Abstract
Aflatoxins are highly toxic contaminants synthesized by several toxigenic strains of Aspergillus as secondary metabolites. The biosynthesis of aflatoxins is a complicated process involving a chain of reactions which are catalyzed by various enzymes encoded by genes present on aflatoxin cluster. The genetic variations among different fungal strains can impact the final compound being produced. This chapter focuses on the biosynthetic pathway for aflatoxin production starting from acetate and finishing with the production of aflatoxin. The role of various genes and their encoded enzymes at every reaction has been described. Furthermore, the impact of different factors such as light, temperature, water activity, oxidative stress, carbon sources, nitrogen sources, and pH on aflatoxin biosynthesis has also been described. AFB1 is the most common and toxic aflatoxin being consumed through various sources. Inside the human or animal body, AFB1 is metabolized to different forms, making it either highly toxic or less toxic depending on the metabolism channel. The metabolism of AFB1 has also been covered in this chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
2.1 Biosynthesis
Biosynthesis is an enzyme-catalyzed, multistep process involving the conversion of substrates into highly complex compounds within living organisms. During this process, the simpler compounds are modified either through conversion into other compounds or through joining together in order to form macromolecules. The biosynthesis of metabolites is carried out through certain metabolic pathways. Aflatoxins are synthesized by certain fungal species as secondary metabolites through a chain of reactions. The study regarding the biosynthesis of secondary metabolites generally involves the identification of reactions and their sequence through which the cells convert the primary metabolites into the final molecule. Additionally, the features regulating these processes are also identified. Even though a great variation may exist in chemical structures of metabolites, the biosynthesis of most secondary microbial metabolites may occur only through a certain number of biosynthetic pathways.
The biosynthetic pathway elucidation is a multistep procedure which may involve the identification of the primary metabolite from which the final molecule is being made and isolation of intermediate compounds formed along the pathway in order to hypothesize the sequence of reactions based on their chemical structures. The identification of enzymes involved at each point during the biosynthetic pathway and their isolation is also important in the understanding of this process. Furthermore, in order to completely understand the biosynthetic process, identification of regulatory factors and biosynthetic genes is also important.
2.2 Aflatoxin Biosynthesis
A complicated biosynthetic pathway consisting of at least 27 reactions catalyzed by different enzymes is involved in the production of aflatoxins (Roze et al. 2013; Yu 2012). The genes containing the codes for enzymes involved in the AF biosynthetic pathway are grouped in a cluster. The expression of these genes is controlled by two cluster-specific regulatory genes, namely, aflR and aflS (Chang 2003; Price et al. 2006). Additionally, the synthesis of AFs can also be influenced by environmental stimuli such as oxidative stress, nutrient sources, pH, and light which may initiate complex mechanisms through activation of various cell signaling pathways, hence modifying the expression of the genes playing a role in the production of toxin (Affeldt et al. 2014; Klich 2007; Montibus et al. 2015).
The DNA information in aspergilli is structured in eight chromosomes, and the genes resulting in AF synthesis are situated in the 54th cluster, 75-kb region of the fungal genome on chromosome III, 80 kb from the telomere of chromosome 3 (Georgianna and Payne 2009). This cluster consists of 30 genes (Fig. 2.1) and is mainly controlled by the regulatory genes aflR and aflS (Chang 2003; Price et al. 2006). The gene cluster involved in AF production has been extensively analyzed in A. flavus as well as in A. parasiticus. The studies have revealed the homology of the clustered genes among these species to be between 90 and 99% (Yu et al. 2000b). Due to the presence of these changes, one of the major differences existing among A. flavus and A. parasiticus is their ability to synthesize B- and G-type aflatoxins. The A. flavus majorly synthesizes aflatoxins B1 and B2, while A. parasiticus possesses the ability to produce both B and G types of aflatoxin. The functional genes responsible for the synthesis of G-type aflatoxin include aflF, aflU, and nadA which, respectively, encode for an aryl alcohol dehydrogenase, a cytochrome P450 monooxygenase, and an oxidase (Ehrlich et al. 2004; Ehrlich et al. 2008). Furthermore, the elucidation of the aflatoxin-producing cluster has also been aided by studies on A. nidulans which has the capability to produce sterigmatocystin, an intermediate metabolite produced along the AF biosynthetic pathway. The A. parasiticus and A. nidulans exhibit the homology between 55 and 75%.
Gene cluster for aflatoxin biosynthesis. This figure is adopted from the work of Yu (2012)
Birch (1967) first proposed the formation of aflatoxins through a polyketide pathway, and extensive studies have shown the involvement of at least 27 enzymatic reactions till now. Moreover, extensive studies have shown that along with other enzymes, cytochrome P450 also exhibits an important part in the production of aflatoxin. These enzymes are responsible for the attachment of functional groups (i.e., acetyl, methyl) during AF biosynthesis (Nelson 2011). Among the biosynthetic pathways for the production of mycotoxins known to date, the aflatoxin gene cluster has the highest number of cytochrome P450 (Roze et al. 2015). Thus owing to the number of oxidative changes involved, AF biosynthesis is among the longest and most complex processes (Minto and Townsend 1997).
2.3 Aflatoxin Biosynthetic Pathway
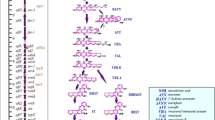
Aflatoxin biosynthetic pathway comprises of cascade of reactions catalyzed by various enzymes (Fig. 2.2).
Aflatoxin biosynthesis pathway. This figure is adopted from work of Gacem and El Hadj-Khelil (2016)
2.3.1 Acetate to Norsolorinic Acid
Aflatoxins are the polyketide derivative furanocoumarin compounds. The biosynthesis of these substances require the hexanoate starter components (from acetyl-CoA and malonyl-CoA) which act as an initial substrate in a series of reactions ultimately leading to the formation of aflatoxins (Roze et al. 2013). The first step in the biosynthetic pathway for aflatoxins involves the formation of norsolorinic acid (NOR). The NOR is the first stable compound among the aflatoxin precursors (Bennett et al. 1971). The formation of polyketide from a hexanoyl unit is assisted by a couple of fatty acid synthase enzymes and a polyketide synthase enzyme.
The enzymes catalyzing these changes are encoded majorly by four genes. The aflA and aflB were formerly known as fas-2 and fas-1, respectively, because of their involvement in encoding fatty acid synthases. The aflA and aflB synthesize α and β protein subunits which are considered to play an important role in the formation of a polyketide structure from hexanoate units (Roze et al. 2013; Yu et al. 2004a). The gene aflC, also known as pksA, contains the coding regarding the synthesis of polyketide skeletons. The polyketide synthase is involved in the chain elongation that occurs in most of the secondary metabolites that are acetate derivatives. This enzyme has been found to further convert the polyketide structure into norsolorinic acid anthrone (NAA) (Roze et al. 2013). Furthermore, another gene, hypC, is known to encode noranthrone oxidase, a 17-kDa enzyme that has been demonstrated to be involved in the catalytic transition of NAA to NOR (Ehrlich and Yu 2009). The hypC is located in the region between aflC and nor-1. The nor-1 gene, due to its participation in aflatoxin biosynthesis, is also known as aflD.
2.3.2 Norsolorinic Acid to Averantin
The norsolorinic acid is further converted into averantin (AVN). The conversion of NOR into AVN is directed by the aflD (nor-1) gene. The cloning of the aflD gene was achieved through genetic complementation. The aflD gene encodes a ketoreductase that is required in order to convert the 1′-keto group of NOR to the 1′-hydroxyl group of AVN (Zhou and Linz 1999). Earlier it was predicted that the aflE (norA) and aflF (norB) genes present on the AF cluster were associated with this step. The short-chain aryl alcohol dehydrogenases, which may have the tendency to further assist the conversion of NOR to AVN, are encoded by these genes (Cary et al. 1996). However, they have also been shown to take part in other steps involved in the series of reactions catalyzed by enzymes during AFB1 biosynthesis.
2.3.3 Averantin to 5'-Hydroxyaverantin
The earliest proof revealing the conversion of Averantin (AVN) to 5′-hydroxyaverantin (HAVN) was established through the experiments involving radioisotope incorporation (Bennett et al. 1980). The studies showed that in A. parasiticus, the transformation of AVN to averufin (AVF) is accomplished by two enzymatic reactions among which the first reaction involves the conversion of polyketide anthraquinone averantin into HAVN through hydroxylation. This reaction is catalyzed by P450 monooxygenase enzymes (Yabe et al. 1991). The gene ord-1, encoding the P450 monooxygenase, was cloned and disrupted by Yu et al. (1997). The studies of ord-1 mutant in the presence of a substrate proved that HAVN is an intermediate compound formed during the transformation of AVN to AVF. The ord-1 gene that exhibits a high degree of similarity in sequence to A. nidulans stcf (Brown et al. 1996) was renamed as aflG (avnA).
2.3.4 5′-Hydroxyaverantin to Averufin
Averufin (AVF) is among key intermediary compounds formed during the biosynthetic formation of aflatoxins through many studies (Keller et al. 2000). Furthermore, initially the involvement of several metabolites were reported during the transformation of AVN to AVF (Bhatnagar et al. 1992). However, the later studies negated the involvement of some of these metabolites as an intermediate in aflatoxin formation. One of such metabolites was averufanin (AVNN) which proved to be a shunt metabolite instead of being an intermediate in aflatoxin biosynthetic pathway (Sakuno et al. 2003; Yabe and Nakajima 2004).
An alcohol dehydrogenase-encoding gene cluster, aflH (adhA), was characterized by Chang et al. (2000) in A. parasiticus. The experiments showed that adhA deletion mutants resulted in predominant accumulation of HAVN. However, after prolonged growth periods, the mutant strains showed production of AVNN in minor quantities which was observed to be a shunt metabolite. Hence, the transformation of HAVN might be directly into the AVF or indirectly, as a result of the actions of additional cytosolic enzymes. Two cytosolic enzymes along with 5′-oxoaverantin (OAVN), a novel AF intermediate, were described during further studies by Sakuno et al. (2003). Among the series of intermediates formed during aflatoxin biosynthesis, OAVN was observed to be formed during the transformation of HAVN to AVF. The aflH (adhA) gene is responsible for encoding alcohol dehydrogenase enzyme that is involved in catalytic conversion of HAVN to OAVN. However, it was observed that the adhA deletion mutant was leaky, revealing the possible involvement of other genes or enzymes during transformation of OAVN to AVF. The study by Sakuno et al. (2005) showed the association of aflK (vbs) gene with the transformation of OAVN to AVF. Although initially the aflK gene was linked with the transformation of a versiconal compound into versicolorin B only, this was the first time described that the same enzyme can be involved in the catalysis of two reactions during AFB1 biosynthesis. It was further hypothesized that this might have happened as a result of evolution in the gene cluster of AFB1.
2.3.5 Averufin to Versiconal Hemiacetal Acetate
The oxidation of averufin (AVF) transforms it into versiconal hemiacetal acetate (VHA). VHA is known as a precursor to aflatoxin (Fitzell et al. 1977). The gene aflI (avfA) has been shown to take part in the biotransformation of AVF to VHA. Yu et al. (2000b) revealed that when AVF-accumulating mutant, A. parasiticus SRRC 165, was complemented with aflI (avfA) gene from A. flavus, it restored the ability of strain transform AVF to VHA consequently producing aflatoxins, hence confirming the role of aflI (avfA) in this process. However, the precise role of aflI in the AVF oxidation has still not been completely clarified, though it has been commonly projected that the aflI encoded enzyme is involved in the catalysis of the ring closure step during the production of hydroxyversicolorone (HVN).
Furthermore, the gene aflV-encoded enzyme has been shown to be responsible for catalyzing the process of AVF transformation to HVN and the aflW gene product to be involved in conversion of HVN to VHA through a Baeyer-Villiger reaction. The gene aflV (cypX) is involved in encoding P450 microsomal monooxygenase, and aflW (moxY) has been found to encode a cytosolic monooxygenase (Wen et al. 2005).
2.3.6 Versiconal Hemiacetal Acetate to Versiconal
The subsequent phase after the formation of VHA includes its transformation into a versiconal (VAL). Various studies have revealed the involvement of an esterase in this conversion (Kusumoto and Hsieh 1996). The esterase enzyme was identified to have been encoded by aflJ (estA) (Yu et al. 2003). Furthermore, the esterase enzyme has also been isolated from A. parasiticus (Hsieh 1989; Kusumoto and Hsieh 1996). Additionally, the aflJ deletion mutants of A. parasiticus showed the accumulation of some metabolites such as versicolorin A (VERA) and VHA (Chang et al. 2004). In addition, trace amounts of versiconoc acetate (VOAc) along with other downstream metabolites in aflatoxin biosynthetic pathway including VAL and versicolorin B were also accumulated. The enzyme esterase is also known to exhibit involvement in the reversible transformation of VHA to VOAc. Furthermore, a study by Chang et al. (2004) confirmed the participation of esterase encoded by aflJ in transformation of VHA to VAL and VOAc to VOH during the biosynthesis of aflatoxins.
2.3.7 Versiconal to Versicolorin B
The conversion of VAL/VHOH to VERB was identified to have been catalyzed by a cyclase enzyme, named versicolorin B synthase (McGuire et al. 1996; Silva and Townsend 1997). This gene was cloned and named vbs (Silva et al. 1996). It was also observed during these studies that the recombinant proteins of the vbs gene exhibited the cyclase activity. The vbs gene was renamed as aflK due to its involvement in aflatoxin biosynthesis reaction cascade (Yu et al. 2004b). This enzyme is also involved in the conversion of OAVN into AVF as mentioned earlier. Furthermore, the closure of the bisfuran ring is also catalyzed by the aflK-encoded enzyme. The bisfuran ring is known to be responsible for the toxic character of aflatoxins as it binds with DNA after metabolization (Yu et al. 2004b).
2.3.8 Versicolorin B to Versicolorin a
The subsequent transformations of VERB are critical in determining the type of aflatoxin going to be synthesized. The VERB structure contains the tetrahydrofuran ring that is similar to the one present in AFB2/AFG2, hence forming the AFB2/AFG2 as a final product of aflatoxin biosynthetic pathway. Contrarily, the transformation of VERB to versicolorin A (VERA) leads to the formation of AFB1 or AFG1 eventually, as these toxins contain a dihydrobisfuran ring like VERA. The transformation of VERB to VERA involves the desaturation of the bisfuran ring (Yabe et al. 1993). It was identified that the stcL-disrupted A. nidulans did not synthesize sterigmatocystin (ST) compounds, consequently resulting in VERB accumulation (Kelkar et al. 1997). Furthermore, aflL (verB), a homologue of stcL in A. parasiticus and A. flavus, is considered to be involved in the conversion of VERB to VERA as it encodes a cytochrome P450 monooxygenase/desaturase (Kelkar et al. 1997).
2.3.9 Versicolorin a to Demethylsterigmatocystin and Versicolorin B to Dihydrodemethylsterigmatocystin
The biosynthetic pathway of aflatoxins involves the conversion of VERA to demethylsterigmatocystin (DMST) resulting in the formation of AFB1 or AFG1. Additionally, the transformation of VERB to dihydrodemethylsterigmatocystin (DHDMST) eventually leads to formation of AFB2 or AFG2. Henry and Townsend (2005) have described the changes occurring during this chain of reactions in detail.
Four genes have been observed to take part in the VERA to DMST conversion. The ver-1 also known as aflM is a ketoreductase encoding gene that is similar to nor-1. The gene aflM was observed to be involved in the transformation of VERA to an intermediate compound which has still not been isolated. The homologous gene for aflM was also identified in A. nidulans as stcU. Furthermore, another gene, aflN (verA) responsible for coding a cytochrome P450-type monooxygenase, was shown to catalyze the transformation of VERA to another intermediate before converting into DMST. The aflN homologue in A. nidulans has also been identified as stcS (Yu et al. 2004a, b). Additionally, the disruption of stcU and stcS resulted in accumulation of VERA, hence confirming their requirement in the transformation of VERA to DMST ultimately (Keller et al. 1995). However, their exact function remains to be identified. The third gene involved in the transformation of VERA, aflY (hypA), is considered to encode a Baeyer-Villiger monooxygenase which appears to act as a mediator between two hypothetical structures during the transformation of VERA to DMST. The disruption of aflY in A. parasiticus resulted in accumulation of VERA which may suggest that it is involved as a part of the enzyme complex without permitting the development of intermediate compounds. The gene aflX (ordB) is further responsible for coding an oxidoreductase that is involved in the catalysis of oxidative decarboxylation and ring closure of the intermediate formed after aflY-catalyzed oxidation.
2.3.10 Demethylsterigmatocystin to Sterigmatocystin and Dihydrodemethylsterigmatocystin to Dihydrosterigmatocystin
The involvement of O-methyltransferases in aflatoxin biosynthesis was confirmed after studies on purified enzymes. It was revealed that O-methyltransferase I is involved in the catalysis of methyl transfer from S-adenosylmethionine (SAM) to the hydroxyls of DHDMST as well as DMST in order to produce DHST and ST, respectively. The O-methyltransferase I is a 43-kDa enzyme that has been isolated from A. parasiticus. The gene corresponding to this enzyme, dmtA, was isolated from A. flavus, A. parasiticus, and A. sojae and was later named aflO (omtB) (for O-methyltransferase B) (Yu et al. 2000b). The stcP gene in A. nidulans was identified to be homologous to the aflO gene. Furthermore, the disruption of aflO caused failure of DMST transformation to ST.
2.3.11 Sterigmatocystin to O-Methylsterigmatocystin and DHST to Dihydro-O-Methylsterigmatocystin
The O-methyltransferase catalyzes the transformation of ST to OMST and DHST to DHOMST. The gene containing the code for O-methyltransferase was first cloned using A. parasiticus through reverse genetics by producing antibodies against the O-methyltransferase isolated from A. parasiticus (Keller et al. 1993). The gene responsible for these transformations, aflP (omtA), was formerly named as omt-1 and then omtA (Yu et al. 1993). The enzyme O-methyltransferase A is substrate specific; hence it cannot methylate DMST or DHDMST. In addition to A. parasiticus, the genomic sequence of aflP was cloned from A. flavus as well (Yu et al. 1995). Furthermore, the homologue for aflP was also identified in different other species of Aspergillus, either aflatoxigenic or non-aflatoxigenic (Klich et al. 1995). However, the orthologue of aflP is absent in A. nidulans which can explain the absence of aflatoxin as an end product and the presence of ST in this particular specie.
2.3.12 Formation of Aflatoxin B and Aflatoxin G
During the final conversions of OMST into AFB1, various genes are involved including aflQ, hypB, aflE, and hypE. The roles of aflQ and hypB have been accurately defined in the final transformations; however partial roles have been reported for various other genes involved in AF biosynthesis. The gene responsible for encoding a P450 monooxygenase, aflQ (ordA), is present adjacent to aflP in the aflatoxin cluster. The expression of this gene ultimately leads to the transformation of OMST into aflatoxin B1 or G1 and DHOMST into aflatoxin B2 or G2 (Ehrlich 2009). The aflQ transforms OMST into 11-hydroxy-O-methylsterigmatocystin (HOMST), a precursor of AFB1 (Zeng et al. 2011). Nevertheless, it is still not clear whether two successive reactions involving monooxygenase are catalyzed by aflQ (ordA) gene product, OrdA, during the later steps of aflatoxin biosynthesis. The hypB gene encodes an oxidase reported to take part in conversion of HOMST into a seven-ring lactone (MW, 370 Da) and is expressed under conditions suitable for aflatoxin production (Ehrlich et al. 2010). This compound is converted into another unknown intermediary product through hydrolytic enzymes which do not belong to the aflatoxin cluster (Ehrlich 2009). The synthesis of G-group aflatoxins has been proposed to involve additional enzyme(s) (Yu et al. 1998). Further studies showed that cytochrome P450 monooxygenase encoded by cypA gene is involved in the formation of G-type aflatoxins (Ehrlich et al. 2004). Furthermore, the nadA gene that was earlier considered as a member of sugar utilization cluster (Yu et al. 2000a) was also shown to be a member of the adjoining aflatoxin cluster through microarray studies and participated in the formation of AFG1/AFG2 (Yu et al. 2011). The disruption of nadA gene showed that a recently observed AF intermediate, NADA (formed after OMST), is converted into G1 type of aflatoxin through a cytosolic enzyme named NadA. Initially, the gene aflE (norA), homologous to aflD in the aflatoxin cluster, was believed to be involved in the conversion of NOR to AVN; however, further studies depicted the involvement of aflE in mainly final two transformations during formation of AFB1. Although the role of aflE in AF biosynthetic pathway has been confirmed as its absence resulted in accumulation of deoxyaflatoxin, its exact position on AF gene cluster is still not known. Furthermore, the hypB was also predicted for its involvement in one of the steps exhibiting oxidation during the transformation of OMST to AFs. The involvement of hypE (aflLa) during the last steps of AFB1 biosynthesis was also proposed because the disruption of hypE led to the production of an intermediary compound prior to the formation of deoxyAFB1 synthesis. The hypE depicts homologies with several bacterial enzymes, and its participation along AF enzymatic pathway was suggested in combination with P450 monooxygenase (Ehrlich 2009). A. flavus is involved in the production of only B1 and B2 aflatoxins, while A. parasiticus is capable of producing aflatoxins B1, B2, G1, and G2. The presence of the intact nadA and norB genes has only been shown in A. parasiticus which is the G-group producer. The data proposes that norB is responsible for encoding another enzyme which is predominantly involved in the formation of AFG1 and AFG2 (Ehrlich et al. 2008).
2.3.13 Formation of Aflatoxin M
The aflatoxins M1 and M2 are the products of AFB1 and AFB2 biologically converted within mammals. These products were initially separated from bovine milk (Garrido et al. 2003). After aflatoxins enter a mammalian (humans or animals) body, the liver cytochrome P450 enzymes metabolize them, consequently converting them to a reactive epoxide intermediate or hydroxylated aflatoxins M1 and M2. The epoxide intermediate is a more toxic compound exhibiting higher carcinogenicity, while aflatoxins M1 and AFM2 are less harmful metabolites of aflatoxins B1 and AFB2, respectively. Some recent studies involving feeding of aspertoxin (12c-hydroxy-OMST) (Yabe et al. 2012) revealed that A. parasiticus also exhibits production of some minor aflatoxins including M1, M2, GM1, and GM2. Furthermore, feeding of O-methylsterigmatocystin (OMST) to A. parasiticus resulted in production of AFM1 and AFGM1 along with AFB1 and AFG1, while feeding with DHOMST resulted in the production of aflatoxins M2 and GM2 in addition to B2 and G2. This revealed that OrdA is responsible for catalyzing the reaction involving 12c-hydroxylation resulting in transformation of OMST to aspertoxin and also the subsequent transformation from aspertoxin to AFM1. In this scenario, the AFB1 is not a precursor of AFM1.
2.4 Factors Affecting Aflatoxin Biosynthesis
The aflatoxin biosynthesis is a complicated process based on a chain of reactions catalyzed by various enzymes coded by different genes. The synthesis of aflatoxins is affected by different nutritional and environmental factors which are discussed below.
2.4.1 Oxidative Stress
The aflatoxins are considered to be biosynthesized by A. parasiticus and A. flavus as a response by their cells against oxidative stress. A study showed that the aflatoxigenic strains of A. parasiticus require more oxygen as compared to non-aflatoxigenic strains during the phase of their active growth (Jayashree and Subramanyam 2000). It was further shown that aflatoxin synthesis in A. parasiticus is prompted by an increase in oxidative stress. The higher oxygen demand may consequently cause an increase in reactive oxygen species (ROS), hence increasing oxidative stress (Walsh et al. 2011). The relation of oxidative stress with aflatoxin synthesis was further confirmed as its alleviation resulted in a decrease in aflatoxin production (Huang et al. 2009). The microarray analysis revealed the downregulation of all the genes of A. flavus involved in AF biosynthesis after treatment with an antioxidant caffeic acid (Kim et al. 2008). Additionally, the toxigenic A. parasiticus possesses higher antioxidant activities by enzymes like superoxide dismutase in comparison to its nontoxigenic forms (Narasaiah et al. 2006). Catalases (CAT), superoxide dismutases (SOD), and glutathione peroxidase (GPX) play critical roles in the defense system of fungal strains against reactive oxygen species (ROS), hence facilitating the cellular defense to cope with oxidative stress. Among these, SOD provides frontline defense through conversion of the free radicals into H2O2 and O2. Afterward, peroxidases and catalases assist the transformation of H2O2 into H2O and/or O2 in to H2O, respectively (Weydert and Cullen 2010). Aflatoxin production is considered to perform as a secondary defense mechanism after the primary mechanism that involves antioxidant enzymes (Hong et al. 2013). Furthermore, the AFB1 biosynthesis is promoted in the availability of acetate units, and the beta-oxidation of fatty acids results in their degradation into acetate units among fungal strains (Maggio-Hall and Keller 2004), hence linking this pathway indirectly to AFB1 production.
2.4.2 Carbon
The synthesis of secondary metabolites is well known for being dependent upon the availability of carbon and its source. Sugars are considered to be a favorable source of carbon aflatoxin biosynthesis as they produce the polyketide starter units (Davis and Diener 1968; Maggio-Hall et al. 2005). Various studies have shown the presence of simple sugars to be related to higher levels of aflatoxin synthesis in A. parasiticus, A. flavus, and A. nidulans (Bhatnagar et al. 2006; Calvo et al. 2002; Liu et al. 2016). In contrast, d-glucal (a glucose derivative), when used as the principle sugar source in the medium, was able to inhibit the aflatoxin production because d-glucal is not metabolized by fungi (Zhang et al. 2014). The carbon source utilization among Aspergillus is mediated by a sugar cluster which contains four genes grouped in a 7.5-kB cluster. This cluster of genes is positioned next to aflatoxin gene cluster in A. flavus and A. parasiticus (Bhatnagar et al. 2006; Yu et al. 2000a). Many genes present in the aflatoxin cluster have CreA-binding sites close to their promoter regions, which may be a basis of forming its relation with regulation of AF production (Georgianna and Payne 2009). CreA is a transcription factor that in combination with genes is involved in the process of carbon catabolic repression (CCR). The aspergilli use CCR as a strategic mechanism in order to preserve energy and regulate the carbon catabolism for using the most favorable carbon source (Deepika et al. 2016; Ruijter and Visser 1997). Furthermore, CreA also plays an important role in several other functions in addition to AFB1 production in A. flavus (Fasoyin et al. 2018).
2.4.3 Nitrogen
Among Aspergillus species, the nitrogen sources are regulated by nitrogen metabolite repression mechanism. The nitrogen sources have been found to affect the synthesis of aflatoxin and ST in various ways (Calvo et al. 2002). In this perspective, AreA plays a critical role as it modulates the genes responsible for utilization of substitute nitrogen sources. Different nitrogen sources may affect the AF production differently as some substrates, such as asparagine, glutamate, and ammonium salts, support the AF synthesis, while some substrates like sodium nitrate and tryptophan do not favor AF production (Yu 2012). It is because the nitrogen source media govern the under- and over-expression of the areA gene, consequently resulting in a higher or lower production levels of aflatoxin (Fasoyin et al. 2019).
2.4.4 pH
The lower pH levels are generally associated with high contents of aflatoxin production in A. flavus. A study showed a tenfold increase in AF production when the pH was reduced to 4.0, while an increase in pH caused a decrease in AF synthesis (Cotty 1988). For A. parasiticus, a decrease in pH level to <6.0 was found to stimulate the AFB synthesis, whereas the higher pH level favored the synthesis of G-type aflatoxin (Buchanan and Ayres 1975). The aflM expression is higher in acidic media as compared to that in neutral or alkaline media. Additionally, the fungal growth results in pH reduction, consequently enhancing AF production with time (Keller et al. 1997).
2.4.5 Light
The stimulus of light exerts a high impact on adaptation as well as survival of fungal strains. Light may have an impact on growth and morphological features of the fungus, consequently affecting the production of secondary metabolites as well (Rangel et al. 2015). The light induces a strong “velvet comlex” in aspergilli that is governed by a global regulator veA that controls a number of genetic elements including photoreceptors (Purschwitz et al. 2008). VeA regulates half of the gene clusters involved in secondary metabolite formation (28 out of the 56), including the AF gene cluster (Cary et al. 2015). The veA is an essential gene for the production of AFB1 in A. flavus (Duran et al. 2009). The null mutants of veA and IaeA did not exhibit aflR expression (Amaike and Keller 2009). Deletion of veA in A. parasiticus resulted in the absence of versicolorin A, an aflatoxin intermediate, hence confirming its role in aflR/aflS expression (Calvo et al. 2004).
2.4.6 Temperature
Temperature is among the major influencing factors affecting aflatoxin synthesis. It is associated with promoting the expression of the structural biosynthesis genes (aflD and aflO). However, it does not induce expression of regulatory genes (aflR and aflS) (Gallo et al. 2016). A study by OBrian et al. (2007) showed that AF biosynthetic genes were expressed more under the temperature of 28 °C as compared to higher temperature of 37 °C, but the aflatoxin pathway regulatory genes, aflR and aflS, did not show difference at these two temperatures. Furthermore, the genes involved in aflatoxin biosynthesis were found to be downregulated when exposed to elevated temperatures of 42 °C (Liu et al. 2017).
2.4.7 Water Activity
The water activity (a w) may also affect aflatoxin biosynthesis. All the genes involved in AFB1 biosynthesis and laeA were found to be inhibited under the a w of 0.99 as compared to the a w of 0.96, while highest expression was observed at a w 0.92 (Liu et al. 2017).
2.5 Metabolism of Aflatoxin B1
The metabolism of aflatoxin B1 occurs through oxidative reactions catalyzed by the members of cytochrome P450 (CYP450) supergene family of isoenzymes. The CYP450 enzyme family is composed of hemoproteins and electron carriers that, during the cellular respiration, catalyze or enhance the oxidation-reduction reactions (Lamb et al. 2009). Earlier, it had been considered that CYP450 specifically originated from the liver, but later studies showed that they are distributed throughout the body (Ding and Kaminsky 2003). However, xenobiotics are mainly metabolized in the liver (Shimada et al. 1994). The isoforms of CYP450 involved in AFB1 metabolism in the body include CYP1A1, CYP1A2, CYP3A4, CYP2Cs, CYP3A5, and CYP3A7 (Shimada et al. 1994). The glutathione S-transferase (GST) and AFB1-aldehyde reductase also catalyze the AFB1 metabolism resulting in the formation of reactive metabolites, among which some can be used as the biomarkers for AF exposure (Bbosa et al. 2013). The metabolites of AFB1 formed by actions of different CYP450 isoenzymes vary in their carcinogenic potential. The toxic impacts of AFB1 are associated with activation as well as detoxification rate at the primary and secondary levels of metabolism (Neal et al. 1987). Furthermore, the fate of aflatoxin B1 metabolism varies among and within humans and animals. In addition, the activation rate of aflatoxins varies among children and adults belonging to the same species as well, consequently affecting their resistance toward AFB1 toxicity (Ramsdell and Eaton 1990). Additionally, AFB1 metabolism varies among humans belonging to different regions of the world. The main pathways involved in AFB1 metabolism include O-dealkylation, ketoreduction, epoxidation, and hydroxylation (Fig. 2.3). These reactions can result in formation of either highly toxic metabolite (AFBO and AFM1) or relatively nontoxic compounds (AFP1, AFQ1, or AFB2a) (Wu et al. 2009).
AFB1 metabolic pathway mediated by CYP450. This figure is adopted from the work of Rushing and Selim (2019)
2.5.1 Aflatoxin B1–8,9-Epoxide
The AFB1 is metabolized to B1–8,9-epoxide (AFBO) through the help of enzyme system P450 in the liver. The AFBO has two isomers, endo-8,9-epoxide and exo-8,9-epoxide (Raney et al. 1992a). The isoenzymes, CYP3A4 and CYP1A2, are primarily responsible for this conversion. The CYP1A2 acts as a primary producer of AFBO when AFB1 is in lower concentrations. In contrast, at higher AFB1 concentrations, CYP3A4 majorly produces AFBO resulting in the formation of exo AFBO isomers only (Ueng et al. 1995). Also, the CYP1A2 produces more exo isomers as compared to CYP3A4 at lower AFB1 levels (Gallagher et al. 1996). This intermediate exhibits extremely electrophilic character, allowing it to instantly react with amines of proteins as well as of nucleic acids. It reacts with DNA and attaches with N7 position of guanine, consequently forming AFB1-N7-guanine adduct imparting AFB1-exo-8,9-epoxide highly carcinogenic character (Johnson and Guengerich 1997).
2.5.2 AFQ1
AFQ1 is a relatively nontoxic metabolite of AFB1 formed through hydroxylation mediated solely by CYP3A4 (Kamdem et al. 2006). It was first observed in monkey liver microsomal preparations exposed to AFB1. Generally, AFQ1 is produced in much higher amounts in comparison to AFM1; however, rat microsomes were observed to produce lower amounts (Masri et al. 1974). It was shown that AFQ1 occurred in humans frequently in amounts ranging from 1 to 11% of initial AFB1 amounts (Yourtee et al. 1987). However, the potential of AFQ1 to bind with DNA is significantly lower as compared to that of AFBO, hence making it a detoxification product of AFB1 in comparison to AFM1 which is toxic in nature (Raney et al. 1992b). Furthermore, another study showed the presence of AFQ1 in levels higher than AFM1 and AFB1-N7-guanine after monitoring the urinary and fecal samples. Moreover, the concentrations were found to be higher in fecal matter in comparison to urine, making it a potential biomarker source for evaluation of AFB1 exposure (Mykkänen et al. 2005). Even though AFQ1 is one of the most abundant metabolites of AFB1, it is seldom used as a biomarker for AFB1 exposure assessment.
2.5.3 AFP1
AFP1 is also a detoxification metabolite of AFB1 produced by hydroxylation through P450 enzymes including CYP2A13, CYP2A3, and CYP321A1 (He et al. 2006; Niu et al. 2008). Studies have shown the presence of this metabolite in urine of individuals exposed to AFB1 and those who had developed hepatocellular carcinoma (HCC) probably as a consequence of AFB1 exposure (Ross et al. 1992).
2.5.4 Aflatoxicol
In contrast to other metabolites of AFB1, aflatoxicol (AFL) is found in cytosolic fractions of liver preparations. The formation of AFL is mediated by NADPH reductase, typically in the cytosol (Partanen et al. 2010). Unlike AFP1 and AFQ1, AFL retains its DNA-binding capacity, consequently retaining its toxic nature. Therefore, it is not considered as a detoxification product of AFB1. Furthermore, the AFL acts as a reservoir for AFB1 as it has a tendency to be converted back to AFB1 though enzyme actions, which further enhances the toxic effects (Partanen et al. 2010). The AFL is the only AFB1 metabolite that can be transferred through the placenta of a human and can be formed from placenta as well. This metabolite has been observed in human urine as well as in breast milk of individuals exposed to AFB1 (Kussak et al. 1998).
2.5.5 AFH1
AFH1 resembles AFL structurally with an additional hydroxyl group at the terminal cyclopentenone ring. Two enzyme systems are involved in the metabolic conversion of AFB1 to AFH1, namely, the microsomal hydroxylase and cytoplasmic reductase. However, it is not clear whether the AFH1 is formed through hydroxylation of AFL or reduction of AFQ1.
2.5.6 AFB2a
AFB2a was initially characterized as a product of AFB1 formed as a result of acid catalysis. The mild acidic conditions promote the addition of waster across the 8,9-double bond to form the hemiacetal ring. This nonenzymatic transformation was observed in acidic media of molds that had been added with AFB1 (Ciegler et al. 1966). It has been shown that AFB2a possesses lower toxicity as compared to AFB1 due to lower DNA-binding capacity, hence making it a detoxification product of AFB1. However, it has a unique tendency to bind with cellular proteins which can contribute to other cellular toxicities. In addition, this binding usually occurs with primary amines in alkaline conditions. The binding may take place on phosphoethanolamine head groups of phospholipids. It is one-of-a-kind structurally characterized aflatoxin-lipid adduct till now (Rushing and Selim 2017).
2.6 Conclusion
Aflatoxins are highly toxic compounds produced by various fungal strains belonging to Aspergillus. These toxins are produced as secondary metabolites through a complex biosynthetic pathway which has been under investigation for decades. The aflatoxin biosynthesis involves a number of reactions catalyzed by different enzymes which consequently produce intermediate compounds, ultimately forming aflatoxins. The aflatoxin formation is regulated by expression of genes at different steps in the chain of reactions. Studies on AF biosynthesis have helped reveal the type of enzymes and the genes responsible for encoding these enzymes taking part in the biosynthesis of aflatoxins. The production of aflatoxin is affected by several factors such as light, temperature, pH, water activity, and nutrient sources. The knowledge about the impact of these factors on gene expression can help propose effective strategies to prevent contamination of food as well as feed commodities with aflatoxins. Additionally, comprehensive understanding about AFB1 metabolism can also help identify different biomarkers in order to assess the exposure of the population to aflatoxins. This can aid in determining an accurate estimate regarding the threat of aflatoxins being faced by populations of different regions around the world.
References
Affeldt KJ, Carrig J, Amare M, Keller NP (2014) Global survey of canonical Aspergillus flavus G protein-coupled receptors. MBio:5
Amaike S, Keller NP (2009) Distinct roles for VeA and LaeA in development and pathogenesis of aspergillus flavus. Eukaryot Cell 8:1051–1060
Bbosa GS, Kitya D, Ogwal-Okeng J (2013) Aflatoxins metabolism, effects on epigenetic mechanisms and their role in carcinogenesis. Health 5. https://hdl.handle.net/123456789/169
Bennett J, Lee L, Shoss S, Boudreaux G (1980) Identification of averantin as an aflatoxin B1 precursor placement in the biosynthetic pathway. Appl Environ Microbiol 39:835–839
Bennett JW, Lee LS, Vinnett C (1971) The correlation of aflatoxin and norsolorinic acid production. J Am Oil Chem Soc 48:368
Bhatnagar D, Cary JW, Ehrlich K, Yu J, Cleveland TE (2006) Understanding the genetics of regulation of aflatoxin production and aspergillus flavus development. Mycopathologia 162:155–166
Bhatnagar D, Ehrlich K, Cleveland T (1992) Oxidation-reduction reactions in biosynthesis of secondary metabolites. Handb Appl Mycol 5:255–286
Birch A (1967) Biosynthesis of polyketides and related compounds. Science 156:202–206
Brown D, Yu J, Kelkar H, Fernandes M, Nesbitt T, Keller N, Adams T, Leonard T (1996) Twenty-five coregulated transcripts define a sterigmatocystin gene cluster in aspergillus nidulans. PNAS 93:1418–1422
Buchanan R, Ayres J (1975) Effect of initial pH on aflatoxin production. Appl Microbiol 30:1050–1051
Calvo AM, Bok J, Brooks W, Keller NP (2004) veA is required for toxin and sclerotial production in aspergillus parasiticus. Appl Environ Microbiol 70:4733–4739
Calvo AM, Wilson RA, Bok JW, Keller NP (2002) Relationship between secondary metabolism and fungal development. Microbiol Mol Biol Rev 66:447–459
Cary J, Han Z, Yin Y, Lohmar J, Shantappa S, Harris-Coward P, Mack B, Ehrlich K, Wei Q, Arroyo-Manzanares N (2015) Transcriptome analysis of aspergillus flavus reveals veA-dependent regulation of secondary metabolite gene clusters including the novel aflavarin cluster. Eukaryot Cell 14:983–997
Cary JW, Wright M, Bhatnagar D, Lee R, Chu FS (1996) Molecular characterization of an aspergillus parasiticus dehydrogenase gene norA located on the aflatoxin biosynthesis gene cluster. Appl Environ Microbiol 62:360–366
Chang PK (2003) The aspergillus parasiticus protein AFLJ interacts with the aflatoxin pathway-specific regulator AFLR. Mol Genet Genomics 268:711–719
Chang PK, Yabe K, Yu J (2004) The aspergillus parasiticus estA-encoded esterase converts versiconal hemiacetal acetate to versiconal and versiconol acetate to versiconol in aflatoxin biosynthesis. Appl Environ Microbiol 70:3593–3599
Chang PK, Yu J, Ehrlich KC, Boue SM, Montalbano BG, Bhatnagar D, Cleveland TE (2000) adhA in aspergillus parasiticus is involved in conversion of 5′-hydroxyaverantin to averufin. Appl Environ Microbiol 66:4715–4719
Ciegler A, Lillehoj E, Peterson R, Hall H (1966) Microbial detoxification of aflatoxin. Appl Microbiol 14:934–939
Cotty P (1988) Aflatoxin and sclerotial production by Aspergillus flavus: influence of pH. Growth 4:11
Davis ND, Diener UL (1968) Growth and aflatoxin production by aspergillus parasiticus from various carbon sources. Appl Microbiol 16:158–159
Deepika V, Murali T, Satyamoorthy K (2016) Modulation of genetic clusters for synthesis of bioactive molecules in fungal endophytes: a review. Microbiol Res (Pavia) 182:125–140
Ding X, Kaminsky LS (2003) Human extrahepatic cytochromes P450: function in xenobiotic metabolism and tissue-selective chemical toxicity in the respiratory and gastrointestinal tracts. Annu Rev Pharmacol Toxicol 43:149–173
Duran R, Cary J, Calvo A (2009) The role of veA in Aspergillus flavus infection of peanut, corn and cotton. Open Mycol J. https://doi.org/10.2174/1874437000903010027
Ehrlich KC (2009) Predicted roles of the uncharacterized clustered genes in aflatoxin biosynthesis. Toxins 1:37–58
Ehrlich KC, Chang PK, Yu J, Cotty PJ (2004) Aflatoxin biosynthesis cluster gene cypA is required for G aflatoxin formation. Appl Environ Microbiol 70:6518–6524
Ehrlich KC, Li P, Scharfenstein L, Chang PK (2010) HypC the anthrone oxidase involved in aflatoxin biosynthesis. Appl Environ Microbiol 76:3374–3377
Ehrlich KC, Scharfenstein LL, Montalbano BG, Chang PK (2008) Are the genes nadA and norB involved in formation of aflatoxin G1? Int J Mol Sci 9:1717–1729
Ehrlich KC, Yu J (2009) Aflatoxin-like gene clusters and how they evolved. In: Mycotoxins in food, feed and bioweapons. Springer, Berlin, Heidelberg
Fasoyin OE, Wang B, Qiu M, Han X, Chung KR, Wang S (2018) Carbon catabolite repression gene creA regulates morphology, aflatoxin biosynthesis and virulence in aspergillus flavus. Fungal Genet Biol 115:41–51
Fasoyin OE, Yang K, Qiu M, Wang B, Wang S, Wang S (2019) Regulation of morphology, aflatoxin production, and virulence of aspergillus flavus by the major nitrogen regulatory gene areA. Toxins 11:718
Fitzell DL, Singh R, Hsieh DP, Motell EL (1977) Nuclear magnetic resonance identification of versiconal hemiacetal acetate as an intermediate in aflatoxin biosynthesis. J Agric Food Chem 25:1193–1197
Gacem MA, El Hadj-Khelil AO (2016) Toxicology, biosynthesis, bio-control of aflatoxin and new methods of detection. Asian Pac J Trop Biomed 6:808–814
Gallagher EP, Kunze KL, Stapleton PL, Eaton DL (1996) The kinetics of aflatoxin B1Oxidation by human cDNA-expressed and human liver microsomal cytochromes P450 1A2 and 3A4. Toxicol Appl Pharmacol 141:595–606
Gallo A, Solfrizzo M, Epifani F, Panzarini G, Perrone G (2016) Effect of temperature and water activity on gene expression and aflatoxin biosynthesis in aspergillus flavus on almond medium. Int J Food Microbiol 217:162–169
Garrido N, Iha M, Santos Ortolani M, Duarte Favaro R (2003) Occurrence of aflatoxins M1 and M2 in milk commercialized in Ribeirão Preto-SP, Brazil. Food Addit Contam A 20:70–73
Georgianna DR, Payne GA (2009) Genetic regulation of aflatoxin biosynthesis: from gene to genome. Fungal Genet Biol 46:113–125
He XY, Tang L, Wang SL, Cai QS, Wang JS, Hong JY (2006) Efficient activation of aflatoxin B1 by cytochrome P450 2A13, an enzyme predominantly expressed in human respiratory tract. Int J Cancer 118:2665–2671
Henry KM, Townsend CA (2005) Ordering the reductive and cytochrome P450 oxidative steps in demethylsterigmatocystin formation yields general insights into the biosynthesis of aflatoxin and related fungal metabolites. J Am Chem Soc 127:3724–3733
Hong SY, Roze LV, Wee J, Linz JE (2013) Evidence that a transcription factor regulatory network coordinates oxidative stress response and secondary metabolism in aspergilli. Microbiology 2:144–160
Hsieh D (1989) Potential human health hazards of mycotoxins. Bioactive Mol 10:69–80
Huang JQ, Jiang HF, Zhou YQ, Lei Y, Wang SY, Liao BS (2009) Ethylene inhibited aflatoxin biosynthesis is due to oxidative stress alleviation and related to glutathione redox state changes in aspergillus flavus. Int J Food Microbiol 130:17–21
Jayashree T, Subramanyam C (2000) Oxidative stress as a prerequisite for aflatoxin production by aspergillus parasiticus. Free Radic Biol Med 29:981–985
Johnson WW, Guengerich FP (1997) Reaction of aflatoxin B1 exo-8, 9-epoxide with DNA: kinetic analysis of covalent binding and DNA-induced hydrolysis. Proc Natl Acad Sci 94:6121–6125
Kamdem LK, Meineke I, Gödtel-Armbrust U, Brockmöller J, Wojnowski L (2006) Dominant contribution of P450 3A4 to the hepatic carcinogenic activation of aflatoxin B1. Chem Res Toxicol 19:577–586
Kelkar HS, Skloss TW, Haw JF, Keller NP, Adams TH (1997) Aspergillus nidulans stcL encodes a putative cytochrome P-450 monooxygenase required for bisfuran desaturation during aflatoxin/sterigmatocystin biosynthesis. J Biol Chem 272:1589–1594
Keller N, Dischinger H, Bhatnagar D, Cleveland T, Ullah A (1993) Purification of a 40-kilodalton methyltransferase active in the aflatoxin biosynthetic pathway. Appl Environ Microbiol 59:479–484
Keller NP, Nesbitt C, Sarr B, Phillips TD, Burow GB (1997) pH regulation of sterigmatocystin and aflatoxin biosynthesis in aspergillus spp. Phytopathology 87:643–648
Keller NP, Segner S, Bhatnagar D, Adams TH (1995) stcS, a putative P-450 monooxygenase, is required for the conversion of versicolorin a to sterigmatocystin in aspergillus nidulans. Appl Environ Microbiol 61:3628–3632
Keller NP, Watanabe CM, Kelkar HS, Adams TH, Townsend CA (2000) Requirement of monooxygenase-mediated steps for sterigmatocystin biosynthesis by aspergillus nidulans. Appl Environ Microbiol 66:359–362
Kim JH, Yu J, Mahoney N, Chan KL, Molyneux RJ, Varga J, Bhatnagar D, Cleveland TE, Nierman WC, Campbell BC (2008) Elucidation of the functional genomics of antioxidant-based inhibition of aflatoxin biosynthesis. Int J Food Microbiol 122:49–60
Klich M, Yu J, Chang PK, Mullaney E, Bhatnagar D, Cleveland T (1995) Hybridization of genes involved in aflatoxin biosynthesis to DNA of aflatoxigenic and non-aflatoxigenic aspergilli. Appl Microbiol Biotechnol 44:439–443
Klich MA (2007) Environmental and developmental factors influencing aflatoxin production by aspergillus flavus and aspergillus parasiticus. Mycoscience 48:71–80
Kussak A, Andersson B, Andersson K, Nilsson CA (1998) Determination of aflatoxicol in human urine by immunoaffinity column clean-up and liquid chromatography. Chemosphere 36:1841–1848
Kusumoto KI, Hsieh DP (1996) Purification and characterization of the esterases involved in aflatoxin biosynthesis in aspergillus parasiticus. Can J Microbiol 42:804–810
Lamb DC, Lei L, Warrilow AG, Lepesheva GI, Mullins JG, Waterman MR, Kelly SL (2009) The first virally encoded cytochrome p450. J Virol 83:8266–8269
Liu J, Sun L, Zhang N, Zhang J, Guo J, Li C, Rajput SA, Qi D (2016) Effects of nutrients in substrates of different grains on aflatoxin B1 production by aspergillus flavus. Biomed Res Int 2016. https://doi.org/10.1155/2016/7232858
Liu X, Guan X, Xing F, Lv C, Dai X, Liu Y (2017) Effect of water activity and temperature on the growth of aspergillus flavus, the expression of aflatoxin biosynthetic genes and aflatoxin production in shelled peanuts. Food Control 82:325–332
Maggio-Hall LA, Wilson RA, Keller NP (2005) Fundamental contribution of β-oxidation to polyketide mycotoxin production in planta. Mol Plant Microbe Infect 18:783–793
Maggio-Hall LA, Keller NP (2004) Mitochondrial β-oxidation in aspergillus nidulans. Mol Microbiol 54:1173–1185
Masri MS, Haddon WF, Lundin RE, Hsieh DP (1974) Aflatoxin Q1. Newly identified major metabolite of aflatoxin B1 in monkey liver. J Agric Food Chem 22:512–515
Mcguire SM, Silva JC, Casillas EG, Townsend CA (1996) Purification and characterization of versicolorin B synthase from aspergillus parasiticus. Catalysis of the stereodifferentiating cyclization in aflatoxin biosynthesis essential to DNA interaction. Biochemistry 35:11470–11486
Minto RE, Townsend CA (1997) Enzymology and molecular biology of aflatoxin biosynthesis. Chem Rev 97:2537–2556
Montibus M, Pinson-Gadais L, Richard-Forget F, Barreau C, Ponts N (2015) Coupling of transcriptional response to oxidative stress and secondary metabolism regulation in filamentous fungi. Crit Rev Microbiol 41:295–308
Mykkänen H, Zhu H, Salminen E, Juvonen RO, Ling W, Ma J, Polychronaki N, Kemiläinen H, Mykkänen O, Salminen S (2005) Fecal and urinary excretion of aflatoxin B1 metabolites (AFQ1, AFM1 and AFB-N7-guanine) in young Chinese males. Int J Cancer 115:879–884
Narasaiah KV, Sashidhar R, Subramanyam C (2006) Biochemical analysis of oxidative stress in the production of aflatoxin and its precursor intermediates. Mycopathologia:162–179
Neal G, Nielsch U, Judah D, Hulbert P (1987) Conjugation of model substrates or microsomally-activated aflatoxin B1 with reduced glutathione, catalysed by cytosolic glutathione-S-transferases in livers of rats, mice and Guinea pigs. Biochem Pharmacol 36:4269–4276
Nelson DR (2011) Progress in tracing the evolutionary paths of cytochrome P450 Proteins and proteomics. https://doi.org/10.1016/j.bbapap.2010.08.008
Niu G, Wen Z, Rupasinghe SG, Zeng RS, Berenbaum MR, Schuler MA (2008) Aflatoxin B1 detoxification by CYP321A1 in Helicoverpa zea. Arch Insect Biochem 69:32–45
Obrian G, Georgianna D, Wilkinson J, Yu J, Abbas H, Bhatnagar D, Cleveland T, Nierman W, Payne G (2007) The effect of elevated temperature on gene transcription and aflatoxin biosynthesis. Mycologia 99:232–239
Partanen HA, El-Nezami HS, Leppänen JM, Myllynen PK, Woodhouse HJ, Vähäkangas KH (2010) Aflatoxin B1 transfer and metabolism in human placenta. Toxicol Sci 113:216–225
Price MS, Yu J, Nierma WC, Kim HS, Pritchard B, Jacobus CA, Bhatnagar D, Cleveland TE, Payne GA (2006) The aflatoxin pathway regulator AflR induces gene transcription inside and outside of the aflatoxin biosynthetic cluster. FEMS Microbiol Lett 255:275–279
Purschwitz J, Müller S, Kastner C, Schöser M, Haas H, Espeso EA, Atoui A, Calvo AM, Fischer R (2008) Functional and physical interaction of blue-and red-light sensors in aspergillus nidulans. Curr Biol 18:255–259
Ramsdell HS, Eaton DL (1990) Species susceptibility to aflatoxin B1 carcinogenesis: comparative kinetics of microsomal biotransformation. Cancer Res 50:615–620
Raney KD, Coles B, Guengerich FP, Harris TM (1992a) The endo-8, 9-epoxide of aflatoxin B1: a new metabolite. Chem Res Toxicol 5:333–335
Raney KD, Shimada T, Kim DH, Groopman JD, Harris TM, Guengerich FP (1992b) Oxidation of aflatoxins and sterigmatocystin by human liver microsomes: significance of aflatoxin Q1 as a detoxication product of aflatoxin B1. Chem Res Toxicol 5:202–210
Rangel DE, Alder-Rangel A, Dadachova E, Finlay RD, Kupiec M, Dijksterhuis J, Braga GU, Corrochano LM, Hallsworth JE (2015) Fungal stress biology: a preface to the fungal stress responses special edition. Curr Genet 61:231–238
Ross RK, Yu M, Henderson B, Yuan JM, Qian GS, Tu JT, Gao YT, Wogan G, Groopman J (1992) Urinary aflatoxin biomarkers and risk of hepatocellular carcinoma. Lancet 339:943–946
Roze LV, Hong SY, Linz JE (2013) Aflatoxin biosynthesis: current frontiers. Annu Rev Food Sci Technol 4:293–311
Roze LV, Laivenieks M, Hong SY, Wee J, Wong SS, Vanos B, Awad D, Ehrlich KC, Linz JE (2015) Aflatoxin biosynthesis is a novel source of reactive oxygen species—a potential redox signal to initiate resistance to oxidative stress? Toxins 7:1411–1430
Ruijter GJ, Visser J (1997) Carbon repression in aspergilli. FEMS Microbiol Lett 151:103–114
Rushing BR, Selim MI (2017) Structure and oxidation of pyrrole adducts formed between aflatoxin B2a and biological amines. Chem Res Toxicol 30:1275–1285
Rushing BR, Selim MI (2019) Aflatoxin B1: a review on metabolism, toxicity, occurrence in food, occupational exposure, and detoxification methods. Food Chem Toxicol 124:81–100
Sakuno E, Wen Y, Hatabayashi H, Arai H, Aoki C, Yabe K, Nakajima H (2005) Aspergillus parasiticus cyclase catalyzes two dehydration steps in aflatoxin biosynthesis. Appl Environ Microbiol 71: 2999–3006
Sakuno E, Yabe K, Nakajima H (2003) Involvement of two cytosolic enzymes and a novel intermediate, 5′-oxoaverantin, in the pathway from 5′-hydroxyaverantin to averufin in aflatoxin biosynthesis. Appl Environ Microbiol 69:6418–6426
Shimada T, Yamazaki H, Mimura M, Inui Y, Guengerich FP (1994) Interindividual variations in human liver cytochrome P-450 enzymes involved in the oxidation of drugs, carcinogens and toxic chemicals: studies with liver microsomes of 30 Japanese and 30 Caucasians. J Pharmacol Exp Ther 270:414–423
Silva JC, Minto RE, Barry Iii CE, Holland KA, Townsend CA (1996) Isolation and characterization of the Versicolorin B synthase gene from aspergillus parasiticus: expansion of the aflatoxin b1 biosynthetic gene cluster. J Biol Chem 271:13600–13608
Silva JC, Townsend CA (1997) Heterologous expression, isolation, and characterization of versicolorin B synthase from aspergillus parasiticus: a key enzyme in the aflatoxin B1 biosynthetic pathway. J Biol Chem 272:804–813
Ueng YF, Shimada T, Yamazaki H, Guengerich FP (1995) Oxidation of aflatoxin B1 by bacterial recombinant human cytochrome P450 enzymes. Chem Res Toxicol 8:218–225
Walsh MC, Buzoianu SG, Gardiner GE, Rea MC, Gelencsér E, Jánosi A, Epstein MM, Ross RP, Lawlor PG (2011) Fate of transgenic DNA from orally administered Bt MON810 maize and effects on immune response and growth in pigs. PLoS One 6:271–277
Wen Y, Hatabayashi H, Arai H, Kitamoto HK, Yabe K (2005) Function of the cypX and moxY genes in aflatoxin biosynthesis in aspergillus parasiticus. Appl Environ Microbiol 71:3192–3198
Weydert CJ, Cullen JJ (2010) Measurement of superoxide dismutase, catalase and glutathione peroxidase in cultured cells and tissue. Nat Protoc 5:51–66
Wu Q, Jezkova A, Yuan Z, Pavlikova L, Dohnal V, Kuca K (2009) Biological degradation of aflatoxins. Drug Metab Rev 41:1–7
Yabe K, Ando Y, Hamasaki T (1991) A metabolic grid among versiconal hemiacetal acetate, versiconol acetate, versiconol and versiconal during aflatoxin biosynthesis. Microbiology 137:2469–2475
Yabe K, Chihaya N, Hatabayashi H, Kito M, Hoshino S, Zeng H, Cai J, Nakajima H (2012) Production of M−/GM-group aflatoxins catalyzed by the OrdA enzyme in aflatoxin biosynthesis. Fungal Genet Biol 49:744–754
Yabe K, Matsuyama Y, Ando Y, Nakajima H, Hamasaki T (1993) Stereochemistry during aflatoxin biosynthesis: conversion of norsolorinic acid to averufin. Appl Environ Microbiol 59:2486–2492
Yabe K, Nakajima H (2004) Enzyme reactions and genes in aflatoxin biosynthesis. Appl Microbiol Biotechnol 64:745–755
Yourtee D, Bean T, Kirk-Yourtee C (1987) Human aflatoxin B1 metabolism: an investigation of the importance of aflatoxin Q1 as a metabolite of hepatic post-mitochondrial fraction. Toxicol Lett 38:213–224
Yu J (2012) Current understanding on aflatoxin biosynthesis and future perspective in reducing aflatoxin contamination. Toxin 4:1024–1057
Yu J, Bhatnagar D, Cleveland TE (2004a) Completed sequence of aflatoxin pathway gene cluster in aspergillus parasiticus. FEBS Lett 564:126–130
Yu J, Cary J, Bhatnagar D, Cleveland T, Keller N, Chu F (1993) Cloning and characterization of a cDNA from aspergillus parasiticus encoding an O-methyltransferase involved in aflatoxin biosynthesis. Appl Environ Microbiol 59:3564–3571
Yu J, Chang PK, Bhatnagar D, Cleveland TE (2000a) Cloning of a sugar utilization gene cluster in aspergillus parasiticus. Biochim Biophys Acta (BBA)-Gene Struct Exp 1493:211–214
Yu J, Chang PK, Bhatnagar D, Cleveland TE (2003) Cloning and functional expression of an esterase gene in aspergillus parasiticus. Mycopathologia 156:227–234
Yu J, Chang PK, Cary JW, Bhatnagar D, Cleveland TE (1997) avnA, a gene encoding a cytochrome P-450 monooxygenase, is involved in the conversion of averantin to averufin in aflatoxin biosynthesis in aspergillus parasiticus. Appl Environ Microbiol 63:1349–1356
Yu J, Chang PK, Ehrlich KC, Cary JW, Bhatnagar D, Cleveland TE, Payne GA, Linz JE, Woloshuk CP, Bennett JW (2004b) Clustered pathway genes in aflatoxin biosynthesis. Appl Environ Microbiol 70:1253–1262
Yu J, Chang PK, Ehrlich KC, Cary JW, Montalbano B, Dyer JM, Bhatnagar D, Cleveland TE (1998) Characterization of the critical amino acids of anAspergillus parasiticus cytochrome P-450 monooxygenase encoded by ordA that is involved in the biosynthesis of aflatoxins B1, G1, B2, and G2. Appl Environ Microbiol 64:4834–4841
Yu J, Chang PK, Payne GA, Cary JW, Bhatnagar D, Cleveland TE (1995) Comparison of the omtA genes encoding O-methyltransferases involved in aflatoxin biosynthesis from aspergillus parasiticus and A. flavus. Gene 163:121–125
Yu J, Fedorova ND, Montalbano BG, Bhatnagar D, Cleveland TE, Bennett JW, Nierman WC (2011) Tight control of mycotoxin biosynthesis gene expression in aspergillus flavus by temperature as revealed by RNA-Seq. FEMS Microbiol Lett 322:145–149
Yu J, Woloshuk CP, Bhatnagar D, Cleveland TE (2000b) Cloning and characterization of avfA and omtB genes involved in aflatoxin biosynthesis in three aspergillus species. Gene 248:157–167
Zeng H, Hatabayashi H, Nakagawa H, Cai J, Suzuki R, Sakuno E, Tanaka T, Ito Y, Ehrlich KC, Nakajima H (2011) Conversion of 11-hydroxy-O-methylsterigmatocystin to aflatoxin G 1 in aspergillus parasiticus. Appl Microbiol Biotechnol 90:635–650
Zhang JD, Han L, Yan S, Liu CM (2014) The non-metabolizable glucose analog D-glucal inhibits aflatoxin biosynthesis and promotes kojic acid production in aspergillus flavus. BMC Microbiol 14:1–9
Zhou R, Linz JE (1999) Enzymatic function of the Nor-1 protein in aflatoxin biosynthesis in aspergillus parasiticus. Appl Environ Microbiol 65:5639–5641
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Ethics declarations
The authors declare no conflict of interest.
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Muaz, K. et al. (2021). Aflatoxin Biosynthesis. In: Hakeem, K.R., Oliveira, C.A.F., Ismail, A. (eds) Aflatoxins in Food. Springer, Cham. https://doi.org/10.1007/978-3-030-85762-2_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-85762-2_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-85761-5
Online ISBN: 978-3-030-85762-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)