Abstract
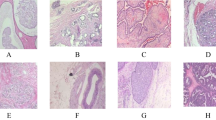
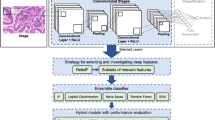
Histopathological image analysis of biopsy sample provides an accurate diagnosis method for cancer. Usually Pathologists examine the microscopic images of biopsy sample manually for the detection and grading of cancer. Automation in this field helps the pathologists to take a second opinion before confirming with their findings. We propose an effective method of automated cancer detection by combining the effect of transfer learning and ensemble learning. Six pre-trained models such as Densenet121, Resnet 50, Xception, EfficientNet B7, MobileNetV2, and VGG19 are used for preparing an ensemble model. A dataset contains 5547 H&E stained histopathological images of malignant and benign tissues are used to train and validate each models individually and obtained an accuracy of 77.9%, 79%, 79.8%, 78.3%, 77%, and 76% respectively. Based on the accuracy, best performing three models Resnet50, Xception, and EfficientnetB7 are selected to form an ensemble model. Then the layers and weights of these models are freezed and the output layers are concatenated to make an ensemble model. New dense layers are added to the ensembled model to provide a single output for binary classification. The model is compiled by an Adam optimizer with a learning rate of 0.001. The images are again applied to this ensemble model to classify the malignant and benign tissues and obtained an accuracy of 96% and precision of 96%.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Boyle, P., Levin, B.: World Cancer report 2008: IARC Press. International Agency for Research on Cancer (2008)
https://www.who.int/nmh/publications/ncd_report_chapter1.pdf
Xie, J., Liu, R., Luttrell, J., Zhang, C.: Deep learning based analysis of histopathological images of breast cancer. Front. Genet. 10(80), 1–19 (2019)
Bello, M., Nápoles, G., Sánchez, R., Bello, R., Vanhoof, R.: Deep neural network to extract high-level features and labels in multi-label classification problems, Neurocomputing, 413, 259–270 (2020). ISSN 0925–2312
Hussain, M., Bird, J.J., Faria, D.R.: A study on CNN transfer learning for image classification. In: Lotfi, A., Bouchachia, H., Gegov, A., Langensiepen, C., McGinnity, M. (eds.) UKCI 2018. AISC, vol. 840, pp. 191–202. Springer, Cham (2019). https://doi.org/10.1007/978-3-319-97982-3_16
Nanni, L., Ghidoni, S., Brahnam, S.: Ensemble of convolutional neural networks for bioimage classification, Appl. Comput. Inf. (2020). ISSN 22108327
Howard, A.G., et al.: MobileNets: efficient convolutional neural networks for mobile vision applications (2017). https://arxiv.org/abs/1704.04861
Chollet, F.: Xception: deep learning with depthwise separable convolutions. In: 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 1800–1807. Honolulu, HI, USA (2017)
https://towardsdatascience.com/understanding-and-coding-a-resnet-in-keras-446d7ff84d33
Huang, G., Liu, Z., Weinberger, K.Q.: Densely connected convolutional networks. https://arxiv.org/abs/1608.06993
Tan, M., Le, Q.V.: EfficientNet: rethinking model scaling for convolutional neural networks. In: Proceedings of International Conference on Machine Learning (ICML) (2019)
Huang, F., Xie, G., Xiao, R.: Research on ensemble learning. In: Proceedings of the 2009 International Conference on Artificial Intelligence and Computational Intelligence. vol. 3. IEEE Computer Society (2009)
Opitz, D., Maclin, R.: Popular ensemble methods: an empirical study. J. Artif. Intell. Res. 11, 169–198 (1999)
Zhu, Y., Brettin, T., Evrard, Y.A., et al.: Ensemble transfer learning for the prediction of anti-cancer drug response. Sci. Rep. 10, 18040 (2020). https://doi.org/10.1038/s41598-020-74921-0
Xue, D., et al.: An application of transfer learning and ensemble learning techniques for cervical histopathology image classification. IEEE Access 8 (2020)
Kandaswamy, C., Silva, L.M., Alexandre, L.A., Santos, J.M.: Deep transfer learning ensemble for classification. In: Rojas, I., Joya, G., Catala, A. (eds.) IWANN 2015. LNCS, vol. 9094, pp. 335–348. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-19258-1_29
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Devassy, B.R., Antony, J.K. (2022). Histopathological Image Classification Using Ensemble Transfer Learning. In: Misra, R., Shyamasundar, R.K., Chaturvedi, A., Omer, R. (eds) Machine Learning and Big Data Analytics (Proceedings of International Conference on Machine Learning and Big Data Analytics (ICMLBDA) 2021). ICMLBDA 2021. Lecture Notes in Networks and Systems, vol 256. Springer, Cham. https://doi.org/10.1007/978-3-030-82469-3_18
Download citation
DOI: https://doi.org/10.1007/978-3-030-82469-3_18
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-82468-6
Online ISBN: 978-3-030-82469-3
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)