Abstract
Spinocerebellar ataxia type 1 (SCA1) is one of the intractable neurodegenerative diseases caused by the mutation of Ataxin-1 (Atxn1) gene. By various comprehensive approaches, we have newly discovered key molecules such as PQBP1, VCP, HMGB1, RpA1, and YAP/YAPdeltaC that mediate the SCA1 pathology from the gene mutation to phenotypes. The functions of these molecules are involved in transcription, RNA splicing, and DNA damage repair, and their functional impairments contribute to neurodegeneration via multiple pathways. Based on the knowledge, we have also developed gene therapies using adeno-associated virus and other disease-modifying therapies. In this review, we focus on summarizing our original works for understanding and conquering SCA1.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
- Comprehensive analysis
- New pathology
- Target genes
- Impairment of DNA damage repair
- RNA splicing
- Gene therapy
- Disease-modifying therapy
1 Discovery of Novel SCA1 Pathologies by Comprehensive Analyses
Spinocerebellar ataxia type 1 (SCA1) is a neurodegenerative disease which mainly affects Purkinje cells in the cerebellum and motor neurons in the spinal cord. Since the discovery of causative gene, Ataxin-1 (Atxn1), more than 20 years ago, substantial amount of knowledge about the mechanism has been accumulated. Identification of Atxn1-interacting factors such as capicua (CIC) (Lam et al., 2006) and RNA-binding motif protein 17 (RBM17) as binding proteins to Atxn1 (Lim et al., 2008) indicates the involvement in transcription and splicing in the SCA1 pathology (Zoghbi & Orr, 2009).
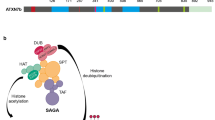
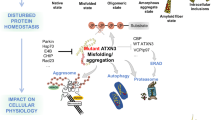
Meanwhile, by employing comprehensive analyses, our group has identified key molecules that mediate functional dysregulation caused by mutant Atxn1 protein. First, by yeast two-hybrid screening, we found six clones that interact with polyglutamine (polyQ) tract sequences. One of them was the already known molecule VCP (TERA/p97/VCP), and the others were novel genes such as PQBP1 (polyglutamine-binding protein 1) (Imafuku et al., 1998; Waragai et al., 1999). Functions of PQBP1 have been identified during the last 20 years by our group and other groups (Okazawa, 2018). PQBP1 is involved in transcriptional regulation, RNA splicing, RNA stress, and DNA damage repair (Waragai et al., 1999; Okazawa et al., 2002; Kunde et al., 2011; Mizuguchi et al., 2014; Ito et al., 2015b; Wan et al., 2015; Morchikh et al., 2017) and determines gene expression profiles related to neural stem cell proliferation, neuronal cilia, neurite extension, and synapse function (Ikeuchi et al., 2013; Ito et al., 2015b; Li et al., 2013; Okazawa et al., 2001; Okazawa et al., 2002; Wang et al., 2013; Waragai et al., 1999). Interaction of mutant Atxn1 with PQBP1 basically impairs such multiple functions in neurons including Purkinje cells and spinal motor neurons (Fig. 24.1).
The discovery of HMGB1 (Qi et al., 2007) and VCP (Imafuku et al., BBRC 1998; Waragai et al., Hum Mol Genet 1999) led to identifying that impairment of DNA damage repair is another key pathological event in SCA1 (Ito et al., 2015a; Qi et al., 2007). Comprehensive proteome analysis revealed that HMGB1/2 were decreased in Purkinje cells of SCA1 model mice and that mutant Atxn1 interact with HMGB1/2 to impair the DNA damage repair function (Qi et al., 2007). HMGB1/2 are known to regulate the unwinding or folding DNA structures, and their functional inhibition in nuclei of neurons leads to inhibition of nuclear functions (Alessandra & Bianchi, 2003; Travers, 2003). As mentioned above, VCP was discovered as a binding protein to polyQ tract sequence (Imafuku et al., 1998). Several studies revealed involvement of VCP in DNA damage repair (Acs et al., 2011; Meerang et al., 2011), and interaction of mutant Atxn1 with VCP also leads to impairment of DNA damage repair (Fujita et al., 2013).
2 Gene Therapy of SCA1 with HMGB1
We performed a proteome analysis of soluble nuclear proteins from neurons expressing mutant polyQ protein and revealed that HMGB1/2 are decreased in cerebellar neurons expressing mutant Atxn1, which is a mutation of the causative gene of SCA1 and in cerebral neurons expressing huntingtin (Htt), which is a mutation of the causative gene of Huntington’s disease (Qi et al., 2007). This result showed that HMGB1/2 proteins were commonly reduced in vulnerable neurons. In addition, similar reductions were observed in vulnerable neurons even before the onset of transgenic mice (R6/2 mice) and knock-in mice (Atxn1-154Q/2Q-KI mice). HMGB1/2 are one of the most abundant proteins in the nucleus, and these are known as essential DNA structural proteins that unwind DNA from histone complex or bent DNA (Alessandra & Bianchi, 2003; Travers, 2003).
Mutant Ataxin-1 or huntingtin binds to HMGB1 and impairs the functions directly or promotes the degradation (Qi et al., 2007). Due to the deficiency of HMGB1, nuclear and possibly cytoplasmic functions including DNA damage repair are impaired (Fig. 24.1) (Qi et al., 2007). Supplementation of HMGB1 suppressed neuronal cell death in Drosophila SCA1 model, in which mutant Atxn1 causes degeneration of photoreceptor cells of the complex eye (Qi et al., 2007).
Based on the results from Drosophila model, we moved to Atxn1-KI mouse model. We first mated HMGB1-Tg mice with Atxn1-KI mice of the same background C57BL/6 and generated double transgenic mice (Atxn1-KI;HMGB1 mice), and we tested their motor dysfunction and lifespan. In the rotarod test, SCA1 model mice showed motor dysfunction from 5 weeks of age, which continued to decline. On the other hand, in the double transgenic mouse, the shortened rotarod stay time was improved from 7 weeks of age, and the improvement was sustained at least until 21 weeks of age (Ito et al., 2015a). Nuclear DNA damage in Purkinje cells of the mutant Atxn1-KI mice was recovered in the double transgenic mouse. The 50% survival duration was extended from 217 days to 282 days (+30%), and the maximum survival duration was increased from 274 days to 360 days (Ito et al., 2015a).
Moreover, gene therapy using an adeno-associated virus (AAV) vector was effective. HMGB1 was widely and highly expressed in the cerebellar neurons especially in Purkinje cells by a single injection onto the cerebellum surface of 5-week-old SCA1 model mice. Similar to the results in the double transgenic mice, improvement in motor function was observed at 9 and 13 weeks of age in the gene therapy experiment using the AAV vector. Furthermore, the lifespan of mutant Atxn1-KI mice was extended from 217 days to 365.5 days (nearly +70%), and the maximum was extended from 274 days to 448 days (Fig. 24.2) (Ito et al., 2015a). In addition, it was revealed that HMGB1 also enhances mitochondrial DNA damage repair, suggesting that HMGB1 suppresses neurodegeneration by repairing both nuclear and mitochondrial DNA (Ito et al., 2015a).
Gene therapy of HMGB1 in SCA1 model mice. HMGB1 is decreased in Purkinje cells and other neurons of mutant Atxn1-KI mice and also human patients. AAV-HMGB1 gene therapy recovers the deficiency of HMGB1 and elongates the lifespan of the model mice. Similar approaches can be used to recover deficiency of other target molecules
3 Gene Therapy of SCA1 with RpA1
In addition to the proteome analysis revealing the role of HMGB1/2 in SCA1 pathology, we found from multiple omics analyses that other molecules such as TERA/VCP/p97 and Ku70 involved in DNA damage repair are functionally impaired in polyglutamine disease (Enokido et al., 2010; Fujita et al., 2013; Qi et al., 2007). These proteins are basically involved in non-homologous end joining of DNA double-strand break repair among various types of DNA damage repair. Therefore, we further asked which type of DNA damage repair most significantly contributes to the SCA1 pathology (Barclay et al., 2014). From gene screens with Drosophila SCA1 models, we identified that RpA1, a protective protein for naked single-strand DNA in various types of DNA damage, has the largest therapeutic effect on the lifespan-shortening by mutant Atxn1 expression in motor neurons (Barclay et al., 2014). Furthermore, an immunoprecipitation (IP) assay revealed that RpA1 binds to Ataxin-1 and mutant Ataxin-1 binds to RpA1 more strongly (Barclay et al., 2014).
Therefore, we developed gene therapy of mutant Atxn1-KI mice with AAV-RpA1 at the timing of onset, which induced significant improvement of motor function lasting over 50 weeks after injection (Taniguchi et al., 2016). DNA double-strand break, which is the final outcome of various forms of DNA damage, was also ameliorated in cerebellar neurons, and the abnormal patterns of gene expression were also partially corrected (Taniguchi et al., 2016).
4 Gene Therapy with PQBP1
Though we have not examined the effect of AAV-PQBP1 on mutant Atxn1-KI model mice, we have unexpectedly experienced the examination in Alzheimer’s disease (AD) mouse models (Tanaka et al., 2018). We reached to the conclusion that PQBP1 is also involved in the AD pathology as follows. First, we performed comprehensive phosphoproteome analysis and revealed that SRRM2 phosphorylation occurs at the earliest stage before extracellular Abeta aggregation (Tagawa et al., 2015). The phosphorylation of SRRM2 inhibits its interaction with a chaperone protein TCP1alpha and prevents nuclear translocation of SRRM2 (Tanaka et al., 2018). Given that SRRM2 functions as a scaffold protein in the nucleus, the major target of SRRM2 scaffolding PQBP1 was decreased in the nucleus. The reduction of PQBP1 in the nucleus disturbs proper splicing of synapse-related hnRNA and decreases their mRNA (Ito et al., 2015b), which is just like neurons in patients of Renpenning syndrome (Kalscheuer et al., 2003; Lenski et al., 2004; Lubs et al., 2006; Okazawa, 2018; Stevenson et al., 2005). Therefore, we performed gene therapy against AD model mice with AAV-PQBP1 and found the rescue of phenotypes as expected (Tanaka et al., 2018). The similar approach will be feasible in therapeutics for the SCA1 pathology.
5 Developmental Pathology of SCA1
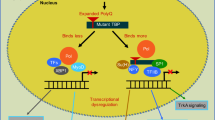
Previous studies have accumulated a great deal of knowledge about pathological conditions caused by gene mutations. However, the relationship between timing and the major pathology is still not unclear, which slows down the progress of disease-modifying therapy. In a previous study, we discovered transcriptional repression-induced atypical cell death (TRIAD) due to RNA polymerase II inhibition and identified a molecule YAPdeltaC that regulates TRIAD (Hoshino et al., 2006). To elucidate the function of YAPdeltaC in SCA1, we used the Tet-ON system for YAPdeltaC expression and examined how the time-specific expression of YAPdeltaC affects the symptoms and survival of Ataxin-1-KI mice. Unexpectedly, expression of YAPdeltaC during development remarkably extended the lifespan, while the expression of YAPdeltaC from adulthood (from 8 weeks old) was less effective (Fujita et al., 2017).
We revealed that YAPdeltaC is a transcriptional co-factor that enhances RORalpha function that regulates gene expression in cerebellar neurons during development collaborating with normal Ataxin-1. On the other hand, mutant Ataxin-1 inhibits the interaction between YAPdeltaC and RORalpha and inhibits gene expression required for maturation of cerebellar neurons (Fujita et al., 2017). These results indicate the role of mutant Atxn1 during the development and the developmental pathology influencing the adult pathology beyond a long time interval.
6 Future Prospects
Various comprehensive analyses of our group unraveled novel pathologies of SCA1 such as impairments of DNA damage repair (HMGB1, Ku70, VCP, RpA1, PQBP1) and RNA splicing (PQBP1) and transcription (HMGB1, PQBP1, YAP/YAPdeltaC). A single gene mutation of Atxn1 leads to phenotypes via multiple pathways, which might be “gain of function” or “loss of function.” From the viewpoint of mutant Atxn1, it could be described as “gain of function” when mutant Atxn1 deprives some target proteins to inclusion body. However, it could be also described as “loss of function” from the aspect of normal Atxn1. For instance, normal Atxn1 have physiological interaction with YAP/YAPdeltaC or PQBP1 in transcription and splicing regulation. Therefore, prevention of these target proteins from the functional complex by mutant Atxn1 is similar to loss of function of normal Atxn1.
Recently gene-based therapies like gene therapy, antisense oligonucleotide, and genome editing are developing rapidly. Basically, gene-based therapies are classified into two types, upregulation and downregulation of the target gene. Antisense oligonucleotide, therefore, is used for knockdown of mutant Atxn1. However, it has an unexpected pitfall that depletion of Atxn1 might increase the risk of Alzheimer’s disease (Crespo-Barreto et al., 2010; Matilla et al., 1998; Suh et al., 2019), and the concern of off-target effect cannot be completely excluded. Actually, currently ongoing clinical trials of antisense therapy targeting selectively mutant Htt mRNA might accelerate brain atrophy of Huntington’s disease patients judging from their ventricular size at clinical trial phase 2 (Supplementary Results of ref. (Suh et al., 2019) (Tabrizi et al., 2019). Therefore, upregulating the target whose activity is lost in the pathology of SCA1 might be safer for human patients than downregulating the causative gene.
References
Acs, K., Luijsterburg, M. S., Ackermann, L., Salomons, F. A., Hoppe, T., & Dantuma, N. P. (2011). The AAA-ATPase VCP/p97 promotes 53BP1 recruitment by removing L3MBTL1 from DNA double-strand breaks. Nature Structural & Molecular Biology, 18, 1345–1350.
Alessandra, A., & Bianchi, M. E. (2003). HMGB proteins and gene expression. Current Opinion in Genetics & Development, 2, 170–178.
Barclay, S. S., Tamura, T., Ito, H., et al. (2014). Systems biology analysis of Drosophila in vivo screen data elucidates core networks for DNA damage repair in SCA1. Human Molecular Genetics, 23, 1345–1364.
Crespo-Barreto, J., Fryer, J. D., Shaw, C. A., Orr, H. T., & Zoghbi, H. Y. (2010). Partial loss of ataxin-1 function contributes to transcriptional dysregulation in spinocerebellar ataxia type 1 pathogenesis. PLoS Genetics, 6, 1–17.
Enokido, Y., Tamura, T., Ito, H., et al. (2010). Mutant huntingtin impairs Ku70-mediated DNA repair. The Journal of Cell Biology, 189, 425–443.
Fujita, K., Mao, Y., Uchida, S., et al. (2017). Developmental YAPdeltaC determines adult pathology in a model of spinocerebellar ataxia type 1. Nature Communications, 8, 1864.
Fujita, K., Nakamura, Y., Oka, T., et al. (2013). A functional deficiency of TERA/VCP/p97 contributes to impaired DNA repair in multiple polyglutamine diseases. Nature Communications, 4, 1816.
Hoshino, M., Qi, M. L., Yoshimura, N., et al. (2006). Transcriptional repression induces a slowly progressive atypical neuronal death associated with changes of YAP isoforms and p73. The Journal of Cell Biology, 172, 589–604.
Ikeuchi, Y., delaTorre-Ubieta, L., Matsuda, T., Steen, H., Okazawa, H., & Bonni, A. (2013). The XLID protein PQBP1 and the GTPase dynamin 2 define a signaling link that orchestrates ciliary morphogenesis in postmitotic neurons. Cell Reports, 4, 879–889.
Imafuku, I., Waragai, M., Takeuchi, S., Kanazawa, I., Kawabata, M., Mouradian, M. M., & Okazawa, H. (1998). Polar amino acid-rich sequences bind to polyglutamine tracts. Biochemical and Biophysical Research Communications, 253, 16–20.
Ito, H., Fujita, K., Tagawa, K., et al. (2015a). HMGB1 facilitates repair of mitochondrial DNA damage and extends the lifespan of mutant ataxin-1 knock-in mice. EMBO Molecular Medicine, 7, 78–101.
Ito, H., Shiwaku, H., Yoshida, C., et al. (2015b). In utero gene therapy rescues microcephaly caused by Pqbp1-hypofunction in neural stem progenitor cells. Molecular Psychiatry, 20, 459–471.
Kalscheuer, V. M., Freude, K., Musante, L., et al. (2003). Mutations in the polyglutamine binding protein 1 gene cause X-linked mental retardation. Nature Genetics, 35, 313–315.
Kunde, S. A., Musante, L., Grimme, A., Fischer, U., Müller, E., Wanker, E. E., & Kalscheuer, V. M. (2011). The X-chromosome-linked intellectual disability protein PQBP1 is a component of neuronal RNA granules and regulates the appearance of stress granules. Human Molecular Genetics, 20, 4916–4931.
Lam, Y. C., Bowman, A. B., Jafar-Nejad, P., et al. (2006). ATAXIN-1 interacts with the repressor Capicua in its native complex to cause SCA1 neuropathology. Cell, 127, 1335–1347.
Lenski, C., Abidi, F., Meindl, A., Gibson, A., Platzer, M., Frank Kooy, R., Lubs, H. A., Stevenson, R. E., Ramser, J., & Schwartz, C. E. (2004). Novel truncating mutations in the polyglutamine tract binding protein 1 gene (PQBP1) cause Renpenning syndrome and X-linked mental retardation in another family with microcephaly. American Journal of Human Genetics, 74, 777–780.
Li, C., Ito, H., Fujita, K., Shiwaku, H., Qi, Y., Tagawa, K., Tamura, T., & Okazawa, H. (2013). Sox2 transcriptionally regulates Pqbp1, an intellectual disability-microcephaly causative gene, in neural stem progenitor cells. PLoS One, 8, 1–15.
Lim, J., Crespo-Barreto, J., Jafar-Nejad, P., Bowman, A. B., Richman, R., Hill, D. E., Orr, H. T., & Zoghbi, H. Y. (2008). Opposing effects of polyglutamine expansion on native protein complexes contribute to SCA1. Nature, 452, 713–718.
Lubs, H., Abidi, F. E., Echeverri, R., Holloway, L., Meindl, A., Stevenson, R. E., & Schwartz, C. E. (2006). Golabi-Ito-hall syndrome results from a missense mutation in the WW domain of the PQBP1 gene. Journal of Medical Genetics, 43, e30.
Matilla, A., Roberson, E. D., Banfi, S., Morales, J., Armstrong, D. L., Burright, E. N., Orr, H. T., Sweatt, J. D., Zoghbi, H. Y., & Matzuk, M. M. (1998). Mice lacking ataxin-1 display learning deficits and decreased hippocampal paired-pulse facilitation. The Journal of Neuroscience, 18, 5508–5516.
Meerang, M., Ritz, D., Paliwal, S., Garajova, Z., Bosshard, M., Mailand, N., Janscak, P., Hubscher, U., Meyer, H., & Ramadan, K. (2011). The ubiquitin-selective segregase VCP/p97 orchestrates the response to DNA double-strand breaks. Nature Cell Biology, 13, 1376–1382.
Mizuguchi, M., Obita, T., Serita, T., Kojima, R., Nabeshima, Y., & Okazawa, H. (2014). Mutations in the PQBP1 gene prevent its interaction with the spliceosomal protein U5-15kD. Nature Communications, 5, 3822.
Morchikh, M., Cribier, A., Raffel, R., Amraoui, S., Cau, J., Severac, D., Dubois, E., Schwartz, O., Bennasser, Y., & Benkirane, M. (2017). HEXIM1 and NEAT1 long non-coding RNA form a multi-subunit complex that regulates DNA-mediated innate immune response. Molecular Cell, 67, 387–399.e5.
Okazawa, H. (2018). PQBP1, an intrinsically disordered/denatured protein at the crossroad of intellectual disability and neurodegenerative diseases. Neurochemistry International, 119, 17–25.
Okazawa, H., Rich, T., Chang, A., et al. (2002). Interaction between mutant ataxin-1 and PQBP-1 affects transcription and cell death. Neuron, 34, 701–713.
Okazawa, H., Sudol, M., & Rich, T. (2001). PQBP-1 (Np/PQ): A polyglutamine tract-binding and nuclear inclusion-forming protein. Brain Research Bulletin, 56, 273–280.
Qi, M. L., Tagawa, K., Enokido, Y., et al. (2007). Proteome analysis of soluble nuclear proteins reveals that HMGB1/2 suppress genotoxic stress in polyglutamine diseases. Nature Cell Biology, 9, 402–414.
Stevenson, R. E., Bennett, C. W., Abidi, F., Kleefstra, T., Porteous, M., Simensen, R. J., Lubs, H. A., Hamel, B. C. J., & Schwartz, C. E. (2005). Renpenning syndrome comes into focus. American Journal of Medical Genetics, 134(A), 415–421.
Suh, J., Romano, D. M., Nitschke, L., et al. (2019). Loss of Ataxin-1 potentiates Alzheimer’s pathogenesis by elevating cerebral BACE1 transcription. Cell, 178, 1159–1175.e17.
Tabrizi, S. J., Leavitt, B. R., Landwehrmeyer, G. B., et al. (2019). Targeting huntingtin expression in patients with Huntington’s disease. The New England Journal of Medicine, 380, 2307–2316.
Tagawa, K., Homma, H., Saito, A., et al. (2015). Comprehensive phosphoproteome analysis unravels the core signaling network that initiates the earliest synapse pathology in preclinical Alzheimer’s disease brain. Human Molecular Genetics, 24, 540–558.
Tanaka, H., Kondo, K., Chen, X., et al. (2018). The intellectual disability gene PQBP1 rescues Alzheimer’s disease pathology. Molecular Psychiatry, 23, 2090–2110.
Taniguchi, J. B., Kondo, K., Fujita, K., et al. (2016). RpA1 ameliorates symptoms of mutant ataxin-1 knock-in mice and enhances DNA damage repair. Human Molecular Genetics, 25, 4432–4447.
Travers, A. A. (2003). Priming the nucleosome: A role for HMGB proteins? EMBO Reports, 4, 131–136.
Wan, D., Zhang, Z. C., Zhang, X., Li, Q., & Han, J. (2015). X chromosome-linked intellectual disability protein PQBP1 associates with and regulates the translation of specific mRNAs. Human Molecular Genetics, 24, 4599–4614.
Wang, Q., Moore, M. J., Adelmant, G., Marto, J. A., & Silver, P. A. (2013). PQBP1, a factor linked to intellectual disability, affects alternative splicing associated with neurite outgrowth. Genes & Development, 27, 615–626.
Waragai, M., Lammers, C. H., Takeuchi, S., Imafuku, I., Udagawa, Y., Kanazawa, I., Kawabata, M., Mouradian, M. M., & Okazawa, H. (1999). PQBP-1, a novel polyglutamine tract-binding protein, inhibits transcription activation by Brn-2 and affects cell survival. Human Molecular Genetics, 8, 977–987.
Zoghbi, H. Y., & Orr, H. T. (2009). Pathogenic mechanisms of a polyglutamine-mediated neurodegenerative disease, spinocerebellar ataxia type 1. The Journal of Biological Chemistry, 284, 7425–7429.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Okazawa, H., Tanaka, H. (2021). Molecular Dissection and Therapeutic Application of SCA1 Pathologies Revealed by Comprehensive Approaches. In: Mizusawa, H., Kakei, S. (eds) Cerebellum as a CNS Hub. Contemporary Clinical Neuroscience. Springer, Cham. https://doi.org/10.1007/978-3-030-75817-2_24
Download citation
DOI: https://doi.org/10.1007/978-3-030-75817-2_24
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-75816-5
Online ISBN: 978-3-030-75817-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)