Abstract
The preimplantation mammalian embryo is a simplistic, self-contained, and a superior model for investigating the inherent complexities of cell fate decision mechanisms. All mammals begin their humble journey from a single-cell fertilized zygote contained within a proteinaceous coat called the zona pellucida. The zygote embarks on a series of well-orchestrated events, beginning with the activation of embryonic genome, transition from meiotic to mitotic divisions, spatial organization of the cells, timely differentiation into committed trophectoderm (TE) and primitive endoderm (PrE), and ultimately escape from zona pellucida for implantation into the uterus. The entire development of preimplantation embryo can be studied in vitro using a minimalistic and defined culture system. The ease of culture along with the ability to manipulate gene expression and image the embryos makes them an ideal model system for investigation into the first two of several cell fate decisions made by the embryo that result in a pluripotent epiblast (EPI) and differentiated TE and PrE lineages. This chapter reviews our latest knowledge of preimplantation embryo development, setting the stage for understanding placental development in subsequent chapters in this Book.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
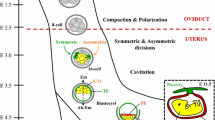
The developing mammalian embryo is a unique and valuable model system for studying cell fate decisions. One of the fundamental questions in biology is how a relatively simple single-cell zygote achieves the seemingly impossible task of generating a complex multicellular being. The embryonic cells are tasked with having to make orderly cell fate decisions in a spatiotemporal manner, differentiate into specialized cell types, self-organize into higher order tissues and organ systems, and ultimately generate a functional being. In this regard, the developing mammalian embryo, a “self-contained” system serves as a unique model system for in vitro investigation into genetic, epigenetic, and mechanochemical forces that are at the basis for regulating cell fate (White et al. 2018). Early stages of mammalian embryo development, from maturation of oocytes to blastocyst, can be recapitulated in vitro using a simplistic culture system without the need for external maternal cues (Whitten and Biggers 1968). The recent discovery of genome editors now expands the range of mammalian species (beyond mice) available for genetic manipulation and for comparative studies. Advances in sequencing technologies, including the ability to sequence single cells, now allow us to capture genetic and epigenetic signature at a higher resolution within the embryos. Finally, breakthroughs in live cell imaging permit tracking of respective morphological fates in vitro (Nagy et al. 2003; Brinster 1963). Cumulatively, the advantages offered by the use of the embryo as a model system and numerous technological advantages are facilitating major breakthroughs in our understanding of first two cell fate decisions: (1) the emergence of trophectoderm (TE) and inner cell mass (ICM) and (2) the differentiation of ICM into pluripotent epiblast (EPI) and committed primitive endoderm (PrE). In this chapter, key landmark events in the development of blastocyst are discussed (Fig. 1). These include awakening of the oocyte by sperm penetration and subsequent transition from meiotic to mitotic divisions (Fig. 2a–f), reductional cleavage divisions resulting in the progressive decline in the size of the embryonic cells and the emergence of heterogeneities in the cleaved blastomeres (Fig. 2g), self-organization of cleaved blastomeres into inside and outside cells (Fig. 2h), compaction of the outer cells and the erasure of distinct cell boundaries (Fig. 2h–j), development of trophectoderm and fluid-filled blastocyst cavity (Fig. 2k), generation of extraembryonic endoderm, and ultimately hatching and implantation of the embryo (Fig. 2l–n). Among the mammalian embryos, the mouse zygote has been studied extensively and will be the basis for discussion below. Several recent articles on mammalian blastocyst development have been published, and the readers are encouraged to peruse the literature for additional information (White et al. 2018; White and Plachta 2020; Zhang and Hiiragi 2018; Leung and Zernicka-Goetz 2015).
Mouse embryo preimplantation development. The embryo undergoes successive and timely cleavage divisions resulting in an increased number of totipotent blastomeres with a progressively smaller cell size. In the compact morula, these blastomeres become self-organized into inside and outside cells. By embryonic day (E) 2.5 late eight-cell stage, the boundaries between the outside cells begin to disappear by “compaction.” By E3.5, the embryo transforms into a fluid-filled blastocyst with outside trophectoderm (TE) and inner cell mass (ICM). By E4.5, the ICM cells differentiate further into primitive endoderm cells (PrE) lining the blastocoel cavity and pluripotent epiblast (EPI) cells at embryonic pole. TE marks the first visual and deterministic cell fate decision followed by the development of PrE making the second fate decision. In the figure, cell lineages are marked by distinctive colors to visually differentiate them
Key landmark events leading to blastocyst development. (a) Oocyte following LH surge resume meiosis I. Actin microfilaments self-organize into meiotic spindles to one side of the embryo resulting from asymmetrical cortical softening. (b) Completion of meiosis I results in extrusion of one half of the chromosomes. (c) Extruded chromosomes with small amounts of cytoplasm result in the formation of residual cell, first polar body. Cytoplasmic streaming holds the now metaphase II in place and is arrested at this stage. (d) Penetration of sperm following fertilization results in the completion of meiosis II and extrusion of second polar body. (e) The now haploid oocyte genome and sperm genomes undergo decondensation, accumulate nuclear membrane, and generate male and female pronuclei. (f) The now fertilized zygote begins transition to mitotic cleavage divisions. The maternally deposited messages mediate the transition until zygotic genome activation that takes place at two-cell stage in mouse embryos. (g) Asymmetries in epigenetic signatures among blastomeres arise at four-cell stage. Two of the four blastomeres express high levels of H3R26 methylation. (h) By late eight-cell stage apicobasal polarization emerges in the outside cells. On the apical surface: microvilli develop on the contact-free cell surface. Actin filaments and actin-binding protein Ezrin localize to the apical surface alongside Par3/Par6/aPkc apical polarization complex. On the basal surface, E-cadherin accumulates at the junctional complex alongside Par1/Jam1 and Na/K ATPase. (i) By 16-cell stage post-compaction embryo, Hippo signaling is fixed. In the outside cells, Hippo signaling is inactive, resulting in the upregulation of deterministic TE-specification gene, Cdx2. In the inside cells, hippo signaling is activated, resulting in the loss of Cdx2 and upregulation of pluripotency gene, Pou5f1. (j) The actin rings in the outside cells stabilize and mature junctions in a zipper-like mechanism. (k) Activation of sodium pumps resulting in pumping of sodium ions and diffusion of water resulting in progressive accumulation of fluids and fusion of microlumens to form blastocyst cavity. (l) Inside pluripotent cells differentiate into Nanog +ve EPI cells (red) and Gata6 +ve PrE cells (yellow) in a characteristic salt-and-pepper fashion. (m) The PrE cells via a combination of apoptosis and migration, resulting in the lining of PrE cells towards the blastocyst cavity. The blastocyst via a combination of mechanical forces resulting from expansion of blastocyst and expression of proteases result in the thinning and eventual rupture of zona pellucida. (n) Expanded blastocyst escapes from zona pellucida in a process known as zona hatching. (The composite figure has been generated from White et al. (2018) with permission from Elsevier Journal)
2 Setting the Stage: Fertilization Awakens the Oocyte and Jumpstarts Embryonic Development
Embryonic development begins with the fusion of a haploid sperm with the oocyte. The timeline for sperm and egg development is heterogeneous in both sexes. Both gametes start their journey as primordial germ cells (PGC)—a small cluster of cells in the primitive streak at the posterior end of the gastrulating embryo (Ohinata et al. 2009; Saitou and Yamaji 2012; Irie et al. 2014; Ginsburg et al. 1990). Upregulation of a triad of transcription factors Prdm1 (Ohinata et al. 2005), Prdm14 (Yamaji et al. 2008, 2013; Ma et al. 2011; Grabole et al. 2013), and Tfap2c (Schafer et al. 2011; Weber et al. 2010) in a subset of precursor mesodermal cells results in the suppression of somatic program, epigenetic erasure, and reactivation of pluripotency, ultimately giving rise to PGC. The PGC then make their orderly journey to their eventual resting place, the genital ridge—a mesenchyme abutting the mesonephric system. Key differences between the sexes emerge here. In males, upregulation of Nanos2 prevents progression of male germ cells to meiosis, which is resumed postnatally (Suzuki and Saga 2008). In females, the germ cells enter into prophase of meiosis I and are arrested at the diplotene stage (dictyate stage) at the time of birth and undergo sequential albeit discontinuous progression to a haploid stage. The oocyte maintains close contact with the nurse cells—the cumulus cells via transzonal cellular projections that traverse the width of zona pellucida and provide nutritional and transcriptional support, and regulate meiosis (Barrett and Albertini 2010). Postnatally, a surge in luteinizing hormone (LH) will result in the loss of cumulus-oocyte contacts, breakdown of the nuclear membrane, and resumption of meiosis (Bury et al. 2017). In what could be considered as a first departure from symmetry, the cortex of the oocyte softens at one-end promoting migration and assembly of meiotic spindle, followed by cortical thickening to hold the spindle in place (Fig. 2a) (Chaigne et al. 2013). The oocytes lack centrosomes; therefore, the first few divisions are dependent on self-organization of actin filaments (Fig. 2a) (Dumont et al. 2007; Yi et al. 2013). Asymmetrical positioning of the spindle ensures generation and expulsion of a smaller sister cell (polar body) encompassing one half of the chromosomes and a smaller volume of cytoplasm following meiosis I (Fig. 2b, c) (Yi et al. 2011; Mogessie and Schuh 2017). This ensures that bulk of cytoplasm with transcriptional and translational machinery was preserved in the oocyte for progression to mitotic divisions. Following the first meiotic division culminating in generation of the first polar body, the meiotic spindle is assembled in a similar fashion with polymerization of actin filaments, cytoplasmic streaming (Fig. 2d) (Yi et al. 2011), and is arrested at metaphase stage of meiosis II again, awaiting activation by sperm. Following entry of sperm into the oocyte, a sperm-borne oocyte-activating factor (PLC-zeta) triggers reactivation and resumption of meiosis and expulsion of second polar body (Fig. 2d, e) (Clift and Schuh 2013). This marks the first keystone event.
3 Cleavage, Imprinting Erasure, and Embryonic Genome Activation
Following fertilization, the maternal and paternal chromosomes undergo decondensation, and assemble nuclear membranes forming male and female pronuclei (Fig. 2e). The next immediate hurdle is the transition of oocyte from meiotic to mitotic divisions when the now zygotic genome is transcriptionally silent. The maternally deposited transcripts guide this transition and predominantly code for protein transportation, localization, and cell cycle genes (Fig. 2f) (Li and Albertini 2013). Paternal and maternal DNA undergo histone exchange, global DNA methylation, gamete-specific imprinting erasure, and activation of retrotransposons, culminating in the attainment of a totipotent genome (Habibi and Stunnenberg 2017; Jachowicz et al. 2017; Burton and Torres-Padilla 2014). Even though the mammalian embryo does not go through a predetermined cell fate specification as seen in Drosophila, C. elegans, Xenopus, and other lower phyla, it is now clear that subtle differences between blastomeres emerge as early as two-cell stage and major differences are evident by four-cell stage (Fig. 2g) (Tabansky et al. 2013; Fujimori et al. 2003; Piotrowska-Nitsche et al. 2005; Littwin and Denker 2011; Antczak and Van Blerkom 1997). Following DNA replication and first mitotic cleavage division, differences in allocation of maternal transcripts between the two blastomeres, especially in the localization of ribosomal RNAs, are evident at two-cell stage (Piotrowska et al. 2001; Piotrowska and Zernicka-Goetz 2001). Likewise, differences in expression of epigenetic modifiers Prdm14 (Burton et al. 2013) and Carm1 (Torres-Padilla et al. 2007) were reported by the four-cell stage, with two of the four cells expressing high levels of Prdm14 and Carm1. Additionally, differences in H3 methylation H3R17 and H3R26 emerge in blastomeres at the four-cell stage, which tend to bias contribution to ICM (Fig. 2g) (Torres-Padilla et al. 2007; Burton and Torres-Padilla 2014). Carm1 regulates H3R26 methylation and increases expression of Sox2, ultimately biasing the cells to pluripotent ICM fate. It is not clear, what regulates increased expression of Carm1 at the four-cell stage. Prior studies that reported no differences in transcriptional profile among blastomeres until the 16-cell stage in the mouse embryo were clearly constrained by the depth and limitations of sequencing (Dietrich and Hiiragi 2008; Guo et al. 2010; Ralston and Rossant 2008; Wicklow et al. 2014; Alarcon and Marikawa 2005; Motosugi et al. 2005; Hiiragi and Solter 2004). In summary, though the blastomeres are overtly homogenous and seemingly identical morphologically, heterogeneities are built into blastomeres by the four-cell stage, setting the stage for lineage specification (Tabansky et al. 2013; Fujimori et al. 2003). Regardless, the fate of the blastomeres remains flexible and is only finalized at the blastocyst stage to accommodate for stochastic errors that are to be expected in the developing embryo.
4 Compaction: First Visual Departure
Following a series of mitotic cleavage events that lead to a progressively smaller and morphologically indistinguishable blastomeres within the zona pellucida, the first overt morphological change that disrupts the uniformity is the compaction of blastomeres soon after the eight-cell stage (Ziomek and Johnson 1980). Blastomeres on the outside establish apicobasal polarity with the accumulation of microvilli, actin-binding protein Ezrin, and actomyosin complex on the contact-free apical cell surface (Fig. 2h) (Ducibella et al. 1977; Louvet et al. 1996; Vinot et al. 2005). On the basolateral surface, E-cadherin accumulates in the junctional complexes (Vestweber et al. 1987; Ziomek and Johnson 1980). It is yet unclear as to what triggers apical polarity at the cell surface, but preliminary reports suggest that actomyosin complex is accumulated at the apical cortex by phospholipase C-mediated hydrolysis of phosphoinol phosphate 2 and activation of protein kinase C (Zhu et al. 2017). Activated protein kinase activates RhoA, which in turn triggers polarization of the actin network, accumulation of the Par3-Par6-aPKC complex (Vinot et al. 2005), and ultimately the formation of an apical domain. In the cell-cell contact basolateral surface, accumulation of Par1 (Vinot et al. 2005), Jam-1 (Fig. 2h) (Thomas et al. 2004), and Na+/K+ ATPase (Watson and Kidder 1988) was seen alongside E-cadherin (Fig. 2k). Accumulation of E-cadherin at the basolateral surface increases the surface area of cell contacts, resulting in a gradual flattening of the outer blastomeres and erasure of distinct cell boundaries, and establishment of a primitive epithelium-like structure in the embryo (Fig. 2j) (Johnson 2009; Ducibella and Anderson 1975). It was long believed that accumulation of E-cadherin at junctional complexes and an increase in intercellular adhesion were responsible for morphological changes during compaction. However, recent studies cast doubt on this basic assumption as to whether E-cadherin transligation will yield enough forces to achieve cellular deformation (Samarage et al. 2015; Maitre et al. 2012). Rather, the discovery of filopodia—long membrane protrusions emanating from the apical cell surface and adhering via E-cadherin to the apical surface of adjacent cells—to be likely responsible for characteristically distinct compaction event (Fierro-Gonzalez et al. 2013). The filopodia are connected to the cytoskeleton and myosin motor protein, myosin-10 (Fierro-Gonzalez et al. 2013). Laser ablation of the filopodia results in rapid rounding of the cells and loss of compaction, bringing adhesion-based compaction model into question (Maitre et al. 2015). These observations corroborate a mechanochemical model for compaction, whereby the pulsatile actomyosin contractility linking the neighboring cells is redistributed by junctional E-cadherin away from cell-cell contact surface. Future investigations are aimed at understanding how the E-cadherin-dependent filopodia emerge, and the pulsatile actomyosin contractility is initiated and regulated.
5 Specification of Inner Cell Mass (ICM) and Trophectoderm: The First Cell Fate Decision
Following compaction and between 8- and 16-cell late-morula stage embryo, the cells are organized into outer polar and inner apolar cells, which give rise to TE and ICM lineages, respectively. However, the phenotypes and lineage identities at the 16-cell stage are not stable. Isolated apolar blastomeres can become polarized if placed on the outside position (Ziomek et al. 1982); outside cells of the late morula can produce ICM derivatives when aggregated with earlier embryos (Rossant and Vijh 1980), and cells of the ICM can produce trophoblast tissue (Handyside 1978). It is not until blastocyst formation that the TE and ICM lineages are irreversibly determined. Formation of the blastocyst marks the first visual cell fate decision, with TE cells on the outside committed to placental development, and cells on the inside (ICM) committed to the development of fetus and components of extraembryonic membranes. These events as discussed below are dependent on stochastic and deterministic events involving Hippo signaling system.
5.1 Differentially Expressed Transcription Factors Lead to Lineage Specification
At the late 16-cell stage, when the TE and ICM lineages are set-aside, differential expression of specific transcription factors becomes apparent in the mouse embryo. Inner apolar cells show strong expression of Sox family member Sox2 (Avilion et al. 2003), the homeobox gene Nanog (Chambers et al. 2003; Mitsui et al. 2003), and octamer-binding transcription factor gene Pou5f1 (Niwa et al. 2000). Sox2 is the first transcription factor to be specifically upregulated in the inner cells starting at the 16-cell stage (Guo et al. 2010). Nanog and Pou5f1 are found uniformly expressed in all cells, with strong expression evident at the eight-cell stage, and later restricted to the ICM of the blastocyst (Fig. 2l) (Palmieri et al. 1994; Dietrich and Hiiragi 2008).
The caudal-related homeobox gene, Cdx2, and the zinc-finger transcription factor Gata3, have an expression pattern opposite to that of Sox2, Nanog, and Pou5f1. Cdx2 and Gata3 are strongly expressed in the outside polarized cell population (Strumpf et al. 2005; Beck et al. 1995; Ralston et al. 2010). It is speculated that an imbalance between Pou5f1 and Cdx2 initiates a reciprocal inhibition system that results in the restricted pattern (Ralston et al. 2010). To support this, TE-like cells found in Cdx2 null embryos reexpress Pou5f1 (Strumpf et al. 2005), while Pou5f1 null embryos express TE markers in the ICM (Nichols et al. 1998). However, a more recent study shows that Pou5f1 levels remain high during the accumulation of Cdx2 and speculates that Pou5f1 levels therefore do not affect accumulation and maintenance of Cdx2 (Dietrich and Hiiragi 2008). It is still possible however that the reciprocal inhibition pathway is dependent on Cdx2 reaching a certain threshold before it is able to downregulate Pou5f1. Either way, single blastomere RNA sequencing has shown that activation of TE-specific genes, including Cdx2, shows up as the first overt transcriptional difference between blastomeres (Posfai et al. 2017). To summarize, the currently accepted model for lineage specification in mouse embryo combines gene expression patterning and blastomere position. In this model, molecular differences between blastomeres are established during the eight-cell stage, and then a combination of stochastic processes restrict blastomeres with specific molecular profiles to the inside or outside populations (Dietrich and Hiiragi 2008; Lanner 2014). Following this stochastic allocation, a deterministic event led by Hippo signaling establishes the lineage commitment (Fig. 3) (Nishioka et al. 2008).
Hippo signaling in the mouse embryo. Hippo signaling regulates lineage specification in the early mouse embryo. In the outside cells, Hippo signaling is active. Development of polarization complex (Par3/Par6/aPKC) in the outside cells results in tethering of junction-associate protein angiomoitin (Amot) to the apical side and subsequent lack of Lats1/2 activation. Yap protein is available for translocation into the nucleus, which in turn activates Tead4. Tead4 in turn activates trophectoderm gene expression, including Cdx2. In the inside cells, Hippo signaling is active. Lack of polarization complex allows for Amot to activate Lats kinase. Lats kinases phosphorylate Yap and subsequently sequester Yap protein in the cytoplasm. Lack of Yap translocation in the nucleus results in loss of expression of TE specification genes and upregulation of pluripotent genes including Pou5f1 and establishment of pluripotent inner cell mass cells
5.2 Hippo Signaling Pathway and Lineage Specification
The Hippo signaling pathway was first discovered in Drosophila as a tumor suppressor-signaling pathway, but is also conserved in mice and humans (Harvey et al. 2013; Yu and Guan 2013). The Hippo pathway is regulated by a variety of stimuli including cell-cell adhesion mediated by E-cadherin and by cell polarity mediated by angiomotin (Amot) which is thought to be the stimuli that initiates Hippo signaling activity in the mouse embryo (Fig. 3) (Kim et al. 2011; Hirate et al. 2013). The main components of this pathway are the protein kinase Mst1/2 and its co-activator Sav1 that function to activate kinase Lats1/2. In turn, Lats1/2 and its coactivator Mob1a/b phosphorylate Yap1 and Taz, which are transcriptional coactivators of Tead proteins (Yu and Guan 2013). Amot interacts with the E-cadherin-a α/β-catenin complex and serves as a scaffold protein that associates with Yap and Lats proteins (Fig. 3) (Paramasivam et al. 2011). Phosphorylation of Yap1 results in cytoplasmic sequestration and in turn suppresses target gene expression. Therefore, activation of Hippo signaling suppresses expression of Tead target genes, and inactivation of the pathway induces the expression of those target genes.
In the mouse embryo, Hippo signaling is an upstream regulator of Cdx2 expression and is directly involved in ICM vs. TE lineage specification (Yagi et al. 2007). In the inside cells of the mouse embryo destined to become ICM, Hippo signaling is activated as a result of increased cell-cell contacts (Nishioka et al. 2009). This activation is mediated by Lats1 and Lats2 kinase, which phosphorylates and sequesters Yap1 in the cytoplasm, preventing it from coactivating Tead4, a transcription factor that directly regulates Cdx2 expression (Nishioka et al. 2009; Hirate et al. 2013). Transient upregulation of Lats2 in the mouse embryo results in reduced nuclear accumulation of Yap1, downregulation of Cdx2, and failure to form a blastocoel because of the absence of TE cells (Fig. 3) (Nishioka et al. 2009). This phenotype is also a characteristic of Tead4 null embryos, indicating that aberrant overexpression of Hippo component Lats2 suppresses differentiation to the TE lineage via its control of Tead4 activation (Yagi et al. 2007). In the outside cells destined to become TE, Hippo signaling is inactive, which allows Yap1 to translocate into the nucleus, activate Tead4, and upregulate Cdx2 (Fig. 3) (Nishioka et al. 2008, 2009). Null mutation of either Lats1 or Lats2 does not appear to disrupt normal development during the preimplantation stages (McPherson et al. 2004; St John et al. 1999; Yabuta et al. 2007). However, overexpression of a catalytically inactive form of Lats2 (Lats kinase dead, or Lats-KD) is able to dominantly inhibit both Lats1 and Lat2, resulting in an embryo that expresses Cdx2 in inside cells (Nishioka et al. 2009). Furthermore, a double knockout of both Lats1 and Lats2 exhibits the same phenotype, strongly suggesting that Lats1/2 kinase is necessary to activate expression of the TE lineage marker, Cdx2 (Nishioka et al. 2009).
The importance of Hippo signaling in TE vs. ICM lineage specification has been confirmed in several mouse models: double knockdown and knockout of Lats1 and Lats2 genes (Lorthongpanich et al. 2013; Nishioka et al. 2009), overexpression of a catalytically inactive form of Lats2 (Lats2 kinase dead) (Nishioka et al. 2009), and depletion of Amot and Amot12 (Hirate et al. 2013) all show nuclear accumulation of Yap1, strong expression of Cdx2 in both outside and inside cells, failure to develop ICM-derived tissues, and TE-like blastomeres that populate both the inner and outer cell positions. Overexpression of Lats2, overexpression of a dominant negative form of Yap (Nishioka et al. 2009) (dnYap), and Tead4 null embryos (Nishioka et al. 2008, 2009; Yagi et al. 2007; Ralston et al. 2010) show cytoplasmic accumulation of Yap1, reduced expression of Cdx2, and failure of embryonic cells to differentiate into trophectoderm.
5.3 Hippo Signaling as It Relates to Cell Polarity and Cell Position
Apicobasal polarity in mouse embryos is dictated by the PAR-aPKC system (Vinot et al. 2005). This system involves a set of evolutionarily conserved proteins that include PDZ-domain-containing scaffold proteins (PARs) and atypical protein kinase C (aPKC) which dictate polarity in a variety of both invertebrate and vertebrate species (Fig. 3) (Suzuki and Ohno 2006). As discussed above, polarization begins at the eight-cell stage in the mouse embryo and is marked by apically restricted microvilli (experimentally visualized with phosphorylated-Ezrin) (Louvet et al. 1996) and accumulation of PARD6b and later aPKCζ at the apical surface (Vinot et al. 2005). Establishment of polarity in turn has been shown to directly suppress Hippo signaling activity in embryos (Hirate et al. 2013). Disruption of the PAR-aPKC system via injection of RNAi constructs in mouse zygotes causes Hippo pathway activation as evidenced by disruption of apical localization of Amot, exclusion of Yap1 from the nucleus, and TE development failure (Hirate et al. 2013). However, when the same embryos are dissociated, Hippo signaling is not activated despite the disruption of polarity (Hirate et al. 2013). This suggests that activation of Hippo signaling is not only dependent on the apolar status of individual blastomeres but is also dependent on cell-cell adhesion. Furthermore, in apolar cells where the PAR-aPKC system is inactive, Amot is found associated with basolateral adherens junctions via binding to the E-cadherin complex (Hirate et al. 2013). Activation of Amot at adherens junctions is thought to potentiate the activation/function of Lats kinases and results in Hippo pathway activation (Hirate et al. 2013). Together, these observations indicate that Hippo signaling responds to a combination of inputs: cell-cell adhesion and cell polarization. In the inner cells, Hippo signaling is activated by increased cell-cell contacts whereas in the outer cells, cell-cell contact-dependent Hippo signaling is suppressed by the polarization status of the outside cells mediated by the PAR-aPKC system (Sasaki 2015). To relate this finding to TE-specific expression of Cdx2, a study analyzing localization of aPKC in Cdx2 mutant embryos showed that aPKC localization to the apical surface was not affected by the lack of Cdx2. This suggests that cell polarization is independent of Cdx2, and Cdx2 upregulation is genetically downstream of cell polarization (Ralston and Rossant 2008).
From these studies, progression of events can be summarized as follows: (1) cell polarization begins at the eight-cell stage, (2) from the 16-cell stage onwards final polarity status in individual blastomeres is established, (3) Hippo signaling is suppressed in the polarized outer cells and is activated by cell-cell adhesion in the apolar inner cells (Fig. 3), and (4) this leads to lineage specific gene expression to stabilize the TE and ICM fates in the blastocyst (Fig. 3). Hippo signaling is therefore a key driver for segregation of ICM and TE lineages; however, it is still unknown at what time point Hippo signaling is sufficient to establish cell fate (Posfai et al. 2017).
6 Blastocyst Maturation and Hatching
Following the stabilization of Hippo signaling and the establishment of outer TE cells and ICM cells, the embryo transitions to a fluid-filled blastocyst stage. Blastocyst cavity is formed by active transport of sodium ions across the TE cells which create an osmotic gradient and an influx of liquid to create a fluid-filled blastocyst (Fig. 2k) (Aziz and Alexandre 1991; Benos et al. 1985; Watson and Barcroft 2001). The expanding blastocyst cavity is stabilized by tight junctions between the outer cells resulting in the generation of first embryonic epithelium the TE (Wang et al. 2008). Recent studies have highlighted the emergence of atypical apical actin rings that extend to the junctions between the outer cells and subsequent stabilization of adherens and tight junctions in a zipper-like mechanism in the TE (Fig. 2j) (Zenker et al. 2018). The expanding blastocyst cavity results in marginalization of ICM cells towards one end of the blastocyst (embryonic pole). The pluripotent ICM cells express Pou5f1, Sox2, Nanog, and Gata6 (Fig. 2l) (Ohnishi et al. 2014). Soon after, a reciprocal expression is established between Nanog expressing EPI cells producing FGF4 in an Sox2/Pou5f1-dependent fashion and Gata6 and Ffgr1/2 expressing FGF4-responsive PrE cells—in a characteristic “salt-and-pepper” fashion (Fig. 2l) (Chazaud et al. 2006). Following the establishment of two distinct populations, the PrE cells line up the blastocyst cavity through a combination of migration, apoptosis, and adhesion (Fig. 2m) (Saiz and Plusa 2013). The blastocyst at this stage will need to escape from zona pellucida to initiate implantation with uterine epithelium. The blastocyst escapes from zona in still a poorly understood process but likely includes a combination of mechanical forces from an expanding blastocyst, and proteolytic activity (Fig. 2m) (Seshagiri et al. 2009; Perona and Wassarman 1986). The escape of blastocyst (Fig. 2n) from zona pellucida will herald an implantation phase, and subsequent development is contingent on maternal inputs.
7 Concluding Remarks
More than one-third of pregnancies are lost in the preimplantation phase of embryo development, and close to 50% of human pregnancies fail during the first few weeks of pregnancy (Hyde and Schust 2015). The prevalence is especially worse for recipients in ovum donation programs, and in patients undergoing in vitro fertilization (McDonald et al. 2009; Margalioth et al. 2006; Abdalla et al. 1998). These huge rates in embryonic losses are also true in livestock, where embryonic and preimplantation loses remain major contributors of infertility (Berg et al. 2010). Majority of our current understanding on blastocyst development comes from studies in rodent models. Traditionally, this may have to do with ease of husbandry and the ability to perform sophisticated genetic modifications. However, key differences in the expression of lineage specification genes in non-rodent models remain and necessitate investigation into alternative model species. For example, there is a lack of mutually exclusive and antagonistic expression of POU5F1 and CDX2 in TE and ICM cells in embryos from livestock, a hallmark of rodent embryos (Rossant 2011). As discussed in subsequent chapters in this book, key differences also emerge post-hatching in the implantation phase between mouse, human, and other livestock species. In this regard, comparative studies involving livestock and other non-rodent model systems will likely contribute to a greater understanding of lineage specification, and in bridging gaps left by the rodent models. Livestock offer several key advantages, including a well-established culture system, unlimited supply of oocytes (slaughterhouse), and in the post-CRISPR and genome editing era, an ability to perform genetic modification. Ultimately, cross species investigations will unlock conserved mechanisms, highlight key differences, and will have a greater impact on animal and human health.
Abbreviations
- Amot:
-
Angiomotin
- aPKC:
-
Atypical protein kinase C
- dnYap:
-
Dominant-negative Yap
- EPI:
-
Epiblast
- ICM:
-
Inner cell mass
- Lats-KD:
-
Lats kinase dead
- LH:
-
Luteinizing hormone
- Na+/K+ ATPase:
-
Sodium potassium ATPase
- PAR:
-
PDZ-domain-containing scaffold protein
- PGC:
-
Primordial germ cell
- PLC:
-
Phospholipase C
- PrE:
-
Primitive endoderm
- TE:
-
Trophectoderm
References
Abdalla HI, Billett A, Kan AK, Baig S, Wren M, Korea L, Studd JW (1998) Obstetric outcome in 232 ovum donation pregnancies. Br J Obstet Gynaecol 105(3):332–337. https://doi.org/10.1111/j.1471-0528.1998.tb10096.x
Alarcon VB, Marikawa Y (2005) Unbiased contribution of the first two blastomeres to mouse blastocyst development. Mol Reprod Dev 72(3):354–361. https://doi.org/10.1002/mrd.20353
Antczak M, Van Blerkom J (1997) Oocyte influences on early development: the regulatory proteins leptin and STAT3 are polarized in mouse and human oocytes and differentially distributed within the cells of the preimplantation stage embryo. Mol Hum Reprod 3(12):1067–1086. https://doi.org/10.1093/molehr/3.12.1067
Avilion AA, Nicolis SK, Pevny LH, Perez L, Vivian N, Lovell-Badge R (2003) Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev 17(1):126–140. https://doi.org/10.1101/gad.224503
Aziz M, Alexandre H (1991) The origin of the nascent blastocoele in preimplantation mouse embryos ultrastructural cytochemistry and effect of chloroquine. Roux Arch Dev Biol 200(2):77–85. https://doi.org/10.1007/BF00637187
Barrett SL, Albertini DF (2010) Cumulus cell contact during oocyte maturation in mice regulates meiotic spindle positioning and enhances developmental competence. J Assist Reprod Genet 27(1):29–39. https://doi.org/10.1007/s10815-009-9376-9
Beck F, Erler T, Russell A, James R (1995) Expression of Cdx-2 in the mouse embryo and placenta: possible role in patterning of the extra-embryonic membranes. Dev Dyn 204(3):219–227
Benos DJ, Biggers JD, Balaban RS, Mills JW, Overstrom EW (1985) Developmental aspects of sodium-dependent transport processes of preimplantation rabbit embryos. Soc Gen Physiol Ser 39:211–235
Berg DK, van Leeuwen J, Beaumont S, Berg M, Pfeffer PL (2010) Embryo loss in cattle between days 7 and 16 of pregnancy. Theriogenology 73(2):250–260. https://doi.org/10.1016/j.theriogenology.2009.09.005
Brinster RL (1963) A method for in vitro cultivation of mouse ova from two-cell to blastocyst. Exp Cell Res 32:205–208. https://doi.org/10.1016/0014-4827(63)90093-4
Burton A, Torres-Padilla ME (2014) Chromatin dynamics in the regulation of cell fate allocation during early embryogenesis. Nat Rev Mol Cell Biol 15(11):723–734. https://doi.org/10.1038/nrm3885
Burton A, Muller J, Tu S, Padilla-Longoria P, Guccione E, Torres-Padilla ME (2013) Single-cell profiling of epigenetic modifiers identifies PRDM14 as an inducer of cell fate in the mammalian embryo. Cell Rep 5(3):687–701. https://doi.org/10.1016/j.celrep.2013.09.044
Bury L, Coelho PA, Simeone A, Ferries S, Eyers CE, Eyers PA, Zernicka-Goetz M, Glover DM (2017) Plk4 and Aurora A cooperate in the initiation of acentriolar spindle assembly in mammalian oocytes. J Cell Biol 216(11):3571–3590. https://doi.org/10.1083/jcb.201606077
Chaigne A, Campillo C, Gov NS, Voituriez R, Azoury J, Umana-Diaz C, Almonacid M, Queguiner I, Nassoy P, Sykes C, Verlhac MH, Terret ME (2013) A soft cortex is essential for asymmetric spindle positioning in mouse oocytes. Nat Cell Biol 15(8):958–966. https://doi.org/10.1038/ncb2799
Chambers I, Colby D, Robertson M, Nichols J, Lee S, Tweedie S, Smith A (2003) Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113(5):643–655
Chazaud C, Yamanaka Y, Pawson T, Rossant J (2006) Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev Cell 10(5):615–624. https://doi.org/10.1016/j.devcel.2006.02.020
Clift D, Schuh M (2013) Restarting life: fertilization and the transition from meiosis to mitosis. Nat Rev Mol Cell Biol 14(9):549–562. https://doi.org/10.1038/nrm3643
Dietrich JE, Hiiragi T (2008) Stochastic processes during mouse blastocyst patterning. Cells Tissues Organs 188(1–2):46–51. https://doi.org/10.1159/000118783
Ducibella T, Anderson E (1975) Cell shape and membrane changes in the eight-cell mouse embryo: prerequisites for morphogenesis of the blastocyst. Dev Biol 47(1):45–58. https://doi.org/10.1016/0012-1606(75)90262-6
Ducibella T, Ukena T, Karnovsky M, Anderson E (1977) Changes in cell surface and cortical cytoplasmic organization during early embryogenesis in the preimplantation mouse embryo. J Cell Biol 74(1):153–167. https://doi.org/10.1083/jcb.74.1.153
Dumont J, Million K, Sunderland K, Rassinier P, Lim H, Leader B, Verlhac MH (2007) Formin-2 is required for spindle migration and for the late steps of cytokinesis in mouse oocytes. Dev Biol 301(1):254–265. https://doi.org/10.1016/j.ydbio.2006.08.044
Fierro-Gonzalez JC, White MD, Silva JC, Plachta N (2013) Cadherin-dependent filopodia control preimplantation embryo compaction. Nat Cell Biol 15(12):1424–1433. https://doi.org/10.1038/ncb2875
Fujimori T, Kurotaki Y, Miyazaki J, Nabeshima Y (2003) Analysis of cell lineage in two- and four-cell mouse embryos. Development 130(21):5113–5122. https://doi.org/10.1242/dev.00725
Ginsburg M, Snow MH, McLaren A (1990) Primordial germ cells in the mouse embryo during gastrulation. Development 110(2):521–528
Grabole N, Tischler J, Hackett JA, Kim S, Tang F, Leitch HG, Magnusdottir E, Surani MA (2013) Prdm14 promotes germline fate and naive pluripotency by repressing FGF signalling and DNA methylation. EMBO Rep 14(7):629–637. https://doi.org/10.1038/embor.2013.67
Guo G, Huss M, Tong GQ, Wang C, Li Sun L, Clarke ND, Robson P (2010) Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev Cell 18(4):675–685. https://doi.org/10.1016/j.devcel.2010.02.012
Habibi E, Stunnenberg HG (2017) Transcriptional and epigenetic control in mouse pluripotency: lessons from in vivo and in vitro studies. Curr Opin Genet Dev 46:114–122. https://doi.org/10.1016/j.gde.2017.07.005
Handyside AH (1978) Time of commitment of inside cells isolated from preimplantation mouse embryos. J Embryol Exp Morphol 45:37–53
Harvey KF, Zhang X, Thomas DM (2013) The Hippo pathway and human cancer. Nat Rev Cancer 13(4):246–257. https://doi.org/10.1038/nrc3458
Hiiragi T, Solter D (2004) First cleavage plane of the mouse egg is not predetermined but defined by the topology of the two apposing pronuclei. Nature 430(6997):360–364. https://doi.org/10.1038/nature02595
Hirate Y, Hirahara S, Inoue K, Suzuki A, Alarcon VB, Akimoto K, Hirai T, Hara T, Adachi M, Chida K, Ohno S, Marikawa Y, Nakao K, Shimono A, Sasaki H (2013) Polarity-dependent distribution of angiomotin localizes Hippo signaling in preimplantation embryos. Curr Biol 23(13):1181–1194. https://doi.org/10.1016/j.cub.2013.05.014
Hyde KJ, Schust DJ (2015) Genetic considerations in recurrent pregnancy loss. Cold Spring Harb Perspect Med 5(3):a023119
Irie N, Tang WW, Azim Surani M (2014) Germ cell specification and pluripotency in mammals: a perspective from early embryogenesis. Reprod Med Biol 13(4):203–215. https://doi.org/10.1007/s12522-014-0184-2
Jachowicz JW, Bing X, Pontabry J, Boskovic A, Rando OJ, Torres-Padilla ME (2017) LINE-1 activation after fertilization regulates global chromatin accessibility in the early mouse embryo. Nat Genet 49(10):1502–1510. https://doi.org/10.1038/ng.3945
Johnson MH (2009) From mouse egg to mouse embryo: polarities, axes, and tissues. Annu Rev Cell Dev Biol 25:483–512. https://doi.org/10.1146/annurev.cellbio.042308.113348
Kim NG, Koh E, Chen X, Gumbiner BM (2011) E-cadherin mediates contact inhibition of proliferation through Hippo signaling-pathway components. Proc Natl Acad Sci U S A 108(29):11930–11935. https://doi.org/10.1073/pnas.1103345108
Lanner F (2014) Lineage specification in the early mouse embryo. Exp Cell Res 321(1):32–39. https://doi.org/10.1016/j.yexcr.2013.12.004
Leung CY, Zernicka-Goetz M (2015) Mapping the journey from totipotency to lineage specification in the mouse embryo. Curr Opin Genet Dev 34:71–76. https://doi.org/10.1016/j.gde.2015.08.002
Li R, Albertini DF (2013) The road to maturation: somatic cell interaction and self-organization of the mammalian oocyte. Nat Rev Mol Cell Biol 14(3):141–152. https://doi.org/10.1038/nrm3531
Littwin T, Denker HW (2011) Segregation during cleavage in the mammalian embryo? A critical comparison of whole-mount/CLSM and section immunohistochemistry casts doubts on segregation of axis-relevant leptin domains in the rabbit. Histochem Cell Biol 135(6):553–570. https://doi.org/10.1007/s00418-011-0816-0
Lorthongpanich C, Messerschmidt DM, Chan SW, Hong W, Knowles BB, Solter D (2013) Temporal reduction of LATS kinases in the early preimplantation embryo prevents ICM lineage differentiation. Genes Dev 27(13):1441–1446. https://doi.org/10.1101/gad.219618.113
Louvet S, Aghion J, Santa-Maria A, Mangeat P, Maro B (1996) Ezrin becomes restricted to outer cells following asymmetrical division in the preimplantation mouse embryo. Dev Biol 177(2):568–579
Ma Z, Swigut T, Valouev A, Rada-Iglesias A, Wysocka J (2011) Sequence-specific regulator Prdm14 safeguards mouse ESCs from entering extraembryonic endoderm fates. Nat Struct Mol Biol 18(2):120–127. https://doi.org/10.1038/nsmb.2000
Maitre JL, Berthoumieux H, Krens SF, Salbreux G, Julicher F, Paluch E, Heisenberg CP (2012) Adhesion functions in cell sorting by mechanically coupling the cortices of adhering cells. Science 338(6104):253–256. https://doi.org/10.1126/science.1225399
Maitre JL, Niwayama R, Turlier H, Nedelec F, Hiiragi T (2015) Pulsatile cell-autonomous contractility drives compaction in the mouse embryo. Nat Cell Biol 17(7):849–855. https://doi.org/10.1038/ncb3185
Margalioth EJ, Ben-Chetrit A, Gal M, Eldar-Geva T (2006) Investigation and treatment of repeated implantation failure following IVF-ET. Hum Reprod 21(12):3036–3043. https://doi.org/10.1093/humrep/del305
McDonald SD, Han Z, Mulla S, Murphy KE, Beyene J, Ohlsson A (2009) Preterm birth and low birth weight among in vitro fertilization singletons: a systematic review and meta-analyses. Eur J Obstet Gynecol Reprod Biol 146(2):138–148. https://doi.org/10.1016/j.ejogrb.2009.05.035
McPherson JP, Tamblyn L, Elia A, Migon E, Shehabeldin A, Matysiak-Zablocki E, Lemmers B, Salmena L, Hakem A, Fish J, Kassam F, Squire J, Bruneau BG, Hande MP, Hakem R (2004) Lats2/Kpm is required for embryonic development, proliferation control and genomic integrity. EMBO J 23(18):3677–3688. https://doi.org/10.1038/sj.emboj.7600371
Mitsui K, Tokuzawa Y, Itoh H, Segawa K, Murakami M, Takahashi K, Maruyama M, Maeda M, Yamanaka S (2003) The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113(5):631–642
Mogessie B, Schuh M (2017) Actin protects mammalian eggs against chromosome segregation errors. Science 357(6353). https://doi.org/10.1126/science.aal1647
Motosugi N, Bauer T, Polanski Z, Solter D, Hiiragi T (2005) Polarity of the mouse embryo is established at blastocyst and is not prepatterned. Genes Dev 19(9):1081–1092. https://doi.org/10.1101/gad.1304805
Nagy A, Gertsenstein M, Vintersten K, Behringer R (2003) Manipulating the mouse embryo: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Nichols J, Zevnik B, Anastassiadis K, Niwa H, Klewe-Nebenius D, Chambers I, Scholer H, Smith A (1998) Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95(3):379–391
Nishioka N, Yamamoto S, Kiyonari H, Sato H, Sawada A, Ota M, Nakao K, Sasaki H (2008) Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech Dev 125(3–4):270–283. https://doi.org/10.1016/j.mod.2007.11.002
Nishioka N, Inoue K, Adachi K, Kiyonari H, Ota M, Ralston A, Yabuta N, Hirahara S, Stephenson RO, Ogonuki N, Makita R, Kurihara H, Morin-Kensicki EM, Nojima H, Rossant J, Nakao K, Niwa H, Sasaki H (2009) The Hippo signaling pathway components Lats and Yap pattern Tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev Cell 16(3):398–410. https://doi.org/10.1016/j.devcel.2009.02.003
Niwa H, Miyazaki J, Smith AG (2000) Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat Genet 24(4):372–376
Ohinata Y, Payer B, O’Carroll D, Ancelin K, Ono Y, Sano M, Barton SC, Obukhanych T, Nussenzweig M, Tarakhovsky A, Saitou M, Surani MA (2005) Blimp1 is a critical determinant of the germ cell lineage in mice. Nature 436(7048):207–213. https://doi.org/10.1038/nature03813
Ohinata Y, Ohta H, Shigeta M, Yamanaka K, Wakayama T, Saitou M (2009) A signaling principle for the specification of the germ cell lineage in mice. Cell 137(3):571–584. https://doi.org/10.1016/j.cell.2009.03.014
Ohnishi Y, Huber W, Tsumura A, Kang M, Xenopoulos P, Kurimoto K, Oles AK, Arauzo-Bravo MJ, Saitou M, Hadjantonakis AK, Hiiragi T (2014) Cell-to-cell expression variability followed by signal reinforcement progressively segregates early mouse lineages. Nat Cell Biol 16(1):27–37. https://doi.org/10.1038/ncb2881
Palmieri SL, Peter W, Hess H, Scholer HR (1994) Oct-4 transcription factor is differentially expressed in the mouse embryo during establishment of the first two extraembryonic cell lineages involved in implantation. Dev Biol 166(1):259–267
Paramasivam M, Sarkeshik A, Yates JR 3rd, Fernandes MJ, McCollum D (2011) Angiomotin family proteins are novel activators of the LATS2 kinase tumor suppressor. Mol Biol Cell 22(19):3725–3733. https://doi.org/10.1091/mbc.E11-04-0300
Perona RM, Wassarman PM (1986) Mouse blastocysts hatch in vitro by using a trypsin-like proteinase associated with cells of mural trophectoderm. Dev Biol 114(1):42–52. https://doi.org/10.1016/0012-1606(86)90382-9
Piotrowska K, Zernicka-Goetz M (2001) Role for sperm in spatial patterning of the early mouse embryo. Nature 409(6819):517–521. https://doi.org/10.1038/35054069
Piotrowska K, Wianny F, Pedersen RA, Zernicka-Goetz M (2001) Blastomeres arising from the first cleavage division have distinguishable fates in normal mouse development. Development 128(19):3739–3748
Piotrowska-Nitsche K, Perea-Gomez A, Haraguchi S, Zernicka-Goetz M (2005) Four-cell stage mouse blastomeres have different developmental properties. Development 132(3):479–490. https://doi.org/10.1242/dev.01602
Posfai E, Petropoulos S, de Barros FRO, Schell JP, Jurisica I, Sandberg R, Lanner F, Rossant J (2017) Position- and Hippo signaling-dependent plasticity during lineage segregation in the early mouse embryo. Elife 6. https://doi.org/10.7554/eLife.22906
Ralston A, Rossant J (2008) Cdx2 acts downstream of cell polarization to cell-autonomously promote trophectoderm fate in the early mouse embryo. Dev Biol 313(2):614–629. https://doi.org/10.1016/j.ydbio.2007.10.054
Ralston A, Cox BJ, Nishioka N, Sasaki H, Chea E, Rugg-Gunn P, Guo G, Robson P, Draper JS, Rossant J (2010) Gata3 regulates trophoblast development downstream of Tead4 and in parallel to Cdx2. Development 137(3):395–403. https://doi.org/10.1242/dev.038828
Rossant J (2011) Developmental biology: a mouse is not a cow. Nature 471(7339):457–458. https://doi.org/10.1038/471457a
Rossant J, Vijh KM (1980) Ability of outside cells from preimplantation mouse embryos to form inner cell mass derivatives. Dev Biol 76(2):475–482
Saitou M, Yamaji M (2012) Primordial germ cells in mice. Cold Spring Harb Perspect Biol 4(11). https://doi.org/10.1101/cshperspect.a008375
Saiz N, Plusa B (2013) Early cell fate decisions in the mouse embryo. Reproduction 145(3):R65–R80. https://doi.org/10.1530/REP-12-0381
Samarage CR, White MD, Alvarez YD, Fierro-Gonzalez JC, Henon Y, Jesudason EC, Bissiere S, Fouras A, Plachta N (2015) Cortical tension allocates the first inner cells of the mammalian embryo. Dev Cell 34(4):435–447. https://doi.org/10.1016/j.devcel.2015.07.004
Sasaki H (2015) Position- and polarity-dependent Hippo signaling regulates cell fates in preimplantation mouse embryos. Semin Cell Dev Biol 47-48:80–87. https://doi.org/10.1016/j.semcdb.2015.05.003
Schafer S, Anschlag J, Nettersheim D, Haas N, Pawig L, Schorle H (2011) The role of BLIMP1 and its putative downstream target TFAP2C in germ cell development and germ cell tumours. Int J Androl 34(4 Pt 2):e152–158; discussion e158–159. https://doi.org/10.1111/j.1365-2605.2011.01167.x
Seshagiri PB, Sen Roy S, Sireesha G, Rao RP (2009) Cellular and molecular regulation of mammalian blastocyst hatching. J Reprod Immunol 83(1–2):79–84. https://doi.org/10.1016/j.jri.2009.06.264
St John MA, Tao W, Fei X, Fukumoto R, Carcangiu ML, Brownstein DG, Parlow AF, McGrath J, Xu T (1999) Mice deficient of Lats1 develop soft-tissue sarcomas, ovarian tumours and pituitary dysfunction. Nat Genet 21(2):182–186. https://doi.org/10.1038/5965
Strumpf D, Mao CA, Yamanaka Y, Ralston A, Chawengsaksophak K, Beck F, Rossant J (2005) Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 132(9):2093–2102
Suzuki A, Ohno S (2006) The PAR-aPKC system: lessons in polarity. J Cell Sci 119(Pt 6):979–987. https://doi.org/10.1242/jcs.02898
Suzuki A, Saga Y (2008) Nanos2 suppresses meiosis and promotes male germ cell differentiation. Genes Dev 22(4):430–435. https://doi.org/10.1101/gad.1612708
Tabansky I, Lenarcic A, Draft RW, Loulier K, Keskin DB, Rosains J, Rivera-Feliciano J, Lichtman JW, Livet J, Stern JN, Sanes JR, Eggan K (2013) Developmental bias in cleavage-stage mouse blastomeres. Curr Biol 23(1):21–31. https://doi.org/10.1016/j.cub.2012.10.054
Thomas FC, Sheth B, Eckert JJ, Bazzoni G, Dejana E, Fleming TP (2004) Contribution of JAM-1 to epithelial differentiation and tight-junction biogenesis in the mouse preimplantation embryo. J Cell Sci 117(Pt 23):5599–5608. https://doi.org/10.1242/jcs.01424
Torres-Padilla ME, Parfitt DE, Kouzarides T, Zernicka-Goetz M (2007) Histone arginine methylation regulates pluripotency in the early mouse embryo. Nature 445(7124):214–218. https://doi.org/10.1038/nature05458
Vestweber D, Gossler A, Boller K, Kemler R (1987) Expression and distribution of cell adhesion molecule uvomorulin in mouse preimplantation embryos. Dev Biol 124(2):451–456. https://doi.org/10.1016/0012-1606(87)90498-2
Vinot S, Le T, Ohno S, Pawson T, Maro B, Louvet-Vallee S (2005) Asymmetric distribution of PAR proteins in the mouse embryo begins at the 8-cell stage during compaction. Dev Biol 282(2):307–319. https://doi.org/10.1016/j.ydbio.2005.03.001
Wang H, Ding T, Brown N, Yamamoto Y, Prince LS, Reese J, Paria BC (2008) Zonula occludens-1 (ZO-1) is involved in morula to blastocyst transformation in the mouse. Dev Biol 318(1):112–125. https://doi.org/10.1016/j.ydbio.2008.03.008
Watson AJ, Barcroft LC (2001) Regulation of blastocyst formation. Front Biosci 6:D708–D730. https://doi.org/10.2741/watson
Watson AJ, Kidder GM (1988) Immunofluorescence assessment of the timing of appearance and cellular distribution of Na/K-ATPase during mouse embryogenesis. Dev Biol 126(1):80–90. https://doi.org/10.1016/0012-1606(88)90241-2
Weber S, Eckert D, Nettersheim D, Gillis AJ, Schafer S, Kuckenberg P, Ehlermann J, Werling U, Biermann K, Looijenga LH, Schorle H (2010) Critical function of AP-2 gamma/TCFAP2C in mouse embryonic germ cell maintenance. Biol Reprod 82(1):214–223. https://doi.org/10.1095/biolreprod.109.078717
White MD, Plachta N (2020) Specification of the first mammalian cell lineages in vivo and in vitro. Cold Spring Harb Perspect Biol 12(4):a035634. https://doi.org/10.1101/cshperspect.a035634
White MD, Zenker J, Bissiere S, Plachta N (2018) Instructions for assembling the early mammalian embryo. Dev Cell 45(6):667–679. https://doi.org/10.1016/j.devcel.2018.05.013
Whitten WK, Biggers JD (1968) Complete development in vitro of the pre-implantation stages of the mouse in a simple chemically defined medium. J Reprod Fertil 17(2):399–401. https://doi.org/10.1530/jrf.0.0170399
Wicklow E, Blij S, Frum T, Hirate Y, Lang RA, Sasaki H, Ralston A (2014) HIPPO pathway members restrict SOX2 to the inner cell mass where it promotes ICM fates in the mouse blastocyst. PLoS Genet 10(10):e1004618. https://doi.org/10.1371/journal.pgen.1004618
Yabuta N, Okada N, Ito A, Hosomi T, Nishihara S, Sasayama Y, Fujimori A, Okuzaki D, Zhao H, Ikawa M, Okabe M, Nojima H (2007) Lats2 is an essential mitotic regulator required for the coordination of cell division. J Biol Chem 282(26):19259–19271. https://doi.org/10.1074/jbc.M608562200
Yagi R, Kohn MJ, Karavanova I, Kaneko KJ, Vullhorst D, DePamphilis ML, Buonanno A (2007) Transcription factor TEAD4 specifies the trophectoderm lineage at the beginning of mammalian development. Development 134(21):3827–3836. https://doi.org/10.1242/dev.010223
Yamaji M, Seki Y, Kurimoto K, Yabuta Y, Yuasa M, Shigeta M, Yamanaka K, Ohinata Y, Saitou M (2008) Critical function of Prdm14 for the establishment of the germ cell lineage in mice. Nat Genet 40(8):1016–1022. https://doi.org/10.1038/ng.186
Yamaji M, Ueda J, Hayashi K, Ohta H, Yabuta Y, Kurimoto K, Nakato R, Yamada Y, Shirahige K, Saitou M (2013) PRDM14 ensures naive pluripotency through dual regulation of signaling and epigenetic pathways in mouse embryonic stem cells. Cell Stem Cell 12(3):368–382. https://doi.org/10.1016/j.stem.2012.12.012
Yi K, Unruh JR, Deng M, Slaughter BD, Rubinstein B, Li R (2011) Dynamic maintenance of asymmetric meiotic spindle position through Arp2/3-complex-driven cytoplasmic streaming in mouse oocytes. Nat Cell Biol 13(10):1252–1258. https://doi.org/10.1038/ncb2320
Yi K, Rubinstein B, Unruh JR, Guo F, Slaughter BD, Li R (2013) Sequential actin-based pushing forces drive meiosis I chromosome migration and symmetry breaking in oocytes. J Cell Biol 200(5):567–576. https://doi.org/10.1083/jcb.201211068
Yu FX, Guan KL (2013) The Hippo pathway: regulators and regulations. Genes Dev 27(4):355–371. https://doi.org/10.1101/gad.210773.112
Zenker J, White MD, Gasnier M, Alvarez YD, Lim HYG, Bissiere S, Biro M, Plachta N (2018) Expanding actin rings zipper the mouse embryo for blastocyst formation. Cell 173(3):776–791.e717. https://doi.org/10.1016/j.cell.2018.02.035
Zhang HT, Hiiragi T (2018) Symmetry breaking in the mammalian embryo. Annu Rev Cell Dev Biol 34:405–426. https://doi.org/10.1146/annurev-cellbio-100617-062616
Zhu M, Leung CY, Shahbazi MN, Zernicka-Goetz M (2017) Actomyosin polarisation through PLC-PKC triggers symmetry breaking of the mouse embryo. Nat Commun 8(1):921. https://doi.org/10.1038/s41467-017-00977-8
Ziomek CA, Johnson MH (1980) Cell surface interaction induces polarization of mouse 8-cell blastomeres at compaction. Cell 21(3):935–942. https://doi.org/10.1016/0092-8674(80)90457-2
Ziomek CA, Johnson MH, Handyside AH (1982) The developmental potential of mouse 16-cell blastomeres. J Exp Zool 221(3):345–355
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Telugu, B.P., Pence, L. (2021). Development of Pre-implantation Mammalian Blastocyst. In: Geisert, R.D., Spencer, T. (eds) Placentation in Mammals. Advances in Anatomy, Embryology and Cell Biology, vol 234. Springer, Cham. https://doi.org/10.1007/978-3-030-77360-1_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-77360-1_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-77359-5
Online ISBN: 978-3-030-77360-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)