Abstract
Unlike the intermediate filament- and septin-based cytoskeletons which are apolar structures, the microtubule (MT) and actin cytoskeletons are polarized structures in mammalian cells and tissues including the testis, most notable in Sertoli cells. In the testis, these cytoskeletons that stretch across the epithelium of seminiferous tubules and lay perpendicular to the basement membrane of tunica propria serve as tracks for corresponding motor proteins to support cellular cargo transport. These cargoes include residual bodies, phagosomes, endocytic vesicles and most notably developing spermatocytes and haploid spermatids which lack the ultrastructures of motile cells (e.g., lamellipodia, filopodia). As such, these developing germ cells require the corresponding motor proteins to facilitate their transport across the seminiferous epithelium during the epithelial cycle of spermatogenesis. Due to the polarized natures of these cytoskeletons with distinctive plus (+) and minus (−) end, directional cargo transport can take place based on the use of corresponding actin- or MT-based motor proteins. These include the MT-based minus (−) end directed motor proteins: dyneins, and the plus (+) end directed motor proteins: kinesins, as well as the actin-based motor proteins: myosins, many of which are plus (+) end directed but a few are also minus (−) end directed motor proteins. Recent studies have shown that these motor proteins are essential to support spermatogenesis. In this review, we briefly summarize and evaluate these recent findings so that this information will serve as a helpful guide for future studies and for planning functional experiments to better understand their role mechanistically in supporting spermatogenesis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
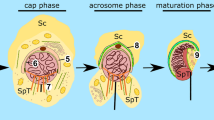
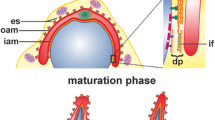
The blood-testis barrier (BTB) in the adult mammalian testis is a unique blood-tissue barrier which restricts paracellular (between cells; i.e., gate-keeper function of the BTB) and transcellular (across cells; i.e., fence function of the BTB) transport (or diffusion) of water, electrolytes, nutrients, cytokines and biomolecules including paracrine and autocrine factors between adjacent Sertoli cells at the base of the seminiferous tubules, also known as the Sertoli cell barrier [1,2,3,4,5,6] (Fig. 1). Interestingly, microvessels found in the interstitial space between seminiferous tubules contribute relatively little barrier function to the BTB in the testis of rodents, primates and humans (Fig. 1) [5, 9]. The BTB also divides the seminiferous epithelium into the basal and the adluminal (apical) compartments as noted in Fig. 1. As such, meiosis I/II and all the cellular events pertinent to post-meiotic development take place behind the BTB in a specialized microenvironment (Fig. 1), whereas mitotic proliferation of spermatogonia and differentiation/transformation of type A and type B spermatogonia to earlier spermatocytes take place in the basal compartment [10,11,12]. The BTB is a highly dynamic blood-tissue barrier since preleptotene spermatocytes, once derived from type B spermatogonia in the basal compartment rodents, are to be transported across the BTB in late Stage VII through early Stage IX of the epithelial cycle while differentiating into leptotene spermatocytes, which can be transformed into zygotene and pachytene spermatocytes to prepare for meiosis. Studies have shown that the BTB in the rodent testis is constituted by the actin-based tight junction (TJ) between adjacent Sertoli cells, reinforced by a testis-specific actin-rich adherens junction (AJ) type called basal ectoplasmic specialization (ES), and supported by the actin-based gap junction, but also intermediate filament-based desmosome [13,14,15,16,17,18,19]. Once haploid spermatids are formed through meiosis, they are also being transported across the seminiferous epithelium in the adluminal compartment before fully developed step 19, 16, and 12 spermatids in the testis of rats, mice, and humans, respectively, are transformed to spermatozoa via spermiogenesis [12, 14, 20] as these cells are lacking the ultrastructures found in motile cells, namely the lamellipodia and filopodia [21]. Spermatozoa are then line-up at the edge of the seminiferous tubule lumen to undergo spermiation in Stage VIII of the epithelial cycle in rodents versus VI in humans, respectively, which is composed of a tightly regulated series of biochemical and cellular events involving multiple signal and regulatory proteins [13, 22,23,24]. The testicular sperm emptied into the epididymis are then undergo another series of maturation processes, rendering them capable of fertilizing the egg.
Schematic drawing of the cross-section of a typical Stage VII seminiferous tubule from adult rat testes. The seminiferous epithelium across the tubule is constituted by adjacent Sertoli cells which, in turn, support germ cells at different stages of their development during spermatogenesis as noted herein with a Sertoli:germ ratio of about 1:30–50 [7, 8] (left panel). Between adjacent Sertoli cells near the basement membrane are the specialized junctions, namely the actin-based tight junction (TJ), basal ES (ectoplasmic specialization, a testis-specific adherens junction type) and gap junction, which together with the intermediate filament-based desmosome constitute the blood-testis barrier (BTB). The BTB also divides the seminiferous epithelium in the rat testis and other mammalian testes into the basal and the adluminal (apical) compartments, such that meiosis I/II and all subsequent events of post-meiotic spermatid development take place behind the BTB. The most notable structural features are the microtubule (MT)-based and actin-based tracks that stretch across the seminiferous epithelium. These tracks support the corresponding MT- and actin-based motor proteins (see the insets on the right panel) to provide cellular transport of cargoes as discussed in this review. For F-actin, besides serving as the track-like structures to support cellular transport, actin filaments that lay parallel to the Sertoli cell plasma membrane are assembled as bundles which appeared as aggregates of rod-like structures in cross-sections of the testis, both at the Sertoli-spermatid site called apical ES and also at the Sertoli cell-cell interface at the BTB called basal ES. The ES is not only an important ultrastructure to support spermatid and Sertoli cell adhesion, they are crucial to support preleptotene spermatocyte transport across theFig. 1 (continued) BTB (for basal ES) but also spermatid transport across the epithelium (for apical ES). These germ cells, namely spermatocytes and developing spermatids are the cargoes which are to be transported “directionally”, either to be base or to the adluminal edge of the seminiferous epithelium, due to the polarized nature of the actin- and MT-based cytoskeletons through the MT- or actin-dependent motor proteins. For instance, dynein 1 moves cargoes to the minus (−) end of MTs, and kinesin 15 to the plus (+) end of MTs; whereas myosin VIIa moves cargoes to the plus (+) end of actin filaments and myosin VI to the minus (−) end of actin filaments. In brief, the actin- and MT-based tracks found in Sertoli cells work in concert to support the directional\transport of germ cells across the seminiferous epithelium using the corresponding actin- and MT-based motor proteins. Even though germ cells located outside the Sertoli cell actin- and MT-cytoskeletons, the ES provides the means by which these germ cells anchor tightly onto the Sertoli cell cytoskeleton-based tracks to facilitate their transport across the epithelium. Through these actions of corresponding motor proteins, proper germ cell and cargo transports can take place across the seminiferous epithelium during the epithelial cycle to support spermatogenesis
Studies have shown that BTB dynamics that support preleptotene spermatocyte transport across the immunological barrier, and the subsequent haploid spermatid transport across the seminiferous epithelium, are tightly regulated cellular events. These involve several biologically active peptides released at the basement membrane but also at the Sertoli-spermatid adhesion site known as the apical ES via proteolytic cleavage of the structural proteins at these two sites, namely the F5-, the NC1- and the LG3/4/5-peptide [9, 25,26,27]. These bioactive peptides, in turn, are working in concert with a number of signaling proteins such as mTORC1/rpS6/Akt1/2 and FAK-Y407, and cytoskeletal regulatory proteins such as Arp3, Eps8, +TIPs and −TIPs to modulate BTB and ES dynamics [9, 27, 28]. The ultrastructures and the biomolecules that support germ cell transport are the actin- and MT-based cytoskeletons, as well as the corresponding actin- and MT-based motor proteins. In brief, motor proteins are the “vehicles” that carry the “cargoes”, namely preleptotene spermatocytes and spermatids, utilizing the corresponding actin or microtubule (MT)-based cytoskeletons as tracks to transport developing germ cells and other organelles (e.g., residual bodies, phagosomes, cell vacuoles, endocytic vesicles) to their corresponding “destination” across the seminiferous epithelium (Fig. 1). Furthermore, this requires intricate involvement of both actin- and MT-based cytoskeletons to support cargo transport across the seminiferous epithelium. However, much of this information remains unknown. In this review, we provide a timely discussion on latest findings in this area of research regarding the role of motor proteins in supporting cargo transport across the seminiferous epithelium using the rat testis as a study model. We also highlight some of the specific research areas that deserve attentions in future studies, which should be helpful to understand the underlying mechanism(s) of idiopathic male infertility.
Sertoli Cell Cytoskeletons in the Testis
In the seminiferous epithelium of adult rodent testes, similar to other mammalian organs, the two prominent cytoskeletons are the intrinsically polarized actin- and microtubule (MT)-based cytoskeletons which are composed of globular subunits of actin and α-tubulin/β-tubulin oligomers, respectively (Fig. 1) [29,30,31,32,33]. These polarized structures also serve as tracks to support specific motor proteins for directional transport of cargoes across the seminiferous epithelium. On the other hand, the intermediate filament-based cytoskeleton constituted by vimentin [16, 34] and the septin-based cytoskeleton [35] are both apolar structures, thus, they do not support motor proteins for directional cargo transport along their filaments.
Actin-Based Cytoskeleton
A functional actin-based track is composed of linear actin filaments (i.e., filamentous actin, F-actin) derived from polymerized globulin (G)-actin subunits, with the fast-growing barbed (+) end near the base of the seminiferous epithelium, closest to the basement membrane, and the slow-growing pointed (−) end near the seminiferous tubule lumen (Fig. 1) [36, 37]. In brief, polymerization of a linear actin filament occurs by incoming ATP-bound G-actin subunits at the fast-growing barbed (+) end involving actin nucleation proteins (e.g., formin 1, spire 1). The ATP-bound G-actin subunits are rapidly dephosphorylated to ADP-bound G-actin and are all found at the slow-growing pointed (−) end near the tubule lumen (Fig. 1) [37, 38]. The actin-based tracks are most notable in late Stage VIII of the epithelial cycle that stretch across the seminiferous epithelium and align perpendicular to the basement membrane [38] (Fig. 1). However, F-actin are also prominently noted at the apical ES and basal ES/BTB wherein the actin filaments are aligned parallel to the Sertoli cell plasma membrane and appear as bundled structures in cross-sections of the tubules. As such, these actin filaments appear as “rod-like” structures in cross-sections of the tubules at the apical ES and basal ES/BTB sites, thereby reinforcing cell adhesion (Fig. 1). ES in the testis also plays a crucial role to support germ cell transport as preleptotene spermatocytes (at the basal ES) and developing spermatids (at the apical ES) tightly anchored onto the actin filament bundles at the ES, and with the MTs located nearby [18, 33], which are located in close proximity to the plasma membrane of the Sertoli cell. Thus, these cells are separated only by their apposing Sertoli cell-cell or Sertoli-germ cell plasma membranes [3, 39]. Thus, even though these germ cells, namely preleptotene spermatocytes or haploid elongate spermatids, are located “outside” the Sertoli actin filament and MT networks, they are anchor onto these cytoskeletons through the unusual adhesion of ES between these adjacent cells, which are considered as cargoes to the Sertoli cell at the site. Due to this intrinsic polarized nature of the actin filaments, the actin-based plus (+) end-directed motor protein myosin VIIa, and the actin-based minus (−) end-directed myosin VI are capable of moving cargoes either to the base or to the tubule lumen across the epithelium, respectively (Fig. 1).
MT-Based Cytoskeleton
Microtubules (MTs) are also polarized ultrastructures in which a microtubule is composed of 13 laterally associated protofilaments of α- and β-tubulin heterodimers, with a hollow lumen wherein the plus (+) fast growing end is near the basement membrane and the minus (−) slow growing end near the tubule lumen (Fig. 1) [40,41,42,43]. Due to the intrinsic polarized nature of MTs, the MT-based minus (−) end-directed motor protein dynein 1 and the plus (+) end-directed motor protein kinesins (e.g., kinesin 15) can move cargoes to the corresponding minus or plus end of MTs, respectively [44,45,46,47].
Motor Proteins
Motor proteins are a class of molecular motors that bind to either microtubule (MT)- or actin-based tracks. They are capable of converting chemical energy through the hydrolysis of ATP to generate the mechanical force necessary to transport cargoes along the track across cell cytoplasm. Herein, we discuss several motor proteins that have been studied in the testis pertinent to support spermatogenesis. Besides serving as an update, this summary also provides the basis for future studies regarding the role of motor proteins in supporting germ cell and cargo transport across the seminiferous epithelium.
MT-Based Motor Proteins: Dynein and Kinesin
Dynein
Dynein is a family of motor proteins that use MT-based track in retrograde sliding movement towards the minus (−) ends of microtubules [47, 48]. In brief, a dynein motor protein transports cargoes towards the center of the cell or seminiferous tubule lumen in the testis. There are two major classes of dyneins, cytoplasmic and axonemal dyneins, which are classified according to their function and structure differences. Dynein 1 is a cytoplasmic dynein of about 1.5 megadaltons (MDa) (Fig. 2; Fig. 3A), involved in intracellular transport, mitosis, cell polarization and directional cargo transport. For instance, dynein 1 carries the cargo (e.g., spermatid) by “walking” along the MT-track in the Sertoli cell. Even though spermatids locate outside the Sertoli cell, but they are tightly anchored onto the MT-track in the Sertoli cell at the apical ES (or preleptotene spermatocyte anchored onto the MT-track in the Sertoli cell at the basal ES), which is a known adhesion ultrastructure that supports spermatid or preleptotene spermatocyte transport [3, 17]. There are 15 types of axonemal dyneins to support ciliary (e.g., dynein 2) and flagellar movement [48,49,50,51] such as sperm flagella that confers sperm progressive motility. Axonemal dyneins support the beating of flagella and cilia through rapid and efficient sliding movements of MTs [52]. In this context, it is of interest to note that mechanical movement of hair cells in cochlea is supported by the motor protein prestin [53, 54] which is different from the dynein family motor proteins. A functional dynein motor protein is considerably larger and more complex than kinesin or myosin motors, and it is composed of two heavy chains and a variable number of associated intermediate chains, light intermediate chains and light chains (Fig. 3A). For instance, dynein 1 is a dimeric protein composed of two identical heavy chains with a large molecular mass (Mr) of 500 kDa each. Each HC, in turn, binds to a light intermediate chain (LIC), an intermediate chain (IC), and three light chains (LCs) of LC7, LC8, and Tctex 1 (Fig. 3A). Thus, dynein 1 is a dimer of dimers. Each heavy chain is composed of three functional domains: a coiled-coil stalk with MT binding domain (MTBD) containing a globular motor head at the C-terminus, an AAA+ ring containing six AAA+ modules that organized into a doughnut-like structure, and a cargo-binding tail at by N-terminus (Figure 3A). The AAA+ ring can hydrolyze ATP hence converting chemical energy into mechanical force to support cargo transport [59]. In the testis, dynein 1 interacts with a protein complex called dynactin and cargo adaptor to form a functional motor protein called the dynein-dynactin-adaptor complex that supports spermatid transport on MT-based cytoskeleton. Dynein I also transports various cellular cargoes along MT towards the minus (−) end of MT tracks [60]. Cargoes transported by cytoplasmic dynein include endosomes [61], lysosomes [62], phagosomes [63], melanosomes [64], peroxisomes [65], lipid droplets [66], mitochondria [67] and vesicles from the endoplasmic reticulum (ER) destined for the Golgi [68]. These cargo transports hence regulate the intracellular function of cells and tissues through different cell signaling pathways. In the rat testis, dynein 1 is necessary to confer Sertoli cell TJ-permeability barrier function since its knockdown by RNAi perturbs the TJ-barrier function due to gross defects of F-actin and microtubules (MTs) across the Sertoli cell cytosol wherein both cytoskeletons become extensively truncated [69]. These defects, in turn, perturb the distribution of BTB-associated proteins at the site, including the cell adhesion complexes CAR/ZO-1 and N-cadherin/β-catenin [69]. Furthermore, dynein 1 knockdown also perturbs the polymerization activities of F-actin and MTs [69], possibly due to defects in transporting machineries (e.g., actin or MT polymerization proteins) necessary to support cytoskeletal nucleation. More important, the loss of dynein 1 function by RNAi also perturbs the BTB function in vivo since the barrier no longer restricts the diffusion of small molecular biotin across the immunological barrier [69]. Multiple defective sperms are also noted in the epididymis including extensive defects in spermatid heads, tail, and sperm morphology due to defects of intracellular trafficking to support the assembly of essential cellular components during spermiogenesis [69]. The importance of dynein-based motor proteins is also noted in Table 1 since its KO in mice led to embryonic lethality.
Schematic illustration on the functional domains of the microtubule-based motor proteins dyneins and kinesins, and actin-based motor proteins myosins. The different functional domains of motor proteins dyneins, kinesins and myosins are noted in corresponding panels. This figure was prepared based on earlier reports [55,56,57,58]. Abbreviations: DYNC1H1, dynein cytoplasmic 1 heavy chain 1; KIF, kinesin; MT, microtubule; SH3, SRC homology 3 domain; IQ motif, isoleucine and glutamine motif is a basic unit containing about 23 amino acids; PEST motif, a motif rich in proline (P), glutamic acid (E), serine (S) and threonine (T); PH domain, pleckstrin homology domain; MyTH4 domain, Myosin Tail Homology 4 domain; FERM domain, F for 4.1 protein, E for ezrin, R for radixin and M for moesin
Kinesin
Kinesin is a group of related motor proteins that use MT track in anterograde movement, to transport cargoes towards the plus (+) ends of MTs [96,97,98] (Fig. 2). In brief, a kinesin motor protein transports cargoes away from the center of the cell, usually to cell peripheries to support cell homeostasis, or to the base of the seminiferous epithelium in the testis (Fig. 1). Kinesin superfamily members in humans and rodents are organized into 14 families [99, 100]. A functional kinesin motor protein is a tetrameric protein, comprised of two heavy chains and two light chains (Fig. 3A). Each heavy chain has a globular motor head where microtubule binding and ATP hydrolysis take place at its N-terminal region, which in turn generate the energy via ATPase that converts chemical energy into mechanical force to elicit cargo transport. The head region is connected by a short neck linker to a long intertwined coiled-coil stalk, to be followed by the tail at its C-terminal region (Fig. 3A). A light chain associates with a tail which serve as the adapter for binding to a cargo while moving along the MT track towards the MT plus (+) end to facilitate cargo (e.g., spermatid, residual body, phagosome) transport [49, 97, 101] (Table 1). Kinesins typically move cargoes in the direction of MT plus (+) end on MT tracks, such that cargo is transported from the center of the cell to its periphery (i.e., anterograde movement). However, some kinesins (members of the kinesin-5 family), such as kinesin-14, move cargoes to the MT minus (−) end along the MT tracks wherein the motor region is located at the C-terminal region of the heavy chain [102]. On the other hand, kinesin-5 Cin8 (members of the kinesin-5 family) is a bidirectional kinesin which can move a cargo towards the microtubule minus (−) end when works alone but to the plus (+) end in an ensemble with a team of motors [103]. Emerging evidence has shown that kinesins are crucial to support tumorigenesis. For instance, KIF18A promotes invasion and metastasis by activating Akt and MMP-7/MMP-9-related signaling pathways [104] whereas kinesins also support proliferation, cell differentiation, aggressiveness and epithelial-mesenchymal transition of tumor cells [105,106,107,108,109]. A recent report has demonstrated the importance of kinesin-9 in conferring progressive motility in mouse spermatozoa since a deletion of 16 bp nucleotides of the Kif9 gene in mice (Kif9−16/−16) using CRISPR/Cas9 led to defects in flagellar movement of sperm tails [110]. Studies have also shown that kinesin-7 CENP-E is crucial to support chromosome alignment and genome stability of spermatogenic cells (e.g., spermatogonia and spermatocytes) during mitosis and meiosis [111], whereas kinesin-5 Eg5 supports spindle assembly and chromosome alignment of mouse spermatocytes [112]. Nonetheless, much work is needed to better understand the role of kinesins in supporting spermatogenesis in the testis. However, as noted in Table 1, deletion of one of the several kinesins in mice led to embryonic lethality, illustrating the physiological significance of kinesin-based motor proteins in supporting cellular function.
Schematic illustrations on the structural components of the functional motor proteins dynein 1, kinesin 15 and myosin VIIa. (A) A functional dynein motor protein (e.g., dynein 1) complex is composed of the dynein, dynactin and the cargo adaptor (left panel). The dynein motor protein consists of two monomers. Each monomer is composed of a heavy chain (HC) motor and several other subunits: an intermediate chain, a light intermediate chain, and three light chains called LC7 (light chain 7) LC8 and Tctex. Each HC has the N-terminus at the tail and the C-terminal motor unit contains six AAA (ATPase Associated with Cellular Activities) domains, AAA1 to AAA6, and organized into a ring-like structure, which in turn connects to the microtubule binding domain (MTBD) in the motor head at the C-terminus which also binds onto the microtubule. AAA1Fig. 3 (continued) is the major site of ATP hydrolysis with other AAA sites play the regulatory roles. AAA1 converts the chemical energy (from ATP hydrolysis via ATPase) to mechanical force which is transmitted to the HC tail at the N-terminal region. Dynein 1 interacts with its cofactor called dynactin (which also composed of multiple subunits as earlier reviewed [47]) to form the functional dynein-dynactin complex. This complex in turn interacts with the cargo adaptor to form a functional motor protein to support cargo transport. On the right panel is the kinesin motor protein (e.g., kinesin 15) which is also a dimeric protein, composed of two monomers. Each monomer has a heavy chain (HC) with its N-terminal region contains the motor head which is the site for ATP hydrolysis to generate the chemical energy to be transmitted to the mechanical energy via the tail to propel cargo transport at the C-terminal region. The motor head at the N-terminal region also contains the microtubule binding domain. This is followed by the α-helical coiled-coil domain that constitutes the stalk and ends with the C-terminal tail of cargo binding. (B) A functional actin-based plus (+) end directed motor protein myosin (e.g., myosin VIIa) is also a dimeric protein comprised of two monomers. Each monomer has a heavy chain (HC) that begins with the motor head at its N-terminal region which contains the ATP hydrolysis motor domain and the actin-binding domain. This is followed by the neck region that transmits the chemical energy derived from ATP hydrolysis at the motor head to the tail cargo binding site through the coiled-coil domain in the tail. The neck region has a pair of light chains which facilitates the transmission of chemical energy to the cargo propelling mechanical force at the tail cargo binding site. The C-terminal tail region contains the FERM (F, 4.1 protein; E, ezrin; R, radixin; M, moesin), MyTH4 Myosin tail homology 4) and SH3 (SRC homolog 3) domains, and the globular tail domain (GTD) at the C-terminus to support cargo binding
F-actin-Based Motor Proteins: Myosins
Myosins
Myosins are the only known actin-based motor proteins in mammalian cells and tissues including the testis [47, 113]. There are 18 classes of myosin superfamily members known to date based on phylogenetic analysis of their motor domain, and at least 40 myosin genes have been identified [57, 114]. By converting chemical energy via hydrolysis of ATP at the myosin motor head to mechanical energy, which in turn is used to propel cargo to be transported along the actin-based tracks, which are most notable in late Stage VIII tubules across the seminiferous epithelium in the testis [38]. Besides the regular myosins noted in mammalian cells, there is an emerging long-tailed unconventional class of myosins, namely myosin 1E and myosin 1F [115]. In general, each myosin has a Mr of 520 kDa, consisting of six subunits: two 220 kDa heavy chains, and two pairs of light chains (20 kDa for each light chain) (Fig. 3) [116]. Thus, there are two monomers in a functional myosin motor protein, with each monomer consists of a heavy chain and a pair of light chains to a total of three subunits. Each heavy chain, in turn, can be divided into distinctive head, neck and tail domains (Fig. 3B). The globular head domain interacts with actin filaments (i.e., actin-based track) though its actin binding site at the N-terminal region which also contains the ATPase site, capable of hydrolyzing ATP to convert the chemical energy to mechanical energy to propel cargo transport. The neck region of each heavy chain serves as a linker, which also transduces force generated by the catalytic motor domain at the head region. The neck region also provides the binding site for a pair of light chains, which are distinct protein subunits that interact with the neck region (Fig. 3B). The C-terminal tail contains a relatively long α-helical coiled-coil domain and at its C-terminal region, it contains the sequential SH3 (SRC homology 3), MyTH4 (myosin tail homology 4), FERM (F, 4.1 protein; E, ezrin; R, radixin; M, moesin) domains and the globular tail domain (GTD) at its C-terminus. GTD domain is supported by the FERM, MyTH4 and SH3 domains, and GTD also recognizes different cargoes through direct interactions or mediated through adaptor proteins, such as vezatin in the testis [117] (Fig. 3B). Most myosins (e.g., myosin VIIa) walk along actin filaments to the actin plus (+) end, but Myosin VI moves cargoes to the minus (−) end of actin tracks [113]. Myosin VIIa is a member of the myosin superfamily found in testis and other tissues [118] In testes, actin filament bundles constitute the ectoplasmic specialization, which also serve as the attachment site for cell adhesion protein complexes (e.g., N-cadherin-β-catenin, occludin-ZO-1, nectin-afadin). It also supports the transport of spermatids or organelles (i.e., cargoes) by serving as the track [12, 18]. Studies of myosin VIIa in the testis have shown that the knockdown of myosin VIIa in the testis in vivo by RNAi perturbs the organization of F-actin, but also MT tracks, across the seminiferous epithelium wherein these cytoskeletal tracks are extensively truncated [119]. These disruptive changes are likely the results of a considerably reduction in actin and MT polymerization activity in Sertoli cells [119] due to defects in intracellular protein trafficking. These defects also lead to formation of multiple defective sperms with gross changes in their morphology including round-shaped epididymal sperm heads, consistent presence of cytoplasmic droplets in the head region, and structural defects of sperm necks [119]. These findings are also consistent with earlier reports which have shown that KO of myosin motor proteins lead to embryonic fatality in mice (Table 1), and its mutation or genetic variations in humans also lead to defects in brain and heart development due to defects in intracellular protein trafficking.
Concluding Remarks and Future Perspectives
Herein, we summarize findings regarding the role of MT- and actin-based motor proteins in supporting mammalian spermatogenesis. As seen in studies using genetic models through gene deletion in mice (Table 1), and genetic mutations or gene variants in humans (Table 2), embryonic lethality (in mice) and serious pathological conditions (in humans) are noted in Tables 1 and 2, illustrating the significance of these motor proteins in cells and tissues, besides the testis. However, there are main questions remain. For instance, what are the biomolecules that trigger the use of specific plus (+) end or minus (−) directed motor proteins to initiate cargo transport of germ cells or other organelles to support spermatogenesis through different epithelial cycles? What is the mechanism(s) in place that selects the use of actin- or MT-based tracks or both? How does actin- and MT-based cytoskeletons coordinate with each other to streamline the transport of cargoes using their tracks to support spermatogenesis? It is now known that the several locally produced biomolecules, namely the F5-, NC1- and LG3/4/5-peptide, that regulate spermatogenesis exert their regulatory effects through their corresponding downstream signaling molecules on cytoskeletal organization. What is the mechanism(s) by which these biomolecules select the appropriate cytoskeleton, namely the F-actin or MT cytoskeleton, to execute their function? The answers to many of these questions will be helpful to understand and better manage unexplained male infertility. In brief, an intensive race is on to search for answers to some of these questions in the years to come, such as the role of many genes known to regulaste spermatogenesis to support motor protein function [191]. It is likely that the use of scRNA-seq and scATAC-seq coupled with transcriptome profiling and bioinformatics analyses will provide many of the missing information in this race to tackle male infertility (or fertility) in the years to come.
References
Cheng, C. Y., & Mruk, D. D. (2012). The blood-testis barrier and its implication in male contraception. Pharmacological Reviews, 64, 16–64.
Franca, L. R., Auharek, S. A., Hess, R. A., Dufour, J. M., & Hinton, B. T. (2012). Blood-tissue barriers: Morphofunctional and immunological aspects of the blood-testis and blood-epididymal barriers. Advances in Experimental Medicine and Biology, 763, 237–259.
Mruk, D. D., & Cheng, C. Y. (2004b). Sertoli-sertoli and sertoli-germ cell interactions and their significance in germ cell movement in the seminiferous epithelium during spermatogenesis. Endocrine Reviews, 25, 747–806.
Pelletier, R. M. (2011). The Blood-Testis Barrier: The Junctional Permeability, The Proteins And The Lipids. Progress in Histochemistry and Cytochemistry, 46, 49–127.
Setchell, B. P. (2008). Blood-testis barrier, functional and transport proteins and spermatogenesis. Advances in Experimental Medicine and Biology, 636, 212–233.
Stanton, P. G. (2016). Regulation of the blood-testis barrier. Seminars in Cell & Developmental Biology, 59, 166–173.
Weber, J. E., Russell, L. D., Wong, V., & Peterson, R. N. (1983). Three dimensional reconstruction of a rat stage V sertoli cell: II. Morphometry of sertoli-sertoli and sertoli-germ cell relationships. The American Journal of Anatomy, 167, 163–179.
Wong, V., & Russell, L. D. (1983). Three-dimensional reconstruction of a rat stage V sertoli cell: I. Methods, basic configuration, and dimensions. The American Journal of Anatomy, 167, 143–161.
Mao, B., Bu, T., Mruk, D., Li, C., Sun, F., & Cheng, C. Y. (2020a). Modulating the blood-testis barrier towards increasing drug delivery. Trends in Pharmacological Sciences, 41, 691–700.
Hermo, L., Pelletier, R. M., Cyr, D. G., & Smith, C. E. (2010b). Surfing the wave, cycle, life history, and genes/proteins expressed by testicular germ cells. Part 1: Background to spermatogenesis, spermatogonia, and spermatocytes. Microscopy Research and Technique, 73, 241–278.
Hermo, L., Pelletier, R. M., Cyr, D. G., & Smith, C. E. (2010c). Surfing the wave, cycle, life history, and genes/proteins expressed by testicular germ cells. Part 2: Changes in spermatid organelles associated with development of spermatozoa. Microscopy Research and Technique, 73, 279–319.
Hess, R. A., & De Franca, L. R. (2008). Spermatogenesis and cycle of the seminiferous epithelium. Advances in Experimental Medicine and Biology, 636, 1–15.
Cheng, C. Y., & Mruk, D. D. (2015). Biochemistry of sertoli cell/germ cell junctions, germ cell transport, and spermiation in the seminiferous epithelium. In M. D. Griswold (Ed.), Sertoli cell biology (2nd ed., pp. 333–383). Elsevier. https://doi.org/10.1016/B978-0-12-417047-6.00012.0
Hermo, L., Pelletier, R. M., Cyr, D. G., & Smith, C. E. (2010d). Surfing the wave, cycle, life history, and genes/proteins expressed by testicular germ cells. Part 5: Intercellular junctions and contacts between germ cells and sertoli cells and their regulatory interactions, testicular cholesterol, and genes/proteins associated with more than one germ cell generation. Microscopy Research and Technique, 73, 409–494.
Li, M. W. M., Mruk, D. D., & Cheng, C. Y. (2012). Gap junctions and blood-tissue barriers. Advances in Experimental Medicine and Biology, 763, 260–280.
Lie, P. P. Y., Cheng, C. Y., & Mruk, D. D. (2011). The biology of the desmosome-like junction: A versatile anchoring junction and signal transducer in the seminiferous epithelium. International Review of Cell and Molecular Biology, 286, 223–269.
Mruk, D. D., & Cheng, C. Y. (2004a). Cell-cell interactions at the ectoplasmic specialization in the testis. Trends in Endocrinology and Metabolism, 15, 439–447.
Vogl, A. W., Vaid, K. S., & Guttman, J. A. (2008). The sertoli cell cytoskeleton. Advances in Experimental Medicine and Biology, 636, 186–211.
Wong, E. W. P., Mruk, D. D., & Cheng, C. Y. (2008). Biology and regulation of ectoplasmic specialization, an atypical adherens junction type, in the testis. Biochimica et Biophysica Acta (BBA) - Biomembranes, 1778, 692–708.
Xiao, X., Mruk, D. D., Wong, C. K. C., & Cheng, C. Y. (2014). Germ cell transport across the seminiferous epithelium during spermatogenesis. Physiology, 29, 286–298.
Schaks, M., Giannone, G., & Rottner, K. (2019). Actin dynamics in cell migration. Essays in Biochemistry, 63, 483–495.
O’donnell, L. (2014). Mechanisms of spermiogenesis and spermiation and how they are disturbed. Spermatogenesis, 4, E979623.
O’donnell, L., Nicholls, P. K., O’bryan, M. K., Mclachlan, R. I., & Stanton, P. G. (2011). Spermiation: The process of sperm release. Spermatogenesis, 1, 14–35.
Qian, X., Mruk, D. D., Cheng, Y. H., Tang, E. I., Han, D., Lee, W. M., Wong, E. W., & Cheng, C. Y. (2014). Actin binding proteins, spermatid transport and spermiation. Seminars in Cell & Developmental Biology, 30, 75–85.
Li, H., Liu, S., Wu, S., Li, L., Ge, R., & Cheng, C. Y. (2020). Bioactive fragments of laminin and collagen chains: Lesson from the testis. Reproduction, 159, R111–R123.
Liu, S. W., Li, H. T., Ge, R. S., & Cheng, C. Y. (2021). Nc1-peptide derived from collagen A3 (Iv) chain is a blood-tissue barrier regulator: Less from the testis. Asian Journal of Andrology, 23, 123. https://doi.org/10.4103/Aja.Aja_44_20
Wu, S., Yan, M., Ge, R., & Cheng, C. Y. (2020). Crosstalk between sertoli and germ cells in male fertility. Trends in Molecular Medicine, 26, 215–231.
Mao, B. P., Ge, R., & Cheng, C. Y. (2020b). Role of microtubule +Tips and -Tips in spermatogenesis - Insights from studies of toxicant models. Reproductive Toxicology, 91, 43–52.
Dogterom, M., & Koenderink, G. H. (2019). Actin–microtubule crosstalk in cell biology. Nature Reviews Molecular Cell Biology, 20, 38–54.
Dunleavy, J. E. M., O’bryan, M. K., Stanton, P. G., & O’donnell, L. (2019). The cytoskeleton in spermatogenesis. Reproduction, 157, R53–R72.
Lie, P. P. Y., Mruk, D. D., Lee, W. M., & Cheng, C. Y. (2010). Cytoskeletal dynamics and spermatogenesis. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 365, 1581–1592.
O’donnell, L., & O’bryan, M. K. (2014). Microtubules and spermatogenesis. Seminars in Cell & Developmental Biology, 30, 45–54.
Tang, E. I., Mruk, D. D., & Cheng, C. Y. (2016). Regulation of microtubule (Mt)-based cytoskeleton in the seminiferous epithelium during spermatogenesis. Seminars in Cell & Developmental Biology, 59, 35–45.
Nekrasova, O., & Green, K. J. (2013). Desmosome Assembly And Dynamics. Trends in Cell Biology, 23, 537–546.
Lin, Y. H., Kuo, Y. C., Chiang, H. S., & Kuo, P. L. (2011). The role of the septin family in spermiogenesis. Spermatogenesis, 1, 298–302.
Campellone, K. G., & Welch, M. D. (2010). A Nucleator arms race: Cellular control of actin assembly. Nature Reviews. Molecular Cell Biology, 11, 237–251.
Suarez, C., & Kovar, D. R. (2016). Internetwork competition for monomers governs actin cytoskeleton organization. Nature Reviews. Molecular Cell Biology, 17, 799–810.
Wang, L., Yan, M., Wu, S., Wu, X., Bu, T., Wong, C. K. C., Ge, R., Sun, F., & Cheng, C. Y. (2020b). Actin binding proteins, actin cytoskeleton and spermatogenesis - Lesson from toxicant models. Reproductive Toxicology, 96, 76–89.
Lee, N. P. Y., & Cheng, C. Y. (2004). Ectoplasmic specialization, a testis-specific cell-cell actin-based adherens junction type: Is this a potential target for male contraceptive development. Human Reproduction Update, 10, 349–369.
Borisy, G., Heald, R., Howard, J., Janke, C., Musacchio, A., & Nogales, E. (2016). Microtubules: 50 Years on from the discovery of tubulin. Nature Reviews. Molecular Cell Biology, 17, 322–328.
Brouhard, G. J., & Rice, L. M. (2018). Microtubule dynamics: An interplay of biochemistry and mechanics. Nature Reviews. Molecular Cell Biology, 19, 451–463.
Tang, E. I., Mruk, D. D., Lee, W. M., & Cheng, C. Y. (2015). Cell-cell interactions, cell polarity, and the blood-testis barrier. In K. Ebnet (Ed.), Cell polarity 1 (pp. 303–326). Springer International Publishing. https://doi.org/10.1007/978-3-319-14463-4_13
Wang, L., Yan, M., Wu, S., Mao, B., Wong, C. K. C., Ge, R., Sun, F., & Cheng, C. Y. (2020a). Microtubule cytoskeleton and spermatogenesis - Lesson from studies of toxicant models. Toxicological Sciences, 177, 305. https://doi.org/10.1093/Toxsci/Kfaa109. PMID:32647867.
Kikkawa, M. (2013). Big steps toward understanding dynein. The Journal of Cell Biology, 202, 15–23.
Mallik, R., Rai, A. K., Barak, P., Rai, A., & Kunwar, A. (2013). Teamwork in microtubule motors. Trends in Cell Biology, 23, 575. https://doi.org/10.1016/J.Tcb.2013.06.003
Reck-Peterson, S. L., Redwine, W. B., Vale, R. D., & Carter, A. P. (2018). The cytoplasmic dynein transport machinery and its many cargoes. Nature Reviews. Molecular Cell Biology, 19, 382–398.
Wen, Q., Tang, E. I., Xiao, X., Gao, Y., Chu, D. S., Mruk, D. D., Silvestrini, B., & Cheng, C. Y. (2016). Transport of germ cells across the seminiferous epithelium during spermatogenesis-The involvement of both actin- and microtubule-based cytoskeletons. Tissue Barriers, 4, E1265042.
Roberts, A. J., Kon, T., Knight, P. J., Sutoh, K., & Burgess, S. A. (2013). Functions And Mechanics Of Dynein Motor Proteins. Nature Reviews. Molecular Cell Biology, 14, 713–726.
Hirokawa, N. (1998). Kinesin and dynein superfamily proteins and the mechanism of organelle transport. Science, 279, 519–526.
Schroer, T. A., Steuer, E. R., & Sheetz, M. P. (1989). Cytoplasmic dynein is a minus end-directed motor for membranous organelles. Cell, 56, 937–946.
Vale, R. D., Reese, T. S., & Sheetz, M. P. (1985). Identification of a novel force-generating protein, kinesin, involved in microtubule-based motility. Cell, 42, 39–50.
Mallik, R., & Gross, S. P. (2004). Molecular motors: Strategies to get along. Current Biology, 14, R971–R982.
Dallos, P., & Fakler, B. (2002). Prestin, a new type of motor protein. Nature Reviews. Molecular Cell Biology, 3, 104–111.
Oliver, D., He, D. Z., Klöcker, N., Ludwig, J., Schulte, U., Waldegger, S., Ruppersberg, J. P., Dallos, P., & Fakler, B. (2001). Intracellular Anions As The Voltage Sensor Of Prestin, The Outer Hair Cell Motor Protein. Science, 292, 2340–2343.
Edwards, M., Zwolak, A., Schafer, D. A., Sept, D., Dominguez, R., & Cooper, J. A. (2014). Capping protein regulators fine-tune actin assembly dynamics. Nature Reviews. Molecular Cell Biology, 15, 677–689.
Hoang, H. T., Schlager, M. A., Carter, A. P., & Bullock, S. L. (2017). Dync1h1 mutations associated with neurological diseases compromise processivity of dynein-dynactin-cargo adaptor complexes. Proceedings of the National Academy of Sciences of the United States of America, 114, E1597–E1606.
Odronitz, F., & Kollmar, M. (2007). Drawing the tree of eukaryotic life based on the analysis of 2,269 manually annotated myosins from 328 species. Genome Biology, 8, R196.
Parsons, J. T., Horwitz, A. R., & Schwartz, M. A. (2010). Cell adhesion: Integrating cytoskeletal dynamics and cellular tension. Nature Reviews. Molecular Cell Biology, 11, 633–643.
Neuwald, A. F., Aravind, L., Spouge, J. L., & Koonin, E. V. (1999). Aaa+: A class of chaperone-like atpases associated with the assembly, operation, and disassembly of protein complexes. Genome Research, 9, 27–43.
Asai, D. J., & Koonce, M. P. (2001). The dynein heavy chain: Structure, mechanics and evolution. Trends in Cell Biology, 11, 196–202.
Driskell, O. J., Mironov, A., Allan, V. J., & Woodman, P. G. (2007). Dynein is required for receptor sorting and the morphogenesis of early endosomes. Nature Cell Biology, 9, 113–120.
Jordens, I., Fernandez-Borja, M., Marsman, M., Dusseljee, S., Janssen, L., Calafat, J., Janssen, H., Wubbolts, R., & Neefjes, J. (2001). The Rab7 effector protein rilp controls lysosomal transport by inducing the recruitment of dynein-dynactin motors. Current Biology, 11, 1680–1685.
Blocker, A., Severin, F. F., Burkhardt, J. K., Bingham, J. B., Yu, H., Olivo, J. C., Schroer, T. A., Hyman, A. A., & Griffiths, G. (1997). Molecular requirements for bi-directional movement of phagosomes along microtubules. The Journal of Cell Biology, 137, 113–129.
Gross, S. P., Tuma, M. C., Deacon, S. W., Serpinskaya, A. S., Reilein, A. R., & Gelfand, V. I. (2002). Interactions and regulation of molecular motors in xenopus melanophores. The Journal of Cell Biology, 156, 855–865.
Kural, C., Kim, H., Syed, S., Goshima, G., Gelfand, V. I., & Selvin, P. R. (2005). Kinesin and dynein move a peroxisome in vivo: A tug-of-war or coordinated movement? Science, 308, 1469–1472.
Gross, S. P., Welte, M. A., Block, S. M., & Wieschaus, E. F. (2000). Dynein-mediated cargo transport in vivo. A switch controls travel distance. The Journal of Cell Biology, 148, 945–956.
Pilling, A. D., Horiuchi, D., Lively, C. M., & Saxton, W. M. (2006). Kinesin-1 and dynein are the primary motors for fast transport of mitochondria in drosophila motor axons. Molecular Biology of the Cell, 17, 2057–2068.
Presley, J. F., Cole, N. B., Schroer, T. A., Hirschberg, K., Zaal, K. J., & Lippincott-Schwartz, J. (1997). Er-To-Golgi transport visualized in living cells. Nature, 389, 81–85.
Wen, Q., Tang, E. I., Lui, W. Y., Lee, W. M., Wong, C. K. C., Silvestrini, B., & Cheng, C. Y. (2018). Dynein 1 supports spermatid transport and spermiation during spermatogenesis in the rat testis. American Journal of Physiology. Endocrinology and Metabolism, 315, E924–E948.
Neesen, J., Kirschner, R., Ochs, M., Schmiedl, A., Habermann, B., Mueller, C., Holstein, A. F., Nuesslein, T., Adham, I., & Engel, W. (2001). Disruption of an inner arm dynein heavy chain gene results in asthenozoospermia and reduced ciliary beat frequency. Human Molecular Genetics, 10, 1117–1128.
Adams, D., Baldock, R., Bhattacharya, S., Copp, A. J., Dickinson, M., Greene, N. D., Henkelman, M., Justice, M., Mohun, T., Murray, S. A., Pauws, E., Raess, M., Rossant, J., Weaver, T., & West, D. (2013). Bloomsbury report on mouse embryo phenotyping: Recommendations from the impc workshop on embryonic lethal screening. Disease Models & Mechanisms, 6, 571–579.
Ibanez-Tallon, I., Gorokhova, S., & Heintz, N. (2002). Loss of function of axonemal dynein Mdnah5 causes primary ciliary dyskinesia and hydrocephalus. Human Molecular Genetics, 11, 715–721.
Harada, A., Takei, Y., Kanai, Y., Tanaka, Y., Nonaka, S., & Hirokawa, N. (1998). Golgi vesiculation and lysosome dispersion in cells lacking cytoplasmic dynein. The Journal of Cell Biology, 141, 51–59.
Takeda, S., Yonekawa, Y., Tanaka, Y., Okada, Y., Nonaka, S., & Hirokawa, N. (1999). Left-right asymmetry and kinesin superfamily protein Kif3a: New insights in determination of laterality and mesoderm induction by Kif3a-/- mice analysis. The Journal of Cell Biology, 145, 825–836.
Nonaka, S., Tanaka, Y., Okada, Y., Takeda, S., Harada, A., Kanai, Y., Kido, M., & Hirokawa, N. (1998). Randomization of left-right asymmetry due to loss of nodal cilia generating leftward flow of extraembryonic fluid in mice lacking kif3b motor protein. Cell, 95, 829–837.
Tanaka, Y., Kanai, Y., Okada, Y., Nonaka, S., Takeda, S., Harada, A., & Hirokawa, N. (1998). Targeted disruption of mouse conventional kinesin heavy chain, Kif5b, results in abnormal perinuclear clustering of mitochondria. Cell, 93, 1147–1158.
Castillo, A., & Justice, M. J. (2007). The kinesin related motor protein, Eg5, is essential for maintenance of pre-implantation embryogenesis. Biochemical and Biophysical Research Communications, 357, 694–699.
Chauviere, M., Kress, C., & Kress, M. (2008). Disruption of the mitotic kinesin Eg5 gene (Knsl1) results in early embryonic lethality. Biochemical and Biophysical Research Communications, 372, 513–519.
Ueno, H., Huang, X., Tanaka, Y., & Hirokawa, N. (2011). Kif16b/Rab14 molecular motor complex is critical for early embryonic development by transporting Fgf receptor. Developmental Cell, 20, 60–71.
Liu, X. S., Zhao, X. D., Wang, X., Yao, Y. X., Zhang, L. L., Shu, R. Z., Ren, W. H., Huang, Y., Huang, L., Gu, M. M., Kuang, Y., Wang, L., Lu, S. Y., Chi, J., Fen, J. S., Wang, Y. F., Fei, J., Dai, W., & Wang, Z. G. (2010). Germinal cell aplasia in Kif18a mutant male mice due to impaired chromosome congression and dysregulated Bubr1 and Cenp-E. Genes & Cancer, 1, 26–39.
Kannan, M., Bayam, E., Wagner, C., Rinaldi, B., Kretz, P. F., Tilly, P., Roos, M., Mcgillewie, L., Bar, S., Minocha, S., Chevalier, C., Po, C., Sanger Mouse Genetics, P., Chelly, J., Mandel, J. L., Borgatti, R., Piton, A., Kinnear, C., Loos, B., … Yalcin, B. (2017). Wd40-repeat 47, a microtubule-associated protein, is essential for brain development and autophagy. Proceedings of the National Academy of Sciences of the United States of America, 114, E9308–E9317.
Muhia, M., Thies, E., Labonte, D., Ghiretti, A. E., Gromova, K. V., Xompero, F., Lappe-Siefke, C., Hermans-Borgmeyer, I., Kuhl, D., Schweizer, M., Ohana, O., Schwarz, J. R., Holzbaur, E. L. F., & Kneussel, M. (2016). The kinesin Kif21b regulates microtubule dynamics and is essential for neuronal morphology, synapse function, and learning and memory. Cell Reports, 15, 968–977.
Gromova, K. V., Muhia, M., Rothammer, N., Gee, C. E., Thies, E., Schaefer, I., Kress, S., Kilimann, M. W., Shevchuk, O., Oertner, T. G., & Kneussel, M. (2018). Neurobeachin and the kinesin Kif21b are critical for endocytic recycling of nmda receptors and regulate social behavior. Cell Reports, 23, 2705–2717.
Morikawa, M., Tanaka, Y., Cho, H. S., Yoshihara, M., & Hirokawa, N. (2018). The molecular motor Kif21b mediates synaptic plasticity and fear extinction by terminating Rac1 activation. Cell Reports, 23, 3864–3877.
Wang, L., Tanaka, Y., Wang, D., Morikawa, M., Zhou, R., Homma, N., Miyamoto, Y., & Hirokawa, N. (2018). The atypical kinesin Kif26a facilitates termination of nociceptive responses by sequestering focal adhesion kinase. Cell Reports, 24, 2894–2907.
Zhou, R., Niwa, S., Homma, N., Takei, Y., & Hirokawa, N. (2009). Kif26a is an unconventional kinesin and regulates gdnf-ret signaling in enteric neuronal development. Cell, 139, 802–813.
Krendel, M., Kim, S. V., Willinger, T., Wang, T., Kashgarian, M., Flavell, R. A., & Mooseker, M. S. (2009). Disruption of myosin 1e promotes podocyte injury. Journal of the American Society of Nephrology, 20, 86–94.
Conti, M. A., Even-Ram, S., Liu, C., Yamada, K. M., & Adelstein, R. S. (2004). Defects in cell adhesion and the visceral endoderm following ablation of nonmuscle myosin heavy chain II-A in mice. The Journal of Biological Chemistry, 279, 41263–41266.
Tullio, A. N., Accili, D., Ferrans, V. J., Yu, Z. X., Takeda, K., Grinberg, A., Westphal, H., Preston, Y. A., & Adelstein, R. S. (1997). Nonmuscle myosin II-B is required for normal development of the mouse heart. Proceedings of the National Academy of Sciences of the United States of America, 94, 12407–12412.
Ma, X., Bao, J., & Adelstein, R. S. (2007). Loss of cell adhesion causes hydrocephalus in nonmuscle myosin II-B-ablated and mutated mice. Molecular Biology of the Cell, 18, 2305–2312.
Abouhamed, M., Grobe, K., San, I. V., Thelen, S., Honnert, U., Balda, M. S., Matter, K., & Bahler, M. (2009). Myosin Ixa regulates epithelial differentiation and its deficiency results in hydrocephalus. Molecular Biology of the Cell, 20, 5074–5085.
Heimsath, E. G., Jr., Yim, Y. I., Mustapha, M., Hammer, J. A., & Cheney, R. E. (2017). Myosin-X knockout is semi-lethal and demonstrates that myosin-X functions in neural tube closure, pigmentation, hyaloid vasculature regression, and filopodia formation. Scientific Reports, 7, 17354.
Horsthemke, M., Nutter, L. M. J., Bachg, A. C., Skryabin, B. V., Honnert, U., Zobel, T., Bogdan, S., Stoll, M., Seidl, M. D., Muller, F. U., Ravens, U., Unger, A., Linke, W. A., Van Gorp, P. R. R., De Vries, A. A. F., Bahler, M., & Hanley, P. J. (2019). A novel isoform of myosin 18a (Myo18agamma) is an essential sarcomeric protein in mouse heart. The Journal of Biological Chemistry, 294, 7202–7218.
Ajima, R., Akazawa, H., Kodama, M., Takeshita, F., Otsuka, A., Kohno, T., Komuro, I., Ochiya, T., & Yokota, J. (2008). Deficiency of Myo18b in mice results in embryonic lethality with cardiac myofibrillar aberrations. Genes to Cells, 13, 987–999.
Lelli, A., Michel, V., Boutet De Monvel, J., Cortese, M., Bosch-Grau, M., Aghaie, A., Perfettini, I., Dupont, T., Avan, P., El-Amraoui, A., & Petit, C. (2016). Class III Myosins shape the auditory hair bundles by limiting microvilli and stereocilia growth. The Journal of Cell Biology, 212, 231–244.
Fourriere, L., Jimenez, A. J., Perez, F., & Boncompain, G. (2020). The role of microtubules in secretory protein transport. Journal of Cell Science, 133, jcs237016.
Miki, H., Okada, Y., & Hirokawa, N. (2005). Analysis of the kinesin superfamily: Insights into structure and function. Trends in Cell Biology, 15, 467–476.
Webb, S., Mukhopadhyay, A. G., & Roberts, A. J. (2020). Intraflagellar transport trains and motors: Insights from structure. Seminars in Cell & Developmental Biology, 107, 82.
Dagenbach, E. M., & Endow, S. A. (2004). A new kinesin tree. Journal of Cell Science, 117, 3–7.
Lawrence, C. J., Dawe, R. K., Christie, K. R., Cleveland, D. W., Dawson, S. C., Endow, S. A., Goldstein, L. S., Goodson, H. V., Hirokawa, N., Howard, J., Malmberg, R. L., Mcintosh, J. R., Miki, H., Mitchison, T. J., Okada, Y., Reddy, A. S., Saxton, W. M., Schliwa, M., Scholey, J. M., … Wordeman, L. (2004). A standardized kinesin nomenclature. The Journal of Cell Biology, 167, 19–22.
Hirokawa, N., & Noda, Y. (2008). Intracellular transport and kinesin superfamily proteins, kifs: Structure, function, and dynamics. Physiological Reviews, 88, 1089–1118.
Yamada, M., Tanaka-Takiguchi, Y., Hayashi, M., Nishina, M., & Goshima, G. (2017). Multiple kinesin-14 family members drive microtubule minus end-directed transport in plant cells. The Journal of Cell Biology, 216, 1705–1714.
Roostalu, J., Hentrich, C., Bieling, P., Telley, I. A., Schiebel, E., & Surrey, T. (2011). Directional switching of the kinesin Cin8 through motor coupling. Science, 332, 94–99.
Luo, W., Liao, M., Liao, Y., Chen, X., Huang, C., Fan, J., & Liao, W. (2018). The role of kinesin Kif18a in the invasion and metastasis of hepatocellular carcinoma. World Journal of Surgical Oncology, 16, 36.
Huang, Y., Wang, H., Lian, Y., Wu, X., Zhou, L., Wang, J., Deng, M., & Huang, Y. (2018). Upregulation of kinesin family member 4a enhanced cell proliferation via activation of Akt signaling and predicted a poor prognosis in hepatocellular carcinoma. Cell Death & Disease, 9, 141.
Moamer, A., Hachim, I. Y., Binothman, N., Wang, N., Lebrun, J. J., & Ali, S. (2019). A role for kinesin-1 subunits Kif5b/Klc1 in regulating epithelial mesenchymal plasticity in breast tumorigenesis. eBioMedicine, 45, 92–107.
Wang, J., Cui, F., Wang, X., Xue, Y., Chen, J., Yu, Y., Lu, H., Zhang, M., Tang, H., & Peng, Z. (2015). Elevated kinesin family member 26b is a prognostic biomarker and a potential therapeutic target for colorectal cancer. Journal of Experimental & Clinical Cancer Research, 34, 13.
Wang, Q., Zhao, Z. B., Wang, G., Hui, Z., Wang, M. H., Pan, J. F., & Zheng, H. (2013). High expression of Kif26b in breast cancer associates with poor prognosis. PLoS One, 8, E61640.
Yu, Y., Wang, X. Y., Sun, L., Wang, Y. L., Wan, Y. F., Li, X. Q., & Feng, Y. M. (2014). Inhibition of Kif22 suppresses cancer cell proliferation by delaying mitotic exit through upregulating Cdc25c expression. Carcinogenesis, 35, 1416–1425.
Miyata, H., Shimada, K., Morohoshi, A., Oura, S., Matsumura, T., Xu, Z., Oyama, Y., & Ikawa, M. (2020). Testis-enriched kinesin Kif9 is important for progressive motility in mouse spermatozoa. The FASEB Journal, 34, 5389–5400.
She, Z. Y., Yu, K. W., Zhong, N., Xiao, Y., Wei, Y. L., Lin, Y., Li, Y. L., & Lu, M. H. (2020a). Kinesin-7 Cenp-E regulates chromosome alignment and genome stability of spermatogenic cells. Cell Death Discovery, 6, 25.
She, Z. Y., Zhong, N., Yu, K. W., Xiao, Y., Wei, Y. L., Lin, Y., Li, Y. L., & Lu, M. H. (2020b). Kinesin-5 Eg5 is essential for spindle assembly and chromosome alignment of mouse spermatocytes. Cell Div, 15, 6.
Hartman, M. A., & Spudich, J. A. (2012). The myosin superfamily at a glance. Journal of Cell Science, 125, 1627–1632.
Foth, B. J., Goedecke, M. C., & Soldati, D. (2006). New insights into myosin evolution and classification. Proceedings of the National Academy of Sciences of the United States of America, 103, 3681–3686.
Navinés-Ferrer, A., & Martín, M. (2020). Long-tailed unconventional class I myosins in health and disease. International Journal of Molecular Sciences, 21, 2555.
Tamura, K., Iwabuchi, K., Fukao, Y., Kondo, M., Okamoto, K., Ueda, H., Nishimura, M., & Hara-Nishimura, I. (2013). Myosin Xi-I links the nuclear membrane to the cytoskeleton to control nuclear movement and shape in arabidopsis. Current Biology, 23, 1776–1781.
Hyenne, V., Harf, J. C., Latz, M., Maro, B., Wolfrum, U., & Simmler, M. C. (2007). Vezatin, a ubiquitous protein of adherens cell-cell junctions, is exclusively expressed in germ cells in mouse testis. Reproduction, 133, 563–574.
Velichkova, M., Guttman, J., Warren, C., Eng, L., Kline, K., Vogl, A. W., & Hasson, T. (2002). A human homologue of drosophila kelch associates with myosin-VIIa in specialized adhesion junctions. Cell Motility and the Cytoskeleton, 51, 147–164.
Wen, Q., Wu, S., Lee, W. M., Wong, C. K. C., Lui, W. Y., Silvestrini, B., & Cheng, C. Y. (2019). Myosin VIIa supports spermatid/organelle transport and cell adhesion during spermatogenesis in the rat testis. Endocrinology, 160, 484–503.
Poirier, K., Lebrun, N., Broix, L., Tian, G., Saillour, Y., Boscheron, C., Parrini, E., Valence, S., Pierre, B. S., Oger, M., Lacombe, D., Genevieve, D., Fontana, E., Darra, F., Cances, C., Barth, M., Bonneau, D., Bernadina, B. D., N’guyen, S., … Chelly, J. (2013). Mutations in Tubg1, Dync1h1, Kif5c and Kif2a cause malformations of cortical development and microcephaly. Nature Genetics, 45, 639–647.
Weedon, M. N., Hastings, R., Caswell, R., Xie, W., Paszkiewicz, K., Antoniadi, T., Williams, M., King, C., Greenhalgh, L., Newbury-Ecob, R., & Ellard, S. (2011). Exome sequencing identifies a Dync1h1 mutation in a large pedigree with dominant axonal charcot-marie-tooth disease. American Journal of Human Genetics, 89, 308–312.
Harms, M. B., Ori-Mckenney, K. M., Scoto, M., Tuck, E. P., Bell, S., Ma, D., Masi, S., Allred, P., Al-Lozi, M., Reilly, M. M., Miller, L. J., Jani-Acsadi, A., Pestronk, A., Shy, M. E., Muntoni, F., Vallee, R. B., & Baloh, R. H. (2012). Mutations in the tail domain of Dync1h1 cause dominant spinal muscular atrophy. Neurology, 78, 1714–1720.
Vissers, L. E., De Ligt, J., Gilissen, C., Janssen, I., Steehouwer, M., De Vries, P., Van Lier, B., Arts, P., Wieskamp, N., Del Rosario, M., Van Bon, B. W., Hoischen, A., De Vries, B. B., Brunner, H. G., & Veltman, J. A. (2010). A de novo paradigm for mental retardation. Nature Genetics, 42, 1109–1112.
Willemsen, M. H., Vissers, L. E., Willemsen, M. A., Van Bon, B. W., Kroes, T., De Ligt, J., De Vries, B. B., Schoots, J., Lugtenberg, D., Hamel, B. C., Van Bokhoven, H., Brunner, H. G., Veltman, J. A., & Kleefstra, T. (2012). Mutations in Dync1h1 cause severe intellectual disability with neuronal migration defects. Journal of Medical Genetics, 49, 179–183.
Chen, Y., Xu, Y., Li, G., Li, N., Yu, T., Yao, R. E., Wang, X., Shen, Y., & Wang, J. (2017). Exome sequencing identifies de novo Dync1h1 mutations associated with distal spinal muscular atrophy and malformations of cortical development. Journal of Child Neurology, 32, 379–386.
Dagoneau, N., Goulet, M., Genevieve, D., Sznajer, Y., Martinovic, J., Smithson, S., Huber, C., Baujat, G., Flori, E., Tecco, L., Cavalcanti, D., Delezoide, A. L., Serre, V., Le Merrer, M., Munnich, A., & Cormier-Daire, V. (2009). Dync2h1 mutations cause asphyxiating thoracic dystrophy and short rib-polydactyly syndrome, type III. American Journal of Human Genetics, 84, 706–711.
El Hokayem, J., Huber, C., Couve, A., Aziza, J., Baujat, G., Bouvier, R., Cavalcanti, D. P., Collins, F. A., Cordier, M. P., Delezoide, A. L., Gonzales, M., Johnson, D., Le Merrer, M., Levy-Mozziconacci, A., Loget, P., Martin-Coignard, D., Martinovic, J., Mortier, G. R., Perez, M. J., … Cormier-Daire, V. (2012). Nek1 and Dync2h1 are both involved in short rib polydactyly majewski type but not in beemer langer cases. Journal of Medical Genetics, 49, 227–233.
Schmidts, M., Arts, H. H., Bongers, E. M., Yap, Z., Oud, M. M., Antony, D., Duijkers, L., Emes, R. D., Stalker, J., Yntema, J. B., Plagnol, V., Hoischen, A., Gilissen, C., Forsythe, E., Lausch, E., Veltman, J. A., Roeleveld, N., Superti-Furga, A., Kutkowska-Kazmierczak, A., … Mitchison, H. M. (2013). Exome sequencing identifies Dync2h1 mutations as a common cause of asphyxiating thoracic dystrophy (Jeune Syndrome) without major polydactyly, renal or retinal involvement. Journal of Medical Genetics, 50, 309–323.
Merrill, A. E., Merriman, B., Farrington-Rock, C., Camacho, N., Sebald, E. T., Funari, V. A., Schibler, M. J., Firestein, M. H., Cohn, Z. A., Priore, M. A., Thompson, A. K., Rimoin, D. L., Nelson, S. F., Cohn, D. H., & Krakow, D. (2009). Ciliary abnormalities due to defects in the retrograde transport protein Dync2h1 in short-rib polydactyly syndrome. American Journal of Human Genetics, 84, 542–549.
Zuccarello, D., Ferlin, A., Cazzadore, C., Pepe, A., Garolla, A., Moretti, A., Cordeschi, G., Francavilla, S., & Foresta, C. (2008). Mutations in dynein genes in patients affected by isolated non-syndromic asthenozoospermia. Human Reproduction, 23, 1957–1962.
Zariwala, M. A., Leigh, M. W., Ceppa, F., Kennedy, M. P., Noone, P. G., Carson, J. L., Hazucha, M. J., Lori, A., Horvath, J., Olbrich, H., Loges, N. T., Bridoux, A. M., Pennarun, G., Duriez, B., Escudier, E., Mitchison, H. M., Chodhari, R., Chung, E. M., Morgan, L. C., … Knowles, M. R. (2006). Mutations of Dnai1 in primary ciliary dyskinesia: Evidence of founder effect in a common mutation. American Journal of Respiratory and Critical Care Medicine, 174, 858–866.
Blackstone, C. (2018). Converging cellular themes for the hereditary spastic paraplegias. Current Opinion in Neurobiology, 51, 139–146.
Lee, J. R., Srour, M., Kim, D., Hamdan, F. F., Lim, S. H., Brunel-Guitton, C., Decarie, J. C., Rossignol, E., Mitchell, G. A., Schreiber, A., Moran, R., Van Haren, K., Richardson, R., Nicolai, J., Oberndorff, K. M., Wagner, J. D., Boycott, K. M., Rahikkala, E., Junna, N., … Michaud, J. L. (2015). De novo mutations in the motor domain of Kif1a cause cognitive impairment, spastic paraparesis, axonal neuropathy, and cerebellar atrophy. Human Mutation, 36, 69–78.
Hirokawa, N., & Tanaka, Y. (2015). Kinesin Superfamily Proteins (Kifs): Various functions and their relevance for important phenomena in life and diseases. Experimental Cell Research, 334, 16–25.
Caballero Oteyza, A., Battaloglu, E., Ocek, L., Lindig, T., Reichbauer, J., Rebelo, A. P., Gonzalez, M. A., Zorlu, Y., Ozes, B., Timmann, D., Bender, B., Woehlke, G., Zuchner, S., Schols, L., & Schule, R. (2014). Motor protein mutations cause a new form of hereditary spastic paraplegia. Neurology, 82, 2007–2016.
Cogne, B., Latypova, X., Senaratne, L. D. S., Martin, L., Koboldt, D. C., Kellaris, G., Fievet, L., Le Meur, G., Caldari, D., Debray, D., Nizon, M., Frengen, E., Bowne, S. J., Lives, C., Cadena, E. L., Daiger, S. P., Bujakowska, K. M., Pierce, E. A., Gorin, M., … Isidor, B. (2020). Mutations in the kinesin-2 motor Kif3b cause an autosomal-dominant ciliopathy. American Journal of Human Genetics, 106, 893–904.
Dimassi, S., Labalme, A., Ville, D., Calender, A., Mignot, C., Boutry-Kryza, N., De Bellescize, J., Rivier-Ringenbach, C., Bourel-Ponchel, E., Cheillan, D., Simonet, T., Maincent, K., Rossi, M., Till, M., Mougou-Zerelli, S., Edery, P., Saad, A., Heron, D., Des Portes, V., … Lesca, G. (2016). Whole-exome sequencing improves the diagnosis yield in sporadic infantile spasm syndrome. Clinical Genetics, 89, 198–204.
Meier, N., Bruder, E., Lapaire, O., Hoesli, I., Kang, A., Hench, J., Hoeller, S., De Geyter, J., Miny, P., Heinimann, K., Chaoui, R., Tercanli, S., & Filges, I. (2019). Exome sequencing of fetal anomaly syndromes: Novel phenotype-genotype discoveries. European Journal of Human Genetics, 27, 730–737.
Reid, E., Kloos, M., Ashley-Koch, A., Hughes, L., Bevan, S., Svenson, I. K., Graham, F. L., Gaskell, P. C., Dearlove, A., Pericak-Vance, M. A., Rubinsztein, D. C., & Marchuk, D. A. (2002). A kinesin heavy chain (Kif5a) mutation in hereditary spastic paraplegia (Spg10). American Journal of Human Genetics, 71, 1189–1194.
Goizet, C., Boukhris, A., Mundwiller, E., Tallaksen, C., Forlani, S., Toutain, A., Carriere, N., Paquis, V., Depienne, C., Durr, A., Stevanin, G., & Brice, A. (2009). Complicated forms of autosomal dominant hereditary spastic paraplegia are frequent in Spg10. Human Mutation, 30, E376–E385.
Crimella, C., Baschirotto, C., Arnoldi, A., Tonelli, A., Tenderini, E., Airoldi, G., Martinuzzi, A., Trabacca, A., Losito, L., Scarlato, M., Benedetti, S., Scarpini, E., Spinicci, G., Bresolin, N., & Bassi, M. T. (2012). Mutations in the motor and stalk domains of Kif5a in spastic paraplegia type 10 and in axonal charcot-marie-tooth type 2. Clinical Genetics, 82, 157–164.
Liu, Y. T., Laura, M., Hersheson, J., Horga, A., Jaunmuktane, Z., Brandner, S., Pittman, A., Hughes, D., Polke, J. M., Sweeney, M. G., Proukakis, C., Janssen, J. C., Auer-Grumbach, M., Zuchner, S., Shields, K. G., Reilly, M. M., & Houlden, H. (2014). Extended phenotypic spectrum of Kif5a mutations: From spastic paraplegia to axonal neuropathy. Neurology, 83, 612–619.
Brenner, D., Yilmaz, R., Muller, K., Grehl, T., Petri, S., Meyer, T., Grosskreutz, J., Weydt, P., Ruf, W., Neuwirth, C., Weber, M., Pinto, S., Claeys, K. G., Schrank, B., Jordan, B., Knehr, A., Gunther, K., Hubers, A., Zeller, D., … German, A. L. S. N. M. N. D. N. E. T. (2018). Hot-spot Kif5a mutations cause familial ALS. Brain, 141, 688–697.
Duis, J., Dean, S., Applegate, C., Harper, A., Xiao, R., He, W., Dollar, J. D., Sun, L. R., Waberski, M. B., Crawford, T. O., Hamosh, A., & Stafstrom, C. E. (2016). Kif5a mutations cause an infantile onset phenotype including severe myoclonus with evidence of mitochondrial dysfunction. Annals of Neurology, 80, 633–637.
Rydzanicz, M., Jagla, M., Kosinska, J., Tomasik, T., Sobczak, A., Pollak, A., Herman-Sucharska, I., Walczak, A., Kwinta, P., & Ploski, R. (2017). Kif5a de novo mutation associated with myoclonic seizures and neonatal onset progressive leukoencephalopathy. Clinical Genetics, 91, 769–773.
Filosto, M., Piccinelli, S. C., Palmieri, I., Necchini, N., Valente, M., Zanella, I., Biasiotto, G., Lorenzo, D. D., Cereda, C., & Padovani, A. (2018). A novel mutation in the stalk domain of Kif5a causes a slowly progressive atypical motor syndrome. Journal of Clinical Medicine, 8, 17.
Konjikusic, M. J., Yeetong, P., Boswell, C. W., Lee, C., Roberson, E. C., Ittiwut, R., Suphapeetiporn, K., Ciruna, B., Gurnett, C. A., Wallingford, J. B., Shotelersuk, V., & Gray, R. S. (2018). Mutations in kinesin family member 6 reveal specific role in ependymal cell ciliogenesis and human neurological development. PLoS Genetics, 14, E1007817.
Westland, R., Verbitsky, M., Vukojevic, K., Perry, B. J., Fasel, D. A., Zwijnenburg, P. J., Bokenkamp, A., Gille, J. J., Saraga-Babic, M., Ghiggeri, G. M., D’agati, V. D., Schreuder, M. F., Gharavi, A. G., Van Wijk, J. A., & Sanna-Cherchi, S. (2015). Copy number variation analysis identifies novel cakut candidate genes in children with a solitary functioning kidney. Kidney International, 88, 1402–1410.
Sleiman, P. M. A., March, M., Nguyen, K., Tian, L., Pellegrino, R., Hou, C., Dridi, W., Sager, M., Housawi, Y. H., & Hakonarson, H. (2017). Loss-of-function mutations in Kif15 underlying a braddock-carey genocopy. Human Mutation, 38, 507–510.
Alsahli, S., Arold, S. T., Alfares, A., Alhaddad, B., Al Balwi, M., Kamsteeg, E. J., Al-Twaijri, W., & Alfadhel, M. (2018). Kif16b is a candidate gene for a novel autosomal-recessive intellectual disability syndrome. American Journal of Medical Genetics. Part A, 176, 1602–1609.
Yamada, K., Andrews, C., Chan, W. M., Mckeown, C. A., Magli, A., De Berardinis, T., Loewenstein, A., Lazar, M., O’keefe, M., Letson, R., London, A., Ruttum, M., Matsumoto, N., Saito, N., Morris, L., Del Monte, M., Johnson, R. H., Uyama, E., Houtman, W. A., … Engle, E. C. (2003). Heterozygous mutations of the kinesin Kif21a in congenital fibrosis of the extraocular muscles type 1 (Cfeom1). Nature Genetics, 35, 318–321.
Asselin, L., Rivera Alvarez, J., Heide, S., Bonnet, C. S., Tilly, P., Vitet, H., Weber, C., Bacino, C. A., Baranano, K., Chassevent, A., Dameron, A., Faivre, L., Hanchard, N. A., Mahida, S., Mcwalter, K., Mignot, C., Nava, C., Rastetter, A., Streff, H., … Godin, J. D. (2020). Mutations in the Kif21b kinesin gene cause neurodevelopmental disorders through imbalanced canonical motor activity. Nature Communications, 11, 2441.
Nibbeling, E. A. R., Duarri, A., Verschuuren-Bemelmans, C. C., Fokkens, M. R., Karjalainen, J. M., Smeets, C., De Boer-Bergsma, J. J., Van Der Vries, G., Dooijes, D., Bampi, G. B., Van Diemen, C., Brunt, E., Ippel, E., Kremer, B., Vlak, M., Adir, N., Wijmenga, C., Van De Warrenburg, B. P. C., Franke, L., … Verbeek, D. S. (2017). Exome sequencing and network analysis identifies shared mechanisms underlying spinocerebellar ataxia. Brain, 140, 2860–2878.
Wojcik, M. H., Okada, K., Prabhu, S. P., Nowakowski, D. W., Ramsey, K., Balak, C., Rangasamy, S., Brownstein, C. A., Schmitz-Abe, K., Cohen, J. S., Fatemi, A., Shi, J., Grant, E. P., Narayanan, V., Ho, H. H., & Agrawal, P. B. (2018). De Novo variant in Kif26b is associated with pontocerebellar hypoplasia with infantile spinal muscular atrophy. American Journal of Medical Genetics. Part A, 176, 2623–2629.
Martinsson, T., Oldfors, A., Darin, N., Berg, K., Tajsharghi, H., Kyllerman, M., & Wahlstrom, J. (2000). Autosomal dominant myopathy: Missense mutation (Glu-706 --> Lys) in the myosin heavy chain IIa gene. Proceedings of the National Academy of Sciences of the United States of America, 97, 14614–14619.
Chong, J. X., Burrage, L. C., Beck, A. E., Marvin, C. T., Mcmillin, M. J., Shively, K. M., Harrell, T. M., Buckingham, K. J., Bacino, C. A., Jain, M., Alanay, Y., Berry, S. A., Carey, J. C., Gibbs, R. A., Lee, B. H., Krakow, D., Shendure, J., Nickerson, D. A., University Of Washington Center For Mendelian, G, & Bamshad, M. J. (2015). Autosomal-dominant multiple pterygium syndrome is caused by mutations in Myh3. American Journal of Human Genetics, 96, 841–849.
Carapito, R., Goldenberg, A., Paul, N., Pichot, A., David, A., Hamel, A., Dumant-Forest, C., Leroux, J., Ory, B., Isidor, B., & Bahram, S. (2016). Protein-altering Myh3 variants are associated with a spectrum of phenotypes extending to spondylocarpotarsal synostosis syndrome. European Journal of Human Genetics, 24, 1746–1751.
Scala, M., Accogli, A., De Grandis, E., Allegri, A., Bagowski, C. P., Shoukier, M., Maghnie, M., & Capra, V. (2018). A novel pathogenic Myh3 mutation in a child with sheldon-hall syndrome and vertebral fusions. American Journal of Medical Genetics. Part A, 176, 663–667.
Cameron-Christie, S. R., Wells, C. F., Simon, M., Wessels, M., Tang, C. Z. N., Wei, W., Takei, R., Aarts-Tesselaar, C., Sandaradura, S., Sillence, D. O., Cordier, M. P., Veenstra-Knol, H. E., Cassina, M., Ludwig, K., Trevisson, E., Bahlo, M., Markie, D. M., Jenkins, Z. A., & Robertson, S. P. (2018). Recessive spondylocarpotarsal synostosis syndrome due to compound heterozygosity for variants in Myh3. American Journal of Human Genetics, 102, 1115–1125.
Toydemir, R. M., Rutherford, A., Whitby, F. G., Jorde, L. B., Carey, J. C., & Bamshad, M. J. (2006). Mutations in embryonic myosin heavy chain (Myh3) cause freeman-sheldon syndrome and sheldon-hall syndrome. Nature Genetics, 38, 561–565.
Tajsharghi, H., Kimber, E., Kroksmark, A. K., Jerre, R., Tulinius, M., & Oldfors, A. (2008). Embryonic myosin heavy-chain mutations cause distal arthrogryposis and developmental myosin myopathy that persists postnatally. Archives of Neurology, 65, 1083–1090.
Poetter, K., Jiang, H., Hassanzadeh, S., Master, S. R., Chang, A., Dalakas, M. C., Rayment, I., Sellers, J. R., Fananapazir, L., & Epstein, N. D. (1996). Mutations in either the essential or regulatory light chains of myosin are associated with a rare myopathy in human heart and skeletal muscle. Nature Genetics, 13, 63–69.
Olson, T. M., Karst, M. L., Whitby, F. G., & Driscoll, D. J. (2002). Myosin light chain mutation causes autosomal recessive cardiomyopathy with mid-cavitary hypertrophy and restrictive physiology. Circulation, 105, 2337–2340.
Richard, P., Charron, P., Carrier, L., Ledeuil, C., Cheav, T., Pichereau, C., Benaiche, A., Isnard, R., Dubourg, O., Burban, M., Gueffet, J. P., Millaire, A., Desnos, M., Schwartz, K., Hainque, B., Komajda, M., & Project, E. H. F. (2003). Hypertrophic cardiomyopathy: Distribution of disease genes, spectrum of mutations, and implications for a molecular diagnosis strategy. Circulation, 107, 2227–2232.
Jay, A., Chikarmane, R., Poulik, J., & Misra, V. K. (2013). Infantile hypertrophic cardiomyopathy associated with a novel Myl3 mutation. Cardiology, 124, 248–251.
Ching, Y. H., Ghosh, T. K., Cross, S. J., Packham, E. A., Honeyman, L., Loughna, S., Robinson, T. E., Dearlove, A. M., Ribas, G., Bonser, A. J., Thomas, N. R., Scotter, A. J., Caves, L. S., Tyrrell, G. P., Newbury-Ecob, R. A., Munnich, A., Bonnet, D., & Brook, J. D. (2005). Mutation in myosin heavy chain 6 causes atrial septal defect. Nature Genetics, 37, 423–428.
Carniel, E., Taylor, M. R., Sinagra, G., Di Lenarda, A., Ku, L., Fain, P. R., Boucek, M. M., Cavanaugh, J., Miocic, S., Slavov, D., Graw, S. L., Feiger, J., Zhu, X. Z., Dao, D., Ferguson, D. A., Bristow, M. R., & Mestroni, L. (2005). Alpha-myosin heavy chain: A sarcomeric gene associated with dilated and hypertrophic phenotypes of cardiomyopathy. Circulation, 112, 54–59.
Holm, H., Gudbjartsson, D. F., Sulem, P., Masson, G., Helgadottir, H. T., Zanon, C., Magnusson, O. T., Helgason, A., Saemundsdottir, J., Gylfason, A., Stefansdottir, H., Gretarsdottir, S., Matthiasson, S. E., Thorgeirsson, G. M., Jonasdottir, A., Sigurdsson, A., Stefansson, H., Werge, T., Rafnar, T., … Stefansson, K. (2011). A rare variant in Myh6 is associated with high risk of sick sinus syndrome. Nature Genetics, 43, 316–320.
Fananapazir, L., Dalakas, M. C., Cyran, F., Cohn, G., & Epstein, N. D. (1993). Missense mutations in the beta-myosin heavy-chain gene cause central core disease in hypertrophic cardiomyopathy. Proceedings of the National Academy of Sciences of the United States of America, 90, 3993–3997.
Rayment, I., Holden, H. M., Sellers, J. R., Fananapazir, L., & Epstein, N. D. (1995). Structural interpretation of the mutations in the beta-cardiac myosin that have been implicated in familial hypertrophic cardiomyopathy. Proceedings of the National Academy of Sciences of the United States of America, 92, 3864–3868.
Bundgaard, H., Havndrup, O., Andersen, P. S., Larsen, L. A., Brandt, N. J., Vuust, J., Kjeldsen, K., & Christiansen, M. (1999). Familial hypertrophic cardiomyopathy associated with a novel missense mutation affecting the Atp-binding region of the cardiac beta-myosin heavy chain. Journal of Molecular and Cellular Cardiology, 31, 745–750.
Blair, E., Redwood, C., De Jesus Oliveira, M., Moolman-Smook, J. C., Brink, P., Corfield, V. A., Ostman-Smith, I., & Watkins, H. (2002). Mutations of the light meromyosin domain of the beta-myosin heavy chain rod in hypertrophic cardiomyopathy. Circulation Research, 90, 263–269.
Erdmann, J., Daehmlow, S., Wischke, S., Senyuva, M., Werner, U., Raible, J., Tanis, N., Dyachenko, S., Hummel, M., Hetzer, R., & Regitz-Zagrosek, V. (2003). Mutation spectrum in a large cohort of unrelated consecutive patients with hypertrophic cardiomyopathy. Clinical Genetics, 64, 339–349.
Van Driest, S. L., Jaeger, M. A., Ommen, S. R., Will, M. L., Gersh, B. J., Tajik, A. J., & Ackerman, M. J. (2004). Comprehensive analysis of the beta-myosin heavy chain gene in 389 unrelated patients with hypertrophic cardiomyopathy. Journal of the American College of Cardiology, 44, 602–610.
Hougs, L., Havndrup, O., Bundgaard, H., Kober, L., Vuust, J., Larsen, L. A., Christiansen, M., & Andersen, P. S. (2005). One third of danish hypertrophic cardiomyopathy patients with Myh7 mutations have mutations [corrected] in Myh7 rod region. European Journal of Human Genetics, 13, 161–165.
Veugelers, M., Bressan, M., Mcdermott, D. A., Weremowicz, S., Morton, C. C., Mabry, C. C., Lefaivre, J. F., Zunamon, A., Destree, A., Chaudron, J. M., & Basson, C. T. (2004). Mutation of perinatal myosin heavy chain associated with a carney complex variant. The New England Journal of Medicine, 351, 460–469.
Seri, M., Cusano, R., Gangarossa, S., Caridi, G., Bordo, D., Lo Nigro, C., Ghiggeri, G. M., Ravazzolo, R., Savino, M., Del Vecchio, M., D’apolito, M., Iolascon, A., Zelante, L. L., Savoia, A., Balduini, C. L., Noris, P., Magrini, U., Belletti, S., Heath, K. E., … Martignetti, J. A. (2000). Mutations in Myh9 result in the may-hegglin anomaly, and fechtner and Sebastian syndromes. The May-Heggllin/Fechtner Syndrome Consortium. Nature Genetics, 26, 103–105.
Heath, K. E., Campos-Barros, A., Toren, A., Rozenfeld-Granot, G., Carlsson, L. E., Savige, J., Denison, J. C., Gregory, M. C., White, J. G., Barker, D. F., Greinacher, A., Epstein, C. J., Glucksman, M. J., & Martignetti, J. A. (2001). Nonmuscle myosin heavy chain IIa mutations define a spectrum of autosomal dominant macrothrombocytopenias: May-Hegglin anomaly and fechtner, sebastian, epstein, and alport-like syndromes. American Journal of Human Genetics, 69, 1033–1045.
Kunishima, S., Matsushita, T., Kojima, T., Amemiya, N., Choi, Y. M., Hosaka, N., Inoue, M., Jung, Y., Mamiya, S., Matsumoto, K., Miyajima, Y., Zhang, G., Ruan, C., Saito, K., Song, K. S., Yoon, H. J., Kamiya, T., & Saito, H. (2001). Identification of six novel Myh9 mutations and genotype-phenotype relationships in autosomal dominant macrothrombocytopenia with leukocyte inclusions. Journal of Human Genetics, 46, 722–729.
Seri, M., Savino, M., Bordo, D., Cusano, R., Rocca, B., Meloni, I., Di Bari, F., Koivisto, P. A., Bolognesi, M., Ghiggeri, G. M., Landolfi, R., Balduini, C. L., Zelante, L., Ravazzolo, R., Renieri, A., & Savoia, A. (2002). Epstein Syndrome: Another renal disorder with mutations in the nonmuscle myosin heavy chain 9 gene. Human Genetics, 110, 182–186.
Arrondel, C., Vodovar, N., Knebelmann, B., Grunfeld, J. P., Gubler, M. C., Antignac, C., & Heidet, L. (2002). Expression of the nonmuscle myosin heavy chain IIa in the human kidney and screening for Myh9 mutations in epstein and fechtner syndromes. Journal of the American Society of Nephrology, 13, 65–74.
Deutsch, S., Rideau, A., Bochaton-Piallat, M. L., Merla, G., Geinoz, A., Gabbiani, G., Schwede, T., Matthes, T., Antonarakis, S. E., & Beris, P. (2003). Asp1424asn Myh9 mutation results in an unstable protein responsible for the phenotypes in May-Hegglin Anomaly/Fechtner syndrome. Blood, 102, 529–534.
Seri, M., Pecci, A., Di Bari, F., Cusano, R., Savino, M., Panza, E., Nigro, A., Noris, P., Gangarossa, S., Rocca, B., Gresele, P., Bizzaro, N., Malatesta, P., Koivisto, P. A., Longo, I., Musso, R., Pecoraro, C., Iolascon, A., Magrini, U., … Savoia, A. (2003). Myh9-related disease: May-Hegglin Anomaly, Sebastian Syndrome, Fechtner Syndrome, and Epstein Syndrome are not distinct entities but represent a variable expression of a single illness. Medicine (Baltimore), 82, 203–215.
Mhatre, A. N., Kim, Y., Brodie, H. A., & Lalwani, A. K. (2003). Macrothrombocytopenia and progressive deafness is due to a mutation in Myh9. Otology & Neurotology, 24, 205–209.
Lalwani, A. K., Goldstein, J. A., Kelley, M. J., Luxford, W., Castelein, C. M., & Mhatre, A. N. (2000). Human nonsyndromic hereditary deafness Dfna17 is due to a mutation in nonmuscle myosin Myh9. American Journal of Human Genetics, 67, 1121–1128.
Tuzovic, L., Yu, L., Zeng, W., Li, X., Lu, H., Lu, H. M., Gonzalez, K. D., & Chung, W. K. (2013). A human de novo mutation in myh10 phenocopies the loss of function mutation in mice. Rare Diseases, 1, E26144.
Hamdan, F. F., Srour, M., Capo-Chichi, J. M., Daoud, H., Nassif, C., Patry, L., Massicotte, C., Ambalavanan, A., Spiegelman, D., Diallo, O., Henrion, E., Dionne-Laporte, A., Fougerat, A., Pshezhetsky, A. V., Venkateswaran, S., Rouleau, G. A., & Michaud, J. L. (2014). De novo mutations in moderate or severe intellectual disability. PLoS Genetics, 10, E1004772.
Zhu, L., Vranckx, R., Khau Van Kien, P., Lalande, A., Boisset, N., Mathieu, F., Wegman, M., Glancy, L., Gasc, J. M., Brunotte, F., Bruneval, P., Wolf, J. E., Michel, J. B., & Jeunemaitre, X. (2006). Mutations in myosin heavy chain 11 cause a syndrome associating thoracic aortic aneurysm/aortic dissection and patent ductus arteriosus. Nature Genetics, 38, 343–349.
Donaudy, F., Snoeckx, R., Pfister, M., Zenner, H. P., Blin, N., Di Stazio, M., Ferrara, A., Lanzara, C., Ficarella, R., Declau, F., Pusch, C. M., Nurnberg, P., Melchionda, S., Zelante, L., Ballana, E., Estivill, X., Van Camp, G., Gasparini, P., & Savoia, A. (2004). Nonmuscle myosin heavy-chain gene Myh14 is expressed in cochlea and mutated in patients affected by autosomal dominant hearing impairment (Dfna4). American Journal of Human Genetics, 74, 770–776.
Choi, B. O., Kang, S. H., Hyun, Y. S., Kanwal, S., Park, S. W., Koo, H., Kim, S. B., Choi, Y. C., Yoo, J. H., Kim, J. W., Park, K. D., Choi, K. G., Kim, S. J., Zuchner, S., & Chung, K. W. (2011). A complex phenotype of peripheral neuropathy, myopathy, hoarseness, and hearing loss is linked to an autosomal dominant mutation in Myh14. Human Mutation, 32, 669–677.
Hermo, L., Pelletier, R.-M., Cyr, D. G., & Smith, C. E. (2010a). Surfing the wave, cycle, life history, and genes/proteins expressed by testicular germ cells. Part 3: Developmental changes in spermatid flagellum and cytoplasmic droplet and interaction of sperm with the zona pellucida and egg plasma membrane. Microscopy Research and Technique, 73, 320–363.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
The authors have nothing to declare,
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Wu, S. et al. (2021). Motor Proteins and Spermatogenesis. In: Cheng, C., Sun, F. (eds) Molecular Mechanisms in Spermatogenesis. Advances in Experimental Medicine and Biology, vol 1381. Springer, Cham. https://doi.org/10.1007/978-3-030-77779-1_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-77779-1_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-77778-4
Online ISBN: 978-3-030-77779-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)