Abstract
“COVID-2019,” a recently emerged novel coronavirus disease, is causing serious health issues to the public and becoming more and more fatal every next day. On December 31, 2019, low respiratory infection cases were detected in Wuhan, China, which is in China’s Hubei province. The cases were reported to the WHO Office of China and they could not identify the agents for the cause. The first cases were classified to be “pneumonia of unknown etiology.” The investigation program was initiated by the Chinese Center for Disease Control and Prevention (CDC). The etiology of the disease was attributed to a novel virus of the coronavirus (CoV) family. Dr. Tedros Adhanom Ghebreyesus, WHO Director-General, called the disease caused by this CoV the “COVID-19,” which is an acronym for “coronavirus disease 2019.” It is found that “COVID-19” is caused by bête-coronavirus named “severe acute coronavirus-2” (SARS-CoV-2). It belongs to those virus families that appear as pneumonia in the human body. It affects the lower respiratory tract badly. This virus has been identified as another version of the family of severe acute respiratory syndrome coronavirus (SARS-CoV) and the Middle East respiratory syndrome coronavirus (MERS-CoV) [1, 2]. SARS-CoV-2, SARS-CoV, and MERS-CoV possess similarity with them. They have differences in genotypic and phenotypic structure that guide their pathogenesis. So far, as per the findings, this virus originated in bats. It reached humans through contact with unknown animals. The transmission of this virus among humans is via direct contacts, inhalation of infected droplets, and contaminated hands and surfaces. Some of the symptoms of this disease are cough, sore cough, fever, fatigue, and dyspnea/breathlessness. The remedy of this disease is to diagnose the infection at the initial stage, supportive treatment to survive, self-quarantines, mass-quarantines, etc. This paper presents a systematic review of the origin of coronavirus, its types, transmissions, symptoms, and the current developments in diagnosing testing and vaccine trials.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
According to the study, a rare and unexplained case of pneumonia was reported in Wuhan, China, at the end of December 2019. Wuhan is a Chinese city in the Hubei Province. At first, the symptoms of this illness seemed to be somewhat similar to those of pneumonia. However, experts from the “Chinese Center for Disease Control and Prevention (China CDC)” said that this is a new kind of virus-infected disease. The report was focused on the examination of a number of respiratory samples (throat swab samples). It is caused by a novel coronavirus that resembles the native SARS-CoV and MERS-CoV families [3]. The illness was later dubbed COVID-19 by the World Health Organization (WHO) (coronavirus disease). According to a WHO survey, the pandemic affected 205 countries until April 2, 2020, with 900,306 reported cases and 45,692 deaths [4]. COVID-19 has turned into a global threat to mankind. According to the study, the coronavirus spreads among humans through a variety of routes, including direct contact or inhalation of infected droplets and contact with polluted hands/places/objects/surfaces [3]. However, in some circumstances, the presence of gastrointestinal symptoms or a live infectious virus in the feces may be another way for infections to spread in humans through fecal–oral transmission [5]. The droplets are divided into two groups depending on their size. Respiratory droplets are described as droplets that are larger than 5–10 m in diameter. When the scale is less than 5 m, it is referred to as a droplet nucleus [6]. According to current studies, respiratory droplets are largely responsible for coronavirus transmission in humans [7,8,9]. SARS-CoV-2, SARS-CoV, and MERS-CoV all belong to the Orthocoronavirinae, according to phylogenetic analysis. They are also members of the Coronaviridae family [10]. SARS-CoV and MERS-CoV, on the other hand, are not the same as SARS-CoV-2 [11, 12].
The four genera of Orthocoronavirinae subfamily indulged in affecting living objects, for example, mammals are infected through αCoV and βCoV while δCoV and γ-CoV are the main causes of infection in birds. Recently, two outbreaks were found by CoV in humans. This was the case of viral pneumonia caused by SARS and MERS. The first case of SARS-CoV in China was reported in the year 2002, a new kind of CoV, which spread worldwide very quickly. The mortality rate of this outbreak was 11% [13, 14]. This outbreak fetched the attention of researchers on evolution, replication, transmission, severity, biological structure, and development manner of the disease (i.e., microbial infection, inflammation, malignancy, and tissue breakdown) of CoV. After a few years, another virus-related severe respiratory disease outbreak was reported in the Middle East, in 2012 [15]. The starting point of this outbreak was Saudi Arabia and after that spread to other countries, with 37% fatality rate [16]. As this virus was emerged in the Middle East, it was named MERS-CoV. In both the epidemic cases of virus infection, one thing was very clear that they likely came from bats and reached to humans through any intermediate carrier [17]. In a research, it was found that the camel and the civet, respectively, were used as carriers for MERS-CoV and SARS-CoV [18, 19]. Like SARS-CoV and MERS-CoV, the newly introduced SARS-CoV-2 is also related to βCoV lineage.
This paper presents the systematic report on origin of disease, epidemiology, genome structure and life cycle, clinical features, diagnosis, isolation, treatment, prevention, societal and economical impact, and global response.
1.1 Classification of Coronavirus (CoVs)
Coronaviridae has a subfamily called Orthocoronavirinae. Coronaviruses (CoVs) are classified as Orthocoronavirinae. Coronaviruses (CoVs) are divided into four classes. Coronaviridae, Arteriviridae, Mesoniviridae, and Roniviridae are the four families. They are members of the Nidovirales order. As seen in Fig. 1, the Orthocoronavirinae subfamily contains four genera: alphacoronavirus (CoV), betacoronavirus (CoV), deltacoronavirus (CoV), and gammacoronavirus (-CoV). Coronaviridae viruses are single-stranded, enveloped, and very large ribonucleic acid (RNA) viruses with a positive single-stranded, enveloped genome. In contrast to other recognized RNA viruses [10], the genome size of Coronaviridae family viruses is very high which ranges from 25,000 to 32,000 base pairs. The diameter of a virion will range from 118 to 136 nm. Coronavirinae and Torovirinae are the two subfamilies of the Coronaviridae family [19].
1.2 Genomic Organization and Structure of Coronavirus (SARS-CoV-2)
The SARS-CoV-2 contains large, non-segmented, positive single-stranded RNA genome of 30 kb in size. The RNA genome contains a 3′ polyadenylated (A) tail with the 5′ capped structure to translate (encode) replicase polyproteins (large and nonstructural proteins), and for this translation, RNA genome acts as an mRNA. In the whole translation process, discontinuous transcription process is used to synthesize the mRNA [20, 21]. To continue the transcription and replication of RNA, it requires multiple stem-loop structures. These structures are contained by untranslated region (UTR) of the 5′ end of genome. Also the transcriptional regulatory sequences (TRSs) are required for the expression of every genes of the structural gene. Additionally, viral RNA imitation and synthesis are executed by RNA structures of 3′ end UTR.
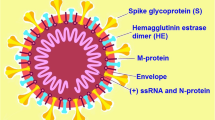
The RNA genome contains open reading frame (ORF), 1a/1b, which further generates five accessory proteins (i.e., ORF9, ORF8, ORF7, ORF6, and ORF3a), four structural proteins (nucleocapsid protein, envelope protein, membrane protein, and spike glycoprotein) (Fig. 2), and 15/16 nonstructural proteins (majority of them are involved in the synthesis of viral RNA) [22, 23]. These proteins support in spreading infection of SARS-CoV-2.
In a recent research, it was found that the coronavirus is spherically shaped and virions are of approximately 125 nm in diameter [24, 25]. Another report says that the diameter of virions varies between 118 and 140 nm [19]. The genome size of this virus lies between 26.4 and 31.7 kb, which is the largest size among all known genomes of RNA viruses [26]. The distinct feature of the coronavirus is its club-shaped spike projections, which originated or emitted from the virion surface. A virion envelop contains nucleocapsids, which come under the category of structural proteins. Presence of the nucleocapsids is very common in all known negatively sensed RNA viruses, but this is very uncommon in positively sensed RNA viruses. Among four structural proteins of coronavirus, spike glycoprotein (S) plays a significant role to host cells in the attachment to itself, and further host proteases (protease like furin) are used to cleave S into two distinct polypeptides, that is, S1 and S2 [27, 28]. For gating access to endoplasmic reticulum (ER), the S protein uses signal sequences of N-terminal. Homotrimers of the S (virus-encoded) protein makes a special spike shape on virus surface [29, 30]. The binding of host receptor and S1 can unstable the prefusion trimer. As a result, highly stable postfusion conformation is formed due to S1 shedding and S2 transition [31]. S1’s receptor binding domain goes through conformational movements to engage a host receptor to cover (down conformation) or uncover (up conformation) the determinants of receptor binding transiently [32]. In this context, receptor’s inaccessible state can be represented by down conformation, while the up conformation represents receptor’s accessible state.
The coronavirus virion contains a very small quantity of envelope protein (E) and protein E follows a common architecture but it is very much divergent in nature [33]. This protein has ion channel activity as well as it contains C-terminal endo-domain and N-terminal ecto-domain. E protein plays the significant role in the facilitation of virus assembly and release. The role of E protein is necessary for pathogenesis [34].
Virion of coronavirus has rich storage of membrane protein (M). M protein is small in size and contains three transmembrane domains to give the shape of the virion [33]. The M protein has the ability to extend between 6 and 8 nm due to the presence of C-terminal endo-domain (small in size) and N-terminal glycosylated ecto-domain (very large in size). In a recent study, it has been found that two distinct conformations may be adopted by M protein to allow it to bind the nucleocapsid and also to boost the membrane curvature [35].
The nucleocapsid protein (N) of coronavirus is a kind of multifunctional protein that plays a vital role in enhancing the effectiveness of assembly and transcription of virus. During the assembly of virion, N protein establishes interaction with the viral M protein. The N protein has two distinct domains, that is, N-terminal domain (NTD) and C-terminal domain (CTD). Both domains are separated by RNA-binding domain. Both domains are separately attached with viral RNA. The optimality of RNA binding depends on the same effort from both domains [36].
Transcriptional regulatory sequence (TRS) and genome packaging signal are the two RNA substrates related to N protein [37, 38]. To bind the RNA-binding domain of C-terminal, genomic packaging signal is used [39].
Hence, detailed understanding of the proteins and genomic structure of coronavirus will help to design and develop vaccines and monoclonal antibody drugs.
2 Manifestations and Epidemiology of SARS-CoV-2
2.1 Origin of SARS-CoV-2
The source of origin plays a significant role in understanding any disease. In the case of COVID-19, primarily the researchers found that a large number of people were infected from the market of live animals (Huanan seafood market) in Wuhan city of China. Initially, the infected people were either shop employees, owner of the shop, or customers to this market. After this incident, the market was closed on January 1, 2020. On the basis of the efforts taken to understand the cause of the spreading of coronavirus, it was found that this virus was transmitted through intermediate carriers in humans. In the primary report of researchers, environmental samples taken from the seafood market depicted the positive evidence of viral RNA, and on that basis, it was recognized that Asian palm civet might be the intermediate host of coronavirus [40]. After several investigations, still there has been doubt about the potential host of the SARS-CoV-2 [41]. Recently, on the basis of several studies, it has been identified that the bats might be the potential host of this virus [11, 12, 42]. Based on the clinical research, it was found that the genetic sequence of coronavirus has more than 95% similarity with the bat coronavirus and more than 70% similarity with SARS-CoV [42, 43]. Figure 3 shows the origin of coronavirus and its spread to humans through potential mediums.
Illustration of the coronavirus’s origin and dissemination into humans through various mediums. The most common way for an infection to spread from an animal to a human is by eating the animal’s meat (primary host) or near contact with an intermediate host. The infection is then transmitted from one sick person to another healthy person. The virus is then spread to the general population through human-to-human contact [44]
A report published in “The Lancet” says that the origin of the coronavirus is the wildlife. Some studies say that the virus had been brought by an infected human in the seafood market and then it spread out in the market environment [45]. Some reports say that there was a conspiracy behind the origin of this deadly virus. In this conspiracy, the role of the Wuhan Institute of Virology has been found suspicious. There is a rumor that this institute is involved in developing human-generated bio-weapon. In another investigation, the virologists stated that this virus is new and has originated from the environment. However, after many investigations, still there is a question about what is the primary source of virus amplification in humans. In this context, more investigations are required to understand the real source of spreading of the virus in humans.
2.2 Symptoms of SARS-CoV-2
Symptoms of COVID-19 infection can take anywhere from 1 to 14 days to appear. Dry cough, respiratory signs (like the flu), acute fever, tiredness, and headache are all typical COVID-19 symptoms [3, 46,47,48]. The disease can manifest itself in a variety of ways, from negative to positive. Good signs indicate the presence of a disease outbreak in this case. A new research looked at 41 patients who were diagnosed with COVID-19 at the outset. Cough, fever, and nausea were the most frequent symptoms in the majority of the patients. Although there was trouble in breathing, headache, hemoptysis, dyspnea, diarrhea, and lymphopenia were all serious and even fatal symptoms in some instances [3]. During a chest CT scan, it was discovered that both the patients had pneumonia. Acute respiratory distress syndrome was seen in almost 29% of the cases, acute heart attack in 12% of the cases, and other infections in 10% of the cases [3].
In another clinical report, patients were found to have unusual symptoms such as acute cardiac injury and respiratory distress syndrome [3]. Gastrointestinal symptoms (i.e., vomiting and diarrhea, and nausea) were observed in the patients during the illness [3, 49, 50]. The first case of the USA was confirmed for respiratory illness followed by vomiting and diarrhea, nausea, and abdominal pain [49].
2.3 Incubation Period of SARS-CoV-2
The time between the symptom onset and getting exposure to the virus is known as the incubation period for SARS-CoV-2. Average time of the incubation period is 5–6 days; however, this period can extend up to 14 days and known as the presymptomatic period. Contagiousness can be spread out by a few of the infected persons. In the earlier studies, the estimated mean incubation period of COVID-19 was 3–7 days within 2–14 days, which shows that this virus had a long transmission period [51, 52]. The transmission period of CoVs was found a little bit different from its counterpart viruses like mean incubation period for SARS-CoVs, it was 5 days within 2–14 days [53], and for MERS-CoVs, it was 5–7 days within 2–14 days [54]. In a recent clinical report on 138 cases, the median incubation period from the initial symptoms to acute respiratory disorder and finally hospitalization was 5 days within the range of 1–10 days, 7 days within the range of 4–8 days, and 8 days within the range of 6–12 days, respectively [55]. In another confirmed case of 425 patients the mean incubation period was 5.2 days (range 4.0–7.0 days) for some patients and for others it was 7.5 days (range 5.0–19 days). The confidence interval (CI) was observed to be 95% for all 425 patients [9]. While the mean incubation period estimated over 1099 patients was 3.0 days, range within 0–24 [56]. In another report, the mean incubation period of the infection was 4.6 days within 3.8–5.8 days with 95% CI and the beginning of 95% disease occurred within 10 days [57, 58]. In comparison to SARS-CoVs and MERS-CoVs, the fatality rate of COVID-19 is less due to the lower incubation period which was estimated to be 3.0 days [55]. Days between 2 and 8 has been estimated as a mean time from primary symptom to admission in hospital [59]. The time duration from the initial symptoms of COVID-19 to death ranged between 6 and 41 days, and the median of the incubation period was 14 days [47]. Age and immune system of the patient decide this period. In critical cases of COVID-19, this can take 3–6 weeks to recover or die out from the disease. Generally, the period is smaller for the patient, whose age is less than 70 years as compared to patients with age greater than 70 years [47]. In symptomatic transmission, the repeated biological samples based on virologic and clinical activities observed that the expulsion and release of CoVs are highest in the upper respiratory tract in the initial phase of the infection of confirmed cases [60,61,62,63], which happened in the first 3 days from the starting of the symptoms [62, 63].
2.4 Transmission of SARS-CoV-2
As per the current studies on the pathophysiological properties of SARS-CoV-2, it had been found that the spreading mechanism of this virus is very uncertain. This study was based on the human-to-human transmission of coronavirus through respiratory droplets [64]. The size of the respiratory droplets lies between 5 and 15 μm, and they remain to sustain in the air for several minutes before dropping on the floor. However, the droplet particles whose size is less than 5 μm remain to sustain in the air up to hours due to their small size and less weight. During this time, if any person comes in the close contact with the infected droplets, then he/she becomes infected.
In the case of symptomatic symptoms, the contagiousness of the viruses related to respiratory infection becomes very high. In symptomatic transmission, a person transmits the infection to another person, while he/she is experiencing the symptoms of coronavirus. The virologic and epidemiologic studies say that the symptomatic transmission occurred through direct contact with the infected patients, or close contact with surfaces and objects contaminated by the virus, or the close contact with respiratory droplets [7,8,9]. In asymptomatic transmission, the virus transmission from the infected person to another healthy person occurs in such a way that it does not develop any symptom of COVID-19. In some laboratory reports, some asymptomatic cases have been confirmed. However, there has not been any confirmed case of asymptomatic transmission found [65].
Generally, the symptomatic patient is more infectious due to the presence of respiratory viruses. However, the evidence of virus transmission is increasing during human-to-human interaction in the incubation period of asymptomatic transmission of COVID-19, and the probability estimation of this period has been between 2 and 10 days [64].
A lot of analysis has been done to understand the source of transmission of SARS-CoV-2 in humans. In spite of respiratory droplets, close contact is also a big source of transmission of SARS-CoV-2. Typically, three organs of human body like mouth, nose, and eyes are responsible for virus outbreak and transmission. Another possibility of infection by aerosol transmission to the people through airborne particles of sneezing, coughing, talking, and breathing, those remain active in the air for a long time. The aerosol has two distinct sizes. The first size varies between 0.25 and 1 μm and the second size is bigger than 2.5 μm. In recent studies on SARS-CoV-2, aerosol remains active in the air for up to 3 hours, which is very much similar to SARS-CoV and MERS-CoV [66].
3 Diagnostic for SARS-CoV-2
The seventh coronavirus, SARS-CoV-2, can infect humans (HCoV). Its infections can be asymptomatic or severe which in case of early diagnosis can aid clinical management and hence control the outbreaks. The diagnostic detects the virus itself or detects the response of the immune system to the infection, that is, antibodies, biomarkers, etc. In Table 1, symptoms of different viral disease are compared. Table 2 mentions the systematic disorders and respiratory disorders caused by COVID-19 infections.
3.1 SARS-CoV-2 RNA Detection
It is based on detecting the unique viral sequences using nucleic acid amplification tests (NAATs), like real-time reverse-transcription polymerase chain reaction abbreviated as rRT-PCR.
Once someone is infected by the virus, it takes some time known as incubation period to develop symptoms (about a week), with a range of 1 and 14 days after exposure and can be detected 1–3 days before the symptom onset in the upper respiratory tract (URT) [9, 51]. In URT, the concentration of the virus is highest at the time of symptom onset and it declines thereafter. The presence of viral RNA is increased during the second week of the illness in the lower respiratory tract (LRT) [67]. In some patients, it is detectable for many days, while in others for several weeks or months but a prolonged presence does not signify a prolonged infectiousness.
Respiratory secretions vary in composition, and sampling adequacy may also vary, resulting in false-negative PCR results. Viral RNA is detected in LRT secretions for the patient who is suspected of SARS-CoV-2 infection and the URT swab is negative. Rectal swabs or feces were positive for SARS-CoV-2 RNA in some of the patients, and some studies suggest that the positivity is prolonged in comparison to that of respiratory specimens. In some cases, SARS-CoV-2 RNA is detected in blood samples, and some studies show that it depends on the severity of the disease, but more studies on this are needed. In oral fluid specimens, that is, induced saliva, the detection rates vary widely when compared with URT specimens of that very same patient, and as of now very limited data is available on detection adequacy of the virus in mouth washes [68]. SARS-CoV-2 was detected in ocular fluids, urine, and semen samples of some of the patients, while positive RNA detection for cerebrospinal fluid and brain tissue has also been reported.
To conclude, SARS-CoV-2 can be detected in respiratory material and at the same time in other body fluids and compartments, but the respiratory samples are taken for diagnostics as the virus is most frequently detected in respiratory material.
Rapid collections of the specimens as well as accurate diagnosis are required to support clinical management of the patients and infection controlling measures to be taken. The trained and competent operators should collect and perform laboratory diagnosis because of the complexity of sampling, analysis done in laboratory, and interpretation of the final results. Those infected with SARS-CoV-2 may or may not develop symptoms, and the most robust evidence of infection may be detected from the fragments of the virus, nucleic acids, and proteins through virological testing. Reports show the cases of infection of COVID-19 with other pathogens either, so if other pathogens test positive, it does not rule out SARS-CoV-2 infection and vice versa.
3.2 Adequate Specimen Collection
While specimen collection, testing, storage, and research, adequate standard operating procedures (SOP) should be there and staff must be trained for all this. Testing combined nasopharyngeal and oropharyngeal swabs has been shown to enhance sensitivity in detection of viruses and the reliability of the result has improved. Swabs from two individuals can be combined, and nasopharyngeal and oropharyngeal swabs may be taken, but some studies show that nasopharyngeal swabs provide results more reliable as compared to oropharyngeal swabs. In mild or asymptomatic cases, the upper respiratory specimens are suitable for the test only in the early-stage of infections. Lower respiratory specimens are collected from patients having a negative URT sampling or if sample collection is delayed. Because of high risk of aerosolization, caution should be exercised and strictly abiding by IPC procedures is required during collection of the samples. Other oral or respiratory fluids sampling procedures must first clear validation test in the laboratory before implementation for the targeted groups of the patients.
3.3 Simplified Specimen Collection
Some studies on combined oropharyngeal and nares/nasal swab, midturbinate or lower nasal or nares swabs or tongue swab by a trained staff or by self-sampling show that these approaches perform well, but their sample sizes are limited [69, 70]. Further assessment as well as validation is required before implementation of these alternatives and to determine for what purpose these collection methods can serve as alternatives. In case of elderly people and young child, oral fluids can be a suitable specimen in schools or nursing homes for mass screening as collecting nasopharyngeal and oropharyngeal swabs are problematic as compared with URT specimens [71, 72].
Oral fluid collection methods have a wide range of the performances compared with naso- and/or oropharyngeal sample collection; they also show variations from posterior oropharyngeal fluids/saliva that may be collected by spitting or drooling, with pipet or special sponges, and gargling with saline solutions is another alternative.
3.4 Serum Specimens
A person in whom COVID-19 infection is strongly suspected but obtained NAAT results are negative, a paired serum specimen can be collected. First specimen may be collected in the acute phase and the other one in the convalescent phase which can be used to test for seroconversion or an increase in antibody titers.
3.5 Fecal Specimens
While testing feces make sure that for this type of sample, the extraction method and NAAT have been validated. If URT and LRT both are tested negative but clinical suspicion on COVID-19 infection is there, NAAT can be considered for fecal specimens [73].
3.6 Postmortem Specimens
The postmortem swab like needle biopsy or specimens from tissues of the dead body from the autopsy may be taken which also includes lung tissues for pathological and microbiological tests [74].
After collection, the specimens must reach the laboratory quickly and a correct specimen handling in the laboratory and during transportation is very essential.
4 Testing of COVID-19 (SARS-CoV-2)
4.1 Nucleic Acid Amplification Test (NAAT)
NAAT such as rRT-PCR should be used to test suspected COVID-19 infections and SARS-CoV-2 genome should be targeted by such tests. Optimal diagnostics has a NAAT assay and minimum of two independent targets on SARS-CoV-2 genome, but a simple algorithm may be adopted in widespread transmission areas. While using one-target assay, a strategy should be in place so as to monitor the mutations which may affect the overall performance. Some of the NAAT systems are having the capacity for fully automated tests by which sample processing, the capacity for RNA extraction, amplification, and reporting is integrated. These systems provide access of tests in areas having limited laboratory capacity with rapid turnaround time. The amplification/detection methods, like isothermal nucleic acid amplification technologies, clustered regularly interspaced short palindromic repeats (CRISPR), and molecular microarray assays are going to be commercialized [75,76,77].
Some negative test results do not actually rule out infection because there are factors that can lead to a negative result even if the person is infected [78, 79]. Such factors are:
-
The specimen of poor quality
-
The specimen was collected from a body compartment which was not containing SARS-CoV-2 at that time
-
Handling or shipping of the specimen was not appropriately done, etc.
Figure 4 is a pictorial representation of diagnostic flow of SARS-CoV-2 infection detection.
4.2 NAAT by Pooling Specimens
Diagnostic capacity can be increased by pooling of samples from different patients to detect SARS-CoV-2 if the testing rate is not meeting the demands [80,81,82]. Specimen pooling may be considered in population groups having very low prevalence of COVID-19 infection. Adequate automation is the key to reliable pooling (software-based algorithms, robotic systems, and laboratory middle-ware which works with sample pooling).
Strategies for pooling the specimens [83, 84] are:
-
The individual specimens are treated to be negative if the result of the pool is negative.
-
Strategies may be different in case of positive pool tests but every specimen must be tested (pool deconvolution) generally to find out positive specimen(s).
-
In matrix pooling, pool is made per row and per column and finally tested using PCR.
4.3 Testing Antibodies
In a response to the infection, the human body produces antibodies which are detected by serological assays. SARS-CoV-2 is a novel pathogen and the understanding of the antibody response is in emerging phase, hence the antibody detection tests should not be used to detect acute infections [85, 86]. Lateral flow antibody detection assays are not suitable for diagnosis because they cannot detect the increase in antibody titers, while (semi)quantitative or quantitative assays can. Antibodies can be detected in the first week of illness in some of the patients while in others with subclinical/mild infection; it can take weeks to develop. The duration of the antibodies generated is still under study, hence serology is not an alternate for virological assays and the presence of antibodies does not ensure that they offer protective immunity. The target viral protein and timing of the testing affect performance of serologic assays in various testing groups (mild disease vs moderate-to-severe disease or young patient vs old patient) [87, 88].
4.4 Antigen Detection (Rapid Diagnostic Tests—RDTs)
The presence of SARS-CoV-2 is tested in viral proteins (antigens) collected from respiratory specimens for rapid (within 30 minutes) diagnostic tests. False-positive (showing a person is infected when actually he is not) results may be obtained if the antibodies also recognize antigens of other human coronaviruses which are non-SARS-CoV-2 viruses. Pairing of NAAT and antigen test validations are being encouraged in clinical studies. High viral loads improve antigen test performance, but antigen RDTs can be implemented in diagnostic algorithms only if the test performance is acceptable.
5 Treatments for SARS-CoV-2
Most of the COVID-19 patients are able to recover at home but at the same time scientists are leaving no stone unturned to develop an effective treatment. Therapies that are being investigated include the drugs that are used to treat autoimmune diseases, antibodies of recovered patients from COVID-19, and additional antiviral drugs.
5.1 Monoclonal Antibodies
FDA (Food and Drug Administration) granted emergency use authorization (EUA) for treatment named as Bamlanivimab which is a monoclonal antibody [89]. It is approved to treat non-hospitalized, recently tested positive for COVID-19 children and adults having mild-to-moderate symptoms but are at risk of severe COVID-19 disease. Within 10 days of symptom development, the patient should be given a single dose of the treatment.
5.2 Chloroquine and Hydroxychloroquine
Chloroquine has been reported as a broad-spectrum antiviral drug which is a commonly used autoimmune disease drug and anti-malarial drug. Chloroquine controls virus infection by interfering in the glycosylation of cellular receptors related to SARS-CoV and increases endosomal pH needed for virus fusion. Chloroquine was recommended for the prevention as well as treatment of COVID-19 pneumonia [90]. Hydroxychloroquine, an analog of chloroquine, has lesser concerns related to drug–drug interactions and has been found to be more effective than chloroquine for Vero cells infected by SARS-CoV-2. Both chloroquine and hydroxychloroquine have immunomodulatory effects and are capable of suppressing the immune response.
5.3 Plasma Transfusion
For SARS treatment, convalescent plasma was administered very early after symptom onset, and the pooled odd of mortality treatment was reduced compared with no therapy. For COVID-19 treatment, the National Health Commission of China appealed convalescent patients for donating blood [42]. Plasma should be collected within 2 weeks after recovery from COVID-19 which will ensure high neutralization; this complexity of obtaining plasma limits its clinical application.
5.4 Corticosteroids
Though corticosteroids were not recommended for viral pneumonia or acute respiratory distress syndrome (ARDS) but in case of severe CARDS such drugs are generally used.
Recently, a large-size RCT (the RECOVERY trial) shows that dexamethasone decreased the death rates by one-third in critical COVID-19 patients [91].
5.5 Vaccines
Vaccines deliver immunogen (specific type of antigen) to train immune system to recognize the pathogens like bacteria or viruses which reduces the risk of infections. Injections in muscles or under skin and oral route are different methods of administering vaccines. More than one dose of vaccine is given to build complete immunity, as a “booster” in case of immunity wears off and against the disease that is a seasonal disease like the yearly flue. Table 3 shows the technologies used for viral vaccines with their advantages and disadvantages.
6 Vaccine Development
Infectious diseases are prevented by vaccines. Some of the diseases that are vaccine preventable are polio, measles, Hepatitis B, influenza, and many more. The ability of the pathogen to spread can be limited by “herd” or “indirect” or “population” immunity developed when most of the people are vaccinated against a disease. When many people have immunity, the people who cannot be vaccinated or have low immunity like young babies are protected by infectious disease. Peptide and nucleic acid like new technologies for human vaccines are being used along with the conventional and well-known technologies to develop the vaccines [91,92,93]. Table 4 shows the mechanism of actions for different types of vaccines, while Table 5 represents the steps taken in effective and safe vaccine development.
As of 2 October 2020, out of 42 candidate vaccines of COVID-19 which are in clinical evaluation, 10 are in Phase-III trials. In preclinical evaluation, 151 candidate vaccines are there [93]. Most of the candidate vaccines are for intramuscular injection and designed for a two-dose schedule. The WHO has launched a coordinated international Phase-III trial of candidate vaccines for speedy evaluation and to ensure that the vaccines are tested in different populations. Table 6 is a list of the COVID-19 vaccines, which are in Phase-III clinical trial, along with the locations of their trials.
As of 4 October 2020, the USA, India, Brazil, Russian Federation, and Colombia are five countries which have highest number of COVID-19 cases. The countries having highest number of deaths are the USA, Brazil, India, Mexico, and the United Kingdom [93].
7 Conclusions
In spite of day and night efforts done by researchers, doctors, and other medical staff, everything about this disease is uncertain, be it the symptoms, the outcomes of the tests, or the success rate of the under trial vaccines. The shortage of resources and facilities as compared to the patients are the main reasons behind the surrender of medical infrastructure of a country. So a collaborative international effort is required to face the COVID-19 pandemic and establish pathways to manage this crisis. To face a pandemic like COVID-19, none of us was prepared so the patients, their relatives, and finally the community must be provided authentic and understandable information to face this devil. This paper presents state-of-the-art research work going on SARS-CoV-2 virus from its origin, its mutations, and diagnosis to clinical trials on vaccine developments.
References
Lu, G., Wang, Q., & Gao, G. F. (2015). Bat-to-human: Spike features determining “host jump” of coronaviruses SARS-CoV, MERS-CoV, and beyond. Trends in Microbiology, 23, 468–478.
Nieto-Torres, J. L., Dediego, M. L., Verdia-Baguena, C., et al. (2014). Severe acute respiratory syndrome coronavirus envelope protein ion channel activity promotes virus fitness and pathogenesis. PLoS Pathogens, 10, e1004077. https://doi.org/10.1371/journal.ppat.1004077.
Huang, C., Wang, Y., Li, X., Ren, L., Zhao, J., Hu, Y., et al. (2020). Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet, 395(10223), 497–506. https://doi.org/10.1016/S0140-6736(20)30183-5.
https://experience.arcgis.com/experience/685d0ace521648f8a5beeeee1b9125cd
Gu, J., Han, B., & Wang, J. (2020). COVID-19: Gastrointestinal manifestations and potential fecal-oral transmission. Gastroenterology, 158(6), 1518–1519.
World Health Organization. (2014). Infection prevention and control of epidemic- and pandemic-prone acute respiratory infections in health care. Geneva: World Health Organization. Available from: https://apps.who.int/iris/bitstream/handle/10665/112656/9789241507134_eng.pdf?sequence=1.
Liu, J., Liao, X., Qian, S., et al. (2020). Community transmission of severe acute respiratory syndrome coronavirus 2, Shenzhen, China, 2020. Emerging Infectious Diseases, 26(6), 1320–1323. https://doi.org/10.3201/eid2606.200239.
Chan, J., Yuan, S., Kok, K., et al. (2020). A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet, 395(10223), 514–523. https://doi.org/10.1016/S0140-6736(20)30154-9.
Li, Q., Guan, X., Wu, P., Wang, X., Zhou, L., Tong, Y., Ren, R., Leung, K. S., Lau, E. H., Wong, J. Y., & Xing, X. (2020). Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. The New England Journal of Medicine, 382, 1199–1207. https://doi.org/10.1056/NEJMoa2001316.
Li, H., Liu, S. M., Yu, X. H., Tang, S. L., & Tang, C. K. (2020). Coronavirus disease 2019 (COVID-19): Current status and future perspective. International Journal of Antimicrobial Agents, 55(5), 105951.
Zhu, N., Zhang, D., Wang, W., Li, X., Yang, B., Song, J., et al. (2020). A novel coronavirus from patients with pneumonia in China, 2019. The New England Journal of Medicine, 382, 727–733.
Lu, R., Zhao, X., Li, J., Niu, P., Yang, B., Wu, H., et al. (2020). Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet, 395(10224), 565–574.
Song, Z., Xu, Y., Bao, L., et al. (2019). From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses, 11(1), 59.
Graham, R. L., Donaldson, E. F., & Baric, R. S. (2013). A decade after SARS: Strategies for controlling emerging coronaviruses. Nature Reviews Microbiology, 11, 836–848.
Zumla, A., Hui, D. S., & Perlman, S. (2015). Middle East respiratory syndrome. Lancet, 386, 995–1007.
Hui, D. S., Azhar, E. I., Kim, Y. J., et al. (2018). Middle East respiratory syndrome coronavirus: Risk factors and determinants of primary, household, and nosocomial transmission. The Lancet Infectious Diseases, 18, e217–e227.
Reusken, C. B., Haagmans, B. L., Muller, M. A., et al. (2013). Middle East respiratory syndrome coronavirus neutralising serum antibodies in dromedary camels: A comparative serological study. The Lancet Infectious Diseases, 13, 859–866.
de Wit, E., van Doremalen, N., Falzarano, D., et al. (2016). SARS and MERS: Recent insights into emerging coronaviruses. Nature Reviews Microbiology, 14, 523–534.
de Groot, R. J., Baker, S. C., Baric, R., Enjuanes, L., Gorbalenya, A. E., Holmes, K. V., Perlman, S., Poon, L., Rottier, P. J. M., Talbot, P. J., & Woo, P. C. Y. (2012). Family Coronaviridae. In Virus taxonomy (pp. 806–828). Amsterdam: Elsevier Academic Press.
Yang, D., & Leibowitz, J. L. (2015). The structure and functions of coronavirus genomic 3′ and 5′ ends. Virus Research, 206, 120–133.
Nakagawa, K., Lokugamage, K. G., & Makino, S. (2016). Viral and cellular mRNA translation in coronavirus-infected cells. In Advances in virus research (Vol. 96, pp. 165–192). Amsterdam: Academic Press.
Ramaiah, A., & Arumugaswami, V. (2020). Insights into cross-species evolution of novel human coronavirus 2019-nCoV and defining immune determinants for vaccine development. bioRxiv, 1–15. https://doi.org/10.1101/2020.01.29.925867.
Wu, A., Peng, Y., Huang, B., et al. (2020). Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host and Microbe, 27(3), 325–328.
Barcena, M., Oostergetel, G. T., Bartelink, W., et al. (2009). Cryo-electron tomography of mouse hepatitis virus: Insights into the structure of the coronavirion. Proceedings of the National Academy of Sciences of the United States of America, 106, 582–587.
Neuman, B. W., Adair, B. D., Yoshioka, C., et al. (2006). Supramolecular architecture of severe acute respiratory syndrome coronavirus revealed by electron cryomicroscopy. Journal of Virology, 80, 7918–7928.
Mousavizadeh, L., & Ghasemi, S. (2021). Genotype and phenotype of COVID-19: Their roles in pathogenesis. Journal of Microbiology, Immunology and Infection, 54(2), 159–163.
Abraham, S., Kienzle, T. E., Lapps, W., et al. (1990). Deduced sequence of the bovine coronavirus spike protein and identification of the internal proteolytic cleavage site. Virology, 176, 296–301.
Luytjes, W., Sturman, L. S., Bredenbeek, P. J., et al. (1987). Primary structure of the glycoprotein E2 of coronavirus MHV-A59 and identification of the trypsin cleavage site. Virology, 161, 479–487.
Beniac, D. R., Andonov, A., Grudeski, E., et al. (2006). Architecture of the SARS coronavirus prefusion spike. Nature Structural & Molecular Biology, 13, 751–752. https://doi.org/10.1038/nsmb1123.
Delmas, B., & Laude, H. (1990). Assembly of coronavirus spike protein into trimers and its role in epitope expression. Journal of Virology, 64, 5367–5375.
Walls, A. C., Xiong, X., Park, Y. J., et al. (2019). Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell, 176, 1026–1039.e1015.
Wrapp, D., Wang, N., Corbett, K. S., et al. (2020). Cryo-EM structure of the 2019-nCoV spike in the refusion conformation. Science, 367(6483), 1260–1263.
Armstrong, J., Niemann, H., Smeekens, S., et al. (1984). Sequence and topology of a model intracellular membrane protein, E1 glycoprotein, from a coronavirus. Nature, 308, 751–752.
Neuman, B. W., Kiss, G., Kunding, A. H., et al. (2011). A structural analysis of M protein in coronavirus assembly and morphology. Journal of Structural Biology, 174, 11–22. https://doi.org/10.1016/j.jsb.2010.11.021.
Godet, M., L’Haridon, R., Vautherot, J. F., et al. (1992). TGEV corona virus ORF4 encodes a membrane protein that is incorporated into virions. Virology, 188, 666–675.
Chang, C. K., Sue, S. C., Yu, T. H., et al. (2006). Modular organization of SARS coronavirus nucleocapsid protein. Journal of Biomedical Science, 13, 59–72. https://doi.org/10.1007/s11373-005-9035-9.
Stohlman, S. A., Baric, R. S., Nelson, G. N., et al. (1988). Specific interaction between coronavirus leader RNA and nucleocapsid protein. Journal of Virology, 62, 4288–4295.
Molenkamp, R., & Spaan, W. J. (1997). Identification of a specific interaction between the coronavirus mouse hepatitis virus A59 nucleocapsid protein and packaging signal. Virology, 239, 78–86.
Kuo, L., & Masters, P. S. (2013). Functional analysis of the murine coronavirus genomic RNA packaging signal. Journal of Virology, 87, 5182–5192. https://doi.org/10.1128/JVI.00100-13.
Kan, B., Wang, M., Jing, H., Xu, H., Jiang, X., Yan, M., et al. (2005). Molecular evolution analysis and geographic investigation of severe acute respiratory syndrome coronavirus-like virus in palm civets at an animal market and on farms. Journal of Virology, 79(18), 11892–11900.
Gralinski, L. E., & Menachery, V. D. (2020). Return of the coronavirus: 2019-nCoV. Viruses, 12(2), 135.
Zhou, P., Yang, X. L., Wang, X. G., Hu, B., Zhang, L., Zhang, W., Si, H. R., Zhu, Y., Li, B., Huang, C. L., & Chen, H. D. (2020). A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature, 579(7798), 270–273. https://doi.org/10.1038/s41586-020-2012-7.
Hui, D. S., Azhar, E. I., Madani, T. A., et al. (2020). The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health – The latest 2019 novel coronavirus outbreak in Wuhan, China. International Journal of Infectious Diseases, 91, 264–266.
Tripathi, A., Jain, A., Mishra, K. K., Pandey, A. B., & Vashist, P. C. (2020). MCNN: A deep learning based rapid diagnosis method for COVID-19 from the X-ray images. Revue d’Intelligence Artificielle, 34(6), 673–682.
Ren, L. L., Wang, Y. M., Wu, Z. Q., Xiang, Z. C., Guo, L., Xu, T., et al. (2020). Identification of a novel coronavirus causing severe pneumonia in human: A descriptive study. Chinese Medical Journal, 133(9), 1015–1024. https://doi.org/10.1097/CM9.0000000000000722.
Wang, W., Tang, J., & Wei, F. (2020). Updated understanding of the outbreak of 2019 novel coronavirus (2019-nCoV) in Wuhan, China. Journal of Medical Virology, 92(4), 441–447. https://doi.org/10.1002/jmv.25689.
Carlos, W. G., Dela Cruz, C. S., Cao, B., Pasnick, S., & Jamil, S. (2020). Novel Wuhan (2019-nCoV) coronavirus. American Journal of Respiratory and Critical Care Medicine, 201(4), 7–8. https://doi.org/10.1164/rccm.2014P7.
Holshue, M. L., DeBolt, C., Lindquist, S., Lofy, K. H., Wiesman, J., Bruce, H., et al. (2020). First case of 2019 novel coronavirus in the United States. The New England Journal of Medicine, 382, 929–936. https://doi.org/10.1056/NEJMoa2001191.
Liu, K., Fang, Y. Y., Deng, Y., Liu, W., Wang, M. F., Ma, J. P., et al. (2020). Clinical characteristics of novel coronavirus cases in tertiary hospitals in Hubei Province. Chinese Medical Journal, 133(9), 1025–1031. https://doi.org/10.1097/CM9.0000000000000744.
Backer, J. A., Klinkenberg, D., & Wallinga, J. (2020). Incubation period of 2019 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20–28 January 2020. Euro Surveillance, 25(5), 2000062.
Lauer, S. A., Grantz, K. H., Bi, Q., et al. (2020). The incubation period of 2019-nCoV from publicly reported confirmed cases: Estimation and application. medRxiv. https://www.medrxiv.org/content/10.1101/2020.02.02.20020016v1 2020
Varia, M., Wilson, S., Sarwal, S., et al. (2003). Investigation of a nosocomial outbreak of severe acute respiratory syndrome (SARS) in Toronto, Canada. CMAJ, 169, 285–292.
Assiri, A., Al-Tawfiq, J. A., Al-Rabeeah, A. A., et al. (2013). Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: A descriptive study. The Lancet Infectious Diseases, 13, 752–761.
Wang, D., Hu, B., Hu, C., et al. (2020). Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA, 323(11), 1061–1069.
Guan, W. J., Ni, Z. Y., Hu, Y., et al. (2020). Clinical characteristics of coronavirus disease 2019 in China. The New England Journal of Medicine, 382, 1708–1720.
Chiu, W. K., Cheung, P. C., Ng, K. L., et al. (2003). Severe acute respiratory syndrome in children: Experience in a regional hospital in Hong Kong. Pediatric Critical Care Medicine, 4, 279–283.
Donnelly, C. A., Ghani, A. C., Leung, G. M., et al. (2003). Epidemiological determinants of spread of causal agent of severe acute respiratory syndrome in Hong Kong. Lancet, 361, 1761–1766.
Leung, G. M., Hedley, A. J., Ho, L.-M., & Chau, P. (2004). The epidemiology of severe acute respiratory syndrome in the 2003 Hong Kong epidemic: An analysis of all 1755 patients. Annals of Internal Medicine, 141, 662–673.
Wang, W., Xu, Y., Ruqin, G., et al. (2020). Detection of SARS-CoV-2 in different types of clinical specimens. JAMA, 323(18), 1843–1844. https://doi.org/10.1001/jama.2020.3786.9.
Lauer, S. A., Grantz, K. H., Bi, Q., et al. (2020). The incubation period of coronavirus disease 2019 (COVID-19) from publicly reported confirmed cases: Estimation and application. Annals of Internal Medicine, 172(9), 577–582. https://doi.org/10.7326/M20-0504.10.
Liu, Y., Yan, L. M., Wan, L., et al. (2020). Viral dynamics in mild and severe cases of CVOID-19. The Lancet Infectious Diseases, 20(6), 656–657. https://doi.org/10.1016/S1473-3099(20)30232-2.
Wolfel, R., Corman, V., Guggemos, W., et al. (2020). Virological assessment of hospitalized cases of coronavirus disease. Nature, 581, 465–469. https://doi.org/10.1101/2020.03.05.20030502.
Centers for Disease Control and Prevention. (2020). 2019 novel coronavirus. https://www.cdc.gov/coronavirus/2019-ncov/about/transmission.html
Weiss, A., Jellingsø, M., & Sommer, M. O. A. (2020). Spatial and temporal dynamics of SARS-CoV-2 in COVID-19 patients: A systematic review and meta-analysis. eBioMedicine, 58, 102916.
Guo, W. L., Jiang, Q., Ye, F., Li, S. Q., Hong, C., Chen, L. Y., & Li, S. Y. (2020). Effect of throat washings on detection of 2019 novel coronavirus. Clinical Infectious Diseases, 71(8), ciaa416. https://doi.org/10.1093/cid/ciaa416.
Vlek, A. L. M., Wesselius, T. S., Achterberg, R., & Thijsen, S. F. T. (2021). Combined throat/nasal swab sampling for SARS-CoV-2 is equivalent to nasopharyngeal sampling. European Journal of Clinical Microbiology & Infectious Diseases, 40(1), 193–195.
LeBlanc, J. J., Heinstein, C., MacDonald, J., Pettipas, J., Hatchette, T. F., & Patriquin, G. (2020). A combined oropharyngeal/nares swab is a suitable alternative to nasopharyngeal swabs for the detection of SARS-CoV-2. Journal of Clinical Virology, 128, 104442.
Tu, Y. P., Jennings, R., Hart, B., Cangelosi, G. A., Wood, R. C., Wehber, K., Verma, P., Vojta, D., & Berke, E. M. (2020). Swabs collected by patients or health care workers for SARS-CoV-2 testing. The New England Journal of Medicine, 383(5), 494–496.
Altamirano, J., Govindarajan, P., Blomkalns, A. L., Kushner, L. E., Stevens, B. A., Pinsky, B. A., & Maldonado, Y. (2020). Assessment of sensitivity and specificity of patient-collected lower nasal specimens for sudden acute respiratory syndrome coronavirus 2 testing. JAMA Network Open, 3(6), e2012005.
Ng, S. C., Chan, F. K., & Chan, P. K. (2020). Screening FMT donors during the COVID-19 pandemic: A protocol for stool SARS-CoV-2 viral quantification. The Lancet. Gastroenterology & Hepatology, 5(7), 642–643.
Tang, J. W., To, K. F., Lo, A. W., Sung, J. J., Ng, H. K., & Chan, P. K. (2007). Quantitative temporal-spatial distribution of severe acute respiratory syndrome-associated coronavirus (SARS-CoV) in post-mortem tissues. Journal of Medical Virology, 79(9), 1245–1253.
Carter, L. J., Garner, L. V., Smoot, J. W., Li, Y., Zhou, Q., Saveson, C. J., Sasso, J. M., Gregg, A. C., Soares, D. J., Beskid, T. R., & Jervey, S. R. (2020). Assay techniques and test development for COVID-19 diagnosis. ACS Central Science, 6(5), 591–605.
World Health Organization. (2020). Diagnostic testing for SARS-CoV-2: Interim guidance, 11 September 2020 (No. WHO/2019-nCoV/laboratory/2020.6). Geneva: World Health Organization.
Esbin, M. N., Whitney, O. N., Chong, S., Maurer, A., Darzacq, X., & Tjian, R. (2020). Overcoming the bottleneck to widespread testing: A rapid review of nucleic acid testing approaches for COVID-19 detection. RNA, 26(7), 771–783.
Zou, L., et al. (2020). SARS-CoV-2 viral load in upper respiratory specimens of infected patients. The New England Journal of Medicine, 382(12), 1177–1179.
Young, B. E., Ong, S. W. X., Kalimuddin, S., Low, J. G., Tan, S. Y., Loh, J., Ng, O. T., Marimuthu, K., Ang, L. W., Mak, T. M., & Lau, S. K. (2020). Epidemiologic features and clinical course of patients infected with SARS-CoV-2 in Singapore. JAMA, 323(15), 1488–1494.
Yelin, I., Aharony, N., Shaer-Tamar, E., Argoetti, A., Messer, E., Berenbaum, D., Shafran, E., Kuzli, A., Gandali, N., Hashimshony, T., & Mandel-Gutfreund, Y. (2020). Evaluation of COVID-19 RT-qPCR test in multi-sample pools. medRxiv.
Sawarkar, S. S., Victor, A., Viotti, M., Haran, S. P., Verma, S., Griffin, D., & Sams, J. (2020). Sample pooling, a population screening strategy for SARS-CoV2 to prevent future outbreak and mitigate the second-wave of infection of the virus. medRxiv.
Abdalhamid, B., Bilder, C. R., McCutchen, E. L., Hinrichs, S. H., Koepsell, S. A., & Iwen, P. C. (2020). Assessment of specimen pooling to conserve SARS CoV-2 testing resources. American Journal of Clinical Pathology, 153(6), 715–718.
Aragón-Caqueo, D., Fernández-Salinas, J., & Laroze, D. (2020). Optimization of group size in pool testing strategy for SARS-CoV-2: A simple mathematical model. Journal of Medical Virology, 92(10), 25929. https://doi.org/10.1002/jmv.25929.
Ben-Ami, R., Klochendler, A., Seidel, M., Sido, T., Gurel-Gurevich, O., Yassour, M., Meshorer, E., Benedek, G., Fogel, I., Oiknine-Djian, E., & Gertler, A. (2020). Large-scale implementation of pooled RNA extraction and RT-PCR for SARS-CoV-2 detection. Clinical Microbiology and Infection, 26(9), 1248–1253.
World Health Organization. (2020, April 8). Advice on the use of point-of-care immunodiagnostic tests for COVID-19. Available from: https://apps.who.int/iris/handle/10665/331713
World Health Organization. (2020, July 27). The unity studies: Early investigations protocols. Available from: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/early-investigations
Okba, N. M., Müller, M. A., Li, W., Wang, C., Geurtsvan Kessel, C. H., Corman, V. M., Lamers, M. M., Sikkema, R. S., de Bruin, E., Chandler, F. D., & Yazdanpanah, Y. (2020). Severe acute respiratory syndrome coronavirus 2−specific antibody responses in coronavirus disease patients. Emerging Infectious Diseases, 26(7), 1478–1488.
Lou, B., Li, T. D., Zheng, S. F., Su, Y. Y., Li, Z. Y., Liu, W., Yu, F., Ge, S. X., Zou, Q. D., Yuan, Q., & Lin, S. (2020). Serology characteristics of SARS-CoV-2 infection since exposure and post symptom onset. European Respiratory Journal, 56(2), 2000763.
https://www.medicinenet.com/monoclonal_antibodies/article.htm#what_are_human_ monoclonal_antibodies
Vincent, M. J., Bergeron, E., Benjannet, S., Erickson, B. R., Rollin, P. E., Ksiazek, T. G., Seidah, N. G., & Nichol, S. T. (2005). Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virology Journal, 2(1), 1–10.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Tripathi, A., Pandey, A.B., Singh, A.K., Jain, A., Tyagi, V., Vashist, P.C. (2022). Diagnosis for COVID-19. In: Pani, S.K., Dash, S., dos Santos, W.P., Chan Bukhari, S.A., Flammini, F. (eds) Assessing COVID-19 and Other Pandemics and Epidemics using Computational Modelling and Data Analysis. Springer, Cham. https://doi.org/10.1007/978-3-030-79753-9_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-79753-9_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-79752-2
Online ISBN: 978-3-030-79753-9
eBook Packages: Computer ScienceComputer Science (R0)