Abstract
Patient-specific implants provide important advantages for patients and medical professionals. The state of the art of cranioplasty implant production is based on the bone structure reconstruction and use of patient’s own anatomical information for filling the bone defect. The present work proposes a two-dimensional investigation of which dataset results in the closest polynomial regression to a gold standard structure combining points of the bone defect region and points of the healthy contralateral skull hemisphere. The similarity measures used to compare datasets are the root mean square error (RMSE) and the Hausdorff distance. The objective is to use the most successful dataset in future development and testing of a semi-automatic methodology for cranial prosthesis modeling. The present methodology was implemented in Python scripts and uses five series of skull computed tomography images to generate phantoms with small, medium and large bone defects. Results from statistical tests and observations made from the mean RMSE and mean Hausdorff distance allow to determine that the dataset formed by the phantom contour points twice and the mirrored contour points is the one that significantly increases the similarity measures.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Bone reconstruction of cranial areas represents a challenge for neurosurgeons and plastic surgeons because of the complex anatomical shapes, unique characteristics of each bone defect, the presence of vital adjacent anatomical structures and the high risk of infection to which the patient is subjected. This procedure is necessary due to trauma, tumor resection, decompressive craniotomy, infections or deformities. It aims to aesthetically restore the affected area and rehabilitate its function, promoting protection to the brain and other vital structures, and the restoration of cerebrospinal fluid dynamics and cerebral blood flow. Consequently, the patient may have his/her self-esteem restored and his/her social life normalized [1,2,3].
The state of the art of cranioplasty implant production process is based on the reconstruction of the bone structure of the patient’s skull from computed tomography images, generating a three-dimensional model. The missing bone segment is virtually modeled based on the patient’s own anatomical information with the ideal geometry for filling the bone defect. The resulting implant can then be produced by subtractive or additive manufacturing [2].
A patient-specific implant provides advantages for patients and medical professionals by reducing surgery time, prosthesis adaptation trauma, patient recovery time, and increasing surgery and implant success rates [4]. Case studies [5, 6] and studies with patient groups [3, 7,8,9] demonstrate the efficiency of the technique.
Currently the three-dimensional modeling process of patient-specific implants is characterized by the use of commercial and user-dependent software, which makes it more expensive and affects its standardization and repeatability [10, 11].
In this context, it is appropriate to propose semi-automatic and automatic methodologies that contribute to this process, automating their critical steps, as performed in the studies by [1, 11, 12]. These studies use information from the morphologically healthy contralateral skull hemisphere in the generation of the segment corresponding to the bone defect.
The present work proposes a two-dimensional investigation of which dataset results in the closest polynomial regression to a gold standard structure combining points of the bone defect region and points of the healthy contralateral hemisphere. Such analysis is not addressed in the cited reference studies and can improve semi-automatic and automatic methodologies results.
2 Materials
Two series of skull computed tomography images were obtained from the Patient Contributed Image Repository website [13] (Table 1). Three other series of skull computed tomography images were obtained from the public database The Cancer Imaging Archive (TCIA) [14], in the HNSCC-3DCT-RT collection [15] (Table 1). Images in DICOM format were unidentified and anonymized prior to making them available to the public domain in both databases. The methodology consists of Python scripts (version 2.7.13, Anaconda 4.4.0 package), using the libraries numpy, skimage and scipy.spatial.
3 Methodology
3.1 Information Extraction from DICOM Images
The identification of the points that delimit the patient’s bone structure (called bone contours) is performed in each image by the measure.find_contours function [16], which is based on the marching squares algorithm (two-dimensional approach of marching cubes algorithm [17]). The function uses a fixed threshold value to determine the interface points between bone and adjacent tissues. Each bone contour is defined as a list of points with x- and y-coordinates in pixels for each slice.
Bone contours of interest are defined as those whose Euclidean distance between the center of bone contour (defined by the arithmetic mean between its points) and the center of image is at most 10% of the smallest image dimension. This criterion is based on the fact that the contour of the skull bone is approximately centered on the image.
In tomographic images of a healthy skull, two bony contours of interest are expected: an outer and an inner contour approximately concentric, being the outer contour longer than the inner one. The contours are structured in a multidimensional array and associated with their image number. Those are defined as the gold standard contours.
It is noteworthy that the x and y coordinates of the obtained contours are organized in such a way that the main axis of the sagittal plane of the patient’s skull is parallel to the x axis, allowing the treatment of each bone contour hemisphere data as a function.
3.2 Phantom Generation
Images above the orbit region and below the skull apex are selected from the image series of the five patients listed in Table I. Each patient’s image series have different image spacing and the skulls have anatomical variations, so the number of images selected to compose each phantom also differ. The number of images selected for patients P1 to P5 are, respectively, 63, 121, 31, 32, 32 images.
From the selected images the bone contours are then obtained in each image according to section A. The threshold value used for all patients is 200 H.U.
Three phantoms are produced for each patient by selecting points on the outer bone contours to simulate large (L), medium (M), and small (S) bone defects, resulting in fifteen phantoms.
For each patient, a point on the series central image (x, y, image number) approximately centered on the left hemisphere of the skull for P1 and P2 and in the right hemisphere for P3, P4, and P5 is defined. This point is set as the center of an ellipsoid with parameters a, b and c referring respectively to the x, y, and z-axes. Ellipsoids with parameters (18, 24, 12), (27, 36, 18) and (36, 48, 24) are used to generate small, medium, and large bone defects, respectively. Each point on the contour of each image is converted to the patient’s coordinate system and checked for externality to the ellipsoid volume. If so, it is maintained.
3.3 Mirroring of Contour Points
In order to use the information from the morphologically healthy contralateral skull hemisphere it is necessary to align it with the bone defect region. Considering that the skull is approximately symmetric, a mirroring operation is convenient. It is necessary to define a contour symmetry axis to perform it. This axis is defined per image by a horizontal vector and a reference point (x\(_{ref}\), y\(_{ref}\)) calculated as the mean between the maximum and minimum values of the x and y-coordinates, respectively, of the gold standard contour points.
The mirroring operation is performed on the phantom contours. The first step consists in reversing the y-coordinate signal of each contour point, mirroring them with respect to the x-axis. Then the points are translated by adding twice y\(_{ref}\) to the y-coordinate of each point in order to spatially overlap them with the original phantom contour points. The resulting contour is called mirrored contour.
3.4 Determination of Contour Points in the Hemisphere of Interest
Since the simulated bone defects in the present study are all lateral, it is convenient to determine the hemisphere that contains the bone defect to process only the contour points that are in the hemisphere of interest. This measure contributes to the computational efficiency of the methodology.
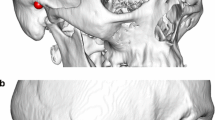
Images 13, 33 and 53 of patient 1 large bone defect phantom (left to right, respectively) (a); Images, 10, 20, 30 of patient 3 medium bone defect phantom (left to right, respectively) (b). Phantom contour points in blue, mirrored contour points in green, reference point represented by the magenta “x” and contour symmetry axis represented by the red dotted line. The bone defect region is highlighted by the black vertical bars, when present
The gold standard contour points, the phantom contour points and the mirrored contour points that have a y-coordinate greater than y\(_{ref}\) for patients 1 and 2 and smaller than y\(_{ref}\) for patients of 3 to 5 are selected for each image.
3.5 Determination of the Polynomial Regression Region in X-Axis
The set of reference points included in the regression affects the morphology of the resulting curve. Therefore, it is important to define reasonable criteria for the region that provides reference points.
This region is determined for each image from the organization of phantom contour points by the x-coordinate in ascending order. The largest interval between the x-coordinates of subsequent points is found. The points whose x-coordinates are these two values are established as the starting (x\(_{ini}\), y\(_{ini}\)) and ending (x\(_{end}\), y\(_{end}\)) points of bone defect.
The starting point of the evaluation region (x\(_{ini\_eval}\), y\(_{ini\_eval}\)) is set as the point with x-coordinate smaller than x\(_{ini}\) and the closest to (x\(_{ini}\), y\(_{ini}\)) with Euclidean distance greater than or equal to 10 mm. The endpoint of the evaluation region (x\(_{end\_eval}\), y\(_{end\_eval}\)) is established as the point with x-coordinate greater than x\(_{end}\) and the closest to (x\(_{end}\), y\(_{end}\)) with Euclidean distance greater than or equal to 10 mm. The prosthesis must overlap with the edge of the bone defect for support, justifying the 10 mm added on each side of the bone defect.
3.6 Dataset for Polynomial Regression
The objective of the present study is to verify which dataset, by combining phantom contour points and mirrored contour points, improves similarity measures between the gold standard contour and the estimated contour. The latter is a polynomial obtained by a fourth-degree two-dimensional polynomial regression using the referred dataset.
The polynomial coefficients are obtained by the numpy.polyfit function [18] (which performs a fourth-degree least squares polynomial regression). It is evaluated according to the evaluation region established in section E using the numpy.polyval function [19]. The x-axis resolution of the evaluation region is 0.2 mm.
The similarity measures used are the root mean square error (RMSE) [20] and the Hausdorff distance (H) [21].
Tests A through D are performed using different datasets as input points to the polynomial fit calculation: only the phantom contour points (Test A); the phantom contour points and the mirrored contour points (Test B); the phantom contour points twice and the mirrored contour points (Test C); the mirrored contour points twice and the phantom contour points (Test D). The use of the same dataset twice aims to influence more strongly the curvature of the resulting curve.
Tests A through D are performed for each bone contour comprising the phantom bone defect. The criterion for identifying the contour with bone defect is a distance between x\(_{ini}\) and x\(_{end}\) greater than 2 mm. Smaller distances indicate that the points are neighbors, with no spacing and therefore no defect between them.
The bone defect starting (x\(_{ini}\), y\(_{ini}\)) and ending (x\(_{end}\), y\(_{end}\)) points delimit the region for calculating the similarity measures. The evaluation region is not appropriate because this set of points comprise the bone defect edges, which are present in the gold standard and therefore in the phantom identically. This would artificially reduce the error between the two datasets.
For each phantom the mean and the standard deviation of each similarity measure are also calculated.
4 Results and Discussion
Figure 1 presents examples of the phantoms, demonstrating the variety of bone defects sizes and series characteristics due to the different number of images and different spacing between images.
Test A (a) to D (d) results for the evaluation region of image 07 of patient 2 small bone defect phantom: in the images on the left, gold standard contour points are in blue, the mirrored contour points are in green and estimated contour points are in red; the image on the right shows the phantom bone contour points in blue and the estimated contour points in red
Figure 2 shows the overlap of phantom contours and the respective mirrored contours in different skull regions for two patients. Good visual overlap of the contours can be observed in the whole skull. This is observed in all phantoms generated despite the anatomical differences between them, indicating convenient positioning of the data for the tests.
The graphical results obtained indicate that the polynomial regression is satisfactorily filling the defect with the desired shape. An example is shown in Fig. 3 for image 7 of patient 2 small bone defect phantom.
Figures 4 and 5 graphically show the results for mean RMSE with respective standard deviation (both in millimeters) and mean Hausdorff distance with respective standard deviation (both in millimeters), respectively, for each phantom and test. The mean similarity measures were calculated between the gold standard contour and the estimated contour in the bone defect region considering all contours that comprise the bone defect of each phantom for each test. Table 2 presents the number of contours considered in the calculation of similarity measures for each phantom.
The mean RMSE and the mean Hausdorff distance varied considerably between patients and between defect sizes in the same patient due to anatomical variations.
The mean RMSE and mean Hausdorff distance increase as the size of the defect increases, which is expected, because the curvature of the phantom contour becomes sharper and potentially more complex, not necessarily corresponding to the characteristic curvature resulting from the fourth-degree polynomial regression. In these cases, the RMSE and Hausdorff distance are decreased by using the mirrored contour, which provides this curvature information.
This fact is demonstrated by the increase of error for test A (which only uses information from the edges of bone defect in the regression) in relation to the other tests as bone defect size increases. In small defects, for 3/5 of the patients the mean RMSE in test A is the smallest among the tests. In medium defects, this occurs for 2/5 of patients. In large defects, for only 1/5 of the patients this is the case. Considering the Hausdorff distance, the cited proportions are similar, respectively, 4/5, 2/5, 1/5.
Considering only tests B, C, and D, in which mirrored bone contour information is used in the regression, the mean RMSE in test C is the lowest in 4/5 of patients with small defect and all five patients with medium and large defect. The cited proportions are identical regarding the Hausdorff distance.
From these observations the following null hypothesis is formulated: there is no statistically significant difference between the RMSE/ Hausdorff distance obtained for test A and for each other test (B, C, D) considering all phantoms.
Since the datasets are paired, it is verified and confirmed that the datasets resulting from the difference between the pairs test A—test B, test A—test C, test A—test D for both similarity measures have approximately normal distributions. That condition allows the calculation of p-values using the t-test (Table 3). If the p-value is smaller than 0.05, the null hypothesis is rejected. The null hypothesis was rejected for all test pairs regarding RMSE values and for test pairs A/B and A/C regarding the Hausdorff distance, indicating that there is significant difference between these test results.
5 Conclusion and Further Work
Combining the statistical tests results and the observations made from the mean RMSE and mean Hausdorff distance, we conclude that the dataset formed by the phantom contour points twice and the mirrored contour points (Test C) is the one that significantly improves the similarity measures. It will be used for further development and testing of a three-dimensional polynomial regression approach for semiautomatic generation of a cranial prosthesis polygonal mesh that complements certain bone defect.
References
Hieu LC et al (2002) Design and manufacturing of cranioplasty implants by 3-axis CNC milling. Technol Health Care 10:413–423
Parthasarathy J (2014) 3D modeling, custom implants and its future perspectives in craniofacial surgery. Ann Maxillofac Surg 4:9–18
Park E et al (2016) Cranioplasty enhanced by three-dimensional printing: custom-made three-dimensional-printed titanium implants for skull defects. J Craniofac Surg 27:943–949
Ventola CL (2014) Medical applications for 3D printing: current and projected uses. Pharm Ther 39:704–711
Jardini A et al (2014) Customised titanium implant fabricated in additive manufacturing for craniomaxillofacial surgery. Virtual Phys Prototyp 9:115–125
Jardini A et al (2014) Cranial reconstruction: 3D biomodel and custom-built implant created using additive manufacturing. J Cranio Maxillofac Surg 42:1877–1884
Cabraja M, Klein M, Lehman TN (2009) Long-term results following titanium cranioplasty of large skull defects. Neurosurg Focus 26:E10
Kim BJ, Hong KS, Park KJ, Park DH, Chung YG, Kang SH (2012) Customized cranioplasty implants using three-dimensional printers and polymethyl-methacrylate casting. J Korean Neurosurg Soci 52:541–546
Rotaru H et al (2012) Cranioplasty with custom made implants: analyzing the cases of 10 patients. J Oral Maxillofac Surg 70:e169–e170
Wehmöller M, Eufinger H, Kruse D, Massberg W (1995) CAD by processing of computed tomography data and CAM of individually designed prostheses. Int J Oral Maxillofac Surg 24:90–97
Gelaude F, Sloten JV, Lauwers B (2006) Automated design and production of cranioplasty plates: outer surface methodology, accuracies and a direct comparison to manual techniques. Comput Aid Des Appl 3:193–202
Hieu LC et al (2002) Design and manufacturing of personalized implants and standardized templates for cranioplasty applications. IEEE Int Conf Ind Technol 2:1025–1030
Patient contributed image repository. Available at: http://www.pcir.org/researchers/downloads_available.html. Accessed 30 Jan 2019
Clark K et al (2013) The cancer imaging archive (TCIA): maintaining and operating a public information repository. J Digital Imag 26:1045–1057
Brejarano T, Couto MDO, Mihaylov IB (2019) Head-and-neck squamous cell carcinoma patients with CT taken during pre-treatment, mid-treatment, and post-treatment (HNSCC-3DCT-RT). The Cancer Imaging Archive. Available at: https://doi.org/10.7937/K9/TCIA.2018.13upr2xf. Accessed 02 Apr 2019
Scikit-image documentation. Function measure.find\_contours. Available at: http://scikit-image.org/docs/0.8.0/api/skimage.measure.find_contours.html#find-contours. Accessed: 27 Apr 2018
Lorensen W, Cline EC (1987) Marching cubes: a high resolution 3D surface construction algorithm. In: SIGGRAPH 87 proceedings, vol 21, pp 163–170
NumPy v1.15 manual. Function numpy.polyfit. Available at: https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.polyfit.html. Accessed: 01 Apr 2019
NumPy v1.15 manual. Function numpy.polyval. Available at: https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.polyval.html#numpy.polyval. Accessed: 01 Apr 2019
Barnston A (1992) Correspondence among the correlation, RMSE, and Heidke verification measures; refinement of the Heidke score notes and correspondence. In: Climate analysis center
Gonçalves GA, Mitishita EA (2016) The use of Hausdorff distance as similarity measure in automatic systems for cartographic update. Bol Ciênc Geod 22:719–735
Acknowledgements
The results published here are in part based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brazil (CAPES)—Finance Code 001.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
The authors declare that they have no conflict of interest.
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this paper
Cite this paper
Garcia, M.G.M., Furuie, S.S. (2022). Regression Approach for Cranioplasty Modeling. In: Bastos-Filho, T.F., de Oliveira Caldeira, E.M., Frizera-Neto, A. (eds) XXVII Brazilian Congress on Biomedical Engineering. CBEB 2020. IFMBE Proceedings, vol 83. Springer, Cham. https://doi.org/10.1007/978-3-030-70601-2_223
Download citation
DOI: https://doi.org/10.1007/978-3-030-70601-2_223
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-70600-5
Online ISBN: 978-3-030-70601-2
eBook Packages: EngineeringEngineering (R0)