Abstract
Satellite DNAs are tandemly repeated sequences organized in large clusters within (peri)centromeric and/or subtelomeric heterochromatin. However, in many species, satellite DNAs are not restricted to heterochromatin but are also dispersed as short arrays within euchromatin. Such genomic organization together with transcriptional activity seems to be a prerequisite for the gene-modulatory effect of satellite DNAs which was first demonstrated in the beetle Tribolium castaneum upon heat stress. Namely, enrichment of a silent histone mark at euchromatic repeats of a major beetle satellite DNA results in epigenetic silencing of neighboring genes. In addition, human satellite III transcripts induced by heat shock contribute to genome-wide gene silencing, providing protection against stress-induced cell death. Gene silencing mediated by satellite RNA was also shown to be fundamental for the early embryonic development of the mosquito Aedes aegypti. Apart from a physiological role during embryogenesis and heat stress response, activation of satellite DNAs in terms of transcription and proliferation can have an evolutionary impact. Spreading of satellite repeats throughout euchromatin promotes the variation of epigenetic landscapes and gene expression diversity, contributing to the evolution of gene regulatory networks and to genome adaptation in fluctuating environmental conditions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
6.1 Introduction
Noncoding repetitive DNAs comprise a considerable portion of most eukaryotic genomes and their function has been studied in diverse model organisms. Among the most intensively investigated noncoding repetitive DNAs are mobile transposable elements (TE) which represent an important source of regulatory sequences (Faulkner et al. 2009; Kapusta et al. 2013). By mediating the distribution of regulatory elements throughout the genome transposons are known to influence the evolution of gene-regulatory networks (Chuong et al. 2017). Recent evidence suggests that TEs can also have potent “epigenetic” effects on the regulation of gene expression and genome evolution (Choi and Lee 2020). The functional significance of another abundant class of noncoding repetitive elements such as satellite DNA, regarding gene expression regulation was also proposed (Ugarković 2005; Pezer et al. 2012) and has been recently experimentally confirmed in different studies (Feliciello et al. 2015a, 2020a; Menon et al. 2014; Joshi and Meller 2017; Halbach et al. 2020). The aim of this review is to present recent findings on the role of satellite DNA in gene expression regulation/modulation and to explain the molecular mechanisms by which satellite DNA affects genes. We focus particularly on the impact of euchromatic satellite DNA repeats dispersed outside of (peri)centromeric regions on adjacent gene expression, as well as the role of satellite transcripts since their importance for gene expression modulation has been reported in different studies. A physiological role of satellite DNAs and their transcripts in the remodeling of global heterochromatin structure and in the modulation of gene expression during development, stress response, and pathological transformation is presented. We also discuss the implication of satellite DNA-mediated gene regulation in the evolution of gene-regulatory networks and on the processes of environmental adaptation.
6.2 Proliferation and Dispersion of Satellite DNA Within Euchromatin
Satellite DNAs are preferentially organized as tandemly repeated sequences assembled in large arrays within gene-poor constitutive heterochromatin in (peri)centromeric and/or telomeric regions. Longer arrays of tandem satellite repeats within euchromatin are generally rare, probably due to the instability caused by intrastrand homologous recombination, although blocks of tandem repeats are found in euchromatin of D. melanogaster (Kuhn et al. 2012) and Triatoma infestans (Pita et al. 2017), while in the beetle Tribolium castaneum some euchromatic satellite DNAs arrays are even composed of higher-order repeats (Vlahović et al. 2017; Pavlek et al. 2015). Bioinformatic analyses of sequenced genomes however have revealed many single repeats or short arrays of satellite DNAs dispersed within euchromatin, in the vicinity of genes, in different insects such as Tribolium castaneum, Drosophila melanogaster, and Locusta migratoria (Ruiz-Ruano et al. 2016; Brajković et al. 2012, 2018; Kuhn et al. 2012). In mammals, single repeats of a major human alpha satellite DNA (Feliciello et al. 2020a, b) as well as of a major mouse satellite DNA (Bulut-Karslioglu et al. 2012) are also found interspersed among genes or within introns. It seems therefore that such mixed organization of satellite DNAs with a majority of the repeats clustered within pericentromeric constitutive heterochromatin combined with single repeats or short multimers dispersed within euchromatin is common for many species. Heterochromatic and euchromatic repeats of the same satellite show different evolutionary dynamics as revealed for Tribolium castaneum satellites, Drosophila 1.688, and human alpha satellite DNAs (Brajković et al. 2012, 2018; de Lima et al. 2020; Feliciello et al. 2020b), suggesting that chromatin domains may influence the evolution of these sequences. While heterochromatic satellite repeats display concerted evolution and a species-specific pattern, euchromatic repeats display high intra- and interspecific divergence. On the other hand, heterochromatic satellites coexisting in different species of the insect genus Pimelia evolve in parallel with fairly similar rates (Bruvo et al. 2003), indicating also the effect of chromatin state on satellite sequence evolution. Human euchromatic alpha satellite repeats have sequence characteristics of (peri)centromeric alpha repeats suggesting heterochromatin as their source but do not exhibit the concerted evolution pattern (Feliciello et al. 2020b). Alpha satellite repeats were continuously inserted within euchromatin throughout primate evolutionary history and stably transmitted to the descendant species, while their sequence divergence generally follows the primate species phylogeny (Feliciello et al. 2020b).
The pattern of dispersion of satellite DNA repeats within euchromatin can be very dynamic, differing significantly among related species as shown for Drosophila X chromosome euchromatin satellite DNAs (Sproul et al. 2020). This suggests that similar to transposable elements, euchromatic satellite repeats can be subjected to cycles of proliferation. Insertional polymorphism of euchromatic satellite repeats detected among populations of the same species or even among individuals within the same population suggests an ongoing movement of these elements within euchromatin and demonstrates the mutational potency of satellite DNAs (Feliciello et al. 2015a, b). Although a novel insertion of satellite repeat within euchromatin in many cases probably does not have an effect on genes, under particular conditions such as heat stress it can modulate the expression of nearby genes by a novel epigenetic mechanism (Feliciello et al. 2015a) which is described in the next sections. Some insertions however can affect proper gene function or even cause disease as demonstrated for the human beta satellite repeats inserted within the splice-acceptor site of a transmembrane serine protease gene which causes childhood-onset deafness (Scott et al. 2001). Mobilization of some transposons in somatic cells can also induce a pathogenic state, e.g., insertion of human L1 retroelement within the APC tumor suppressor gene initiates colorectal cancer (Scott et al. 2016).
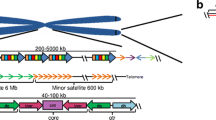
The activation of repetitive elements such as transposons, in terms of transcription and transposition, was intensively studied and was shown to be stress-induced, particularly by heat shock (Ratner et al. 1992; Piacentini et al. 2014; Cavrak et al. 2014; Makarevitch et al. 2015; Ito et al. 2016). In addition, environmental stress is responsible for a significant change in copy number of transposons, as shown in wild barley and in Drosophila (Kalendar et al. 2000; Kim et al. 2014). It has been proposed that heat stress induces modulation of heterochromatin structure which is accompanied by the rearrangement of repeats present therein, in particular of tandem satellite repeats, which are prone to homologous recombination (Fig. 6.1a; Brajković et al. 2012). Intra-chromatid recombination events can give rise to extrachromosomal circular satellite DNAs that are common for diverse eukaryotic organisms including insects, plants, and mammals (Cohen et al. 2006; Navratilova et al. 2008; Cohen and Segal 2009; Paulsen et al. 2018; Sproul et al. 2020). Extrachromosomal satellite DNA circles are proposed to be amplified by rolling circle replication and can be reintegrated within the genome by a random process of site-specific recombination which occurs between short sequence motifs within circularized satellite repeats and homologous motifs at different chromosomal sites, either within euchromatin or heterochromatin (Fig. 6.1a; Feliciello et al. 2006; Brajković et al. 2012). This process can lead to a relatively rapid change in a copy number of particular satellite DNA which can be detected at the population level (Wei et al. 2014; Feliciello et al. 2015b) or even at the individual level (Cardone et al. 1997). In addition, the same process of proliferation can spread satellite repeats to new loci and change the dispersion profiles of satellite DNA within euchromatin. These processes lead to an increase in the genetic variability among individuals within a population as well as between populations (Feliciello et al. 2015a, b).
Models of spreading of satellite DNA repeats in euchromatin. (a) Intra-chromatid recombination of satellite repeats within heterochromatin gives rise to extrachromosomal circular satellite DNAs. Short segments of homology, indicated in yellow, between circularized repeats and target regions in euchromatin are necessary for the insertion by site-specific homologous recombination. (b) Satellite transcripts can be reverse transcribed and by the activity of endonuclease/integrase cDNA is inserted within euchromatin, (c) satellite DNAs can be spread throughout the genome as an integral part of DNA transposons
Some satellite DNAs that are preferentially expressed in cancer such as human satellite II have the ability to reverse-transcribe in cancer cells and through RNA-derived DNA intermediates can expand locally and genome-wide (Bersani et al. 2015). This example of human satellite II shows that similar to retrotransposons, some satellite DNAs can proliferate through RNA intermediates and indicates coupling of satellite DNA transcription and proliferation (Fig. 6.1b). DNA transposons belonging to the Helitron superfamily have a propensity to capture and mobilize flanking DNA sequences (Thomas et al. 2014). Since some satellite repeats are found as integral parts of DNA transposons while some satellite arrays are flanked by Helitron transposons, it was proposed that the spread of satellite repeats throughout the genome can be linked to the process of transposition (Fig. 6.1c; Brajković et al. 2012; Satović et al. 2016; Vojvoda Zeljko et al. 2020). In the human genome, single repeats of a major alpha satellite DNA dispersed within euchromatin are often embedded within abundant retroelements such as Alu, L1, or ERVL-LRTs; however, there is no evidence for such elements playing a role in the spreading of alpha repeats throughout euchromatin (Feliciello et al. 2020b). While segmental duplication can be associated with the dispersion of some alpha repeats, the prevalent mechanism of spreading seems to be mediated by extrachromosomal circles of alpha satellite DNA whose insertion is facilitated by short sequence homology between alpha repeats and their target sequences (Feliciello et al. 2020b). Extrachromosomal circular DNAs (eccDNA) composed of alpha satellite repeats ranging in size from less than 2 kb to over 20 kb are detected in human cells (Cohen et al. 2010), revealing the propensity of tandemly arranged alpha repeats to generate eccDNA. The main mechanisms proposed to be responsible for the proliferation of satellite repeats and their dispersion within euchromatin are shown in Fig. 6.1.
6.3 Satellite DNA Transcription: Heat Stress Activation
Apart from a specific genomic organization of satellite DNA which is characterized by their partial dispersion within euchromatin, transcripts of satellite DNAs have also been proposed to have gene-regulatory potential (Ugarković 2005). Although satellite DNAs are preferentially embedded in constitutive heterochromatin which is considered transcriptionally inert, their transcription was reported in many species belonging to vertebrates, invertebrates, and plants (Ugarković 2005). Transcription of satellite DNAs is often bidirectional and proceeds usually by RNA polymerase II (RNAP II) from internal promoters as shown in mice (Lu and Gilbert 2007), humans (Bury et al. 2020) as well as in insects (Pezer and Ugarković 2008, 2009, 2012). The satellite transcripts fall into two main categories: long noncoding RNAs (>200 nt) and small RNAs (<200 nt; reviewed in Arunkumar and Melters 2020). Among small RNAs, the most represented are small interfering RNAs (siRNAs) which, through an RNA interference mechanism (RNAi) are involved in the epigenetic process of heterochromatin formation in fission yeast, insects as well as in plants and nematodes (Volpe et al. 2002; Pal-Bhadra et al. 2004; Grewal and Elgin 2007; Fagegaltier et al. 2009). In mammals, however, long satellite transcripts play a role in heterochromatin formation, maintenance, and regulation (Saksouk et al. 2015; Johnson et al. 2017). During mitosis, the level of mouse major and minor satellite RNA and of human alpha satellite RNA is regulated by Dicer-mediated cleavage (Fukagawa et al. 2004; Huang et al. 2015) while in meiosis the MIWI protein guided by PIWI-interacting RNAs (piRNAs) together with the endoribonuclease Dicer controls satellite RNA level (Hsieh et al. 2020). In diverse species, from plants, insects to mammals, centromeric satellite transcripts are involved in the recruitment and loading of centromere-specific histone H3 variant CENP-A as well as of CENP-B and CENP-C proteins, which are necessary for centromere organization, maintenance, and function (Bouzinba-Segard et al. 2006; Wong et al. 2007; Rosic et al. 2014; Arunkumar and Melters 2020; Chap. 7 of this book). Controlled expression of (peri)centromeric satellite RNAs, therefore, seems to be essential for ensuring proper kinetochore assembly and faithful chromosome segregation.
Constitutive heterochromatin is sensitive to temperature fluctuations and is dynamically regulated in response to environmental stimuli (Ayoub et al. 1999; Wang et al. 2016). Possible mechanism of temperature-mediated heterochromatin modulation includes stress-response transcription factors involved in heterochromatin assembly. In human cells, heat stress activates heat shock transcription factor 1 (HSF1) which recruits major cellular acetyltransferases to pericentric heterochromatin leading to targeted hyperacetylation (Col et al. 2017), facilitating particularly the transcription of satellite III DNA (Jolly et al. 2004; Rizzi et al. 2004) and satellite II (Tilman et al. 2012) but also, to a lower extent, transcription of a major alpha satellite DNA (Feliciello et al. 2020a). The human alpha satellite transcription seems to be controlled by centromere-nucleolar contacts and when the nucleolus is disrupted alpha satellite transcript levels increase substantially (Bury et al. 2020). The possible damage of nucleolus structure upon heat stress might therefore also influence the activation of alpha satellite transcription. Although human pericentromeric satellite DNAs such as alpha, satellites II and III are heavily methylated no change in methylation was detected upon heat stress (Eymery et al. 2009), confirming that transcription activation is not related to DNA methylation status. In Drosophila, under standard conditions, transcription factor dATF-2 which regulates expression of stress response genes recruits heterochromatin protein 1 (HP1) to pericentromeric heterochromatin regions that contain dATF-2 binding sites. Under stress conditions activated MAP kinase such as p38 phosphorylates dATF-2 which is released from heterochromatin, leading to the abolishment of HP1 and disruption of heterochromatin (Seong et al. 2011). In vivo studies on insect T. castaneum revealed heat-stress induced transcription of a major satellite DNA TCAST1 located within pericentromeric heterochromatic and in centromeric regions, followed by the processing of long satellite transcripts into siRNAs (Pezer and Ugarković 2012; Sermek et al. 2021). Induced satellite DNA transcription is coupled with the almost complete demethylation of satellite DNA suggesting a possible role of DNA methylation in the control of satellite DNA transcription activation upon heat stress (Feliciello et al. 2013). In Arabidopsis specific transcription factors HIT4, MED14, and UVH6 are required for transcriptional activation of heterochromatic DNA. Transposons in particular respond to heat stress and this process is accompanied by heterochromatin decondensation and 3D genome reorganization (Bourguet et al. 2018; Wang et al. 2013; Sun et al. 2020).
6.4 Satellite RNA and Euchromatic Satellite Repeats in Gene Expression Regulation
Since expression of heterochromatic satellite DNAs is induced upon heat stress in different model organisms it was investigated whether this could be linked to modulation of expression of genes located in the vicinity of satellite repeats. In the beetle T. castaneum enhanced heat stress-induced transcription of a major TCAST1 satellite DNA correlates with an increased level of repressive heterochromatin marks H3K9me2/3 on satellite repeats in constitutive heterochromatin as well as on dispersed TCAST1 satellite elements within euchromatin and their proximal regions up to 6 kb from the insertion site (Feliciello et al. 2015a). TCAST1 satellite DNA repeats dispersed within euchromatin, therefore, seem to serve as nucleation sites for transient heterochromatin formation which results in partial suppression of nearby genes upon heat stress, representing the first experimental proof for the gene-modulatory role of a satellite DNA (Feliciello et al. 2015a). In addition, the role of TCAST1-derived siRNAs in transient H3K9me2/3 enrichment at euchromatic and heterochromatic TCAST1 repeats upon heat stress is proposed (Fig. 6.2a). This proposal is consistent with the fact that small RNAs initiate the epigenetic silencing of repetitive DNAs such as satellite DNAs or transposons (TE), and the strength of these epigenetic effects was shown to be positively correlated with the amount of small RNAs targeting some TE families (Lee 2015; Lee and Karpen 2017; Choi and Lee 2020). Since this novel mode of gene expression regulation does not seem to be unique to a specific satellite DNA it is hypothesized that different satellites which are partially dispersed in the vicinity of genes and whose transcription is induced upon heat stress, could influence the expression of associated genes by the same mechanism of temporary “heterochromatinization.” Furthermore, in plants, the strength of epigenetic silencing of a TE family positively correlates with the family copy number (Cheng et al. 2006; Noreen et al. 2007), while in Drosophila, the proportion of TEs with cis-spreading of repressive marks also increases with family copy number (Lee and Karpen 2017). In addition, it could be proposed that the copy number of satellite DNAs which would be related to the level of satellite transcripts might also influence the strength of their epigenetic effects. Apart from copy number, the influence of chromatin state on the expression of transposon-derived small RNAs during embryogenesis was reported in plants (Papareddy et al. 2020). Chromatin organization is also proposed to be responsible for distinct transcription regulation of satellite DNAs in the beetle T. castaneum during embryogenesis and heat stress (Sermek et al. 2021). Namely, transcription of a major TCAST1 satellite DNA which proceeds from heterochromatic loci is specifically induced during these processes. In contrast, the transcription of a minor TCAST2 satellite which proceeds predominantly from euchromatic clusters remains unchanged. Consequently, the levels of the silent histone mark H3K9me3 at minor TCAST2 repeats as well as the expression of nearby genes are not influenced by heat stress (Sermek et al. 2021).
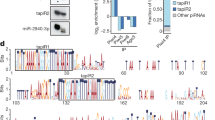
Mechanisms of satellite DNA-mediated gene expression regulation. (a) Heat stress promotes transcription of abundant pericentromeric satellite DNAs: TCAST1 in beetle T. castaneum and alpha satellite DNA in human cells. This is accompanied by increased H3K9me3 levels at euchromatic TCAST1 and alpha satellite repeats, respectively, resulting in partial suppression of nearby genes (Feliciello et al. 2015a, 2020a). Genes associated with satellite repeats are schematically shown: exons are represented by rectangles, satellite elements by arrows, and complex containing satellite RNAs by a circle. (b) In the mosquito Aedes aegypti satellite repeats located at a single euchromatic locus promote sequence-specific gene silencing in trans via the expression of piRNAs which participate in the degradation of maternally inherited transcripts during early embryonal development (Halbach et al. 2020). (c) The transcription of human Satellite III (sat III) loci is induced upon heat stress and satellite 3 transcripts sequester transcription factor CREBBP and splicing regulatory proteins SRSFs. As a consequence, there is a suppression of gene expression (Goenka et al. 2016; Ninomiya et al. 2020)
Recently it was revealed that repeats of a major human alpha satellite DNA located both in heterochromatin and euchromatin have increased H3K9me3 levels upon heat stress (Feliciello et al. 2020a). H3K9me3 enrichment at alpha repeats upon heat stress correlates with the dynamics of alpha satellite DNA transcription activation while spreading of H3K9me3 up to 1–2 kb from the insertion sites reveals that euchromatic alpha repeats act as modulators of local chromatin structure. Aside from satellite DNAs, some transposons in plants (Eichten et al. 2012) and mammals (Rebollo et al. 2011; Liu et al. 2018) reduce expression of neighboring genes by spreading heterochromatin marks, DNA methylation, and/or H3K9me2/3 from the insertion sites. A widespread influence of transposons on H3K9me3 spreading and expression of neighboring genes was also observed in Drosophila (Sienski et al. 2012; Lee and Karpen 2017). All these results suggest that epigenetic effects, in particular H3K9me3 enrichment mediated by siRNAs and piRNAs, respectively, are common for some satellite DNAs and transposons, becoming pronounced upon stress and may affect neighboring gene expression.
While in the beetle T. castaneum and in human cells the major satellites’ repeats dispersed within euchromatin modulate the local chromatin environment in cis inducing neighboring gene silencing, in the mosquito Aedes aegypti evolutionary old and conserved satellite repeats located at a single euchromatic locus promote sequence-specific gene silencing in trans via the expression of abundant PIWI-interacting RNAs (piRNAs). The satellite-derived piRNAs participate in the degradation of maternally inherited transcripts during the maternal-to-zygotic transition and are fundamental to early embryonic development (Halbach et al. 2020; Fig. 6.2b). Satellite DNA-derived siRNAs also play a specific role in gene expression regulation in Drosophila. Namely, short, tandem clusters of 1.688 satellite DNA in the X chromosome euchromatin of D. melanogaster males guide the dosage compensation complex MLS which increases expression of nearby genes and the 1.688 siRNAs play a role in this process (Menon et al. 2014; Joshi and Meller 2017, Chap. 1 of this book). The short euchromatic array of 1.688 satellite on the X chromosome is also shown to promote specific targeting of POF protein which is involved in the global regulation of genes on D. melanogaster chromosome 4 (Kim et al. 2018), while depletion of a large block of pericentromeric 1.688 satellite seems to affect eggshell formation (Ekhteraei-Tousi et al. 2020). Human alpha satellite DNA repeats in addition to primates have been detected as rare, highly conserved elements in evolutionarily distant species such as chicken and zebrafish (Li and Kirby 2003). The presence of several coding mRNAs in human and chick embryos that contain alpha-like satellite repeats as a part of their 5′ or 3′ untranslated regions indicates that their expression could be controlled in trans by alpha satellite RNA (Li and Kirby 2003).
Some satellite DNA repeats are located within introns of particular genes affecting their expression under specific conditions or developmental stages. One such example is the tandem repeats found within the intron of the major histocompatibility complex gene (MHIIβ) in the fish Salvelinus fontinalis which are involved in temperature-dependant modulation of expression of this gene (Croisetière et al. 2010). The minisatellite was proposed to play a role in the regulation of the adaptive immune response but the molecular mechanism behind its gene-modulatory effect was not investigated. In plants such as Arabidopsis thaliana and particularly in rice, introns of many genes contain heterochromatin associated with repetitive elements, mostly transposons (Duan et al. 2017; Espinas et al. 2020). The establishment and maintenance of heterochromatin within introns seem to be critical for transcriptional control of the associated genes which are predominantly required for environmental responses and development (Le et al. 2015; Khan et al. 2013). The transcription of genes with intronic heterochromatin is regulated by an epigenetic mechanism that involves the conserved nuclear protein complex, mutation of which results in severe developmental defects (Duan et al. 2017; Espinas et al. 2020). Introns containing long arrays of satellite DNAs are characteristic for Drosophila Y chromosome genes which are solely expressed during spermatogenesis (Hardy et al. 1981). The gigantic introns of these genes are transcribed in line with their exons; however, their expression requires a unique gene expression program, which acts on both transcription and posttranscriptional processing (Fingerhut et al. 2019). It is proposed that satellite DNA-containing gigantic introns could act in a manner similar to enhancers, recruiting transcriptional machinery to the Y-loop genes, while intron size can play a critical role in the regulation of gene expression (Shaul 2017; Fingerhut et al. 2019).
6.5 “Macroheterochromatin” in Gene Expression Regulation
A “micro-heterochromatin” is formed on some satellite repeats or short arrays dispersed within euchromatin and can affect the expression of genes located in the vicinity (Feliciello et al. 2015a, b, 2020a). On the other hand, a “macro-heterochromatin” is composed of megabase stretches of satellite DNA such as those on the Drosophila Y chromosome and polymorphism in heterochromatic Y chromosomes results in genome-wide gene expression variation (Lemos et al. 2010). It seems that Y chromosome heterochromatin serves as a source of epigenetic variation in natural populations that interacts with chromatin components to modulate the expression of biologically relevant phenotypic variation. Increasing the amount of repeats on the X or Y D. melanogaster chromosome results in a decrease of H3K9me2/3 levels at repeat-rich regions at pericentromeres and the Y chromosome, implying a role for satellite DNA in global chromatin dynamics and redistribution of chromatin regulators across the genome (Brown et al. 2020). Since satellite DNAs are characterized by a rapid copy number change often observed at the intraspecific level (Cardone et al. 1997; Wei et al. 2014; Feliciello et al. 2015b), a significant difference in their amount may contribute to the diversity of expression of genes and repetitive elements among populations and individuals. Analyses of 3D genome structures reveal that pericentromeric heterochromatin spatially contacts distant euchromatin regions enriched for repressive epigenetic marks, such as regions associated with epigenetically silenced transposable elements or other repeats, as shown in D. melanogaster (Lee et al. 2020). It can be proposed that due to such interactions, pericentromeric heterochromatin could impact the expression of distant euchromatic genes which are associated with “H3K9me2/3 islands.” This also indicates that an interplay between satellite DNA repeats located within heterochromatin and euchromatin might be involved in genome-wide gene expression regulation.
6.6 Satellite DNA Role in Stress Response and Environmental Adaptation
Numerous in vitro studies on human cell lines have shown a strong increase of pericentromeric satellite III expression induced by a large number of stressing agents including heat shock, DNA damaging agents, and hyperosmotic stress (reviewed in Vourc'h and Biamonti 2011). While most of these stressing agents act through heat shock transcription factor 1 (HSF1), transcription of satellite III in response to hyperosmotic stress depends on Tonicity Enhancer Binding Protein (TonEBP) which controls genes responsible for the survival of cells subjected to high osmotic pressure (Valgardsdottir et al. 2008). It was proposed that stress-induced activation of satellite III occurred through at least two independent pathways which both lead to the formation of nuclear stress bodies, and is considered to be a part of a general cellular response to stress. Namely, satellite III transcripts recruit critical factors involved in the transcriptional process, contributing to heat-induced transcriptional silencing and seem to be required to provide protection against heat-shock-induced cell death (Goenka et al. 2016; Fig. 6.2c). Satellite III RNA also mediates in the recruitment of a number of RNA binding proteins involved in pre-mRNA processing and participates in the control of gene expression upon heat stress at the level of splicing regulation (Ninomiya et al. 2020). The alteration of the splicing profile is mainly characterized by an increase in intron retention events during the recovery from heat shock. Intron retention prevents the export of the pre-mRNAs from the nucleus resulting in suppression of gene expression at the posttranscriptional level. Expression of centromeric satellites is also strongly induced by genotoxic stress as shown for mouse minor satellite DNA and their accumulation under stress conditions seems to be a conserved feature of the cellular stress response (Hédouin et al. 2017).
In the insect Tribolium castaneum and in human cells activation of (peri)centromeric satellite DNA transcription during heat stress response reinforces “heterochromatinization” and helps heterochromatin recovery (Pezer and Ugarković 2012; Feliciello et al. 2020a). Since heterochromatin is important for genome stability and integrity, satellite DNA transcripts might have a protective effect in stressed cells/organisms. In addition, induced “heterochromatinization” leads to transient suppression of genes located in the vicinity of dispersed TCAST1 satellite elements (Feliciello et al. 2015a), as described previously in this chapter. However, what is the physiological consequence of such transient gene suppression upon heat stress? It is known that after strong heat stress genomes undergo a substantial transcriptional silencing and the role of human satellite III in this process was demonstrated (Goenka et al. 2016). It could be hypothesized that other satellite DNAs contribute to the same process of gene repression which is necessary to protect the cell from stress-induced damage. While human satellite III RNA affects gene expression genome-wide, in the case of TCAST1 satellite expression of genes located in the vicinity of euchromatic TCAST1 repeats is affected. Within genes associated with euchromatic TCAST1 satellite repeats, there is a significant overrepresentation of immunoglobulin-like genes (Brajković et al. 2012). Stress and the immune response are tightly connected in insects and mild physical or thermal stress leads to short-term immune memory (Altincicek et al. 2009; Freitak et al. 2012; Marshall and Sinclair 2012; Eggert et al. 2015). In mammals, genes involved in immunity and stress are more likely to contain transposon sequences within UTRs than other genes (van de Lagemaat et al. 2003). In plants, genes required for environmental response and development are enriched with heterochromatic introns associated with repetitive elements (Duan et al. 2017; Espinas et al. 2020). These data indicate that repetitive elements, either transposons or satellite repeats, seem to be preferentially associated with environment susceptible genes such as stress or immune response genes and might affect their expression under specific conditions. In addition, the high evolutionary dynamics of repetitive elements can promote expression variation and the evolution of associated genes. Differential transcription activation of satellite DNA families by heat stress and clustering of their repeats near some genes, as observed in T. castaneum (Brajković et al. 2018), may facilitate satellite-mediated gene modulatory effects and increase the complexity of the transcriptional response to the environment. Satellite DNA-induced changes of the transcriptome might create a modified gene interaction network with a strong adaptive potential on which natural selection can act (Fig. 6.3). In ectothermic organisms in particular, whose body temperatures conform to ambient temperature, the temperature is one of the principal environmental variables that drive adaptive evolution. It is also important to mention that satellite DNAs are subjected to a high evolutionary turnover, resulting in a rapid change of their copy number (Meštrović et al. 1998) as well as in the emergence of new satellites which could sometimes contribute to the evolution of a novel feature (Ugarković and Plohl 2002). In the New World Monkey genus Aotus the newly amplified satellite DNA builds a centrally located heterochromatin block in the nucleus of the rod cells which is responsible for the evolution of night vision characteristic for species of this genus (Koga et al. 2017). This represents an example of how a newly acquired satellite DNA contributes to the adaptation of its host organism to exploit an ecological niche.
Model explaining the role of satellite DNAs in the adaptation to environmental conditions. Heat stress induces activation of satellite DNAs in terms of transcription and proliferation which can lead to the insertion polymorphism of satellite repeats within euchromatin. This may cause gene expression diversity among individuals upon heat stress, on which natural selection can act, fixing the beneficial mutations and contributing to the adaptation process
6.7 Satellite DNA in Pathological Transformation and Development
Satellite DNA transcription is activated not only by environmental stress but also upon pathological conditions. In epithelial cancers increased satellite DNA transcription is observed (Ting et al. 2011) and it is often associated with a deficiency of tumor suppressor proteins, in particular p53 which restrains the movement of repetitive elements (Wylie et al. 2016). Besides p53, deficiency of the tumor suppressor BRCA1 impairs the integrity of constitutive heterochromatin and induces abnormal transcription of satellite DNA repeats (Zhu et al. 2011). Overexpressed heterochromatic satellite RNAs sequester BRCA1-associated proteins causing destabilization of DNA replication forks, and promote breast cancer formation in mice (Zhu et al. 2018). In mouse K-ras-mutated pancreatic precancerous tissues, transcripts of a major pericentromeric satellite DNA inhibit the DNA-damage repair function of YBX1 protein and accelerate tumor formation, and so act as “intrinsic mutagens” (Kishikawa et al. 2016, 2018). Human satellite II transcripts which are preferentially expressed in cancer cells are immunogenic, able to directly activate the innate immune system to produce cytokines, modulating in this way the immune response against tumor cells (Tanne et al. 2015). In addition, demethylated human satellite II and its transcripts act as molecular sponges and sequester chromatin regulatory proteins into abnormal nuclear bodies in cancer (Hall et al. 2017). Expression of human satellite II is also strongly induced in herpesvirus infected cells by viral proteins, while satellite II transcripts modulate viral protein expression and release of infectious particles, having functionally important consequences for viral replication (Nogalski et al. 2019). Hypomethylation of pericentromeric sequences and subsequent derepression of associated satellite transcripts triggers an interferon response in zebrafish (Rajshekar et al. 2018).
Apart from stress and pathological states, activation of satellite DNA transcription in many organisms is associated with cell cycle progression, development, and differentiation (Probst et al. 2010; Kishi et al. 2012; Park et al. 2018; Ferreira et al. 2020). The bidirectional transcription of murine minor satellite DNA occurs during mitosis and transcripts stabilize the overall kinetochore structure in the G2/M phase (Ferri et al. 2009), while in meiosis transcription mostly occurs during the early-pachytene stage (Hecht 1986). During early mouse embryogenesis, the major pericentromeric satellite RNA modulates the activity of histone methyltransferase SUV39H2 and reduces H3K9me3 levels in zygotes (Burton et al. 2020), while during the midblastula stage a burst of strand-specific transcription of a major pericentromeric satellite DNA is essential for heterochromatin formation and early development progression (Probst et al. 2010; Casanova et al. 2013). In addition, the same satellite is differentially expressed in cells of the developing central nervous system (Rudert et al. 1995) and this satellite RNA whose level is significantly increased during neuronal differentiation (Kishi et al. 2012) is necessary for the correct higher-order organization of pericentromeric heterochromatin (Fioriniello et al. 2020). It is interesting that although the transcription of the major satellite proceeds from both DNA strands only the satellite forward RNA is involved in the initial heterochromatin formation during embryogenesis (Maison et al. 2011) and in pericentromeric heterochromation organization in neurons (Fioriniello et al. 2020). Major and minor mouse satellite RNAs are also involved in the large-scale reorganization of constitutive heterochromatin during muscle differentiation (Park et al. 2018). In chicken and zebrafish, transcription of alphoid repeat sequences displays a specific temporal and spatial expression pattern during embryogenesis (Li and Kirby 2003). In insects, transcription of satellite DNAs is also developmentally regulated being increased during specific stages of embryogenesis as revealed for the major TCAST1 satellite DNA of T. castaneum, and transcripts in the form of TCAST1 piRNAs and siRNAs are proposed to be necessary for initial heterochromatin formation (Pezer and Ugarković 2012; Sermek et al. 2021). In the mosquito Aedes aegypti, piRNAs which derive from conserved euchromatic satellite DNA are necessary for embryonic development (Halbach et al. 2020), while RNA from a simple satellite DNA of D. melanogaster is required for sperm maturation and male fertility (Mills et al. 2019). Examples in other species also indicate that transcription activation of satellite DNAs as well as of some other repetitive families such as LINE1 might be a part of normal developmental and differentiation processes. While pericentromeric satellite DNA transcripts are important for regulation of heterochromatin establishment during early mouse embryogenesis and for heterochromatin remodeling during differentiation (Park et al. 2018; Burton et al. 2020), LINE-1 transcripts are relevant for regulation of global chromatin dynamics: de- and recondensation (Jachowicz et al. 2017), acting as a nuclear scaffold to direct gene expression programs essential for embryo development (Percharde et al. 2018). Transcripts of some satellite DNAs such as FA-SAT DNA are proposed to be important for cell proliferation (Ferreira et al. 2020). FA-SAT DNA is highly conserved in mammals being primarily located at the telomeres and FA-SAT RNA forms a nuclear complex with Pyruvate Kinase Muscle Isozyme protein (PKM2) which seems to participate in cell-cycle progression.
In conclusion, satellite DNA transcripts activated either by environmental stress or during pathological transformation and viral infection, are implicated in immune system modulation and stress responsiveness, although they act through different molecular pathways and mechanisms. On the one hand, satellite transcripts can promote oncogenic processes by inducing mutations (Kishikawa et al. 2016), affecting epigenetic regulators (Hall et al. 2017), enhancing tumor cell proliferation (Nogalski and Shenk 2020), or compromising replication fork stability and genome integrity (Zeller and Gasser 2017; Zhu et al. 2018). Satellite RNAs also provide protection against heat-shock-induced cell death (Goenka et al. 2016) and are a prerequisite for early embryonic development (Probst et al. 2010; Halbach et al. 2020) and cell differentiation (Park et al. 2018). On the other hand, besides playing a physiological role in the modulation of global heterochromatin structure as well as of the gene expression program during development and stress response, activation of satellite DNAs in terms of transcription and proliferation (mobilization) has an evolutionary impact. It generates spreading and insertion polymorphism of euchromatic satellite repeats, causing variation of epigenetic landscapes and gene expression diversity within species (Feliciello et al. 2015a; Fig. 6.3). This variation is additionally enhanced by the propensity of satellites to change copy numbers and to form new satellite families. This gene expression diversity contributes to the evolution of gene regulatory networks, increases the evolvability of species, and could represent a powerful adaptive response of the genome to changing environmental conditions.
References
Altincicek B, Knorr E, Vilcinskas A (2009) Beetle immunity: identification of immune-inducible genes from the model insect Tribolium castaneum. Dev Comp Immunol 32:585–595
Arunkumar G, Melters DP (2020) Centromeric transcription: a conserved Swiss-Army knife. Genes (Basel) 11:E911
Ayoub N, Goldshmidt I, Cohen A (1999) Position effect variegation at the mating-type locus of fission yeast: a cis-acting element inhibits covariegated expression of genes in the silent and expressed domains. Genetics 152(2):495–508
Bersani F, Lee E, Kharchenko PV, Xu AW, Liu M, Xega K, MacKenzie OC, Brannigan BW, Wittner BS, Jung H, Ramaswamy S, Park PJ, Maheswaran S, Ting DT, Haber DA (2015) Pericentromeric satellite repeat expansions through RNA-derived DNA intermediates in cancer. Proc Natl Acad Sci USA 112:15148–15153
Bourguet P, de Bossoreille S, López-González Pouch-Pélissier MN, Gómez-Zambrano Á, Devert A, Pélissier T, Pogorelcnik R, Vaillant I, Mathieu O (2018) A role for MED14 and UVH6 in heterochromatin transcription upon destabilization of silencing. Life Sci Alliance 1:e201800197
Bouzinba-Segard H, Guais A, Francastel C (2006) Accumulation of small murine minor satellite transcripts leads to impaired centromeric architecture and function. Proc Natl Acad Sci USA 103:8709–8714
Brajković J, Feliciello I, Bruvo-Mađarić B, Ugarković Đ (2012) Satellite DNA-like elements associated with genes within euchromatin of the beetle Tribolium castaneum. G3 (Bethesda) 2:931–941
Brajković J, Pezer Ž, Bruvo-Mađarić B, Sermek A, Feliciello I, Ugarković Đ (2018) Dispersion profiles and gene associations of repetitive DNAs in the euchromatin of the beetle Tribolium castaneum. G3 (Bethesda) 8:875–886
Brown EJ, Nguyen AH, Bachtrog D (2020) The Drosophila Y chromosome affects heterochromatin integrity genome-wide. Mol Biol Evol 37:2808–2824
Bruvo B, Pons J, Ugarković D, Juan C, Petitpierre E, Plohl M (2003) Evolution of low-copy number and major satellite DNA sequences coexisting in two Pimelia species-groups (Coleoptera). Gene 17(312):85–94
Bulut-Karslioglu A, Perrera V, Scaranaro M, de la Rosa-Velazquez IA, van de Nobelen S, Shukeir N, Popow J, Gerle B, Opravil S, Pagani M, Meidhof S, Brabletz T, Manke T, Lachner M, Jenuwein T (2012) A transcription factor-based mechanism for mouse heterochromatin formation. Nat Struct Mol Biol 19:1023–1030
Burton A, Brochard V, Galan C, Ruiz-Morales ER, Rovira Q, Rodriguez-Terrones D, Kruse K, Le Gras S, Udayakumar VS, Chin HG, Eid A, Liu X, Wang C, Gao S, Pradhan S, Vaquerizas JM, Beaujean N, Jenuwein T, Torres-Padilla ME (2020) Heterochromatin establishment during early mammalian development is regulated by pericentromeric RNA and characterized by non-repressive H3K9me3. Nat Cell Biol 22:767–778
Bury L, Moodie B, Ly J, McKay LS, Miga KH, Cheeseman IM (2020) Alpha-satellite RNA transcripts are repressed by centromere-nucleolus associations. eLife 9:e59770
Cardone DE, Feliciello I, Marotta M, Rosati C, Chinali G (1997) A family of centromeric satellite DNAs from the European brown frog Rana graeca italica. Genome 40:774–781
Casanova M, Pasternak M, El Marjou F, Le Baccon P, Probst AV, Almouzni G (2013) Heterochromatin reorganization during early mouse development requires a single-stranded noncoding transcript. Cell Rep 4:1156–1167
Cavrak VV, Lettner N, Jamge S, Kosarewicz A, Bayer LM, Mittelsten Scheid O (2014) How a retrotransposon exploits the Plant’s heat stress response for its activation. PLoS Genet 10:e1004115
Cheng C, Daigen M, Hirochika H (2006) Epigenetic regulation of the rice retrotransposon Tos17. Mol Gen Genomics 276:378–390
Choi JY, Lee YCG (2020) Double-edged sword: the evolutionary consequences of the epigenetic silencing of transposable elements. PLoS Genet 16:e1008872
Chuong EB, Elde NC, Feschotte C (2017) Regulatory activities of transposable elements: from conflicts to benefits. Nat Rev Genet 18:71–86
Cohen S, Segal D (2009) Extrachromosomal circular DNA in eukaryotes: possible involvement in the plasticity of tandem repeats. Cytogenet Genome Res 124:327–338
Cohen Z, Bacharach E, Lavi S (2006) Mouse major satellite DNA is prone to eccDNA formation via DNA ligase IV-dependent pathway. Oncogene 25:4515–4524
Cohen S, Agmon N, Sobol O, Segal D (2010) Extrachromosomal circles of satellite repeats and 5S ribosomal DNA in human cells. Mob DNA 1:11
Col E, Hoghoughi N, Dufour S, Penin J, Koskas S, Faure V, Ouzounova M, Hernandez-Vargash H, Reynoird N, Daujat S, Folco E, Vigneron M, Schneider R, Verdel A, Khochbin S, Herceg Z, Caron C, Vourc'h C (2017) Bromodomain factors of BET family are new essential actors of pericentric heterochromatin transcriptional activation in response to heat shock. Sci Rep 7:5418
Croisetière S, Bernatchez L, Belhumeur P (2010) Temperature and length-dependent modulation of the MH class II beta gene expression in brook charr (Salvelinus fontinalis) by a cis-acting minisatellite. Mol Immunol 47:1817–1829
de Lima LG, Hanlon SL, Gerton JL (2020) Origins and evolutionary patterns of the 1.688 satellite DNA family in Drosophila phylogeny. G3 (Bethesda) 10:4129–4146
Duan CG, Wang X, Zhang L, Xiong X, Zhang Z, Tang K, Pan L, Hsu CC, Xu H, Tao WA, Zhang H, Zhu JK (2017) A protein complex regulates RNA processing of intronic heterochromatin-containing genes in Arabidopsis. Proc Natl Acad Sci USA 114:E7377–E7E84
Eggert H, Diddens-de Buhr MF, Kurtz J (2015) A temperature shock can lead to trans-generational immune priming in the red flour beetle, Tribolium castaneum. Ecol Evol 5:1318–1326
Eichten SR, Ellis NA, Makarevitch I, Yeh CT, Gent JI, Guo L, McGinnis KM, Zhang X, Schnable PS, Vaughn MW, Dawe RK, Springer NM (2012) Spreading of heterochromatin is limited to specific families of maize retrotransposons. PLoS Genet 8:e1003127
Ekhteraei-Tousi S, Lewerentz J, Larsson J (2020) Painting of fourth and the X-linked 1.688 satellite in D. melanogaster is involved in chromosome-wide gene regulation. Cells 9:323
Espinas NA, Tu LN, Furci L, Shimajiri Y, Harukawa Y, Miura S, Takuno S, Saze H (2020) Transcriptional regulation of genes bearing intronic heterochromatin in the rice genome. PLoS Genet 16:e1008637
Eymery A, Horard B, El Atifi-Borel M, Fourel G, Berger F, Vitte AL, Van den Broeck A, Brambilla E, Fournier A, Callanan M, Gazzeri S, Khochbin S, Rousseaux S, Gilson E, Vourc'h C (2009) A transcriptomic analysis of human centromeric and pericentric sequences in normal and tumor cells. Nucleic Acids Res 37:6340–6354
Fagegaltier D, Bougé AL, Berry B, Poisot E, Sismeiro O, Coppée JY, Théodore L, Voinnet O, Antoniewski C (2009) The endogenous siRNA pathway is involved in heterochromatin formation in Drosophila. Proc Natl Acad Sci USA 106:21258–21263
Faulkner GJ, Kimura Y, Daub CO, Wani S, Plessy C, Irvine KM, Schroder K, Cloonan N, Steptoe AL, Lassmann T, Waki K, Hornig N, Arakawa T, Takahashi H, Kawai J, Forrest AR, Suzuki H, Hayashizaki Y, Hume DA, Orlando V, Grimmond SM, Carninci P (2009) The regulated retrotransposon transcriptome of mammalian cells. Nat Genet 41:563–571
Feliciello I, Picariell O, Chinali G (2006) Intra-specific variability and unusual organization of the repetitive units in a satellite DNA from Rana dalmatina: molecular evidence of a new mechanism of DNA repair acting on satellite DNA. Gene 383:81–92
Feliciello I, Parazajder J, Akrap I, Ugarković Đ (2013) First evidence of DNA methylation in insect Tribolium castaneum - environmental regulation of DNA methylation within heterochromatin. Epigenetics 8:534–541
Feliciello I, Akrap I, Ugarković Đ (2015a) Satellite DNA modulates gene expression in the beetle Tribolium castaneum after heat stress. PLoS Genet 11:e1005466
Feliciello I, Akrap I, Brajković J, Zlatar I, Ugarković Đ (2015b) Satellite DNA as a driver of population divergence in the red flour beetle Tribolium castaneum. Genome Biol Evol 7:228–239
Feliciello I, Sermek A, Pezer Ž, Matulić M, Ugarković Đ (2020a) Heat stress affects H3K9me3 level at human alpha satellite DNA repeats. Genes 11:663
Feliciello I, Pezer Z, Kordiš D, Bruvo-Mađarić B, Ugarković Đ (2020b) Evolutionary history of alpha satellite DNA repeats dispersed within human genome euchromatin. Genome Biol Evol 12:2125–2138
Ferreira D, Escudeiro A, Adega F, Anjo SI, Manadas B, Chaves R (2020) FA-SAT ncRNA interacts with PKM2 protein: depletion of this complex induces a switch from cell proliferation to apoptosis. Cell Mol Life Sci 77:1371–1386
Ferri F, Bouzinba-Segard H, Velasco G, Hubé F, Francastel C (2009) Noncoding murine centromeric transcripts associate with and potentiate Aurora B kinase. Nucleic Acids Res 37:5071–5080
Fingerhut JM, Moran JV, Yamashita YM (2019) Satellite DNA-containing gigantic introns in a unique gene expression program during Drosophila spermatogenesis. PLoS Genet 15:e1008028
Fioriniello S, Csukonyi E, Marano D, Brancaccio A, Madonna M, Zarrillo C, Romano A, Marracino F, Matarazzo MR, D'Esposito M, Della Ragione F (2020) MeCP2 and major satellite forward RNA cooperate for Pericentric heterochromatin organization. Stem Cell Rep 15:1317–1332
Freitak D, Knorr E, Vogel H, Vilcinskas A (2012) Gender- and stressor-specific microRNA expression in Tribolium castaneum. Biol Lett 8:860–863
Fukagawa T, Nogami M, Yoshikawa M, Ikeno M, Okazaki T, Takami Y, Nakayama T, Oshimura M (2004) Dicer is essential for formation of the heterochromatin structure in vertebrate cells. Nat Cell Biol 6:784–791
Goenka A, Sengupta S, Pandey R, Parihar R, Mohanta GC, Mukerji M, Ganesh S (2016) Human satellite-III non-coding RNAs modulate heat-shock-induced transcriptional repression. J Cell Sci 129:3541–3552
Grewal SI, Elgin SC (2007) Transcription and RNA interference in the formation of heterochromatin. Nature 447:399–406
Halbach R, Miesen P, Joosten J, Taşköprü E, Rondeel I, Pennings B, Vogels CBF, Merkling SH, Koenraadt CJ, Lambrechts L, van Rij RP (2020) A satellite repeat-derived piRNA controls embryonic development of Aedes. Nature 580:274–277
Hall LL, Byron M, Carone DM, Whitfield TW, Pouliot GP, Fischer A, Jones P, Lawrence JB (2017) HSATII DNA and HSATII RNA foci sequester PRC1 and MeCP2 into cancer-specific nuclear bodies. Cell Rep 18:2943–2956
Hardy RW, Tokuyasu KT, Lindsley DL (1981) Analysis of spermatogenesis in Drosophila melanogaster bearing deletions for Y-chromosome fertility genes. Chromosoma 83:593–617
Hecht NB (1986) Regulation of gene expression during mammalian spermatogenesis. In: Experimental approaches to mammalian embryonic development. Cambridge University Press, Cambridge
Hédouin S, Grillo G, Ivkovic I, Velasco G, Francastel C (2017) CENP-A chromatin disassembly in stressed and senescent murine cells. Sci Rep 7:42520
Hsieh CL, Xia J, Lin H (2020) MIWI prevents aneuploidy during meiosis by cleaving excess satellite RNA. EMBO J 17:e103614
Huang C, Wang X, Liu X, Cao S, Shan G (2015) RNAi pathway participates in chromosome segregation in mammalian cells. Cell Discov 1:15029
Ito H, Kim JM, Matsunaga W, Saze H, Matsui A, Endo TA, Harukawa Y, Takagi H, Yaegashi H, Masuta Y, Masuda S, Ishida J, Tanaka M, Takahashi S, Morosawa T, Toyoda T, Kakutani T, Kato A, Seki M (2016) Stress-activated transposon in Arabidopsis induces transgenerational abscisic acid insensitivity. Sci Rep 6:23181
Jachowicz JW, Bing X, Pontabry J, Bošković A, Rando OJ, Torres-Padilla ME (2017) LINE-1 activation after fertilization regulates global chromatin accessibility in the early mouse embryo. Nat Genet 49:1502–1510
Johnson WL, Yewdell WT, Bell JC, McNulty SM, Duda Z, O’Neill RJ, Sullivan BA, Straight AF (2017) RNA-dependent stabilization of SUV39H1 at constitutive heterochromatin. eLife 6:e25299
Jolly C, Metz A, Govin J, Vigneron M, Turner BM, Khochbin S, Vourc'h C (2004) Stress-induced transcription of satellite III repeats. J Cell Biol 164:25–33
Joshi SS, Meller VH (2017) Satellite repeats identify X chromatin for dosage compensation in Drosophila melanogaster males. Curr Biol 27:1393–1402
Kalendar R, Tanskanen J, Immonen S, Nevo E, Schulman AH (2000) Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence. Proc Natl Acad Sci USA 97:6603–6607
Kapusta A, Kronenberg Z, Lynch VJ, Zhuo X, Ramsay L, Bourque G, Yandell M, Feschotte C (2013) Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet 9:e1003470
Khan AR, Enjalbert J, Marsollier AC, Rousselet A, Goldringer I, Vitte C (2013) Vernalization treatment induces site-specific DNA hypermethylation at the VERNALIZATION-A1 (VRN-A1) locus in hexaploid winter wheat. BMC Plant Biol 13:209
Kim YB, Oh JH, McIver LJ, Rashkovetsky E, Michalak K, Garner HR, Kang L, Nevo E, Korol AB, Michalak P (2014) Divergence of Drosophila melanogaster repeatomes in response to a sharp microclimate contrast in evolution canyon, Israel. Proc Natl Acad Sci USA 111:10630–10635
Kim M, Ekhteraei-Tousi S, Lewerentz J, Larsson J (2018) The X-linked 1.688 satellite in Drosophila melanogaster promotes specific targeting by painting of fourth. Genetics 208:623–632
Kishi Y, Kondo S, Gotoh Y (2012) Transcriptional activation of mouse major satellite regions during neuronal differentiation. Cell Struct Funct 37:101–110
Kishikawa T, Otsuka M, Yoshikawa T, Ohno M, Ijichi H, Koike K (2016) Satellite RNAs promote pancreatic oncogenic processes via the dysfunction of YBX1. Nat Commun 7:13006
Kishikawa T, Otsuka M, Suzuki T, Seimiya T, Sekiba K, Ishibashi R, Tanaka E, Ohno M, Yamagami M, Koike K (2018) Satellite RNA increases DNA damage and accelerates tumor formation in mouse models of pancreatic Cancer. Mol Cancer Res 16:1255–1262
Koga A, Tanabe H, Hirai Y, Imai H, Imamura M, Oishi T, Stanyon R, Hirai H (2017) Co-opted megasatellite DNA drives evolution of secondary night vision in Azara’s owl monkey. Genome Biol Evol 9:1963–1970
Kuhn GC, Küttler H, Moreira-Filho O, Heslop-Harrison JS (2012) The 1.688 repetitive DNA of Drosophila: concerted evolution at different genomic scales and association with genes. Mol Biol Evol 29:7–11
Le TN, Miyazaki Y, Takuno S, Saze H (2015) Epigenetic regulation of intragenic transposable elements impacts gene transcription in Arabidopsis thaliana. Nucleic Acids Res 43:3911–3921
Lee YCG (2015) The role of piRNA-mediated epigenetic silencing in the population dynamics of transposable elements in Drosophila melanogaster. PLoS Genet 11:e1005269
Lee YCG, Karpen GH (2017) Pervasive epigenetic effects of Drosophila euchromatic transposable elements impact their evolution. elife 6:e25762
Lee YCG, Ogiyama Y, Martins NMC, Beliveau BJ, Acevedo D, Wu CT, Cavalli G, Karpen GH (2020) Pericentromeric heterochromatin is hierarchically organized and spatially contacts H3K9me2 islands in euchromatin. PLoS Genet 16:e1008673
Lemos B, Branco AT, Hartl DL (2010) Epigenetic effects of polymorphic Y chromosomes modulate chromatin components, immune response, and sexual conflict. Proc Natl Acad Sci USA 107:15826–15831
Li YX, Kirby ML (2003) Coordinated and conserved expression of alphoid repeat and alphoid repeat-tagged coding sequences. Dev Dyn 228:72–81
Liu N, Lee CH, Swigut T, Grow E, Gu B, Bassik MC, Wysocka J (2018) Selective silencing of euchromatic L1s revealed by genome-wide screens for L1 regulators. Nature 553:228–232
Lu J, Gilbert DM (2007) Proliferation- dependent and cell cycle- regulated transcription of mouse pericentromeric heterochromatin. J Cell Biol 179:411–421
Maison C, Bailly D, Roche D, Montes de Oca R, Probst AV, Vassias I, Dingli F, Lombard B, Loew D, Quivy JP, Almouzni G (2011) SUMOylation promotes de novo targeting of HP1α to pericentric heterochromatin. Nat Genet 43:220–227
Makarevitch I, Waters AJ, West PT, Stitzer M, Hirsch CN, Ross-Ibarra J, Springer NM (2015) Transposable elements contribute to activation of maize genes in response to abiotic stress. PLoS Genet 11:e1004915
Marshall KE, Sinclair BJ (2012) The impacts of repeated cold exposure on insects. J Exp Biol 215:1607–1613
Menon DU, Coarfa C, Xiao W, Gunaratne PH, Meller VH (2014) siRNAs from an X-linked satellite repeat promote X-chromosome recognition in Drosophila melanogaster. Proc Natl Acad Sci USA 111:16460–16465
Meštrović N, Plohl M, Mravinac B, Ugarković Đ (1998) Evolution of satellite DNAs from the genus Palorus—experimental evidence for the “library” hypothesis. Mol Biol Evol 15:1062–1068
Mills WK, Lee YCG, Kochendoerfer AM, Dunleavy EM, Karpen GH (2019) RNA from a simple-tandem repeat is required for sperm maturation and male fertility in Drosophila melanogaster. eLife 8:e48940
Navratilova A, Koblizkova A, Macas J (2008) Survey of extrachromosomal circular DNA derived from plant satellite repeats. BMC Plant Biol 8:90
Ninomiya K, Adachi S, Natsume T, Iwakiri J, Terai G, Asai K, Hirose T (2020) LncRNA-dependent nuclear stress bodies promote intron retention through SR protein phosphorylation. EMBO J 39:e102729
Nogalski MT, Shenk T (2020) HSATII RNA is induced via a noncanonical ATM-regulated DNA damage response pathway and promotes tumor cell proliferation and movement. Proc Natl Acad Sci USA 117:31891–31901
Nogalski MT, Solovyov A, Kulkarni AS, Desai N, Oberstein A, Levine AJ, Ting DT, Shenk T, Greenbaum BD (2019) A tumor-specific endogenous repetitive element is induced by herpesviruses. Nat Commun 10:90
Noreen F, Akbergenov R, Hohn T, Richert-Poggeler KR (2007) Distinct expression of endogenous Petunia vein clearing virus and the DNA transposon dTph1 in two Petunia hybrida lines is correlated with differences in histone modification and siRNA production. Plant J 50:219–229
Pal-Bhadra M, Leibovitch BA, Gandhi SG, Chikka MR, Bhadra U, Birchler JA, Elgin SC (2004) Heterochromatic silencing and HP1 localization in Drosophila are dependent on the RNAi machinery. Science 303:669–672
Papareddy RK, Páldi K, Paulraj S, Kao P, Lutzmayer S, Nodine MD (2020) Chromatin regulates expression of small RNAs to help maintain transposon methylome homeostasis in Arabidopsis. Genome Biol 21:251
Park J, Lee H, Han N et al (2018) Long non-coding RNA ChRO1 facilitates ATRX/DAXX-dependent H3.3 deposition for transcription-associated heterochromatin reorganization. Nucleic Acids Res 46:11759–11775
Paulsen T, Kumar P, Koseoglu MM, Dutta A (2018) Discoveries of extrachromosomal circles of DNA in normal and tumor cells. Trends Genet 34:270–278
Pavlek M, Gelfand Y, Plohl M, Meštrović N (2015) Genome-wide analysis of tandem repeats in Tribolium castaneum genome reveals abundant and highly dynamic tandem repeat families with satellite DNA features in euchromatic chromosomal arms. DNA Res 22:387–401
Percharde M, Lin CJ, Yin Y, Guan J, Peixoto GA, Bulut-Karslioglu A, Biechele S, Huang B, Shen X, Ramalho-Santos M (2018) A LINE1-nucleolin partnership regulates early development and ESC identity. Cell 174:391–405
Pezer Ž, Ugarković Ð (2008) RNA Pol II promotes transcription of centromeric satellite DNA in beetles. PLoS One 3:e1594
Pezer Z, Ugarković Đ (2009) Transcription of pericentromeric heterochromatin in beetles – satellite DNAs as active regulatory elements. Cytogenet Genome Res 124:268–276
Pezer Ž, Ugarković Đ (2012) Satellite DNA-associated siRNAs as mediators of heat shock response in insects. RNA Biol 9:587–595
Pezer Ž, Brajković J, Feliciello I, Ugarković Đ (2012) Satellite DNA-mediated effects on genome regulation. Genome Dyn 7:153–169
Piacentini L, Fanti L, Specchia V, Bozzetti MP, Berloco M, Palumbo G, Pimpinelli S (2014) Transposons, environmental changes, and heritable induced phenotypic variability. Chromosoma 123:345–354
Pita S, Panzera F, Mora P, Vela J, Cuadrado Á, Sánchez A, Palomeque T, Lorite P (2017) Comparative repeatome analysis on Triatoma infestans Andean and non-Andean lineages, main vector of Chagas disease. PLoS One 12:e0181635
Probst AV, Okamoto I, Casanova M, El Marjou F, Le Baccon P, Almouzni G (2010) A strand-specific burst in transcription of pericentric satellites is required for chromocenter formation and early mouse development. Dev Cell 19:625–638
Rajshekar S, Yao J, Arnold PK, Payne SG, Zhang Y, Bowman TV, Schmitz RJ, Edwards JR, Goll M (2018) Pericentromeric hypomethylation elicits an interferon response in an animal model of ICF syndrome. eLife 7:e39658
Ratner VA, Zabanov SA, Kolesnikova OV, Vasilyeva LA (1992) Induction of the mobile genetic element Dm-412 transpositions in the Drosophila genome by heat shock treatment. Proc Natl Acad Sci USA 89:5650–5654
Rebollo R, Karimi MM, Bilenky M, Gagnier L, Miceli-Royer K, Zhang Y, Goyal P, Keane TM, Jones S, Hirst M, Lorincz MC, Mager DL (2011) Retrotransposon-induced heterochromatin spreading in the mouse revealed by insertional polymorphisms. PLoS Genet 7:e1002301
Rizzi N, Denegri M, Chiodi I, Corioni M, Valgardsdottir R, Cobianchi F, Riva S, Biamonti G (2004) Transcriptional activation of a constitutive heterochromatic domain of the human genome in response to heat shock. Mol Biol Cell 15:543–551
Rosic S, Kohler F, Erhardt S (2014) Repetitive centromeric satellite RNA is essential for kinetochore formation and cell division. J Cell Biol 207:335–349
Rudert F, Bronner S, Garnier JM, Dollé P (1995) Transcripts from opposite strands of gamma satellite DNA are differentially expressed during mouse development. Mamm Genome 6:76–83
Ruiz-Ruano FJ, López-León MD, Cabrero J, Camacho JP (2016) High-throughput analysis of the satellitome illuminates satellite DNA evolution. Sci Rep 6:28333
Saksouk N, Simboeck E, Déjardin J (2015) Constitutive heterochromatin formation and transcription in mammals. Epigenetics Chromatin 8:3
Satović E, Vojvoda Zeljko T, Luchetti A, Mantovani B, Plohl M (2016) Adjacent sequences disclose potential for intra-genomic dispersal of satellite DNA repeats and suggest a complex network with transposable elements. BMC Genomics 17:997
Scott HS, Kudoh J, Wattenhofer M, Shibuya K, Berry A, Chrast R, Guipponi M, Wang J, Kawasaki K, Asakawa S, Minoshima S, Younus F, Mehdi SQ, Radhakrishna U, Papasavvas MP, Gehrig C, Rossier C, Korostishevsky M, Gal A, Shimizu N, Bonne-Tamir B, Antonarakis SE (2001) Insertion of beta-satellite repeats identifies a transmembrane protease causing both congenital and childhood onset autosomal recessive deafness. Nat Genet 27:59–63
Scott EC, Gardner EJ, Masood A, Chuang NT, Vertino PM, Devine SE (2016) A hot L1 retrotransposon evades somatic repression and initiates human colorectal cancer. Genome Res 26:745–755
Seong KH, Li D, Shimizu H, Nakamura R, Ishii S (2011) Inheritance of stress-induced, ATF-2-dependent epigenetic change. Cell 145:1049–1061
Sermek A, Feliciello I, Ugarković Đ (2021) Distinct regulation of the expression of satellite DNAs in the beetle Tribolium castaneum. Int J Mol Sci 22:296
Shaul O (2017) How introns enhance gene expression. Int J Biochem Cell Biol 91:145–155
Sienski G, Dönertas D, Brennecke J (2012) Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151:964–980
Sproul JS, Khost DE, Eickbush DG, Negm S, Wei X, Wong I, Larracuente AM (2020) Dynamic evolution of Euchromatic satellites on the X chromosome in Drosophila melanogaster and the simulans clade. Mol Biol Evol 37:2241–2256
Sun L, Jing Y, Liu X, Li Q, Xue Z, Cheng Z, Wang D, He H, Qian W (2020) Heat stress-induced transposon activation correlates with 3D chromatin organization rearrangement in Arabidopsis. Nat Commun 11:1886
Tanne A, Muniz LR, Puzio-Kuter A, Leonova KI, Gudkov AV, Ting DT, Monasson R, Cocco S, Levine AJ, Bhardwaj N, Greenbaum BD (2015) Distinguishing the immunostimulatory properties of noncoding RNAs expressed in cancer cells. Proc Natl Acad Sci USA 112:15154–15159
Thomas J, Phillips CD, Baker RJ, Pritham EJ (2014) Rolling-circle transposons catalyze genomic innovation in a mammalian lineage. Genome Biol Evol 6:2595–2610
Tilman G, Arnoult N, Lenglez S, Van Beneden A, Loriot A, De Smet C, Decottignies A (2012) Cancer-linked satellite 2 DNA hypomethylation does not regulate Sat2 non-coding RNA expression and is initiated by heat shock pathway activation. Epigenetics 7:903–913
Ting DT, Lipson D, Paul S, Brannigan BW, Akhavanfard S, Coffman EJ, Contino G, Deshpande V, Iafrate AJ, Letovsky S, Rivera MN, Bardeesy N, Maheswaran S, Haber DA (2011) Aberrant overexpression of satellite repeats in pancreatic and other epithelial cancers. Science 331:593–596
Ugarković Đ (2005) Functional elements residing within satellite DNAs. EMBO Rep 6:1035–1103
Ugarković Đ, Plohl M (2002) Variation in satellite DNA profiles – causes and effects. EMBO J 21:5955–5959
Valgardsdottir R, Chiodi I, Giordano M, Rossi A, Bazzini S, Ghigna C, Riva S, Biamonti G (2008) Transcription of satellite III non-coding RNAs is a general stress response in human cells. Nucleic Acids Res 36:423–434
van de Lagemaat LN, Landry JR, Mager DL, Medstrand P (2003) Transposable elements in mammals promote regulatory variation and diversification of genes with specialized functions. Trends Genet 19:530–536
Vlahović I, Glunčić M, Rosandić M, Ugarković Đ, Paar V (2017) Regular higher order repeat structures in beetle Tribolium castaneum genome. Genome Biol Evol 9:2668–2680
Vojvoda Zeljko T, Pavlek M, Meštrović N, Plohl M (2020) Satellite DNA-like repeats are dispersed throughout the genome of the Pacific oyster Crassostrea gigas carried by Helentron non-autonomous mobile elements. Sci Rep 10:15107
Volpe TA, Kidner C, Hall IM, Teng G, Grewal SI, Martienssen RA (2002) Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297:1833–1837
Vourc'h C, Biamonti G (2011) Transcription of satellite DNAs in mammals. Prog Mol Subcell Biol 51:95–118
Wang LC, Wu JR, Chang WL, Yeh CH, Ke YT, Lu CA, Wu SJ (2013) Arabidopsis HIT4 encodes a novel chromocentre-localized protein involved in the heat reactivation of transcriptionally silent loci and is essential for heat tolerance in plants. J Exp Bot 64:1689–1701
Wang J, Jia ST, Jia S (2016) New insight into the regulation of heterochromatin. Trends Genet 32:284–294
Wei KH, Grenier JK, Barbash DA, Clark AG (2014) Correlated variation and population differentiation in satellite DNA abundance among lines of Drosophila melanogaster. Proc Natl Acad Sci USA 111:18793–18798
Wong LH, Brettingham-Moore KH, Chan L, Quach JM, Anderson MA, Northrop EL, Hannan R, Saffery R, Shaw ML, Williams E, Choo KH (2007) Centromere RNA is a key component for the assembly of nucleoproteins at the nucleolus and centromere. Genome Res 17:1146–1160
Wylie A, Jones AE, D'Brot A, Lu WJ, Kurtz P, Moran JV, Rakheja D, Chen KS, Hammer RE, Comerford SA, Amatruda JF, Abrams JM (2016) p53 genes function to restrain mobile elements. Genes Dev 30:64–77
Zeller P, Gasser SM (2017) The importance of satellite sequence repression for genome stability. Cold Spring Harb Symp Quant Biol 82:15–24
Zhu Q, Pao GM, Huynh AM, Suh H, Tonnu N, Nederlof PM, Gage FH, Verma IM (2011) BRCA1 tumour suppression occurs via heterochromatin-mediated silencing. Nature 477:179–184
Zhu Q, Hoong N, Aslanian A, Hara T, Benner C, Heinz S, Miga KH, Ke E, Verma S, Soroczynski J, Yates JR 3rd, Hunter T, Verma IM (2018) Heterochromatin-encoded satellite RNAs induce breast cancer. Mol Cell 70:842–853
Acknowledgments
This work was supported by Croatian Science Foundation grants IP-2014-09-3733 and IP-2019-04-6915, by donation of “Adris” Foundation to Đ. Ugarković, and by the Italian Ministry of Education, University and Research (MIUR), fund for Investments on Basic Research (FIRB) and the International Staff Mobility Program of University of Naples Federico II to I. Feliciello. We thank Mary Sopta for the critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Feliciello, I., Pezer, Ž., Sermek, A., Bruvo Mađarić, B., Ljubić, S., Ugarković, Đ. (2021). Satellite DNA-Mediated Gene Expression Regulation: Physiological and Evolutionary Implication. In: Ugarković, Ð. (eds) Satellite DNAs in Physiology and Evolution. Progress in Molecular and Subcellular Biology, vol 60. Springer, Cham. https://doi.org/10.1007/978-3-030-74889-0_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-74889-0_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-74888-3
Online ISBN: 978-3-030-74889-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)