradiology
- Autoencoders
- UNET
- Detection
- Classification
- Segmentation
- Prediction
- Improve image quality
Introduction
Artificial intelligence (AI) has dominated medical research and clinical applications in recent years. The use of the term AI dates to many decades ago, first introduced in the 1956 Dartmouth Summer Research Project on Artificial Intelligence. AI generally encompasses all applications regarding computers performing tasks requiring human intelligence and its simulation of learning [1]. Machine learning (ML) is a subtype of AI, where algorithms learn to perform tasks by “training” a large dataset, learning the data’s characteristics without explicit assumptions of the relationships between variables [2,3,4]. Of all ML algorithms, neural networks (NN) have recently gained much interest in radiology, due to their natural affinity for analyzing images. These networks consist of layers of interconnected nodes (“neurons”) that are roughly based on the layered organization of neurons in the brain [5]. With a multilayered NN, “deep” networks can be built, hence the term “deep learning (DL)” when referring to applications which employ this type of ML algorithm [6].
Excitement around NNs was stirred up after a DL-based algorithm won the ImageNet Challenge in 2012 (an annual competition for classification of natural images) and greatly surpassed the performance from past years [7]. This excitement in DL quickly extended to the medical imaging field, and has been attracting immense interest not only because of the advances in ML theory and the development of better algorithms, but also due to the advances in hardware (improved computational resources such as graphics processing units (GPUs) and the accumulation of medical data (the large amount of data is commonly referred to as “big data [8]”) needed to train the algorithms [3]. DL applications have shown great potential in ophthalmology [9], dermatology [10], radiology [11], and pathology [12], to name a few examples. In radiology, some uses of AI include automating time-consuming tasks, solving problems that are intellectually difficult for humans, making diagnoses, and asserting predictions.
We provide a brief overview of AI in neuroradiology by describing key terms, common ML algorithms, basic NN architecture, and a small sampling of applications.
Basic Definitions
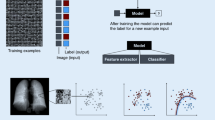
Artificial intelligence (AI) is a field that designs computer systems to perform tasks that mimic human intelligence. Machine learning (ML) is subtype of artificial intelligence that develops algorithms to acquire knowledge and make decisions from the data. Classic ML depends on carefully human-engineered features extracted from input data. For many tasks, however, it is difficult to predetermine which features to extract. To address this problem, representation learning was developed to teach machines to discover not only the mapping from input to output, but also the representation itself. The representation-learning algorithm determines the optimal set of features to best carry out the task. For very complex tasks, a hierarchy of features, from concrete to abstract, local to global, may be needed. Deep learning (DL) provides an elegant solution by using a layered architecture, whereby progressively more complex patterns are extracted as data pass through the layers. Through this tiered processing, simple features (such as intensity, edges, and textures) are conjugated to build more complex features (such as corners, contours, etc.), from which more elaborate structures (such as organs and lesions) are constructed. Similarly, complex abstractions can be formulated upon simpler concrete concepts (Fig. 58.1).
Artificial intelligence methods. Within the subset of machine learning methods, deep learning is usually implemented as a form of supervised learning. Reprinted from “Deep Learning in Neuroradiology”, AJNR Am J Neuroradiol. 2018;39(10):1776–1784, Zaharchuk et al., with permission from WILLIAMS & WILKINS CO.; American Society of Neuroradiology
There are two general methods by which machines learn: “supervised learning” and “unsupervised learning,” which differ in their applications and the input data. In supervised learning, some “ground truth” exists, which is used to train the algorithm. During the training process, the correct answers are known a priori, and the algorithm iteratively makes predictions on the training data and adjusts the parameters to minimize the errors on subsequent iterations. Training continues until the machine achieves a desired level of accuracy or performance plateaus. Common applications of supervised learning include classification and regression. For example, classification algorithms might aim to identify specific tumors as “meningioma,” “astrocytoma,” or “glioblastoma” (multiclass classification) or perhaps predict successful treatment response from radiosurgery (binary classification). The goal of regression techniques is to predict a number or series of numbers (such as biomarkers) from an image, such as the volume of abnormal white matter in a multiple sclerosis patient. Common supervised learning algorithms include linear regression or logistic regression for regression problems; support vector machines (SVM) for classification problems, and K-nearest neighbor and decision trees (including random forest) for both classification and regression problems.
In unsupervised learning, no ground truth images or classifications are provided. They may be unknown, and as such, the procedure can be used to generate hypotheses. In this situation, the algorithm must come up with its own rules to organize images or data. It may use mathematical processes to systematically reduce redundancy, organize data by similarity, or separate into groups based on variability. Common applications of unsupervised learning include clustering (to discover inherent groupings), dimensionality reduction (generalization), and association (pattern search). Some popular examples of unsupervised learning algorithms are: K-means for clustering, principal component analysis (PCA) for dimensionality reduction, and a priori algorithms for association problems.
Machine Learning: Some Basic Terms
Many algorithms use similar approaches such that a brief overview of terminology can be helpful. The following lists of key terms are commonly used in machine learning.
Features are measurable properties or attributes that represent the object of interest. In the case of medical images, features can be the pixel values, curvature, gradient, entropy, etc. Features are often stacked together into a longer feature vector that is used as an input to the ML model. Traditionally, the goal of many imaging researchers has been to create images with desired features, based on their domain knowledge and presumed biological mechanisms. More recently, automated features have been popular, an approach that has been labeled radiomics. With neural networks, features are identified directly from the data without human intervention.
Weights are learnable parameters of the model; in fact, sometimes the words “weights” and “parameters” are used interchangeably. They are usually initialized randomly and are updated during training to optimize the model’s performance. Sometimes, the initial weights can be set based on prior training of a network trained on a similar problem, a method known as “pretraining,” which can reduce training time and improve performance in some situations. In nondeep-learning models, each input feature is multiplied by a weight. In this context, weights represent how much influence a feature or variable has on the output. In neural networks, weights represent the strength of the connection between nodes. The goal of training is to optimize these weights to achieve the best performance. They are then fixed when the model is applied in a production on new, unseen data, a process known as “inference.”
Hyperparameters are the configuration options of the ML model that are selected and usually tuned manually to obtain optimal performance. Learning rate for training a neural network, number of layers, k in k-nearest neighbors, and maximum depth in decision trees are some examples of hyperparameters.
Loss-function is a mathematical expression for evaluating how well the model is fitting the data. The choice of the loss-function is task-dependent. For example, in a regression model to predict treatment response, such as days to progression, the mean-squared error between true and predicted number of days can be used. The larger the difference between the prediction and the truth, the more changes need to be made during the iterative updating of the weights. For binary or multiclass classification, other methods are used, such as cross-entropy.
Gradient descent is an optimization algorithm, which adjusts the parameters in small increments to minimize the loss function. It can be thought of as the algorithm trying to descend the landscape created by the loss function to find the lowest possible loss on the given data, which presumably identifies the model weights that represents the best solution.
Underfitting refers to a model that cannot perform well with training data or new data (Fig. 58.2). Sometimes, this is due to a model that does not have enough parameters to represent the data, suggesting important features for prediction are not being used as inputs to the model.
Illustration of (a) underfitting, (b) best-fitting, and (c) overfitting. (a) Underfitting fails to capture the pattern. (b) Best-fit captures the pattern and is not too inflexible or flexible and is likely to have better accuracy on new, unseen data. (c) Overfitting fits strenuously to the noise of the training data; while it may perform well with the training data, this performance is degraded when applied to new, unseen data
Overfitting occurs when a model learns the training data and all its idiosyncrasies too well, to the extent that it limits the model’s ability to generalize, which result in poor performance on new data (Fig. 58.2). With enough parameters, a model can learn to reproduce the training data exactly, essentially memorizing the particular group of data it is trained on; since new data will necessarily differ, such a solution will show degraded performance on new data the model has never seen (the “test” set). The best way to avoid overfitting is to collect more training examples, though other approaches such as cross-validation, regularization, and dropout can also be used.
K-fold cross validation is a useful procedure to provide a less biased or less optimistic estimate of a model’s performance, which can also reduce overfitting. The dataset is divided into K number of groups/folds, where one group/fold is used as testing set and the remaining k-1 folds are used for training. This process is repeated K times until each fold of the K folds have been used as the testing set. This leads to the creation of K individual models and thus an idea of the sensitivity of the model to different splits of training data. Either the model with the “best” performance can be selected for future predictions or the different models can be used together in consensus for better performance.
Regularization is a technique to reduce overfitting by reducing the complexity of a model. It is based on the idea that smaller values of the parameters tends to minimize the risk of overfitting aspects of the data that are just due to random noise. This is generally accomplished by adding a term to the loss function to penalize large parameter values associated with more complex models. Ridge regression and Lasso are popular regularization methods.
Common Machine Learning Algorithms
Some Common Machine Learning Algorithms
Choosing the appropriate algorithm for the task and the available data is crucial. Below are some common ML algorithms grouped by their functionality (note some algorithms may belong to multiple functional categories) (Fig. 58.3). The most common ML applications in neuroradiology are for classification and regression tasks.
Regression Algorithms
Regression is used for making predictions based on previous observations. Regression algorithms model the relationship between a set of explanatory variables and the outcome variable(s). In radiology, regression models are often used for predicting treatment outcome and risk assessment. Popular regression algorithms include:
Linear Regression methods, the workhorse of statistics, have been co-opted into statistical ML. Linear regression is used when the prediction is continuous and its relationship with the dependent variables is thought to be linear. Multivariate linear regression is used when more than one feature is being used to estimate the final variable of interest (Fig. 58.4).
(a) Simple linear regression model with an equation in the form Y = b0 + b1X, where X is the independent (explanatory) variable and Y is the dependent (output) variable. (b) Instead of fitting a linear line to the explanatory variable (X), logistic regression fits an S-shaped “logistic function” to predict a binary output variable (Y). (c) Multivariate adaptive regression splines model demonstrating a continuous relationship, between the output variable (Y) and the explanatory variable (X), which is different for different ranges of X. For example, there is a positive relationship between Y and X, if X is between 0 and 1 and a negative relationship when X is between 2 and 4
Logistic Regression is used when the prediction is binary (Fig. 58.4). Logistic regression uses the sigmoid function to model the input data, \( g(z)=\frac{1}{1+{e}^{-z}\;} \), and produces an output ranging between a minimum of 0 and a maximum of 1. A threshold is applied to make the binary decision.
Multivariate Adaptive Regression Splines (MARS) is a nonparametric regression method that makes no assumption about the relationship between the predictors and dependent variables (Fig. 58.4). Instead, the relationship between the predictors and dependent variables is derived from the regression data using multiple piecewise linear regression. MARS can derive models even when the relationship between the predictors and the dependent variables is nonmonotonic.
Classification Algorithms
Classification algorithms use supervised learning to separate data into different categories. Popular classification algorithms include:
K-Nearest Neighbor assumes similar data points are close to each other. A new data point is labeled according to the most represented label among “k” number of its nearest neighbors (Fig. 58.5). One concern for these models is that they perform better if there is good balance in the number of examples of each class in the training data. Otherwise, the class with the most examples will tend to dominate the predictions.
Support Vector Machine transforms the seemingly inseparable data into a higher dimensional space and finds a hyperplane that can distinctly classify the data points, with a maximum margin separating the two classes (Fig. 58.6). In SVM, kernels are used to transform the input data into the required format in the higher dimensional space. Choosing the right kernel is a challenge. Some of the kernels used in SVM are linear, nonlinear, polynomial, radial basis function (RBF), and sigmoid.
SVM example. Circles and squares represent different classes. In the standard space (left), the two classes cannot be separated by a linear line. SVM transforms the original data into a different space (right), where they can be separated by an optimal hyperplane (red solid line) with the largest possible margin (black dotted lines)
Decision Trees. are flowchart-like models that can be used for regression and classification problems, with categorical variables or continuous variables. The whole training dataset starts at the root. Different algorithms (e.g., ID3, C4.5, CART, etc.) are available to split the data into subnodes recursively, until leaf/terminal nodes are reached. The goal is to create subnodes that are progressively more homogeneous (pure) (Fig. 58.7). When building the tree, “information gain” and “entropy” are calculated to determine which attribute is used to split each node. Entropy is a measure of the randomness. Information gain (IG) measures how well an attribute separates the data into their target classifications. Mathematically, IG computes the decrease in Entropy after a split based on an attribute (IG = Entropy_before – Entropy_after). Constructing a decision tree is about finding an attribute that returns the lowest entropy and the highest IG. Splitting stops when entropy or IG is zero, or some predetermined criteria (such as maximum depth) is met. To avoid overfitting, the full tree then undergoes pruning., to trim off some branches such that overall accuracy is unaffected. In practice, the training dataset is used to create the tree and the validation dataset is used for trimming.
Clustering Algorithms
Clustering is similar to classification, except that the classes are unknown. Clustering algorithms use unsupervised methods to group data points by their similarity while maximizing the variance between groups. The most popular clustering algorithms are:
K-Means Clustering. that groups similar objects together into clusters (Fig. 58.8). The algorithm starts by guessing the initial centroids for each cluster, and then repeatedly assigns instances to the nearest cluster and re-computes the centroid of that cluster.
The process of assignment and recalculation of the centroids is repeated until the centroids no longer move (i.e., assignment of objects to clusters also stabilizes). This produces a separation of the objects into groups with minimal intracluster distance and maximal intercluster distance.
Hierarchical Clustering is an iterative algorithm that builds a hierarchy of clusters (Fig. 58.9). Initially, each data point is considered as an individual cluster. The similar clusters merge into the same cluster iteratively, until one cluster is formed.
Dimensionality Reduction Algorithms
Dimensional reduction algorithms attempts to summarize and simplify data representation in an unsupervised manner. The goal is to reveal inherent structure within the data. After dimensional reduction, the simplified representation can then be used in a supervised learning method. These algorithms are often used in classification and regression. Principal component analysis is an unsupervised technique, while Linear Discriminant Analysis is a supervised technique. They are common dimensionality reduction techniques used as a preprocessing step in Machine Learning and pattern classification applications.
Principal Component Analysis (PCA) is a mathematical procedure often used to reduce the dimensionality of large data sets. PCA transforms a set of correlated variables to a set of uncorrelated (orthogonal) variables (Fig. 58.10). Dimensionality-reduction is achieved by retaining the dimensions that contains the highest variance (hence, most information), while dropping the dimensions with the lowest variance. It will help us extract essential information from data by reducing the dimensions. PCA captures the most essential information contained in the data using fewer dimensions.
Principal component analysis (PCA) transforms a set of correlated variables to a set of uncorrelated (orthogonal) variables. In this example, the principal component, λ2, captures the maximum variance. To capture information contained in the data using fewer dimensions (dimensionality reduction), the λ1 dimension can be eliminated
Linear Discriminant Analysis is very similar to PCA. In addition to finding the component axes that maximize the variance of the data (PCA), LDA also finds the axes that maximize the separation between multiple classes (Fig. 58.11). LDA transforms the data into a variable space, which minimizes the intraclass variance and maximizes the interclass variance. The features in higher dimension space are then projected onto a lower dimensional space, in order to separate the data into two or more classes.
Ensemble Algorithms
Ensembling is an ML technique that combines several models together to make the final prediction (Fig. 58.12). Typically, ensemble models outperform each constituent model, which is why ensemble models are very powerful and popular. There are three common methods to create ensembles: (1) stacking, (2) bagging, and (3) boosting.
-
1.
Stacking passes the input through several different algorithms in parallel (Fig. 58.13). The corresponding outputs are then as input to the last model, which makes a final decision. The final decision-making step usually uses a regression model.
-
2.
Bagging (aka Bootstrap aggregation) uses the same algorithm and trains it on different subsets of the data (Fig. 58.14). Data in the subsets are random and may repeat. The algorithm is trained on subsets several times and then predicts the final answer by majority voting. The most famous example of bagging is Random Forest, which is bagging on the decision trees.
Random Forest is an ensemble of decision trees for classification or regression tasks (Fig. 58.15). Multiple decision trees are constructed by repeatedly resampling subsets of the training data with replacement. The final consensus prediction of the random forest is determined by polling each decision tree – using either the max-vote (in classification) or mean value (in regression).
-
3.
Boosting uses a series of models that are trained sequentially, to convert weak learners into strong learners, thereby improving the performance. Each subsequent model is designed to correct the errors from its predecessor (Fig. 58.16).
Adaptive Boosting (AdaBoost) is a popular boosting method that uses adaptive weights to force the model to concentrate on difficult cases that are prone to erroneous classification. Subsequent trees are grown to help classify observations that are not misclassified by the previous trees. Predictions of the final ensemble model are the weighted sum of the predictions made by the ensemble of tree models.
Gradient Boosting Machines (GBM) are modern boosting methods that are adapted from AdaBoost. The major difference between AdaBoost and Gradient Boosting Algorithm is how the two algorithms identify and boost the weak learners. GBM uses a gradient-descent-like method to minimize the loss function of the model. Instead of using higher weights to boost weaker learns, as in AdaBoost, GBM adjusts the gradients. The ability to use a customized loss function makes GBM adaptable to a wide range of applications and thus, is widely popular.
EXtreme Gradient Boosting (XGBoost) is a specific implementation of Gradient Boosting, which uses a variety of regularization techniques that reduce overfitting and improve overall performance.
Deep Learning
The classic example of a deep learning model is the neural network (NN), which was inspired by human neural networks.
Each biologic neuron processes and integrates the received stimuli and fires off if the excitation threshold is surpassed, thereby propagating the signal to the downstream neurons. Similarly, an artificial neural network is a computational framework of interconnected neurons (called nodes), arranged in layers (Fig. 58.17). Typically, NN consists of an input layer, one or more interconnected layers of neurons, and an output layer for making predictions. Within a layer, each node processes its input mathematically (applying weights and summing), makes a decision (by applying an activation function), and then passes the output on to the next layer of nodes. Weights (represented by the arrows in Fig. 58.17) connect the nodes in different layers and represent the strength of connections between the nodes.
A neural network consists of an input layer that connects to the input variables, one or more hidden layers, and an output layer that produces the output variables. This example has three hidden layers with five neurons in each layer with two final output classifications. All layers are fully connected. Feature representation gets progressively more complex and abstract as layers get deeper. Within each hidden layer, each node processes its input mathematically (applying weights and summing), makes a decision (by applying an activation function), and then passes the output on to the next layer of nodes
The power of these NNs is in their scalability, which is largely based on their ability to automatically extract relevant features from a labeled dataset, circumventing the need of expert-engineered formulations. Typical NN architectures start with an input layer, where data is turned into features. Next are a few hidden layers, which compute intermediate representations of features. The final layer is the output layer, which produces the results.
Training and Optimizing
As data pass through the multiple layers, a process called “forward propagation,” the NN computes a hierarchy of features (from simple to complex, perceptible to abstract), which are then used to produce the desired output. For each forward propagation of each training data, the performance of the NN is assessed by a loss function, which quantifies the error between the predicted value and the true value. Choosing the right loss function is important. Different loss functions may be selected depending on the task; for example, for binary or multiclass classification, “cross-entropy loss” is commonly used; for segmentation tasks the Dice coefficient [13] may be incorporated in the loss function to assess and reward the algorithm for creating predictions that have high overlap with the ground truth segmentation; for image transformation tasks, mean squared error summed over all voxels could be utilized to compare the similarity of two images [3]. During training, the error calculated by the loss function, is back-propagated through the NN, one layer at a time, and parameters that affect the performance (e.g., the magnitude of the weights at each level) are adjusted accordingly. Typically, this is carried out iteratively by an optimization algorithm such as gradient descent.
NN are ideally trained using large numbers of cases that are divided into three subsets: a training set, a validation set, and a test set. The actual learning process of an ML algorithm requires using a training dataset. After training, the performance of the algorithm is assessed with a set of validation data; this is used to inform the training of the algorithm in later iterations and for selecting the best “hyperparameters,” such as learning rate and prediction thresholds [4]. A test set, which consists of data the algorithm has never seen and is separate from the training and validation sets, is then used to evaluate the final performance of the algorithm [4].
Overfitting and Data Augmentation
Since having a large dataset is crucial for good performance, data augmentation can be performed to increase the size and variety of the dataset. Transformations (e.g., flipping, rotating, skewing, cropping, etc.), modifications of attributes (e.g., orientation, location, size, brightness), and noise can be synthetically applied to the acquired images to artificially generate more training data. Augmentation can potentially improve the robustness of the models, presumably by aiding the NN learn generalized features that are invariant to orientation, noise, etc. Data augmentation should only be applied to the training dataset and not to the validation or test dataset.
Deep learning models have many hyperparameters and even more parameters (e.g., weights and biases). To avoid overfitting, regularization and dropout can be used, although having more training examples is most ideal. Dropout is a regularization method that approximates training many different, slightly modified, smaller NNs in parallel. During training, some nodes (along with their downstream connections) are randomly “dropped” or ignored by the NN. This has the effect of spreading and shrinking the weights, reducing the probability of over-relying on a particular node or a particular feature. Like regularization methods, dropout is effective when there is a limited amount of training data, which makes the model susceptible to overfitting.
Common Deep Learning Algorithms
This is an ever-growing field. Below are a few popular deep learning algorithms used in neuroradiology:
Autoencoder
Autoencoders are a specific type of feedforward neural networks where the generated output image is an improved version of the input image. Autoencoders consist of three components: encoder, code, and decoder (Fig. 58.18). The encoder compresses the input into a lower-dimensional code while the decoder then reconstructs the output from this code. During the encoding step, the autoencoder learns to extract only the important features from the input images and to ignore irrelevant noise. Thus, noise and artifacts are removed when the decoder reconstructs the images. Similar to a U-net, such a method can be used to remove noise from medical images.
Convolutional Neural Network (CNN)
A convolutional neural network (CNN) is a class of NN that is most commonly used for classification and segmentation of both natural and medical images (Fig. 58.19). In traditional NNs, the two-dimensional images are flattened into a long vector of pixel values as input. CNNs can accept nonflattened images and learn the spatial relationship between pixels in a hierarchical manner. The basic CNN has three types of layers:
-
1.
Convolutional layers for extracting feature maps.
-
2.
Pooling layers for trimming down the features.
-
3.
Fully-connected layers for making final predictions.
Convolutional neural network. The input image is submitted to a series of convolutions, producing a stack of features maps containing low-level features. These feature maps are then downsampled by a max pooling layer. Deeper convolution layers produce higher-level global features. Layers of convolutions and max pooling are alternately stacked until the CNN is deep enough to capture the features of the images for the task at hand. Feature maps are then flattened into a single vector for the final classification or regression output-step
The first layer of CNN architecture is the convolution layer, which uses convolution filters (a.k.a. feature detectors or kernels) to extract features from the input image. Filters move across the whole image to detect features by applying a small kernel of weights at each pixel, a mathematic operation called convolution. For each layer, multiple different kernels can be used to learn a wide range of features, such as edges, textures, and other nonlinear representations of the data. Deeper convolution layers assemble lower-level local features into higher-level global features. The filter values are the learnable parameters that are adjusted during training to optimize the extracted features. It is required to put a nonlinear “activation function” at the output of the neuron. Typically the Rectified linear unit (ReLU) is used because it is effective and simple to implement. ReLU outputs the input value for positive inputs and blocks negative inputs, setting the outputs in these cases to zero (Fig. 58.20a). The nonlinear activation functions introduce nonlinearity into the CNNs, so that complex functions can be represented that would not otherwise be possible, making CNNs more powerful than linear classifiers.
(a) ReLU outputs the input value for positive inputs and blocks negative inputs. (b) Leaky ReLU replaces the horizontal component with a function with small nonzero gradient. This is done to mitigate the “dying ReLU” issue. (c) Exponential linear unit (ELU) uses a log curve instead of a straight line for the negative inputs. (d) Scaled exponential linear unit (SELU) is a scaled (α) version of ELU
ReLU is a popular activation function because it is easy to implement. Mathematically, it is defined as y = max(0, x). It is also every effective in removing neurons from the network during the training process. However, the nulled neurons cannot be recovered and are definitively eliminated, which may prevent the network from converging or impair the accuracy. To mitigate this “dying ReLU” problem, variants of the ReLU function were introduced. The leaky ReLU replaces the zero output (for negative inputs) with a function with a small nonzero gradient (Fig. 58.20b). The nonzero gradient will retain the neurons, allow them to recover during training and keep learning. Similar to leaky ReLU, another variant, the Exponential linear unit (ELU) uses a log curve instead of a straight line for the negative inputs (Fig. 58.20c). ELU outperformed all the ReLU variants in the original paper’s experiments. Scaled exponential linear unit (SELU) is a scaled version of ELU with an additional scale parameter, α (Fig. 58.20d).
Pooling layers are introduced between convolution layers to reduce the dimensionality of the feature maps, which also helps with overfitting. Pooling consolidates and generalizes the most important features. Max pooling, which propagates the maximum activation, is often used. Successive pooling operations result in maps with progressively lower resolution, increasingly richer information, and more global representation. After the features are extracted by convolutional layers and consolidated by the pooling layers, they are flattened into a long vector and introduced to one or more fully-connected layers. In the fully-connected layers, all the neurons in one layer are connected to all neurons in the next layer. They are used to generate nonlinear combinations of the learned features, in order to make the final predictions.
UNET
U-Nets
The successive layers of convolution and pooling in CNNs increase abstraction of the feature maps but lose spatial information in the process. Therefore, while CNNs can generate the feature maps to detect or classify a targeted lesion, they cannot locate the lesion within the image for segmentation tasks. U-nets were designed to mitigate this problem. The UNET architecture has three parts (Fig. 58.21):
-
1.
Contracting/Downsampling path.
-
2.
Bottleneck.
-
3.
Expanding/Upsampling path.
UNET architecture. Each gray box corresponds to a multichannel feature map. The number of channels is denoted on top of the box. The x–y size is denoted along the left edge of the box. White boxes represent the copied feature maps from the skipped connections. Different colored arrows represent different operations
There are usually a symmetric number of downsampling and upsampling layers, with extra connections between nodes in shallower layers that skip some deeper layers. Similar to CNNs, the downsampling layers capture the context of the image. Feature maps are generated with successive downsampling, which involves convolution, ReLU, and max pooling steps. The bottleneck layer, consisting of convolutional layers, is added to reduce the number of feature maps. Upsampling layers consist of deconvolution, upsampling, convolution, and ReLU. The expanding path incorporates contextual information (from the contracting path) with localization information (obtained by skip connections) to localize and segment targets within the image.
Generative Adversarial Network (GAN)
Generative adversarial networks (GANs) are used to generate output images that share realistic features with the desired ground truth images [14]. GANs have two submodels: a generator model and a discriminator model (Fig. 58.22). The generator model generates new imaging samples after learning patterns from training images. Many of the methods described above can be used as the generator, such as a U-net. The output produced by a good generator model should be almost indistinguishable from real training images. The discriminator model attempts to distinguish between samples drawn from the training images and those produced by the generator. It receives as input the real and the generated image and trains a network to try to distinguish them from each other. The two models are trained together in an adversarial manner - if the discriminator successfully identifies real and generated samples, the discriminator’s parameters will remain unchanged, but the generator’s parameters will be modified; alternately, if the generator fools the discriminator, the generator’s parameters will remain unchanged, but the discriminator’s parameters will be modified. GANs provide a powerful and clever mechanism for image augmentation and image transformation.
Generative adversarial networks are comprised of a generator and a discriminator. The generated samples from the generator and the real samples are classified as real or fake by the discriminator. The generator is updated based on how well, or not, the generated samples fool the discriminator. The discriminator is updated based how accurately it can classify the samples
Transfer learning is a technique whereby a new model is built upon another neural network model that was previously trained for a similar task. Layers from VGG, GoogLeNet (http://deeplearning.net/tag/googlenet/) or Inception-ResNet (https://keras.rstudio.com/reference/application_inception_resnet_v2.html), trained on large groups of nonmedical images, are often reused in medical imaging models. Transfer learning has the benefit of decreasing the training time for a neural network model and can result in lower generalization error. The weights in reused layers are usually used as the starting point for the training process, and thus may require less training data when compared to models that are built from scratch. Often only some of the deeper layers are re-trained with the new data, as this can frequently lead to better performance.
Model Design and Assessment
For an ML algorithm to be effective, care is needed in selecting the optimal model and cost function, defining the hyperparameters, as well as providing the model with sufficient amounts of training data [3].
Data Preparation and Augmentation
It is standard practice to divide available data into three subsets: a training set, a validation set, and a test set. The actual learning process of an ML algorithm requires using a training dataset. After training, the performance of the algorithm is assessed with a set of validation data; this is used to inform the training of the algorithm in later iterations and for selecting the best “hyperparameters,” such as learning rate and prediction thresholds [4]. A test set, which consists of data the algorithm has never seen and is separate from the training and validation sets, is then used to evaluate the final performance of the algorithm [4]. Random transformations (e.g., flipping, rotating, skewing, dimming, etc.) can be applied to the images, to “augment” the imaging dataset, though these are usually used exclusively in the training set.
Applications in Neuroradiology
In radiology, opportunities exist for AI in all aspects of the imaging life cycle, from protocol automation before acquisition [15], image reconstruction and quality improvement after acquisition [16, 17], to image interpretation [9, 10]. ML can also combine imaging and clinical metadata to predict treatment response or clinical outcome [18]. We shall explore a small sample of AI applications in neuroradiology.
Detection
Critical Findings on Emergent CT
Noncontrast head CT scans are the most commonly ordered studies for emergent diagnosis and they constitute the largest volume of work for neuroradiologists. Automating head CT scan interpretation can streamline the workflow and raise appropriate alerts promptly. Deep learning has been successfully used to detect critical findings such as intracranial hemorrhage, fracture, midline shift, and mass effect on head CTs [19]. Their algorithms achieved an AUC of 0.92 for detecting intracranial hemorrhage, 0.92 for detecting calvarial fractures, 0.93 for detecting midline shift, and 0.86 for detecting mass effect. Different hybrid models were developed and optimized to detect each of the abnormalities. For instance, a modified ResNet18 with five parallel fully connected (FC) layers was used for detecting and distinguishing the types of hemorrhage (intraparenchymal, intraventricular, subdural, extradural, and subarachnoid hemorrhages). The confidences at the slice-level are then combined, using a random forest, to predict the subject-level confidence for the presence of intracranial hemorrhage. A 2D UNET was then used to segment the extent of the hemorrhage. In a similar manner, a modified ResNet18 model was used to detect mass effect and midline shift. A DeepLab-based architecture was used to predict pixel-wise heatmap for acute fractures. These engineered features representative of fractures was used to train a random forest model to predict the presence of a calvarial fracture [19]. Transfer learning has been successful in detecting the presence of hemorrhage on noncontrast brain CT, with accuracies of >98% [20]. These promising performances suggest the potential of using DL to triage head CT scans and prioritize studies with detected critical findings. While this may reduce interpretation time for the flagged studies, it is still unclear if this would have positive effects on patient outcome.
Of all the urgent indications for head CTs, there is nothing that needs more timely accurate diagnosis than acute stroke. There are several commercial software suites that incorporate artificial intelligence for comprehensive acute stroke imaging which includes evaluation of ASPECTS and intracranial hemorrhage on noncontrast CT, large vessel occlusion detection and/or collateral assessment on CTA, and measurement of infarct core and penumbra on CT perfusion. Some software even has emergency activation or mobile-device notification capabilities [21]. In multiple studies [22,23,24] [25], their performance was noninferior to experienced neuroradiologists.
Screening for Aneurysm
Screening for aneurysms is tricky, particularly if they are small. Many computer-assisted algorithms for detection of aneurysms have been designed on different modalities [26,27,28,29]. One of the better models used transfer learning based on ResNet-18 for detecting aneurysms on time-of-flight (TOF) MRA, achieving 91% to 93% sensitivity with detection of more aneurysms than human readers [29]. Digital subtraction angiography (DSA) is the gold standard for diagnosing aneurysms, but can still be challenging if vessels bend and overlap, which can appear similar to aneurysms at certain projections. A two-stage CNN detection system has been used to differentiate vessel overlaps from aneurysms on DSA [28]. The first CNN localizes the ROI to the target vessel (posterior communicating artery) in order to minimize interference from other vessels; the second stage CNN combined frontal and lateral views to detect aneurysms, using a concurrent false-positive suppression algorithm trained to ignore vessel overlaps, achieving an accuracy of 93.5%. In practice, neurointerventionists often use 3D-rotational angiography to help them discern and characterize small aneurysms. 3D-rotational angiography consists of a series of 2D images, taken circumferentially around the head during arterial contrast injection, followed by 3D reconstruction of the vasculature. To simulate this, several 3D-rotational angiography projection images were concatenated onto a single image as an input to a 2D-CNN model [30], and achieved an surprising 99% accuracy in detecting 263 aneurysms.
Classification
Classify Different Tumors and Subtypes
Tumor classification is an essential step to help guide the treatment decision. For decades, the potential for improved classification through various machine learning techniques has been investigated using linear discrimination analysis, support vector machines, decision trees and random forest, radiomics, and shallow neural networks [31]. Today, the automatic classification capability of deep learning methods is getting much attention, and several studies have shown its potential in brain tumor patients. In particular, a new field called Radiomics has been rapidly adopted in the assessment of CNS malignancy. Radiomics is a translational field of research aiming to extract quantitative patterns and interpixel relationships from medical images, that will allow analysis of complex, high-dimensional, quantitative information embedded within the images. Radiomics is often coupled with ML or AI techniques to process the massive amount of data, which typically outperform traditional statistical methods (Fig. 58.23).
Radiomics workflow to classify benign vs. malignant brain masses. A set of features are extracted from the input images and used for training. Various machine learning algorithms are used to classify the images based on the feature vectors. Performance of the ML models is graded according the labels supplied as ground truth
Radiomics with ML is a promising tool for differentiating malignancy from benign tumors, glioblastomas from metastases [32], and classifying metastases by their primary malignancies [33]. Besides using structural features, functional imaging features may also be helpful to classify tumor types. ADC maps, dynamic contrast enhanced permeability maps (K-trans, Kep, Vp, Ve), and dynamic susceptibility contrast perfusion maps (rCBV, rCBF) can be used to differentiate glioblastomas, CNS lymphomas, and metastases [34]. Most studies report similar performance to human reviewers.
Molecular profiling of brain tumors has improved prognosis prediction [35], and is increasingly used in many types of malignancies. Determination of subtypes is most definitive by tissue sampling. Radiogenomics machine-learning is emerging as potential noninvasive alternative to identify surrogate biomarkers that can reflect tumor genomics. For instance, there are at least 4 biologically distinct subgroups identified in medulloblastoma-sonic hedgehog [SHH], wingless-type [WNT], group 3, and group 4, each with prognostic and therapeutic differences. WNT tumors confer more favorable outcomes and better survival. Using MRI–derived radiomic features (such as intensity-based histograms, tumor edge-sharpness, Gabor features, and local area integral invariant features) fed into an SVM, researchers were able to classify SHH, group 3, and group 4 tumors with good accuracy (AUC = 0.79, 0.70, and 0.83, respectively). WNT tumors posed more of a challenge, with AUC ranging from 0.55 to 0.63 [36].
Classify Different Types of Dementia
Besides brain tumors, extensive efforts have been also made to use ML to classify stages along the spectrum of Alzheimer’s disease. Using the ADNI dataset, combined features from MRI and PET were able to distinguish normal control (NC), mild cognitive impairment converters (MCI-C), mild cognitive impairment nonconverters (MCI-NC), and Alzheimer’s disease. A multilevel stacked deep polynomial network was used to classify patients into different binary groups (i.e., AD versus healthy control [NC], or mild cognitive impairment converters [MCI-C] versus nonconverters [MCI-nonconverters]). For distinguishing patients with AD from NCs, they achieved an impressive AUC of 0.97. A lower AUC of 0.80 for predicting MCI converters from nonconverters demonstrated that this is a more difficult task [37]. The flexibility of NNs also allows combination of images with nonimaging data as input. Another study combined similar imaging features with CSF data in the ADNI dataset using a deep-weighted sparse multitask learning framework to improve classification, achieving 95% accuracy in differentiating patients with AD from NCs. Again, multiclass classification was more challenging, achieving an accuracy of 63% for 3 classes (AD, NC, and MCI) and 54% for 4 classes (AD, NC, MCI-C, and MCI-nonconverter) [38].
Segmentation
One of the key advantages of AI-based radiology is the prospect of automatization and standardization of repeated measurements, which is best exemplified by detection and segmentation of lesions. AI-based segmentation is helpful for monitoring disease progression, treatment planning, and volumetric measurements.
Stepping up from detecting the presence of aneurysms, several studies attempted to segment aneurysms using deep learning [39, 40]. Park A et al. [41] proposed a 3D CNN with encoder-decoder architecture to segment the intracranial aneurysms on CTA. Similar to UNet, the model contains skip connections to transmit output directly from the encoder to the decoder. When the model was available to assist the clinicians, their mean sensitivity increased by 0.059 (95% CI, 0.028–0.091; adjusted P = 0.01), mean accuracy increased by 0.038 (95% CI, 0.014–0.062; adjusted P = 0.02), and mean interrater agreement (Fleiss κ) increased by 0.060, from 0.799 to 0.859 (adjusted P = 0.05). Similar performance was achieved in 3D TOF MRA [42] and in DSA [40] with a Dice score coefficient above 0.9.
For optimal management of patients with brain cancer, delineation of initial tumor volume and especially volume change following disease progression or therapy are key neuroradiological tasks. The Response Assessment in Neuro-Oncology (RANO) work group formulated guidelines for assessing treatment response based on size measurements [43]. Several AI approaches have been developed for automatic detection and segmentation of brain tumors [44, 45]. This development is in part attributed to the publicly available Brain Tumor Segmentation (BraTS) dataset [46], and deep learning has shown high potential in detecting and segmenting primary brain tumors in this dataset [47, 48]. Similar AI approaches have been used to segment brain metastases, which may be more challenging due to their size and multiplicity [49,50,51]. Accurate segmentation in addition to segmentation is important because of the high value of stereotactic radiosurgery to treat these lesions. Various neural network architectures were used, including residual networks [52], dense networks [53], U-Nets [54] and V-Nets [55], Pyramid Scene Parsing Nets [56], Feature Pyramid Networks [57], GoogLeNet [58], and the DeepLab_v3 [59]. The latter architecture is currently considered one of the most robust neural networks for image-based semantic segmentation, which represents classification at the image pixel level. The key difference of the DeepLab_v3 approach compared with other architectures is its reliance on atrous (or dilated) convolutions. Consequently, this network has a very large receptive field, thereby incorporating greater spatial context. Such approach may be key for enabling networks to identify local features as well as global contexts, i.e., identifying brain regions, which could enhance the network’s decision-making process on similar local features. Figure 58.24 shows a flowchart of a deep learning segmentation tool based on the DeepLab_v3 architecture.
Diagram showing a DeepLab v3-based segmentation network. In this example, four distinct MR sequences that are commonly used in clinical practice serve as model input; post-Gd T1-weighted inversion recovery prepped fast spoiled gradient-echo (IR-FSPGR), pre- and post-Gd T1-weighted spin echo, and T2-weighted FLAIR imaging. Five contiguous axial slices of each of the four sequences are concatenated in the color-channel dimension to create an input tensor. This tensor is fed into a DeepLab v3-based network to predict the segmentation on the center slice
Stereotactic radiosurgery is also used for treating arteriovenous malformations (AVMs) Traditionally, the lesions are manually segmented for treatment preparation. A 3D V-Net was designed to segment AVMs on postcontrast CT to guide stereotactic radiosurgery. V-Net is a specialized CNN, derived from U-Net, for volumetric (3-D) medical image segmentation. Similar to U-Net, it consists of a contracting (downsampling) path and an expanding (upsampling) path, with skip connections to preserve localization information. More extensive downsampling and upsampling occurs in V-Net, which is accomplished by dividing the contracting path into several stages, each comprising of several 3-D convolutional layers. Whereas U-Net uses max pooling, V-Net uses convolutions for both reducing the resolution and for extracting the most important features, making V-Net more memory efficient. Using manual segmentation by experts as gold standard, the Dice score coefficient of the V-Net model was 0.85 [60].
Prediction
Accurate prediction of outcome is helpful for treatment decisions, especially in the era of “personalized medicine.” Classic prediction methods have been super-dated by ML algorithms which are capable of discovering more complex relations between variables and multivariate interactions.
Prediction in Acute Ischemic Stroke
Many different deep learning models have been used to predict the clinical outcome in acute stroke, such as modified Rankin Scale at 3 months, treatment outcome (good reperfusion), adverse complications (such as hemorrhagic transformation) [61], cognitive performance [62], and hemorrhagic transformation after thrombolysis [63].
As the window for treatment and treatment options for acute stroke broadens, careful selection of appropriate patients is crucial for successful outcomes. Clinical trials using cutoff thresholds of imaging parameters have identified thresholds for ADC (<620 × 10−6 mm2/s) and Tmax (>6 s) as definitions of infarct core and penumbra, respectively. The most common method to select patients for therapy is based on time from presentation (i.e., last seen normal) and penumbra to infarct ratio [64,65,66]. Newer ML models have been built to predict final infarct volume on MRI [67, 68]. Using patients with large vessel occlusion from the Imaging Collaterals in Acute Stroke (iCAS) study and the Diffusion Weighted Imaging Evaluation for Understanding Stroke Evolution Study-2 (DEFUSE-2), a UNET model has been shown to accurately predict final infarct lesions from baseline perfusion-weighted and diffusion-weighted imaging (Fig. 58.25). Even though the model was trained without information about reperfusion status, it was able to predict well in patients with either good or poor perfusion, with better performance than clinically available software packages [69]. In patients with major reperfusion, the UNET model outperformed the clinical thresholding method for Dice coefficient and sensitivity. In patients with minimal reperfusion, the UNET model outperformed the clinical thresholding method in specificity and positive predictive value. The ability to accurately predict final infarct volume from baseline imaging alone, can help guide decision-making, in addition to mismatch profile. In another interesting study, a time-resolved deep-learning model using baseline CTP parameters (cerebral blood volume, time-to-drain) was designed to predict the dynamic progression from penumbra to infarct core over time. Using a multiscale U-Net together with a convolutional auto-encoder, the evolution of the ischemic tissue can be estimated by interpolation [70].
(a) Patient with minimal reperfusion 0% at 24-h. (b) Patient with major reperfusion, 100% at 24-h. Baseline images (DWI, ADC, Tmax, MTT, CBV, CBF) were inputs. Final infarct lesion at 3 to 7 days served as ground truth for the model. The red solid line on the T2-weighted fluid-attenuated inversion recovery images outlines infarct lesions at 3 to 7 days. Numbers after predicted volume (mL) indicate Dice score coefficients. CBF indicates cerebral blood flow; CBV, cerebral blood volume; DSC, Dice score coefficient; DWI, diffusion-weighted imaging; MTT, mean transit time; and Tmax, time to maximum of the residue function. (Reprinted from “Use of Deep Learning to Predict Final Ischemic Stroke Lesions From Initial Magnetic Resonance Imaging”, JAMA Netw Open. 2020;3(3):e200772, Yu et al., with permission under the terms of the CC-BY license, which permits unrestricted use, distribution, and reproduction in any medium)
Other models have incorporated clinical data with imaging data to predict the final outcome. In one study, the addition of clinical data (National Institutes of Health Stroke Scale, age, sex, and time from symptom onset) mildly improved the AUC from 0.85 (imaging data from CT perfusion only) to 0.87 [71]. A novel application that took advantage of the flexibility of NN is demonstrated in a study that trained separate models to predict the outcome based on the treatment strategy. A CNN (CNN+tPA) was trained with patients treated with intravenous recombinant tissue-type plasminogen activator (rtPA) and a separate CNN was trained with patients without rtPA (CNN−tPA). For each test subject, the models would predict the final infarct core if rtPA was administered or withheld, and the treatment effect of rtPA can be estimated by the difference the predicted final infarct core [72]. This study illustrates the potential of using DL to provide recommendations for personalized treatment plans.
Predict Aneurysm Rupture Risk and Outcome
Treatment decisions need to be made for unruptured small aneurysms and SAH patients with multiple aneurysms. Studies have applied machine learning algorithms to predict the outcomes of unruptured aneurysm [73,74,75,76,77,78,79]. Morphological features extracted from DSA can be used for aneurysm stratification [74]. Flatness was the found to be the most important morphological determinant to predict stability of aneurysm; unstable aneurysms were more irregular. Hypertension could influence the morphology of unstable aneurysms [74]. Another study using CNNs to predict rupture risk of small aneurysms (<7 mm diameter) on rotational DSA outperformed human predictions [75].
Predicting complications, such as delayed cerebral ischemia and functional outcome, after aneurysmal rupture could provide guidance for patient care. Efforts have been made to predict delayed cerebral ischemia from a combination of clinical and imaging data with various machine learning algorithms, with modest accuracy [80].
Predict Conversion of MCI to AD
In addition to early diagnosis, ability to predict disease progression can be helpful to patients with debilitating disease such as dementia. Mild cognitive impairment (MCI), which is the clinical precursor of Alzheimer’s disease (AD), has a broadly heterogeneous spectrum with variable rate of progression. Some patients with MCI remain stable over time, while others progress gradually to AD, with approximately 10% to 15% of MCI patients converting to AD each year [81]. Many ML models have been built to predict this conversion. Using data from the Alzheimer’s Disease Neuroimaging Initiative (ADNI), deep learning models based on imaging combined with demographics, neuropsychological (including cognitive assessment, AD assessment scale, memory evaluations), and APOe4 genetic data were studied to predict MCI to AD conversion within 3 years [82]. One such model was able to distinguish the MCI-converters from those with stable MCI with an AUC of 0.925, 86% accuracy, 87.5% sensitivity, and 85% specificity. The model also distinguished patients with AD from healthy controls with 100% accuracy.
Improving Image Quality
There are innovative applications of DL to improve the image quality, reduce acquisition time, and improve the robustness of some advanced CT and MRI techniques.
Image Improvement and Synthesis
For instance, DL can convert images with low-resolution into high-resolution [83], simulate 7 T MR images from data acquired at 3 T [84], and generate normal-dose CT from simulated low-dose CT [85]. By acquiring paired arterial spin-labeling (ASL) CBF images with 2 and 30 min of acquisition time, deep network has been shown to boost the SNR of ASL significantly [86] (Fig. 58.26).
An example of improving the SNR of arterial spin-labeling MR imaging using deep learning. The model is trained using low-SNR ASL images acquired with only a single repetition, while the reference image is a high-SNR ASL image acquired with multiple repetitions (in this case, 6 repetitions). Proton-density-weighted images (acquired routinely as part of the ASL scans for quantitation) and T2-weighted images are also used as inputs to the model to improve performance. The results of passing the low-SNR ASL image through the model are shown on the right, a synthetic image with improved SNR. In this example, the root-mean-squared error (RSME) between the reference image and the synthetic image compared with the original image is reduced nearly three-fold, from 29.3% to 10.8%. Reprinted from “Deep Learning in Neuroradiology”, AJNR Am J Neuroradiol. 2018;39(10):1776–1784, Zaharchuk et al., with permission from WILLIAMS & WILKINS CO.; American Society of Neuroradiology
DL has also been used to create images with different contrast or with features of different modalities, for instance, using DL to generate T1-weighted images from T2-weighted images, or vice versa [87]. The superior soft tissue contrast offered by MRI and the desire to reduce unnecessary radiation dose, makes is attractive to generate synthetic CT from MR images. Synthetic CT has been used to replace CT for radiation therapy [88] and for PET/MR attenuation correction [89].
Dose Reduction and Virtual Contrast Enhancement
The recent concerns over gadolinium deposition in the brain from gadolinium-based contrast agents administration have inspired innovative DL methods to reduce their usage and dosage (Fig. 58.27). Using images acquired with 100% full-dose (0.1 mmol/kg) of gadobenate dimeglumine as target, a DL model was trained to generate full-dose images from 10% low-dose (0.01 mmol/kg) images [90]. Subjects were patients with a variety of pathologies, including gliomas. Compared to the low-dose images, the synthesized full-dose postcontrast images yielded higher image quality with significant improvements (>5 dB PSNR gains and >11.0% improvements in a measure of visual similarity known as the structural similarity index metric [SSIM]). Compared to true full-dose images, the synthesized full-dose images had slightly better motion-artifact suppression, with a nonsignificant reduction in image quality (P = 0.083) and contrast enhancement (P = 0.068).
Another group took this approach to the extreme and used a DL model to predict contrast enhancement from noncontrast MRI images in three groups of subjects: normal subjects, patients with enhancing brain tumors, and patients with nonenhancing brain tumors [91]. Compared with ground truth contrast-enhanced T1-weighted imaging quantitatively, the virtual contrast enhancement yielded a sensitivity of 91.8% and a specificity of 91.2%, AUC of 0.969, a peak signal-to-noise ratio of 23 ± 1 dB, and an SSIM of 0.872 ± 0.031. Qualitatively, the virtual contrast maps for gliomas are blurrier and show less nodular-like ring enhancement, with some false-positive enhancements of nonenhancing gliomas. The ability to synthesize images from ultra-low gadolinium dose, while preserving diagnostic quality, is highly desirable for patients who need imaging repeatedly. These studies show that this is a promising avenue of research using DL.
Dose reduction is also beneficial for positron emission tomography (PET) imaging, which inherently has high radiation exposure. DL has been used to synthesize high-quality virtual 18F-fluorodeoxyglucose (FDG) PET images from low-dose FDG-PET images and the concurrent MR images. A fully convolutional encoder–decoder was trained with low-dose PET images, with 200-fold dose reduction, constructed through undersampling of standard-dose PET images. Both quantitatively and visually, the denoised ultra-low-dose PET images reconstructed with only 0.5% of the standard dose, deliver similar visual quality and diagnostic information as the standard-dose PET images. The addition of MRI images further enhanced the quality of the synthesized images [92]. Another study using a different method to simulate low-dose FDG-PET images achieved similar satisfactory results. Instead of subsampling, low-dose PET images were obtained by acquiring images over a short duration of 3 min (with standard-dose tracer) and the full-dose PET images acquired over the full duration of 12 min served as ground truth. The shorter acquisition time has the additional advantages of reducing motion artifact and improves the efficiency of PET imaging [93].
Besides FDG, DL was also able to reduce radiotracer requirements for amyloid (fluorine 18 [18F]–florbetaben) PET/MRI imaging without sacrificing diagnostic quality [94]. Subsampling one hundredth of the full-dose PET data was used to simulate a low-dose (1%) acquisition to train a CNN model. The synthesized images showed marked improvement on all quality metrics (peak signal-to-noise ratio, SSIM, and root mean square error) compared with the low-dose image. The accuracy for determining amyloid uptake status was high (89%) and similar to intrareader reproducibility of full-dose images (91%). By overcoming the obstacles of high radiation dose, long scan time, and lower SNR, DL is making high quality ultra-low-dose PET images a foreseeable reality.
Reconstruction from Subsampled Diffusion-Weighted Imaging
Neurite orientation dispersion and density imaging (NODDI) is a diffusion-weighted imaging method using models to characterize microstructure of white matter and neurite properties in the brain. NODDI can disentangle crossing fibers and estimate the fiber orientation distribution function (ODF) in each voxel [95]. Similar to DTI, NODDI requires lengthy acquisitions of many (near a hundred) diffusion-weighted images with multiple b-values and orientations [96]. A NN was trained to reconstruct fractional anisotropy and mean diffusivity maps from a small subsets of acquired DTI data, using only 3 to 20 diffusion-encoding directions. The accuracy and precision in DTI reconstruction achieved by the NN was higher than that by conventional reconstructions. The model also performed well in tumor delineation from reconstruction using only three diffusion-encoding directions [97]. A similar DL approach was used to predict tissue property maps, such as neurite dispersion, from subsampled diffusion acquisitions with as few as 8 to 12 diffusion-weighted scans to achieve 12-fold acceleration [98]. With appropriate training in patients, these networks provide clinically meaningful information about tissue microstructure in acute stroke [99] (Fig. 58.28). Fiber tractography can also be improved by directly predicting the fiber ODF in each voxel from undersampled DWI scans with CNNs. Compared with standard acquisitions that use hundreds of gradient directions, the network generates accurate ODFs from as few as 15 gradient directions [100] or 25 DWI scans [101]. The CNNs outperform standard methods in challenging voxels that contain two or even three fiber directions, because they leverage information about the spatial continuity of neighboring voxels in the input data.
Neurite orientation dispersion and density imaging (NODDI) and generalized fractional anisotropy (GFA) parameter maps. Slices showing asymmetries in the brain due to stroke in three participants from the test dataset. Both ODI and GFA parameter maps are displayed for the fully sampled reference images (ref column) as well as the proposed 2D CNN generated images using a dataset undersampled to 24 directions (CNN column). Red arrows highlight the visible asymmetries. (Reprinted from “Simultaneous NODDI and GFA parameter map generation from subsampled q-space imaging using deep learning. Magn Reson Med. 2019; 81: 2399–2411. Gibbons et al., with permission from John Wiley and Sons)
Improve Image Quality in Quantitative Susceptibility Mapping (QSM)
Quantitative susceptibility mapping (QSM) reconstructs tissue magnetic susceptibility in the brain from gradient echo phase MRI and has clinical applications in aging [102] and neurodegeneration [103]. Gold-standard QSM reconstruction requires multiple phase measurements at several tilted head orientations [104]. Deep learning has been used to predict high quality QSM maps from a single orientation phase MRI scan. Models such as QSMnet [105] and DeepQSM [106] have adopted a 3D UNET to generate QSM maps with higher quality and better accuracy than state-of-the-art single orientation methods. This improved performance is evident in higher peak signal-to-noise ratios and reduced normalized root mean squared error, as well as the visible reduction of streak artifacts that contaminate many single-orientation QSM maps. Deep-learning QSM reconstructions take only seconds and are well suited to visualize focal areas of susceptibility abnormalities, e.g., in multiple sclerosis lesions and hemorrhage [105], with high structural similarity to the reference standard (Fig. 58.29).
Quantitative susceptibility mapping using a deep neural network: QSMnet. QSM maps from MEDI and QSMnet are compared for a patient with microbleed (a: red boxes, b: red arrows), a patient with multiple sclerosis lesions (c: blue boxes, d: blue arrows), a patient with large hemorrhage (e: yellow arrows), microbleed (e: red boxes, g: red arrows) and calcification (e: pink boxes, h: pink arrows), and a healthy volunteer with calcification (i: pink boxes, j: pink arrows). The lesions are similarly delineated in both MEDI and QSMnet maps. In f, strong streaking artifacts are observed only in MEDI (green arrows). Note that no lesions except for calcification were observed in the healthy volunteers and, therefore, were trained in QSMnet. (Reprinted from “Quantitative susceptibility mapping using deep neural network: QSMnet, NeuroImage, Volume 179, 2018, Pages 199–206, Yoon et al., with permission from Elsevier)
The accuracy of the final QSM map also depends on preprocessing steps such as receive coil combination and background phase removal. Streamlined pipelines mitigate error propagation from preprocessing by performing multiple necessary steps in a single optimization [107]. Alternatively, CNNs such as SHARQnet have been trained on tens of thousands of synthetic background field examples to accurately remove background phase signal from susceptibility sources with various geometric shapes [108]. General adversarial networks (GAN) have also been used for QSM reconstruction, where the generator network aims to create realistic QSM maps and the discriminator learns to distinguish real and generated images [109]. The GAN architecture reduces residual blurring in the output QSM maps compared to other CNNs and is robust to imperfections in preprocessing steps if the model is trained on high-quality input data.
Reduce Acquisition Time: Magnetic Resonance Fingerprinting
Magnetic resonance fingerprinting (MRF) is a new scanning approach that uses pseudo-random acquisitions (e.g., variable flip angles and repetition times) to obtain unique signal time courses for different tissues [110]. These tissue signatures are then matched to a dictionary of time courses to retrieve multiple corresponding tissue parameters (e.g., quantitative T1 and T2) from a single, rapid scan. Quantitative relaxation parameters offer new insight into subtle pathologies such as differentiating active from inactive lesions in epilepsy [111]. Despite its relative efficiency, MRF requires storage of large dictionaries with over 10,000 entries for matching and is still lengthy to acquire at higher spatial resolutions with whole-brain coverage.
Machine learning methods have been combined with traditional undersampling strategies (e.g., parallel imaging) to further increase the acquisition speed of MRF. The designed CNNs, trained on simulated and actual data, take an input MRF time series and output quantitative T1 and T2 maps. The network parameters are a compact representation of the MRF dictionary, and the CNN inference procedure is 300 to 5000 times faster than typical dictionary matching methods [112, 113]. Combined with parallel imaging, deep learning enables whole-brain T1 and T2 mapping with high spatial resolution (1 mm3 isotropic), in as few as 7 min [114]. This scan time is even faster than conventional T1- or T2-weighted scans at the same resolution.
Challenges Ahead
As powerful as ML algorithms can be, one issue they face is bias, as these algorithms are only as good as the data we feed them. The generalization of trained ML algorithms beyond what they have “seen” in the training data is critical for their increased use. There has been much discussion about this in the computer science field, and is quite important in the field of radiology as well [115]. (See Table 58.1 for key literature in the field of artificial intelligence [17, 36, 50, 69, 91, 99]). The algorithms, once trained, will be representative of the training data and, if trained properly, will perform well on testing data originating from the same distribution as the training data. However, they might perform poorly when applied on data coming from different data sources or patient populations. Discrepant results were also reported with data collected across different scanner models. This and the requirements for well-curated multicenter data for model training are all challenges to overcome before the widespread use of ML-based methods becomes a reality in the clinic.
Another obstacle is the lack of interpretability of the algorithms [116]. With deep neural networks in particular, there is little insight into the inner workings of the models; for instance, they may work well for tumor segmentation and prediction, but precisely how they accomplish these feats are still unclear. The black-box nature of the deep learning algorithms contrast the current radiologic management of patients, where the decision making process is ideally more transparent and traceable. There may also be strong legal and ethical arguments against a decision support system that is based on nontraceable logic. Consequently, there is a need to improve the interpretability of these hidden algorithm structures, which also represents a key step toward accepting this new technology in a routine clinical setting. In order to successfully apply AI tools in a clinical setting, interpretable or explainable solutions would ideally be available for the diagnosis, classification, and response evaluation of patients.
This black-box problem has led to a field of research called “eXplainable AI” or XAI, representing a new set of techniques that attempts to provide an understanding of how input and output data relates to each other. As an example, deep learning models can be made “visible” by introducing decision trees (model regularization) during training. Having regularized models allowing clinical users to step through the inner processes behind the networks’ predictions would represent a key step toward improving interpretability. One approach is to combine deep learning with the novel concept of tree regularization [116], which may have major advantages compared to standard regularization in that it returns a decision tree that best mimics the predictions of the AI-model.
Summary
We are living in the period of the artificial intelligence revolution. AI is rapidly infiltrating and transforming radiology. The small sampling in this chapter highlighted some of the potential directions that can be taken with AI. While it has been speculated that AI will replace human radiologists entirely, it is hard to predict if and when that may happen. AI can advance our diagnostic prowess and refine management decisions. Indeed, AI is a tool to be embraced rather than feared. Working together with well-trained radiologists, AI offers the potential to improve our ability to serve our patients more effectively and more efficiently, with the ultimate goal of alleviating neurological disease.
References
McCarthy J, Minsky ML, Rochester N, Shannon CE. A proposal for the Dartmouth summer research project on artificial intelligence 1955. Available from http://jmc.stanford.edu/articles/dartmouth/dartmouth.pdf.
Esteva A, Robicquet A, Ramsundar B, Kuleshov V, DePristo M, Chou K, et al. A guide to deep learning in healthcare. Nat Med. 2019;25(1):24–9.
Uribe CF, Mathotaarachchi S, Gaudet V, Smith KC, Rosa-Neto P, Benard F, et al. Machine learning in nuclear medicine: part 1-introduction. J Nucl Med. 2019;60(4):451–8.
Zaharchuk G, Gong E, Wintermark M, Rubin D, Langlotz CP. Deep learning in neuroradiology. AJNR Am J Neuroradiol. 2018;39(10):1776–84.
Hopfield JJ. Neural networks and physical systems with emergent collective computational abilities. Proc Natl Acad Sci U S A. 1982;79(8):2554–8.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521(7553):436–44.
Krizhevsky A, Sutskever I, Hinton GE. Imagenet classification with deep convolutional neural networks. In: Neural information processing systems. Nevada: Lake Tahoe; 2012.
President EOot. Big data: Seizing opportunities, preserving values 2014. Available from: https://bigdatawg.nist.gov/pdf/big_data_privacy_report_may_1_2014.pdf.
Ting DSW, Pasquale LR, Peng L, Campbell JP, Lee AY, Raman R, et al. Artificial intelligence and deep learning in ophthalmology. Br J Ophthalmol. 2019;103(2):167–75.
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115–8.
Larson DB, Chen MC, Lungren MP, Halabi SS, Stence NV, Langlotz CP. Performance of a deep-learning neural network model in assessing skeletal maturity on pediatric hand radiographs. Radiology. 2018;287(1):313–22.
Janowczyk A, Madabhushi A. Deep learning for digital pathology image analysis: a comprehensive tutorial with selected use cases. J Pathol Inform. 2016;7:29.
Dice LR. Measures of the amount of ecologic association between species. Ecology. 1945;26(3):297–302.
Goodfellow IJ, Pouget-Abadie J, Mirza M, Xu B, Warde-Farley D, Ozair S, et al. Generative adversarial nets. Adv Neur. 2014;27
Allen B Jr, Seltzer SE, Langlotz CP, Dreyer KP, Summers RM, Petrick N, et al. A road map for translational research on artificial intelligence in medical imaging: From the 2018 national institutes of health/rsna/acr/the academy workshop. J Am Coll Radiol. 2019;16(9 Pt A):1179–89.
Bland J, Mehranian A, Belzunce MA, Ellis S, McGinnity CJ, Hammers A, et al. MR-guided kernel EM reconstruction for reduced dose pet imaging. IEEE Trans Radiat Plasma Med Sci. 2018;2(3):235–43.
Chen KT, Gong E, de Carvalho Macruz FB, Xu J, Boumis A, Khalighi M, et al. Ultra-low-dose (18)f-florbetaben amyloid pet imaging using deep learning with multi-contrast MRI inputs. Radiology. 2019;290(3):649–56.
Xie Y, Jiang B, Gong E, Li Y, Zhu G, Michel P, et al. Journal club: use of gradient boosting machine learning to predict patient outcome in acute ischemic stroke on the basis of imaging, demographic, and clinical information. AJR Am J Roentgenol. 2019;212(1):44–51.
Chilamkurthy S, Ghosh R, Tanamala S, Biviji M, Campeau NG, Venugopal VK, et al. Deep learning algorithms for detection of critical findings in head CT scans: a retrospective study. Lancet. 2018;392(10162):2388–96.
Phong TD, Duong HN, Nguyen HT, Trong NT, Nguyen VH, Hoa TV, et al. Brain hemorrhage diagnosis by using deep learning. In: Proceedings of the 2017 International Conference on Machine Learning and Soft Computing. Ho Chi Minh City, Vietnam: Association for Computing Machinery; 2017. p. 34–9.
Murray NM, Unberath M, Hager GD, Hui FK. Artificial intelligence to diagnose ischemic stroke and identify large vessel occlusions: a systematic review. J Neurointerv Surg. 2020;12(2):156–64.
Guberina N, Dietrich U, Radbruch A, Goebel J, Deuschl C, Ringelstein A, et al. Detection of early infarction signs with machine learning-based diagnosis by means of the Alberta stroke program early CT score (aspects) in the clinical routine. Neuroradiology. 2018;60(9):889–901.
Herweh C, Ringleb PA, Rauch G, Gerry S, Behrens L, Mohlenbruch M, et al. Performance of e-aspects software in comparison to that of stroke physicians on assessing CT scans of acute ischemic stroke patients. Int J Stroke. 2016;11(4):438–45.
Nagel S, Sinha D, Day D, Reith W, Chapot R, Papanagiotou P, et al. E-aspects software is non-inferior to neuroradiologists in applying the aspect score to computed tomography scans of acute ischemic stroke patients. Int J Stroke. 2017;12(6):615–22.
Grunwald IQ, Kulikovski J, Reith W, Gerry S, Namias R, Politi M, et al. Collateral automation for triage in stroke: evaluating automated scoring of collaterals in acute stroke on computed tomography scans. Cerebrovasc Dis. 2019;47(5–6):217–22.
Nakao T, Hanaoka S, Nomura Y, Sato I, Nemoto M, Miki S, et al. Deep neural network-based computer-assisted detection of cerebral aneurysms in MR angiography. J Magn Reson Imaging. 2018;47(4):948–53.
Sichtermann T, Faron A, Sijben R, Teichert N, Freiherr J, Wiesmann M. Deep learning-based detection of intracranial aneurysms in 3d tof-mra. AJNR Am J Neuroradiol. 2019;40(1):25–32.
Duan H, Huang Y, Liu L, Dai H, Chen L, Zhou L. Automatic detection on intracranial aneurysm from digital subtraction angiography with cascade convolutional neural networks. Biomed Eng Online. 2019;18(1):110.
Ueda D, Yamamoto A, Nishimori M, Shimono T, Doishita S, Shimazaki A, et al. Deep learning for mr angiography: automated detection of cerebral aneurysms. Radiology. 2019;290(1):187–94.
Zeng Y, Liu X, Xiao N, Li Y, Jiang Y, Feng J, et al. Automatic diagnosis based on spatial information fusion feature for intracranial aneurysm. IEEE Trans Med Imaging. 2020;39(5):1448–58.
Tandel GS, Biswas M, Kakde OG, Tiwari A, Suri HS, Turk M, et al. A review on a deep learning perspective in brain cancer classification. Cancers (Basel). 2019;11(1):111.
Mohsen H, El-Dahshan E-SA, El-Horbaty E-SM, Salem A-BM. Classification using deep learning neural networks for brain tumors. Future Comput Inform J. 2018;3:68e71.
Kniep HC, Madesta F, Schneider T, Hanning U, Schonfeld MH, Schon G, et al. Radiomics of brain MRI: utility in prediction of metastatic tumor type. Radiology. 2019;290(2):479–87.
Swinburne NC, Schefflein J, Sakai Y, Oermann EK, Titano JJ, Chen I, et al. Machine learning for semi-automated classification of glioblastoma, brain metastasis and central nervous system lymphoma using magnetic resonance advanced imaging. Ann Transl Med. 2019;7(11):232.
Park SH, Won J, Kim SI, Lee Y, Park CK, Kim SK, et al. Molecular testing of brain tumor. J Pathol Transl Med. 2017;51(3):205–23.
Iv M, Zhou M, Shpanskaya K, Perreault S, Wang Z, Tranvinh E, et al. Mr imaging-based radiomic signatures of distinct molecular subgroups of medulloblastoma. AJNR Am J Neuroradiol. 2019;40(1):154–61.
Shi J, Zheng X, Li Y, Zhang Q, Ying S. Multimodal neuroimaging feature learning with multimodal stacked deep polynomial networks for diagnosis of Alzheimer's disease. IEEE J Biomed Health Inform. 2018;22(1):173–83.
Suk HI, Lee SW, Shen D. Alzheimer's disease neuroimaging I. Hierarchical feature representation and multimodal fusion with deep learning for ad/mci diagnosis. NeuroImage. 2014;101:569–82.
Stember JN, Chang P, Stember DM, Liu M, Grinband J, Filippi CG, et al. Convolutional neural networks for the detection and measurement of cerebral aneurysms on magnetic resonance angiography. J Digit Imaging. 2019;32(5):808–15.
Podgorsak AR, Rava RA, Shiraz Bhurwani MM, Chandra AR, Davies JM, Siddiqui AH, et al. Automatic radiomic feature extraction using deep learning for angiographic parametric imaging of intracranial aneurysms. J Neurointerv Surg. 2020;12(4):417–21.
Park A, Chute C, Rajpurkar P, Lou J, Ball RL, Shpanskaya K, et al. Deep learning-assisted diagnosis of cerebral aneurysms using the HeadXNet model. JAMA Netw Open. 2019;2(6):e195600.
Faron A, Sichtermann T, Teichert N, Luetkens JA, Keulers A, Nikoubashman O, et al. Performance of a deep-learning neural network to detect intracranial aneurysms from 3d tof-mra compared to human readers. Clin Neuroradiol. 2020;30(3):591–8.
Lin NU, Lee EQ, Aoyama H, Barani IJ, Barboriak DP, Baumert BG, et al. Response assessment criteria for brain metastases: proposal from the rano group. Lancet Oncol. 2015;16(6):e270–8.
Akkus Z, Galimzianova A, Hoogi A, Rubin DL, Erickson BJ. Deep learning for brain MRI segmentation: state of the art and future directions. J Digit Imaging. 2017;30(4):449–59.
Işın A, Direkoğlu C, Şah M. Review of mri-based brain tumor image segmentation using deep learning methods. Procedia Comput Sci. 2016;102:317–24.
Menze BH, Jakab A, Bauer S, Kalpathy-Cramer J, Farahani K, Kirby J, et al. The multimodal brain tumor image segmentation benchmark (brats). IEEE Trans Med Imaging. 2015;34(10):1993–2024.
Zhao L, Jia K, Deep feature learning with discrimination mechanism for brain tumor segmentation and diagnosis. 2015 International Conference on Intelligent Information Hiding and Multimedia Signal Processing (IIH-MSP); 2015 23–25 Sept. 2015.
Pereira S, Pinto A, Alves V, Silva CA. Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans Med Imaging. 2016;35:1240–51.
Charron O, Lallement A, Jarnet D, Noblet V, Clavier JB, Meyer P. Automatic detection and segmentation of brain metastases on multimodal mr images with a deep convolutional neural network. Comput Biol Med. 2018;95:43–54.
Grovik E, Yi D, Iv M, Tong E, Rubin D, Zaharchuk G. Deep learning enables automatic detection and segmentation of brain metastases on multisequence mri. J Magn Reson Imaging. 2020;51(1):175–82.
Liu Y, Stojadinovic S, Hrycushko B, Wardak Z, Lau S, Lu W, et al. A deep convolutional neural network-based automatic delineation strategy for multiple brain metastases stereotactic radiosurgery. PLoS One. 2017;12(10):e0185844.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. IEEE Conf Comput Vision Pattern Recogn. 2016;2016:770–8.
Huang YH, Feng QJ. Segmentation of brain tumor on magnetic resonance images using 3d full-convolutional densely connected convolutional networks. Nan Fang Yi Ke Da Xue Xue Bao. 2018;38(6):661–8.
Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation. ArXiv. 2015:2015;abs/1505.04597.
Milletari F, Navab N, Ahmadi S-A. V-net: fully convolutional neural networks for volumetric medical image segmentation. In: 2016 Fourth International Conference on 3D Vision (3DV); 2016. p. 565–71.
Zhao H, Shi J, Qi X, Wang X, Jia J. Pyramid scene parsing network. IEEE Conf Comput Vision Pattern Recogn. 2017;2017:6230–9.
Lin T-Y, Dollár P, Girshick R, He K, Hariharan B, Belongie S. Feature pyramid networks for object detection. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2017. p. 936–44.
Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, et al. Going deeper with convolutions. IEEE Conf Comput Vision Pattern Recogn. 2015;2015:1–9.
Chen LC, Papandreou G, Kokkinos I, Murphy K, Yuille AL. DeepLab: semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected CRFs. IEEE Trans Pattern Anal Mach Intell. 2018;40(4):834–48. https://doi.org/10.1109/TPAMI.2017.2699184. Epub 2017 Apr 27. PMID: 28463186.
Wang T, Lei Y, Tian S, Jiang X, Zhou J, Liu T, et al. Learning-based automatic segmentation of arteriovenous malformations on contrast CT images in brain stereotactic radiosurgery. Med Phys. 2019;46(7):3133–41.
van Os HJA, Ramos LA, Hilbert A, van Leeuwen M, van Walderveen MAA, Kruyt ND, et al. Predicting outcome of endovascular treatment for acute ischemic stroke: potential value of machine learning algorithms. Front Neurol. 2018;9:784.
Chauhan S, Vig L, De Grazia DF, M, Corbetta M, Ahmad S, Zorzi M. A comparison of shallow and deep learning methods for predicting cognitive performance of stroke patients from MRI lesion images. Front Neuroinform. 2019;13:53.
Bentley P, Ganesalingam J, Carlton Jones AL, Mahady K, Epton S, Rinne P, et al. Prediction of stroke thrombolysis outcome using CT brain machine learning. Neuroimage Clin. 2014;4:635–40.
Austein F, Riedel C, Kerby T, Meyne J, Binder A, Lindner T, et al. Comparison of perfusion CT software to predict the final infarct volume after thrombectomy. Stroke. 2016;47(9):2311–7.
Straka M, Albers GW, Bammer R. Real-time diffusion-perfusion mismatch analysis in acute stroke. J Magn Reson Imaging. 2010;32(5):1024–37.
Ho KC, Scalzo F, Sarma KV, Speier W, El-Saden S, Arnold C. Predicting ischemic stroke tissue fate using a deep convolutional neural network on source magnetic resonance perfusion images. J Med Imaging (Bellingham). 2019;6(2):026001.
Giacalone M, Rasti P, Debs N, Frindel C, Cho TH, Grenier E, et al. Local spatio-temporal encoding of raw perfusion MRI for the prediction of final lesion in stroke. Med Image Anal. 2018;50:117–26.
Livne M, Boldsen JK, Mikkelsen IK, Fiebach JB, Sobesky J, Mouridsen K. Boosted tree model reforms multimodal magnetic resonance imaging infarct prediction in acute stroke. Stroke. 2018;49(4):912–8.
Yu Y, Xie Y, Thamm T, Gong E, Ouyang J, Huang C, et al. Use of deep learning to predict final ischemic stroke lesions from initial magnetic resonance imaging. JAMA Netw Open. 2020;3(3):e200772-e.
Lucas C, Kemmling A, Bouteldja N, Aulmann LF, Madany Mamlouk A, Heinrich MP. Learning to predict ischemic stroke growth on acute CT perfusion data by interpolating low-dimensional shape representations. Front Neurol. 2018;9:989.
Kasasbeh AS, Christensen S, Parsons MW, Campbell B, Albers GW, Lansberg MG. Artificial neural network computer tomography perfusion prediction of ischemic core. Stroke. 2019;50(6):1578–81.
Nielsen A, Hansen MB, Tietze A, Mouridsen K. Prediction of tissue outcome and assessment of treatment effect in acute ischemic stroke using deep learning. Stroke. 2018;49(6):1394–401.
Zhao X, Gold N, Fang Y, Xu S, Zhang Y, Liu J, et al. Vertebral artery fusiform aneurysm geometry in predicting rupture risk. R Soc Open Sci. 2018;5(10):180780.
Liu Q, Jiang P, Jiang Y, Ge H, Li S, Jin H, et al. Prediction of aneurysm stability using a machine learning model based on pyradiomics-derived morphological features. Stroke. 2019;50(9):2314–21.
Kim HC, Rhim JK, Ahn JH, Park JJ, Moon JU, Hong EP, et al. Machine learning application for rupture risk assessment in small-sized intracranial aneurysm. J Clin Med. 2019;8(5)
Paliwal N, Jaiswal P, Tutino VM, Shallwani H, Davies JM, Siddiqui AH, et al. Outcome prediction of intracranial aneurysm treatment by flow diverters using machine learning. Neurosurg Focus. 2018;45(5):E7.
Liu J, Chen Y, Lan L, Lin B, Chen W, Wang M, et al. Prediction of rupture risk in anterior communicating artery aneurysms with a feed-forward artificial neural network. Eur Radiol. 2018;28(8):3268–75.
Varble N, Tutino VM, Yu J, Sonig A, Siddiqui AH, Davies JM, et al. Shared and distinct rupture discriminants of small and large intracranial aneurysms. Stroke. 2018;49(4):856–64.
Detmer FJ, Luckehe D, Mut F, Slawski M, Hirsch S, Bijlenga P, et al. Comparison of statistical learning approaches for cerebral aneurysm rupture assessment. Int J Comput Assist Radiol Surg. 2020;15(1):141–50.
Ramos LA, van der Steen WE, Sales Barros R, Majoie C, van den Berg R, Verbaan D, et al. Machine learning improves prediction of delayed cerebral ischemia in patients with subarachnoid hemorrhage. J Neurointerv Surg. 2019;11(5):497–502.
Mitchell AJ, Shiri-Feshki M. Temporal trends in the long term risk of progression of mild cognitive impairment: a pooled analysis. J Neurol Neurosurg Psychiatry. 2008;79(12):1386–91.
Spasov S, Passamonti L, Duggento A, Lio P, Toschi N. Alzheimer's disease neuroimaging I. a parameter-efficient deep learning approach to predict conversion from mild cognitive impairment to Alzheimer's disease. NeuroImage. 2019;189:276–87.
Lei Y, Xu D, Zhou Z, Higgins K, Dong X, Liu T, et al. High-resolution CT image retrieval using sparse convolutional neural network. Proc SPIE int soc. Opt Eng. 2018;10573
Bahrami K, Shi F, Zong X, Shin HW, An H, Shen D. Reconstruction of 7t-like images from 3t mri. IEEE Trans Med Imaging. 2016;35(9):2085–97.
Chen H, Zhang Y, Kalra MK, Lin F, Chen Y, Liao P, et al. Low-dose CT with a residual encoder-decoder convolutional neural network. IEEE Trans Med Imaging. 2017;36(12):2524–35.
Gong E, Guo J, Liu J, Fan A, Pauly J, Zaharchuk G. Deep learning and multi-contrast-based denoising for low-snr arterial spin labeling (ASL) mri: SPIE. 2020.
Vemulapalli RNH, Zhou S. Deep networks and mutual information maximization for cross-modal medical image synthesis. In: Zhou S, Greenspan H, Shen D, editors. Deep learning for medical image analysis. London: Elsevier; 2017.
Han X. Mr-based synthetic CT generation using a deep convolutional neural network method. Med Phys. 2017;44(4):1408–19.
Liu F, Jang H, Kijowski R, Bradshaw T, McMillan AB. Deep learning mr imaging-based attenuation correction for pet/mr imaging. Radiology. 2018;286(2):676–84.
Gong E, Pauly JM, Wintermark M, Zaharchuk G. Deep learning enables reduced gadolinium dose for contrast-enhanced brain mri. J Magn Reson Imaging. 2018;48(2):330–40.
Kleesiek J, Morshuis JN, Isensee F, Deike-Hofmann K, Paech D, Kickingereder P, et al. Can virtual contrast enhancement in brain MRI replace gadolinium?: A feasibility study. Investig Radiol. 2019;54(10):653–60.
Xu J, Gong E, Ouyang J, Pauly J, Zaharchuk G. Ultra-low-dose 18f-fdg brain pet/mr denoising using deep learning and multi-contrast information: SPIE. 2020.
Xiang L, Qiao Y, Nie D, An L, Lin W, Wang Q, et al. Deep auto-context convolutional neural networks for standard-dose pet image estimation from low-dose pet/mri. Neurocomputing. 2017;267:406–16.
Chen KT, Gong E, Macruz FBC, Xu J, Boumis A, Khalighi M, et al. Ultra–low-dose 18f-florbetaben amyloid pet imaging using deep learning with multi-contrast MRI inputs. Radiology. 2019;290(3):649–56.
Wedeen VJ, Wang RP, Schmahmann JD, Benner T, Tseng WY, Dai G, et al. Diffusion spectrum magnetic resonance imaging (DSI) tractography of crossing fibers. NeuroImage. 2008;41(4):1267–77.
Zhang H, Schneider T, Wheeler-Kingshott CA, Alexander DC. Noddi: practical in vivo neurite orientation dispersion and density imaging of the human brain. NeuroImage. 2012;61(4):1000–16.
Aliotta E, Nourzadeh H, Sanders J, Muller D, Ennis DB. Highly accelerated, model-free diffusion tensor MRI reconstruction using neural networks. Med Phys. 2019;46(4):1581–91.
Golkov V, Dosovitskiy A, Sperl JI, Menzel MI, Czisch M, Samann P, et al. Q-space deep learning: twelve-fold shorter and model-free diffusion MRI scans. IEEE Trans Med Imaging. 2016;35(5):1344–51.
Gibbons EK, Hodgson KK, Chaudhari AS, Richards LG, Majersik JJ, Adluru G, et al. Simultaneous NODDI and GFA parameter map generation from subsampled q-space imaging using deep learning. Magn Reson Med. 2019;81(4):2399–411.
Koppers S, Haarburger C, Merhof D. Diffusion MRI signal augmentation: From single shell to multi shell with deep learning: Math Vis. 2017:61–70.
Lin Z, Gong T, Wang K, Li Z, He H, Tong Q, et al. Fast learning of fiber orientation distribution function for mr tractography using convolutional neural network. Med Phys. 2019;46(7):3101–16.
Bilgic B, Pfefferbaum A, Rohlfing T, Sullivan EV, Adalsteinsson E. MRI estimates of brain iron concentration in normal aging using quantitative susceptibility mapping. NeuroImage. 2012;59(3):2625–35.
Acosta-Cabronero J, Williams GB, Cardenas-Blanco A, Arnold RJ, Lupson V, Nestor PJ. In vivo quantitative susceptibility mapping (QSM) in Alzheimer's disease. PLoS One. 2013;8(11):e81093.
Liu T, Spincemaille P, de Rochefort L, Kressler B, Wang Y. Calculation of susceptibility through multiple orientation sampling (cosmos): a method for conditioning the inverse problem from measured magnetic field map to susceptibility source image in mri. Magn Reson Med. 2009;61(1):196–204.
Yoon J, Gong E, Chatnuntawech I, Bilgic B, Lee J, Jung W, et al. Quantitative susceptibility mapping using deep neural network: QSMnet. NeuroImage. 2018;179:199–206.
Bollmann S, Rasmussen KGB, Kristensen M, Blendal RG, Ostergaard LR, Plocharski M, et al. DeepQSM—using deep learning to solve the dipole inversion for quantitative susceptibility mapping. NeuroImage. 2019;195:373–83.
Chatnuntawech I, McDaniel P, Cauley SF, Gagoski BA, Langkammer C, Martin A, et al. Single-step quantitative susceptibility mapping with variational penalties. NMR Biomed. 2017;30(4)
Bollmann S, Kristensen MH, Larsen MS, Olsen MV, Pedersen MJ, Ostergaard LR, et al. Sharqnet—sophisticated harmonic artifact reduction in quantitative susceptibility mapping using a deep convolutional neural network. Z Med Phys. 2019;29(2):139–49.
Chen Y, Jakary A, Avadiappan S, Hess CP, Lupo JM. Qsmgan: improved quantitative susceptibility mapping using 3d generative adversarial networks with increased receptive field. NeuroImage. 2019;116389
Ma D, Gulani V, Seiberlich N, Liu K, Sunshine JL, Duerk JL, et al. Magnetic resonance fingerprinting. Nature. 2013;495(7440):187–92.
Ma D, Jones SE, Deshmane A, Sakaie K, Pierre EY, Larvie M, et al. Development of high-resolution 3d mr fingerprinting for detection and characterization of epileptic lesions. J Magn Reson Imaging. 2019;49(5):1333–46.
Cohen O, Zhu B, Rosen MS. Mr fingerprinting deep reconstruction network (drone). Magn Reson Med. 2018;80(3):885–94.
Fang Z, Chen Y, Liu M, Zhan Y, Lin W, Shen D. Deep learning for fast and spatially-constrained tissue quantification from highly-undersampled data in magnetic resonance fingerprinting (mrf). Mach Learn Med Imaging. 2018;11046:398–405.
Chen Y, Fang Z, Hung SC, Chang WT, Shen D, Lin W. High-resolution 3d mr fingerprinting using parallel imaging and deep learning. NeuroImage. 2019;116329
Zou J, Schiebinger L. Ai can be sexist and racist—it’s time to make it fair. Nature. 2018;559:324–6.
Wu M, Hughes MC, Parbhoo S, Zazzi M, Roth V, Doshi-Velez F. Beyond sparsity: tree regularization of deep models for interpretability. In Proceedings of the Thirty-Second AAAI Conference on Artificial Intelligence and Thirtieth Innovative Applications of Artificial Intelligence Conference and Eighth AAAI Symposium on Educational Advances in Artificial Intelligence (AAAI’18/IAAI’18/EAAI’18). AAAI Press, Article 204, 2018;1670–8.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Tong, E. et al. (2023). CNS Machine Learning. In: Faro, S.H., Mohamed, F.B. (eds) Functional Neuroradiology. Springer, Cham. https://doi.org/10.1007/978-3-031-10909-6_58
Download citation
DOI: https://doi.org/10.1007/978-3-031-10909-6_58
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-10908-9
Online ISBN: 978-3-031-10909-6
eBook Packages: MedicineMedicine (R0)