Abstract
The naked mole-rat (Heterocephalus glaber) is the longest-lived rodent, with a maximal reported lifespan of 37 years. In addition to its long lifespan – which is much greater than predicted based on its small body size (longevity quotient of ~4.2) – naked mole-rats are also remarkably healthy well into old age. This is reflected in a striking resistance to tumorigenesis and minimal declines in cardiovascular, neurological and reproductive function in older animals. Over the past two decades, researchers have been investigating the molecular mechanisms regulating the extended life- and health- span of this animal, and since the sequencing and assembly of the naked mole-rat genome in 2011, progress has been rapid. Here, we summarize findings from published studies exploring the unique molecular biology of the naked mole-rat, with a focus on mechanisms and pathways contributing to genome stability and maintenance of proteostasis during aging. We also present new data from our laboratory relevant to the topic and discuss our findings in the context of the published literature.

Photo Credit: Ben Passarelli
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
11.1 Introduction
At 37 years (Lee et al. 2020) – and counting – the naked mole-rat (Heterocephalus glaber; ~45 g) is the longest-lived rodent known (Buffenstein 2005). This species exceeds the longevity of the previous rodent record holder, the much larger (11 kg) African porcupine, Hystrix cristata by more than 10 years, and lives an order of magnitude longer than similar-sized mice (AnAge: (Tacutu et al. 2018)). Based upon the most recent allometric equations, the naked mole-rat exhibits the highest documented longevity quotient (LQ; the ratio of the observed maximal lifespan /predicted maximal lifespan ; LQ = currently ~4.2Footnote 1) for a non-volant mammal (de Magalhaes et al. 2007). Naked mole-rats are subterranean small rodents, native to the Horn of Africa. They are unique among mammals, being one of two eusocial species in which breeding is restricted to one female (‘queen’) and 1–3 males per colony (Jarvis 1981). The majority of the non-breeding subordinates never leave their natal colony and perform many of the maintenance tasks associated with life below ground in arid regions where food is scarce and patchily distributed. Like other eusocial animals, such as wasps and bees, dominant breeders exhibit lower risk of dying than subordinate naked mole-rats, although even the subordinate animals are remarkably long-lived compared to other rodents of comparable size (Ruby et al. 2018).
The naked mole-rat is an attractive, naturally-evolved model to study the molecular mechanisms protecting against aging and age-related diseases, showing negligible physiological decline with advancing age (Buffenstein 2008). Unlike humans that already show declines of cardiac function albeit in the absence of pathology once they are in their thirties (25% maximum lifespan), naked mole-rats maintain cardiac function well into their third decade ~80% maximum lifespan (Grimes et al. 2014), and, strikingly, exhibit few signs of age-associated cardiac pathologies (Buffenstein and Craft 2021). Additionally, these animals are remarkably resistant to tumors – both natural and experimentally induced – with less than 10 observed cases from thousands of necropsies over a 40 year period (Liang et al. 2010; Delaney et al. 2016; Taylor et al. 2017; Seluanov et al. 2018; Delaney et al. 2021). Understanding the molecular mechanisms employed to attenuate physiological declines during aging and protect against tumorigenesis and other chronic diseases could be beneficial in developing therapeutic strategies to promote healthy aging in humans and other animals. These processes include transcriptional, post-transcriptional, translational, and post-translational events, knowledge of which is likely to facilitate a better understanding of cytoprotective mechanisms pivotal to the aging process.
In this review, we summarize some of the unusual molecular biology insights gained from comparative studies using the naked mole-rat that may be pertinent to healthy aging. Additionally, we share some novel and unpublished data from our laboratory, through which we aim to highlight some of the naked mole-rat’s unique features associated with maintaining cellular homeostasis i.e. a stable, optimal internal environment. This chapter is broadly divided into three sections: (a) naked mole-rat genomic stability; (b) insights gleaned from the transcriptome of the naked mole-rat; and (c) maintenance of a stable proteome throughout the long life of this animal. The bulk of this review will deal with the proteome as there is a paucity of genomic and transcriptomic studies on this animal, clearly highlighting pivotal gaps in our knowledge that need to be addressed in the future. Indeed, most of the insights into mechanisms pertinent to the unique biology of naked mole-rats were obtained from protein-based investigations (Fig. 11.1).
Hallmarks of genome and proteome stability in naked mole-rats. Naked mole-rats show negligible aging phenotypes even in their third decade of life as a result of several mechanisms that allow them to maintain healthy, functional genomes and proteomes. These mechanisms, indicated in the accompanying schematic, include elements that are known to be protective in other species as well as some that are unique to this species. Although largely unexplored, transcriptomic evidence points to minimal changes in mRNA abundance between young and old naked mole-rats, suggesting these animals may also have unique mechanisms to maintain a stable transcriptome during aging. Solid green and red arrows represent established/confirmed findings in naked mole-rats; pale red arrows and green boxes with dashed borders indicate our hypotheses based on published observations and our unpublished data; ‘?’ indicates pathways that need further investigation in naked mole-rats
11.2 Maintenance of Genome Stability During Aging
Published genome assemblies and karyotyping data reveal that naked mole-rats have 30 pairs of chromosomes (2n = 58XY) and their genome comprises about 22,561 protein-coding genes (George 1979; Kim et al. 2011; Keane et al. 2014). Of these, approximately 98% could be at least partially annotated based on homology with either human and/or mouse protein coding genes (Kim et al. 2011). In addition, 1779 non-coding sequences have been annotated in the naked mole-rat genome (Keane et al. 2014), yet few studies have explored their functional importance in this species. Nevertheless, the sequencing and assembly of the naked mole-rat genome has, without a doubt, paved the way for comparative transcriptomics and biochemical experiments to discover unique characteristics of this species that might contribute to its extended lifespan , negligible senescence and resistance to cancer.
Genome instability, epigenetic alterations and telomere attrition have long been considered some of the universal hallmarks of aging (Lopez-Otin et al. 2013). Remarkably, accumulating evidence indicates that naked mole-rats maintain genome stability throughout their long lifespan . In this section, we discuss several mechanisms that may allow these animals to maintain a stable genome (Fig. 11.1), including slow organismal growth, tightly controlled cell proliferation rates, and elevated DNA repair pathways.
11.2.1 Unique Features of Genome Maintenance and Cell Division in Naked Mole-Rats
11.2.1.1 DNA Repair and Genome Stability
Naked mole-rats are thought to exhibit a lower mutation rate than observed in mice. This premise is based upon the observation of fewer germline nucleotide substitutions in naked mole-rats (MacRae et al. 2015b) and is consistent with the finding that ~34 DNA repair genes are expressed at higher levels in liver tissue from naked mole-rats in comparison to mice (MacRae et al. 2015a). Among these are genes involved in DNA damage sensing, non-homologous end joining (NHEJ), mismatch repair and homologous recombination repair. Although the expression of genes involved in base excision repair (BER) was found to be comparable between mouse and naked mole-rat in this study, subsequent studies by some of the same authors, as well as unpublished data from the Buffenstein laboratory, demonstrated higher activity of BER as well as nucleotide excision repair (NER) pathways in both liver tissue and primary fibroblasts isolated from naked mole-rats when compared to mice (Evdokimov et al. 2018) (Buffenstein, unpublished data).
Naked mole-rats exposed to 12 Gy irradiation showed less accrual of DNA damage compared to mice – this could be due to the fact that naked mole-rat DNA is more resistant to DNA damage or that these animals have enhanced DNA repair mechanisms. The surviving naked mole-rats from this study are still tumor-free, >10 years after exposure to radiation (Lewis and Buffenstein, unpublished observations). Collectively, these data suggest that the high levels and activity of DNA repair pathways in naked mole-rats play important roles in maintaining low mutagenesis rates, thereby possibly lowering the risk of cancer development.
The tumor suppressor protein p53 also plays an important role in the DNA repair process and in maintaining genome stability. Following DNA damage, p53 modulates chromatin accessibility, transcriptionally upregulates genes involved in NER and BER, and, importantly, induces cell cycle arrest in a p21-dependent manner (Williams and Schumacher 2016). Strikingly, naked mole-rats have very high levels of p53 protein (Lewis et al. 2012) that appears to be remarkably stable and long-lived: in embryonic fibroblasts the half-life of naked mole-rat p53 is 10X greater than that of mouse p53 (Deuker et al. 2020). It is likely that this “ultra-stable p53”, together with elevated levels of DNA repair proteins, play key roles in maintaining genome integrity as well as in preventing tumor formation in naked mole-rats.
Paradoxically, Andziak et al. demonstrated elevated baseline levels of 8-hydroxy-2′-deoxyguanosine (8-OHdG), a marker of oxidative damage to DNA, in kidney and liver tissue from young naked mole-rats when compared to physiologically age-matched mice (Andziak et al. 2006). Similarly, high levels of lipid peroxidation and protein carbonyls were also reported in young naked mole-rats. Unlike in mice, however, these did not accrue during the first 20 years of life, suggesting that naked mole-rats may have a greater tolerance threshold of damage, above which they initiate repair. Additionally, the membrane phospholipid composition of naked mole-rats is markedly different to mice, which could contribute to the observed differences in lipid oxidation levels. For example, naked mole-rat membranes contain 9X less docosahexaenoic acid (DHA) than mice (Hulbert et al. 2006; Mitchell et al. 2007). The presence of measurable levels of damage in cells from healthy animals might also function in actively maintaining various repair processes and thereby ‘priming’ these cells to respond more effectively to acute stressors, as is evident in some other species (Ristow and Schmeisser 2014). Indeed, one could postulate that constitutive maintenance of an active pool of proteins involved in DNA repair could allow for quick and timely repair of any subsequent damage. The observed high levels of oxidized lipids and proteins in naked mole-rats may be attributed to the comparatively hyperoxic laboratory environment that they are forced to contend with from birth. This is a marked contrast to the harsh, hypoxic, hypercapnic gaseous atmospheres below ground, in which they have evolved (Andziak and Buffenstein 2006). The upregulated repair pathways described above likely prevent excessive accumulation of DNA damage throughout their long lives in both the comparatively hyperoxic captive conditions and the highly variable oxidative atmospheres encountered in their natural subterranean habitat.
11.2.1.2 Cell Proliferation
In comparison to other rodents of similar size, naked mole-rats show extremely slow rates of growth and development both in utero and postnatally, continuing somatic growth for at least 1 year after sexual maturity, and brain development for even longer (Jarvis et al. 1991; O’Riain and Jarvis 1998; Orr et al. 2016). Similarly, primary skin fibroblasts isolated from adult naked mole-rats grow three-times slower than their mouse counterparts, with a population doubling time of ~3–4 days compared to ~1 day in mice (Miyawaki et al. 2016). Although this may be explained by the cooler conditions that naked mole-rat cells are routinely cultured at (32–33 °C, corresponding to their normal body temperature versus 37 °C, the normal growth temperature for mouse cells), even when grown at the same temperature as mouse cells (35 °C), growth rates remain slower than observed in mice (Liang et al. 2010) (Lewis, Narayan and Buffenstein, unpublished observations). At 37 °C, naked mole-rat cells initially divide, but subsequently undergo cell cycle arrest. Strikingly, they remain viable for at least 3 weeks at this high temperature following cell cycle arrest (Narayan and Buffenstein, unpublished observations; Fig. 11.2).
Naked mole-rat cells grow to confluence under optimal growth conditions. Like mouse primary skin fibroblasts (a), we find that naked mole-rat skin fibroblasts (b) also grow to confluence under optimal growth conditions. However, under suboptimal growth conditions or in the presence of environmental stressors, the cells undergo growth arrest while often remaining viable for extended periods (c; heat stress at 37 °C for 20 days)
We use the following media formulation: MEM (Thermo Fisher Scientific #11095) supplemented with 10% fetal bovine serum and 1X antibiotic-antimycotic mix (Thermo Fisher Scientific #15240062). Our naked mole-rat cells are cultured on Corning Biocoat collagen-I coated dishes (VWR #62405-617) and maintained at 32 °C, 5% CO2, 3% O2. Media is changed every 1−2 days
Little is known of the mechanisms employed by naked mole-rats to regulate cell cycle and growth rate. One potential factor that has generated considerable attention is the versatile naked mole-rat Ink4 locus. In addition to the cyclin-dependent kinase inhibitor p16INK4a and tumor suppressors p14ARF [p19ARF in mice] and p15INK4b, this locus also generates a fourth product in naked mole-rats that has not been observed in humans and mice. This unusual product is a hybrid of p15INK4b (exon 1) and p16INK4a (exons 2 and 3) that is referred to as pALTINK4a/b (Tian et al. 2015). Induction of pALTINK4a/b, unique to naked mole-rats, was observed in response to stressors such as ultraviolet irradiation and oncogenic stimuli and its expression reportedly allows naked mole-rat cells to arrest more efficiently in comparison to induction of p15INK4b or p16INK4a alone (Tian et al. 2015). The ability of naked mole-rats to induce growth arrest through the induction of an alternatively spliced p15/p16 transcript in addition to the canonical Ink4 locus transcripts may serve to provide an extra layer of cell cycle regulation.
pALTINK4a/b-mediated growth arrest has been attributed to an unusual form of contact inhibition, termed ‘early contact inhibition’ (ECI), wherein cells stop proliferating at relatively low densities (Seluanov et al. 2009). ECI may be due to the secretion of a sticky viscous substance, very high molecular weight hyaluronan, into the cell culture supernatant by naked mole-rat dermal fibroblasts and this is most obvious when media is not changed for several days (Tian et al. 2013). Our laboratory and others have been unable to independently confirm these important findings and attribute this to disparate experimental cell culture protocols. In contrast to those outlined by Seluanov et al. (2009), our approach yields cells proliferating to higher densities than observed in primary mouse cultures (Lewis , Narayan and Buffenstein, unpublished observations, Fig. 11.2). However, we have found that when cells encounter even mild stress (e.g., 21% oxygen, low concentrations of heavy metals, or high temperature) or suboptimal conditions, they stop growing regardless of confluence (Lewis et al. 2012). Further efforts should be directed towards determining the contributions of both ECI and pALTINK4a/b to the observed cancer resistance of naked mole-rats.
Interestingly, studies examining induced pluripotent stem cells (iPSCs) in the naked mole-rat suggest that the Ink4 locus may also be implicated in cancer resistance of iPSCs (Miura et al. 2021). Elevated ARF expression in naked mole-rat iPSCs has been proposed to safeguard against teratoma formation in vivo. In addition, the authors report a disruptive frameshift mutation in embryonic stem (ES) cell-expressed Ras (ERAS) that introduces a premature stop codon and reduced ERAS expression and function in naked mole-rats – these unique features, in concert with other unknown mechanisms, likely make iPSCs from this species remarkably resilient to tumorigenesis (Miyawaki et al. 2016; Suarez-Cabrera et al. 2018).
11.2.1.3 Naked Mole-Rat Epigenome
Epigenetic changes including alterations in DNA methylation and chromatin modifications are widely accepted as hallmarks of aging (Lopez-Otin et al. 2013; Benayoun et al. 2015). These modifications not only sculpt DNA accessibility, thereby modulating downstream gene expression programs, but also function in maintaining genome stability (Benayoun et al. 2015; Dabin et al. 2016). A handful of reports have begun to investigate the epigenome of naked mole-rats and reveal that similar to other mammals, this appears to be subject to regulation (Tan et al. 2017; Lowe et al. 2018, 2020). Epigenetic remodeling driven by expression of reprogramming factors such as OCT4, SOX2, KLF4, and c-MYC has been well characterized. Strikingly, in contrast to mice, one report indicates that naked mole-rat cells are resistant to epigenetic remodeling at specific genes upon expression of classical reprogramming factors (Tan et al. 2017). Inactivation of the tumor suppressor retinoblastoma protein by expression of the SV40 Large T antigen seems necessary to promote epigenetic changes and allows reprograming to iPSCs (Tan et al. 2017). While an independent study described successful reprogramming of naked mole-rat cells by expression of the same classical reprogramming factors, both studies reveal that naked mole-rat iPSCs fail to, or are very inefficient at, forming teratomas in vivo (Miyawaki et al. 2016; Tan et al. 2017). Whether epigenetic mechanisms are at play in preventing tumorigenesis in this context remains to be addressed. Intriguingly, a comparison of histone post-translational modifications by mass spectrometry indicates that naked mole-rat fibroblasts exhibit higher levels of repressive histone 3 lysine 27 trimethylation (H3K27me3) and lower levels of activating H3K27 acetylation, amongst several other epigenome differences, when compared to mouse (Tan et al. 2017). These findings indicate that naked mole-rats harbor differences in both the intensity and patterns of chromatin modifications that may in turn play an important role in epigenetic regulation of gene expression in this species.
In addition to histone modifications, epigenetic changes during aging extend to DNA modifications. During mammalian aging, cells undergo global DNA hypomethylation and also exhibit site-specific DNA hypermethylation primarily at promoter CpG sites of genes involved in differentiation and development (Hannum et al. 2013; Horvath 2013; Benayoun et al. 2015). Specific DNA methylation signatures are considered molecular biomarkers of aging due to their ability to predict chronological age with high confidence in a number of different species including humans, mice, dogs, and whales (Hannum et al. 2013; Horvath 2013; Polanowski et al. 2014; Stubbs et al. 2017; Thompson et al. 2017; Lowe et al. 2018). Interestingly, two studies reveal that DNA methylation-derived ‘epigenetic clocks’ exist in naked mole-rats and can be used to predict naked mole-rat chronological age (Lowe et al. 2018, 2020). In humans and naked mole-rats, the rates of change of age-associated DNA methylation are slower than in mouse, with only 30 aging-associated differentially methylated positions (aDMPs) identified in naked mole-rats, compared with >2800 in mice (Lowe et al. 2018). Using liver tissue from 24 naked mole-rats, 23 aDMP sites were selected to predict age, and when these were measured in liver tissue from a different set of animals aged between 43 and 1,196 weeks (<1 to 23 years), good correlation between the predicted age and actual age was observed (R = 0.88) (Lowe et al. 2020). The aDMP sites could also predict the age of naked mole-rats with high confidence when applied to skin tissue (Lowe et al. 2020). These fascinating studies clearly implicate epigenetic mechanisms of aging in naked mole-rats and highlight the necessity for future functional studies in this organism. In particular, it will be interesting to determine whether these seemingly unique features of the naked mole-rat epigenome, most notably, the observed differences in histone dynamics, play protective roles in genome stability.
11.2.1.4 Telomere Maintenance
Telomeres play a critical role in preserving genome stability by blocking nucleolytic degradation at chromosome ends in addition to inhibiting chromosome fusions and unnecessary DNA recombination and repair events (Blackburn et al. 2015). Telomere attrition is a hallmark of aging and is implicated in both genomic instability and tumorigenesis. In addition to the telomerase ribonucleoprotein complex that maintains telomere length, a telomere-protective protein complex known as shelterin also exists to protect chromosome ends (de Lange 2005). Telomerase activity in a number of different somatic tissues from naked mole-rats was reported to be similar to that of other similarly sized mammals (Seluanov et al. 2007). Additionally, telomere length in naked mole-rat skin fibroblasts cultured ex vivo was found to be in a similar range to that observed in humans, bats and deer mice (Buffenstein, unpublished data), all of which are much shorter than the long telomere lengths observed in inbred laboratory mice. Seluanov et al. report that unlike humans, naked mole-rat telomere length does not appear to change drastically in skin fibroblasts upon extended growth ex vivo i.e. between early and late passage cells (Seluanov et al. 2008), indicating that naked mole-rats may have distinct mechanisms to protect and maintain chromosome ends. In this regard, it is interesting to note that factors involved in telomere maintenance harbor distinct features in this species. For example, RNA-seq data for young (4 years old) and old (20 years old) naked mole-rats indicate that levels of telomerase reverse transcriptase (Tert) is unaltered with age (Kim et al. 2011). These findings are in contrast to studies in mice, where Tert mRNA expression was found to decline with advancing age (Zhang et al. 2018). Although the expression of other telomerase components such as the non-coding telomerase RNA component (Terc) were not reported in this study, some sequence variation has been reported in the naked mole-rat Terc gene, including a SOX17 transcription factor binding site in the promoter region that is not present in humans or guinea pig (Evfratov et al. 2014). Similarly, sequence variations within structural domains of Terc that may impact RNA secondary structure and function have also been reported in naked mole-rats (Evfratov et al. 2014). In addition, genes involved in telomere maintenance and end protection including Tep1 (component of the telomerase complex) and Terf1 (component of the shelterin complex), undergo positive selection in naked mole-rats (Kim et al. 2011). These differences in factors involved in telomere maintenance could represent distinct regulatory mechanisms in naked mole-rats compared to other species. Further investigations into how these factors and possibly additional chromosome end protection pathways promote naked mole-rat genome stability, protection from DNA damage, and extraordinary longevity remain to be addressed.
11.3 Maintenance of Post-transcriptional Gene Regulation During Aging
Post-transcriptional control of gene expression is fundamental to achieving a functional proteome and plays an integral role in maintaining cellular homeostasis (Mata et al. 2005). Indeed, accurate control of mRNA turnover, well-orchestrated pre-mRNA splicing, and translational control mechanisms are all critical for survival, underscored by observations that alterations in these pathways are associated with numerous diseases and aging (Abdelmohsen et al. 2008; Masuda et al. 2009; Smith-Vikos and Slack 2012; Borbolis and Syntichaki 2015; Wei et al. 2015; Steffen and Dillin 2016). Several studies in model systems ranging from worms to mice have demonstrated that the transcriptome is commonly dysregulated with age. In some instances, transcript abundance appears to be significantly more variable in older individuals, with genes from the same functional group often exhibiting expression changes in opposing directions (Southworth et al. 2009; Rangaraju et al. 2015). This phenomenon has been named ‘transcriptional drift’. Suppression of transcriptional drift in the nematode Caenorhabditis elegans gives rise to a more youthful transcriptomic profile and extends lifespan (Rangaraju et al. 2015).
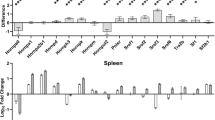
Surprisingly, unlike in other mammals, transcript levels in young (4 years old) and older (20 years old) naked mole-rats were found to be largely similar, particularly in the brain (Kim et al. 2011). In addition, and consistent with this finding, both young and old naked mole-rats were also found to have ~2-fold higher levels relative to young mice of splicing factors in spleen and brain tissue that are maintained throughout their lives. Importantly, unlike mice that show altered splicing patterns with age, alternative splicing of distinct transcripts implicated in aging were well-maintained through old age in naked mole-rats (Lee et al. 2020). Thus, from these limited investigations, it seems likely that naked mole-rats may not display some age-associated alterations at the post-transcriptional level that are commonly observed in other mammals. Systematic investigation into post-transcriptional control of gene expression in this species is warranted and should illuminate how such processes are maintained, as well as how these may contribute to the maintenance of a stable proteome well into old age.
Some additional evidence points towards unique aspects of post-transcriptional RNA processing in naked mole-rats and is evident from findings that these animals harbor a distinct ribosomal RNA (rRNA) cleavage pattern due to the presence of a so-called ‘hidden break’ within its genome. As the most abundant RNA in cells, non-coding rRNA is the backbone of ribosomes and is integral for nucleolar stability (Thomson et al. 2013). Following transcription from ribosome DNA (rDNA), a large precursor rRNA (pre-rRNA) molecule is extensively processed by hundreds of proteins and non-coding RNAs to yield 18S, 28S and 5.8S rRNA, leading to the biogenesis of mature ribosomes in the cytoplasm (Thomson et al. 2013). In naked mole-rats, an insertion of >200 nucleotides was identified within a highly conserved region of rDNA, which consequently led to fragmentation of the mature 28S rRNA into two distinct molecules in this species (Azpurua et al. 2013; Fang et al. 2014). Interestingly, hidden breaks are commonly found in rDNA of protostomes (Natsidis et al. 2019). The main outstanding questions on this topic are: (1) do these hidden breaks harbor any regulatory role(s) affecting rRNA processing or ribosome function; and (2) what cellular machinery is responsible for the recognition and splicing of this region to generate fragmented 28S rRNA. These will be discussed in the section “Unusual ribosome structure”.
While the naked mole-rat non-coding RNA transcriptome remains largely unexplored, one study has shed some light on long non-coding RNAs (lncRNAs). In this study, the authors report >4000 putative lncRNAs in the naked mole-rat and, consistent with other mammals, some of these appear to show tissue-specific expression patterns (Jiang et al. 2016). As our knowledge of the mammalian non-coding genome is rapidly expanding and accumulating evidence links non-coding RNAs to human health (Esteller 2011), future efforts to investigate the naked mole-rat non-coding RNA transcriptome are warranted. Without a doubt, insights into how these animals maintain a healthy, long, cancer-free life could be found by studying lncRNAs and other classes of non-coding RNAs that are unique to naked mole-rats. Additionally, as we enter a new frontier in the study of RNA modifications or the epitranscriptome and its impact on post-transcriptional regulation of gene expression (Frye et al. 2018; Zhang et al. 2019), it seems prudent to extend these studies to naked mole-rats in order to determine their contributions toward maintaining a stable transcriptome in this remarkable mammal.
11.4 Maintenance of a Functional, Stable Proteome During Aging
Loss of protein homeostasis (proteostasis) is observed in several human age-related diseases and “proteostasis collapse” shortens lifespan in several organisms, making it a hallmark of the aging process (Taylor and Dillin 2011; Lopez-Otin et al. 2013). As macromolecules responsible for carrying out most essential cellular processes, proteins must be present: (1) at specific times; (2) in optimal concentrations; (3) in the correctly folded state; and (4) in the correct sub-cellular compartment. This requires tight regulation at the levels of mRNA translation, post-synthesis folding, maintenance of the active conformation, and timely degradation. The machinery responsible for orchestrating these activities is called the ‘Proteostasis Network’ (PN), and its primary function is to maintain proteostasis through a functional, healthy proteome.
The PN comprises >2000 proteins, broadly divided into three categories: protein synthesis machinery, molecular chaperones, and protein degradation machinery (Klaips et al. 2018; Jayaraj et al. 2020). Components of the protein synthesis machinery include the ribosome , RNA-binding proteins, and numerous translation factors. Their primary job is to ensure mRNA transcripts are accurately and efficiently translated into proteins in a temporally and spatially restricted manner. Although some newly synthesized small polypeptides can fold into their native conformation directly on the ribosome , the vast majority of these require other PN components called chaperones to aid in their folding into functional conformations. These chaperones include, for example, various heat-shock proteins (HSPs) and their associated cofactors, as well as T-complex protein-1 ring complex or chaperonin containing TCP-1 (TRiC/CCT) proteins. Additionally, as protein synthesis is error prone, with approximately 1 in 104 amino acids incorrectly incorporated (Zaher and Green 2009), the ribosome-associated quality control (RQC) pathway plays a fundamental role in detecting faulty polypeptides which can be subsequently degraded by the ubiquitin-proteasome system (UPS) (Brandman and Hegde 2016). In addition to removal of nascent polypeptides containing misincorporated amino acids, the UPS also degrades misfolded proteins and maintains proteostasis through the timely degradation of enzymes and other signaling proteins. Although ~80% of all protein degradation is accomplished by the UPS, other PN components, including autophagy and various proteases, also play a critical role in protein degradation (Collins and Goldberg 2017).
What sets naked mole-rats apart from humans, mice and many other animals, are findings suggesting that there is, in fact, no age-dependent decline in the abundance or function of PN components examined in these animals. Indeed, molecular chaperones are maintained at high levels throughout the life of the animal, abundance as well as activity of the protein degradation machinery appears to be unchanged during aging, and, correspondingly, there is no build-up of polyubiquitinated proteins or oxidized proteins with age in naked mole-rats, unlike in other species (Fig. 11.3). In this section, we discuss insights gained, mainly using molecular biology approaches, into some mechanisms responsible for maintaining a functional, stable proteome in naked mole-rats during aging.
The cytosolic proteostasis network in naked mole-rats. A schematic representation of cytosolic proteostasis network components in naked mole-rats is shown. Notably, naked mole-rats have a cleaved 28S rRNA, which might contribute to the observed increase in translational accuracy in this species. Based on our unpublished findings and observations in the literature, we speculate that protein synthesis rates in naked mole-rats are lower than in mice, which, in conjunction with high levels of chaperones such as HSP70, would give rise to fewer misfolded proteins. High levels of HSP90 in these animals could participate in the activation of signaling proteins such as hormone receptors, kinases and transcription factors. Naked mole-rats are able to withstand many cellular stressors, likely due to high levels of the chaperones HSP70 and HSP27. These proteins possibly help to refold misfolded proteins and minimize the formation of toxic protein aggregates. When aggregates are formed, high autophagic activity in naked mole-rats clears these. Misfolded proteins are also targeted for degradation via the ubiquitin-proteasome system, which is facilitated by high levels of CHIP and HSP70, as well as increased proteasome activity in naked mole-rats. Consequently, naked mole-rats show reduced build-up of poly-ubiquitinated proteins. Finally, the 6–12 amino acid peptides produced as a result of proteolytic cleavage in the proteasome are further cleaved into free amino acids through the action of peptidases. Naked mole-rats have lower levels of TPP2 protein than mice; yet, remarkably, they show ~4-fold higher TPP2-specific activity. Solid green and red arrows represent established/confirmed findings in naked mole-rats; pale green and red arrows (with dashed borders) indicate our hypotheses based on published observations as well as our unpublished data
11.4.1 Unique Features of the Proteostasis Network in Naked Mole-Rats
11.4.1.1 Unusual Ribosome Structure
Ribosomes are comprised of two subunits, a large and small subunit (60S and 40S in eukaryotes, respectively), both of which contain numerous ribosomal proteins intertwined with or anchored to the rRNA backbone. Several ribosomal proteins and distinct regions of rRNA are necessary for accurate mRNA decoding and for catalyzing amino acid transfer to the nascent, growing polypeptide chain (Doudna and Rath 2002; Wilson and Doudna Cate 2012; Yusupova and Yusupov 2014). Oddly, as discussed above, the naked mole-rat 28S rRNA harbors a ‘hidden break’ sequence that leads to the fragmentation of the 28S rRNA into two distinct pieces (Azpurua et al. 2013; Fang et al. 2014). This unusual cleavage pattern is shared with two of their South American relatives, the tuco-tuco (Ctenomys brasiliensis) and degu (Octodon degus), but is not seen in the guinea pig (Cavia porcellus) or in the Damaraland mole-rat (Fukomys damarensis) , suggesting that it was a trait present in the common hystricomorph ancestor which was subsequently lost by some species within the clade (Melen et al. 1999; Azpurua et al. 2013). In an attempt to elucidate the consequences of a cleaved 28S rRNA sequence on ribosome structure and mRNA translation, cryo-electron microscopy (cryo-EM) structural studies were conducted on ribosomes isolated from cultured skin fibroblasts of the naked mole-rat, and also on ribosomes from other hystricomorph rodents, the tuco-tuco and guinea pig. Interestingly, the gross structure of the large ribosomal subunit is essentially similar, regardless of whether the 28S rRNA is cleaved (naked mole-rat, tuco-tuco) or not (guinea pig) (Gutierrez-Vargas 2020). However, subtle structural differences in the A-site finger (ASF) and adjacent structures between the guinea pig and the other two species were observed. The length of the ASF reportedly affects frameshift activity in bacteria, with shorter ASFs exhibiting enhanced +1 frameshift activity (Komoda et al. 2006). Thus, the ASF appears to be important for maintenance of the correct reading frame and may contribute to increased translational fidelity or accuracy. While increased rates of translational accuracy, measured using luciferase-based reporter assays, have also been reported in naked mole-rats, further investigations using, for example, novel mass spectrometry-based techniques such as those developed in yeast and E. coli proteomes (Mordret et al. 2019), will enable one to quantitively assess this exciting question. Increased translation accuracy is a hallmark of pristine proteostasis as it leads to the generation of fewer aberrant polypeptides, thereby potentially freeing up PN chaperone and UPS components and allowing them to respond rapidly during stress conditions. Not surprisingly, increased translational accuracy is linked to prolonged longevity (Ke et al. 2017). Similarly, lowering global rates of translation, achieved by either genetic or chemical modulation of the activity of the mammalian target of rapamycin (mTOR) protein complex, a master regulator of protein synthesis, extends lifespan in a variety of models including yeast, worms, flies and mice (Fabrizio et al. 2001; Kapahi et al. 2004; Powers et al. 2006; Hansen et al. 2007; Selman et al. 2009). It has been postulated that under conditions of reduced translation, these models divert energy utilization into repair processes, which possibly leads to an increase in lifespan . The reduction in protein synthesis observed upon mTOR inhibition and corresponding increase in lifespan is likely also associated with increased translation fidelity; indeed, it was recently shown that enhancing the activity of a translation elongation factor kinase, EEF2K, a protein that is otherwise inactivated by mTOR, results in increased translation accuracy and extended lifespan (Xie et al. 2019). It is also possible that selective translation of specific mRNAs, even upon repression of global protein synthesis rates, impacts the aging process.
Contrary to evidence in multiple species linking reduced translation rates to extended lifespan , Azpurua et al. reported similar rates of protein synthesis in naked mole-rat and mouse skin fibroblasts (Azpurua et al. 2013). This finding is hard to resolve with subsequent mass spectrometry data revealing that proteins are generally turned over at slower rates in naked mole-rats than in mice (Swovick et al. 2018), as this would necessitate high rates of degradation in order to maintain steady-state. Protein synthesis and degradation are extremely energy-intensive processes, and increases in translation rates are actively selected against (Wagner 2005). Thus, it seems unlikely that the naked mole-rat will have evolved with high synthesis and degradation rates together with a strikingly low metabolic rate and extended lifespan for an animal of its size (Buffenstein and Yahav 1991; Buffenstein 2005). In support of our hypothesis and preliminary experimental findings that low levels of protein synthesis might occur in the naked mole-rat consistent with other models of longevity (Narayan and Buffenstein, unpublished observations), ribosomal protein abundance was found to be lower in comparison to guinea pigs (Heinze et al. 2018). Moreover, gene set enrichment analysis (GSEA) showed that ‘ribosome ’ (KEGG pathway) and ‘peptide chain elongation’ (Reactome pathway) are both decreased in naked mole-rats (Heinze et al. 2018). Collectively, these disparate studies suggest that rates of protein synthesis and turnover could be lower in naked mole-rats than in shorter-lived rodents, and these, together with high translation accuracy, are likely key contributors in maintaining a stable and functional proteome.
11.4.1.2 High Levels of Molecular Chaperones
Molecular chaperones play a critical role in proteostasis and assist in the attainment of native, functional protein conformations without themselves being present in the final tertiary structure of substrate proteins (Jayaraj et al. 2020). Broadly speaking, chaperones are classified as either “holdases” i.e. proteins that bind to and stabilize proteins, thereby preventing them from aggregating, “foldases” i.e. proteins that actively aid in the folding of proteins thus allowing them to attain their functional conformations, or “disaggregases” (also called “unfoldases”) i.e. proteins that participate in breaking up and resolubilizing aggregates (Barends et al. 2010; Richter et al. 2010). The majority of molecular chaperones are heat shock proteins (HSPs); consequently, the terms ‘chaperone’ and ‘heat shock protein’ are often used interchangeably in scientific literature. HSPs are sub-classified based on their size into functionally similar families: small heat shock proteins (sHSPs), HSP40 proteins or DNAJ proteins, HSP60 proteins or chaperonins, HSP70 proteins, HSP90 proteins and HSP100 (also referred to as HSP110) proteins (Kampinga et al. 2009; Jayaraj et al. 2020).
Several studies using antibody-based methods such as Western Blotting have shown that naked mole-rats have elevated basal expression of many chaperones compared to mice and other rodents (Rodriguez et al. 2014; Pride et al. 2015; Rodriguez et al. 2016). It also appears that, under conditions of cellular stress, naked mole-rats are able to further induce expression of some chaperones to much higher levels than mice (Pride et al. 2015), thereby affording even greater cytoprotection. Indeed, such high levels of HSPs are in keeping with observed positive correlations between basal levels of HSPs and lifespan not only in rodents, but also in other mammals and birds (Salway et al. 2011; Pride et al. 2015; Rodriguez et al. 2016). In this section, we highlight the molecular chaperones that show differential abundance in naked mole-rats relative to short-lived mice. These include components of the sHSP family, HSP70 protein family and HSP90 family. Additionally, by employing antibody-independent technologies to evaluate protein abundance in naked mole-rats relative to other species, we present novel unpublished findings from our laboratory. Our study uses Tandem Mass Tag (TMT)-based quantitative mass spectrometry (Thompson et al. 2003; Pappireddi et al. 2019) to measure chaperone abundance in liver tissue from 16 young, healthy naked mole-rats and 16 young, healthy mice (8 males and 8 females per species). There are major advantages of using mass spectrometry in comparative biology studies. Firstly, unlike antibody-based measurements, mass spectrometry does not rely on the assumption that the antibody binds with equal affinity to the target protein in all species under consideration. This is usually not the case due to differences in epitope sequence and/or differential post-translational modification (PTM) of the epitope. In mass spectrometric measurements, however, we can restrict our analysis to peptides that have identical sequence (and PTM) between the species, which enables direct species comparisons. Moreover, by quantifying multiple sequence-identical peptides from each protein in mass spectrometric studies, there is increased confidence that the measured change is real. For highly conserved proteins like HSPs, these measurements can include many dozens of sequence-identical peptides.
Small Heat Shock Proteins: Levels of HSP27 (HSPB1) have consistently been shown to be elevated in naked mole-rats compared to many other rodents, and the protein appears to be induced to much higher levels in naked mole-rat cells than in mouse cells after heat stress (Pride et al. 2015; Rodriguez et al. 2016). Our mass spectrometry data corroborate these findings: HSP27 levels were generally higher in liver tissue from both female and male naked mole-rats compared to physiologically age-matched C57BL/6 mice (Fig. 11.4).
Differentially expressed molecular chaperones in naked mole-rats in comparison to mice. TMT-based quantitative mass spectrometry was used to measure the abundance of molecular chaperones in liver tissue from young, healthy naked mole-rats and physiologically age-matched C57BL/6 mice (20–60 month old naked mole-rats and 3–5 month old mice; n=8 per sex per species). Shown are the most differentially expressed chaperones in the interspecific comparison. For this comparative analysis, only tryptic peptides that had identical sequences in both species were used (1–3 peptides for HSPB1, 8–13 peptides for HSPA1B, 4–7 peptides for HSPA12A , 2–7 peptides for HSPA12B, 2 peptides for DNAJB2, 3–7 peptides for DNAJC2, 51–61 peptides for HSP90AA1 and 7–11 peptides for TRAP1). Shown is the mean log2 fold change of the naked mole-rat protein relative to mouse, and the upper and lower limits of the 95% confidence intervals for these measurements (error bars)
HSP27 and other sHSPs, such as HSP20 and α-crystallin, are ATP-independent chaperones that function primarily as holdases – they bind to proteins to prevent aggregation and maintain misfolded proteins in folding-competent states for other chaperones like HSP70 (Bakthisaran et al. 2015; Webster et al. 2019). Additionally, cytoprotective sHSPs can have a variety of other benefits including anti-inflammatory properties and the ability to suppress the generation of reactive oxygen species (ROS) (De et al. 2000; Arrigo et al. 2005; Rothbard et al. 2012; Asthana et al. 2014; Bakthisaran et al. 2015). Given the importance of sHSPs in suppression of ROS generation and in preventing build-up of misfolded proteins, the high HSP27 levels observed in naked mole-rats are likely to be crucial in preventing the accumulation of oxidized proteins following stress (or during aging) by providing yet another layer of protection against oxidative stress in this species.
HSP70 and HSP40 protein families: The HSP70 family of proteins includes key chaperones that facilitate the folding of a variety of substrate proteins in an ATP-dependent manner. For reviews of members of this family and their numerous functions please see (Bukau and Horwich 1998; Rosenzweig et al. 2019; Jayaraj et al. 2020). In addition to folding a large proportion of the cellular proteome and preventing protein aggregation, HSP70, working together with HSP40, is critical in protein triage i.e. deciding when a protein cannot be folded and should be targeted for UPS degradation (McDonough and Patterson 2003). It does this in concert with E3 ligase enzymes such as carboxy-terminus of HSC70 interacting protein (CHIP or STUB1), which specifically ubiquitinates misfolded HSP70 and HSP90 client proteins (McDonough and Patterson 2003).
Western blot analysis revealed that the levels of HSP70 family proteins, particularly HSPA1A and/or HSPA1B (collectively known as ‘HSP70’), as well as the associated quality control E3 ligase CHIP, are generally higher in naked mole-rats than in mice, although there are tissue-specific differences (Rodriguez et al. 2014; Pride et al. 2015; Rodriguez et al. 2016). Our quantitative mass spectrometric measurements in liver tissue confirm these findings. Expanding upon this, we find that while HSP70 (HSPA1B) levels are higher in male and female naked mole-rats when compared to mice (Fig. 11.4), HSC70 (HSPA8), BiP (HSPA5) and GRP75 (HSPA9) levels are largely comparable (Narayan and Buffenstein, unpublished observations). Interestingly, HSPA14 and its associated HSP40 protein DNAJC2, both of which are components of the mammalian ribosome-associated complex (mRAC) (Jaiswal et al. 2011) that maintains newly synthesized polypeptides in a folding-competent state, are present at lower levels in naked mole-rats (Fig. 11.4). Lower levels of mRAC may be indicative of less protein misfolding in naked mole-rats possibly arising from reduced rates of protein synthesis.
Our proteomics data also reveal that HSPA12A and HSPA12B, HSP70 family members that have not previously been studied in naked mole-rats, are present at ~2-fold higher levels in naked mole-rat liver tissue relative to mice (Fig. 11.4). These newer members of the HSP70 family harbor several unique functions. For example, when mice genetically engineered to overexpress human HSPA12B and wild-type littermates were subjected to ischemia/reperfusion injury by middle cerebral artery occlusion, animals overexpressing HSPA12B exhibited decreased infarct volumes and showed fewer neurological defects (Ma et al. 2013). These transgenic mice were also protected from lipopolysaccharide (LPS )-induced cardiac dysfunction and inflammation as well as myocardial infarction-induced cardiac damage (Li et al. 2013). High basal levels of these proteins in naked mole-rats may have evolved to protect against variable oxygen availability in the deep nests and superficial burrows (Buffenstein and Craft 2021) and may likely contribute to the extreme tolerance to hypoxia observed in this species (Park et al. 2017).
Levels of many of the twenty-two HSP40 proteins measured in our mass spectrometry study are comparable in naked mole-rats and mice (Narayan and Buffenstein, unpublished observations). The only HSP40 protein that we found to be significantly elevated in both female and male naked mole-rats (~3-4-fold higher) when compared to mice is DNAJB2 (Fig. 11.4). This protein has been shown to play a vital role in preventing the aggregation of superoxide dismutase (SOD1) in a SOD1(G63A) genetically engineered mouse model of amyotrophic lateral sclerosis (ALS) (Novoselov et al. 2013), and has also been shown to strikingly reduce aggregation of Tar DNA-binding protein 43 (TDP-43) (Chen et al. 2016). Like other HSP40 proteins, DNAJB2 (also known as HSJ1A) has a J domain that allows it to bind to HSP70; additionally, it also has ubiquitin interacting motifs (UIM) allowing it to target substrates for proteasomal degradation. Indeed, DNAJB2’s role in preventing SOD1 aggregation has been shown to be UPS-dependent (Novoselov et al. 2013). Thus, the elevated level of DNAJB2 observed in naked mole-rats could play a key role in preventing protein aggregation and maintaining proteostasis through interactions with other chaperones and the UPS.
HSP90 protein family: Unlike other chaperones, HSP90’s primary function appears not to be in folding nascent polypeptides or preventing protein aggregation, but rather in allowing subtle conformational shifts in proteins, helping them to reach their active state whereby they can bind ligands or other proteins (Pearl and Prodromou 2006). Examples include the stabilization and ligand binding of steroid hormone receptors and the folding and activation of various signaling proteins including kinases and transcription factors. Given its role in activating kinases that function in various growth-promoting signaling pathways, HSP90 is an attractive cancer target. Indeed, HSP90 inhibition has been studied as an anti-cancer therapeutic for about three decades, although to date, no drug has been approved for clinical use (Neckers and Workman 2012; Yuno et al. 2018).
Using antibody-based measurements, HSP90 protein levels were shown to be largely comparable in naked mole-rat tissues relative to mice (Rodriguez et al. 2014, 2016). Based on our mass spectrometric data of specific HSP90 family members, mitochondrial HSP90 (TRAP1) levels are ~2-fold lower in naked mole-rats, while HSP90α (HSP90AA1) levels are higher (Fig. 11.4). This is interesting, given elevated HSP90 levels are usually associated with poor prognosis in cancers, yet naked mole-rats have an extremely low incidence of tumors. It is probable that elevated HSP90α in naked mole-rats serves other (beneficial) functions: these could include, for example, stabilizing the high levels of p53 present in naked mole-rats (Walerych et al. 2004; Lewis et al. 2012; Deuker et al. 2020) and/or maintaining p53 in an easily-activatable state and/or assisting HSP70 in protein triage through interactions with the UPS.
In summary, a number of molecular chaperones, notably HSP70, HSPA12A/B , DNAJB2, HSP27 and HSP90α are present at higher levels (2-4 fold) in naked mole-rats relative to mice (Fig. 11.4), and likely play important roles in maintaining a healthy proteome throughout the long life of this animal through efficient protein folding and prevention of misfolded protein aggregates (Fig. 11.3). Consistent with this hypothesis, protein extracts from young and old naked mole-rats were demonstrated to be more resistant to unfolding than mouse proteins in the presence of denaturants such as urea (Perez et al. 2009).
11.4.1.3 Increased Activity of Protein Degradation and Cellular Recycling Machinery
A key function of the PN is to recognize polypeptides or proteins that are aberrantly folded and to eradicate these from the intracellular milieu. Additionally, the PN also participates in maintaining steady state of proteins through turnover of correctly folded proteins in a timely manner. This is particularly important for transcription factors which tend to have numerous downstream targets – if these proteins are not tightly regulated and degraded in a temporal manner, the signaling pathways they control will be chronically “on”, which could be detrimental to cells.
Underscoring the critical role of protein degradation machinery in maintaining proteostasis is the observation that enhancing the function of the proteasome or autophagy through genetic or chemical interventions results in extended health- and life- span in a variety of animal models (Kruegel et al. 2011; Rubinsztein et al. 2011; Vilchez et al. 2012; Pyo et al. 2013; Nguyen et al. 2019). Correspondingly, a common feature of long-lived models of aging is a high basal level of proteasome and/or autophagic activity, and, importantly, sustained activity of these components through life (Kenyon 2010; Vilchez et al. 2014; Labbadia and Morimoto 2015). Like these genetically/chemically-manipulated long-lived experimental models of aging, naked mole-rats show high proteasome activity and elevated autophagy (Rodriguez et al. 2012; Zhao et al. 2014; Triplett et al. 2015), which is discussed in detail below.
Proteasome : When compared to mice, naked mole-rats have higher basal 26S proteasome activity in multiple tissues (Perez et al. 2009; Rodriguez et al. 2012) (Janki, Kramer, Grimes and Buffenstein, unpublished observations). What is unclear in these studies is how this was achieved: whether the observed increase in activity is due to enzyme abundance (i.e. naked mole-rats have more proteasomes than mice), specific activity (i.e. proteasomes in naked mole-rats are more efficient at degrading substrates) or some combination of both. To address this discrepancy, we searched our TMT-based mass spectrometry dataset, generated using liver tissue from young, healthy naked mole-rats and mice (n = 16 per species) for proteasome subunits. We focused on components of the 20S core proteasome and the 19S regulatory particle (also known as PA200), which together form the 26S proteasome . First, we demonstrated that in the same samples used for mass spectrometry, higher proteasome activity was observed in the naked mole-rat samples (2-fold higher than in mice), corroborating published findings (Fig. 11.5a). Similar to the study by Perez (Perez et al. 2009), our mass spectrometry data showed that proteasome abundance was comparable in both species for not only the core proteasome subunits, but also the constituent proteins of the 19S cap (Figs. 11.5b and c). The only exception was PSMB5, one of the catalytic subunits of the 20S core proteasome , which is present at ~50% higher levels in naked mole-rats (Fig. 11.5b).
Higher proteasome activity is observed in tissue extracts from naked mole-rats compared to mice. (a) Proteasome activity was measured in liver extracts prepared from naked mole-rats and physiologically age-matched C57BL/6 mice (20–60 month old naked mole-rats and 3–5 month old mice; n = 8 per sex per species) using the established Suc-LLVY-AMC assay as previously described (Rodriguez et al. 2012). Assays were run in the absence and presence of the proteasome inhibitor epoxomicin (10 uM). The difference between the total activity (without epoxomicin) and activity in the presence of epoxomicin is considered to be proteasome-specific and is displayed on the y-axis (Epoxomicin-corrected chymotryptic (ChT) activity). Shown are the activity measurements for each of the 8 animals in the groups (circles), the mean (thick horizontal line), and standard deviation (error bars). p-values were calculated using unpaired Student’s t-tests. (b and c) TMT-based quantitative mass spectrometry was used to measure the abundance of 20S core proteasome (b) and 19S regulatory particle (c) subunits in the liver extracts prepared above. For this comparative analysis, only tryptic peptides that had identical sequences in both species were used. Shown is the mean log2 fold change of the naked mole-rat protein relative to mouse, and the upper and lower limits of the 95% confidence intervals for these measurements (error bars)
Our data raise the interesting possibility that naked mole-rat proteasomes might have a higher specific activity than mouse proteasomes. This could be due to: (1) higher levels of the catalytic subunit PSMB5 (Fig. 11.5b); (2) differential expression of catalytic immunoproteasome subunits (Rodriguez et al. 2012); (3) the presence of HSP70, which specifically binds to the naked mole-rat proteasome and increases its activity (Rodriguez et al. 2014); (4) possible different proportions of the various regulatory proteasome caps and hybrid combinations of these caps; (5) presence of as yet unknown regulatory proteins in the naked mole-rat proteasome ; (6) presence of specific as yet unknown PTMs in one/many proteasome subunits; and/or (7) the finding that naked mole-rats have fewer poly-ubiquitinated proteins compared to mice (Perez et al. 2009), which frees up the proteasome to degrade the artificial substrate used in these proteasome activity assays. The last point merits some discussion as although the Suc-LLVY-AMC peptide-based proteasome activity assay was originally described to measure the activity of purified proteasomes in vitro, it has since been routinely used to determine maximum proteasome activity in cell and tissue extracts (Rivett et al. 1994; Liggett et al. 2010). This is in spite of previous findings that the artificial substrate, Suc-LLVY-AMC, can be also cleaved by calpains, tripeptidyl peptidase (TPP2 ) and chymotrypsin-like proteases. All of these enzymes are usually present in cell/tissue extracts and show disparate activity under different experimental conditions, thereby complicating data interpretation (Chandu et al. 2003; Giguere and Schnellmann 2008). In addition to the presence of these non-proteasomal proteases, data interpretation in this assay is also complicated by the presence of other proteasome substrates such as oxidized proteins and poly-ubiquitinated proteins in cell extracts. Thus, the artificial substrate must compete with endogenous substrates for proteasome-degradation, and so a higher measured activity in this context could be due to fewer endogenous proteasome substrates in that particular sample, freeing up the proteasome to degrade the artificial substrate. This makes the assay a readout for proteasomal substrate load rather than specific activity. Regardless of these caveats, the finding that “proteasome activity” detected in naked mole-rats – whether due to a lower substrate load in this species i.e. fewer misfolded proteins and polyubiquitinated proteins, or higher specific activity of the proteasome – is greater than mice is striking, as it suggests that proteasomes in this species have a greater capacity to cope with conditions of stress, during which UPS activity is critical for survival.
Proteolysis in the proteasome usually generates peptides that are predominantly about 6–12 amino acids in length (Kisselev et al. 1999; Book et al. 2005). These polypeptides are subsequently processed by various endo- and exo- peptidases that trim them into amino acids. Although generally overlooked in the PN, these peptidases play important roles in preventing the accumulation of toxic intermediate peptides and in facilitating amino acid recycling. A key player here is the protease TPP2 , that physically interacts with the proteasome and may possibly substitute for proteasome functions in some contexts (Glas et al. 1998; Geier et al. 1999; Fukuda et al. 2017). Given the observed high proteasome activity in naked mole-rats, one would expect the activity of downstream proteases like TPP2 to also be high. Unexpectedly, our mass spectrometric data showed that TPP2 protein levels were 3–4X lower in young, healthy naked mole-rats when compared to physiologically age-matched mice (Fig. 11.6a). Strikingly, however, when we measured TPP2 activity in these extracts, we found that in spite of having less TPP2 enzyme, the measured activity in naked mole-rats was 3-5X higher than in mice (Fig. 11.6b). As the artificial peptide substrate used in the TPP2 activity assay may also be cleaved be other cellular peptidases, we performed our assays in the presence and absence of butabindide oxalate (BO), an inhibitor that has been reported to be specific for the membrane-bound form of TPP2 (Rose et al. 1996; Glas et al. 1998; Dasgupta et al. 2014). The difference between measured activity in the absence and presence of BO may therefore be inferred to be TPP2-specific (Fig. 11.6b). As with the proteasome activity assay discussed above, it is possible that the
Naked mole-rats show ~4-fold higher TPP2 activity than mice. (a) TMT-based quantitative mass spectrometry was used to measure the abundance of TPP2 in liver tissue from young, healthy naked mole-rats and physiologically age-matched C57BL/6 mice (20–60 month old naked mole-rats and 3–5 month old mice; n = 8 per sex per species). For this comparative analysis, only tryptic peptides that had identical sequences in both species were used (12–15 peptides). Shown is the mean log2 fold change of the naked mole-rat protein relative to mouse, and the upper and lower limits of the 95% confidence intervals for these measurements (error bars). (b) Liver extracts were prepared from 4 male naked mole-rats (two 2 year old animals and two 7 year old animals) and 4 male C57BL/6 mice (two 4 month old animals and two 12 month old animals) as described previously for proteasome activity assays (Rodriguez et al. 2012) and activity assays were set up in a similar manner except using AAF-AMC (100 uM) as the substrate. Assays were run in the absence and presence of the TPP2-specific inhibitor butabindide oxalate (500 uM). The difference between the total activity (without butabindide) and activity in the presence of butabindide is considered to be TPP2-specific and is displayed on the y-axis (Butabindide-corrected TPP2 activity). Shown is the mean activity for the 4 animals in each group (grey bars) and standard deviation (error bars). p-values were calculated using unpaired Student’s t-tests
observed high TPP2 activity detected in naked mole-rat liver extracts is due to the presence of fewer endogenous TPP2 substrates, rather than higher specific activity. There could also be other TPP2-like proteases unique to the naked mole-rat that contribute to the measured activity. Given these caveats, it would be prudent to measure specific activity of TPP2 using purified protein prepared from naked mole-rats and mice, in order to confirm our preliminary findings based on whole liver extracts. Nonetheless, it is intriguing to speculate that the observed high TPP2 activity in naked mole-rats not only assists in amino acid recycling, but also perhaps compensates for any loss in proteasome function under some stress conditions, as has been observed in EL4 lymphoma cells (Glas et al. 1998; Geier et al. 1999).
Autophagy: Although originally believed to be solely a non-selective process essential for the recycling of nutrients under conditions of stress and starvation, autophagy has since been shown to play a vital role in the selective clearance of misfolded proteins and damaged organelles including mitochondria and peroxisomes, making it an important component of the PN (Olzmann and Chin 2008; Glick et al. 2010). There are numerous reports on the benefits of autophagy in aging: elevated autophagy is a common feature of long-lived models. The mTOR nutrient sensing pathway is a key regulator (inhibitor) of autophagy, and indeed, inhibiting mTOR signaling with rapamycin enhances the rate of autophagy and extends lifespan (Zhang et al. 2014). The lifespan extension benefits of interventions such as caloric restriction (CR) and reduced insulin/IGF1 signaling are also dependent, at least in part, on autophagy (Madeo et al. 2015; Hansen et al. 2018). Additionally, overexpression of some autophagy genes, resulting in increased rates of autophagy, has been shown to extend lifespan (Simonsen et al. 2008; Pyo et al. 2013).
High basal levels of autophagy in naked mole-rats when compared to mice has been shown by two independent groups in skin fibroblasts (Rodriguez et al. 2011; Pride et al. 2015) and in various tissues (Zhao et al. 2014). Measurements were made using biochemical methods (LC3-II/LC3-I ratio) and confirmed using electron microscopy, which revealed the presence of many more autophagosome-like structures in naked mole-rats (Zhao et al. 2014). Perhaps the most striking observation related to autophagy in naked mole-rats is that in the brain, rates of autophagy appear to be similar throughout life – or at least until 24 years of age (Triplett et al. 2015). In keeping with this finding, mTOR signaling remains largely unchanged. This is in stark contrast to mouse brains, where mTOR signaling increases with age (Yang et al. 2012). Autophagic flux also decreases in fibroblasts isolated from older mice (Ott et al. 2016). The sustained high level of autophagy throughout the life of the naked mole-rat is likely critical to maintaining its extended health- and life- span by preventing the accumulation of misfolded proteins, damaged organelles, and protein aggregates in the animal.
11.5 Master Regulators of Cellular Homeostasis and Stress Resistance
Transcription factors including forkhead box O (FOXO) proteins, nuclear factor erythroid 2-related factor 2 (NRF2 or NFE2L2), hypoxia-induced factor 1 α (HIF1α) and heat shock factor 1 (HSF1) have been linked to extended longevity, cellular detoxification and/or stress resistance in a variety of invertebrate and vertebrate models (Kenyon 2010; Leiser and Kaeberlein 2010; Li et al. 2017; Denzel et al. 2019). Of these, NRF2 and HIF1α have been investigated in naked mole-rats in some detail (Fig. 11.7) and are discussed below.
Master regulators of longevity and stress resistance in naked mole-rats. This schematic summarizes known findings on master regulators of aging in naked mole-rats. Signaling through the two major growth/nutrient-sensing pathways implicated in longevity, insulin/IGF1 and mTOR, are decreased in naked mole-rats. These, together with increased signaling through various protective pathways (NRF2, HIF1α and HSF1), may result in stress resistance and longevity through increased expression of antioxidants, chaperones and autophagic and proteasome components. In this schematic, solid green and red arrows represent established/confirmed findings in naked mole-rats; pale green and red arrows (with dashed borders) indicate our hypotheses based on published observations as well as our unpublished data
Although FOXO proteins are essentially unexplored in these animals, transcript levels of insulin/IGF1 signaling components appear to be reduced (Kim et al. 2011). This is an important and pertinent finding, as inhibition of insulin/IGF1 signaling has been shown to induce FOXO expression and extend lifespan in other species (Kenyon 2010). Known FOXO targets include autophagy genes, detoxification enzymes such as catalase and SOD, and UPS genes (Greer and Brunet 2005), many of which are elevated in naked mole-rats. Taken together, these data suggest that FOXO levels and activity are likely elevated in naked mole-rats and may be contributing factors in their remarkable anti-aging phenotype. The insulin/IGF1 signaling pathway has been shown to closely interact with the mTOR nutrient sensing pathway (Yoon 2017), downregulation of which also extends lifespan in a variety of species (Fabrizio et al. 2001; Kapahi et al. 2004; Powers et al. 2006; Hansen et al. 2007; Selman et al. 2009). Strikingly, unlike in mice, where an age-dependent increase in mTOR complex 1 (mTORC1) is observed, mTOR activity remains unchanged during aging in naked mole-rat brains (Yang et al. 2012; Triplett et al. 2015). Thus, reduced signaling through the insulin/IGF1 and mTOR pathways in the naked mole-rat, which are possibly maintained during aging, could be key contributors of healthy aging in this species (Fig. 11.7).
HSF1, the regulator of heat shock protein expression, particularly in response to cellular stress, is also present at high levels in naked mole-rats, and, interestingly, HSF1 protein levels have been shown to positively correlate with lifespan (Rodriguez et al. 2016). Although FOXO and HSF1 act in anti-aging pathways that are largely independent of each other, it has been proposed that the synergistic activation of both these transcription factors could be highly beneficial in reducing proteotoxicity, particularly in the context of neurodegenerative diseases (Perez et al. 2012). The naked mole-rat appears to validate this hypothesis, and further studies to dissect the molecular details and consequences of the activation of these beneficial master regulators of anti-aging mechanisms in naked mole-rats are an exciting research avenue to explore.
11.5.1 Increased NRF2 Signaling and Antioxidant Levels
NRF2 is a transcription factor whose gene targets include a plethora of antioxidants and detoxicants including various glutathione S-transferase (GST) proteins and NAD(P)H:quinone oxidoreductase 1 (NQO1), as well as some HSPs and proteasome subunits (Jaiswal 2000; Reddy et al. 2007). Under non-stress conditions, NRF2 is found in complex with Kelch-like ECH-Associated Protein 1 (KEAP1), which targets it for proteasomal degradation (Itoh et al. 1999). During oxidative stress, ROS oxidize specific cysteine residues on KEAP1, causing a change in conformation that prevents binding to NRF2 (Tong et al. 2006). Consequently, NRF2 is no longer targeted for proteasome-mediated degradation and can translocate to the nucleus to activate its target genes.
Naked mole-rats appears to have ~3-fold more Nrf2 mRNA and NRF2 protein than mice under non-stress conditions (Lewis et al. 2015). This is likely due to the fact that KEAP1 is present at much lower levels in naked mole-rats (Lewis et al. 2015). Consequently, naked mole-rats are in a ‘primed’ phase even under non-stress conditions, ready to respond promptly and efficiently to oxidative stress through high basal levels of antioxidants (Lewis et al. 2012). These high antioxidant levels are likely maintained well into old age in these animals, as, unlike in mice and other species, there is no age-associated increase in oxidative damage i.e. protein carbonyls or lipid peroxidation, in naked mole-rats (Andziak and Buffenstein 2006; Andziak et al. 2006).
11.5.2 Increased HIF1α Levels and Hypoxia Tolerance
Naked mole-rats are extremely hypoxia-tolerant animals, able to survive up to 18 min without oxygen (Park et al. 2017). This appears to be due to a switch to anaerobic glycolysis driven by increased uptake and utilization of fructose during anoxia (Park et al. 2017). Additionally, captive naked mole-rats appear to harbor some other salient features in hypoxia signaling response pathways. For example, the naked mole-rat genome encodes an HIF1α protein containing a T407I amino acid substitution within the Von Hippel-Lindau (VHL) protein interacting domain compared to other mammals (Kim et al. 2011). Furthermore, the naked mole-rat VHL protein also displays amino acid differences (Kim et al. 2011). As VHL is responsible for the ubiquitin-dependent degradation of HIF1α under normoxia, it seems probable these differences may function in adaptation to extremely low oxygen supply, at least in part, by impeding HIF1α degradation. Indeed, in keeping with their evolutionary history of living with hypoxic conditions for millions of years, HIF1α is present at much higher levels in a number of naked mole-rat tissues compared to mice but nevertheless can be further induced following experimental hypoxia (Xiao et al. 2017a). Additionally, isolated primary hepatic stellate cells from the naked mole-rat appear resistant to hypoxia as evidenced by lack of cell death following hypoxia exposure (Xiao et al. 2017a). Collectively, these findings indicate that cell autonomous mechanisms likely function in the naked mole-rat’s adaptation to hypoxic stress.
A study probing the transcriptional response of naked mole-rats to hypoxic exposure in comparison to normoxia found that the differentially expressed transcripts in muscle tissue were enriched for genes involved in cell adhesion, metabolic processes, ion transport , ion transmembrane transport , and cell-cell signaling. Furthermore, KEGG pathway analysis revealed enrichment of genes involved in focal adhesion, MAPK signaling, circadian rhythm, cell cycle, metabolic pathways, and glycine, serine and threonine metabolism (Xiao et al. 2017b). As the role of cell-cell contact and focal adhesion in hypoxia remain poorly explored, future studies investigating the role of these pathways in naked mole-rats may shed light on hypoxia adaptation and mechanisms that protect these animals from prolonged hypoxic stress.
11.6 Concluding Remarks
Likely in response to survival in arid, hypoxic, hypercapnic subterranean environments, the naked mole-rat has evolved an impressive suite of cellular defenses, that, in a predator-free laboratory environment, translate into an unusually long, healthy life. With elevated DNA repair pathways, antioxidants, molecular chaperones and strong cell cycle control mechanisms, the naked mole-rat appears to be primed to sense, and rapidly eliminate, signs of cellular damage. The result is the maintenance of homeostasis throughout the long life of this remarkable animal, through a stable genome and pristine proteome, which manifest as sustained physiological function even in old age. Although tremendous efforts have been made to advance our understanding of naked mole-rat biology, several new avenues are now ready for future exploration, particularly in light of advances in transcriptomic and proteomic methodologies. Such studies will provide unparalleled insights into the molecular mechanisms underlying the extraordinary ability of the naked mole-rat to resist physiological declines and associated pathologies with increasing chronological age.
Notes
- 1.
Predicted maximal longevity in years was based on the allometric equation of de Magalhaes et al. (2007): tmax = 4.88 M0.153 where M is the mass in grams.
References
Abdelmohsen K, Kuwano Y, Kim HH, Gorospe M (2008) Posttranscriptional gene regulation by RNA-binding proteins during oxidative stress: implications for cellular senescence. Biol Chem 389(3):243–255
Andziak B, Buffenstein R (2006) Disparate patterns of age-related changes in lipid peroxidation in long-lived naked mole-rats and shorter-lived mice. Aging Cell 5(6):525–532
Andziak B, O’Connor TP, Qi W, DeWaal EM, Pierce A, Chaudhuri AR, Van Remmen H, Buffenstein R (2006) High oxidative damage levels in the longest-living rodent, the naked mole-rat. Aging Cell 5(6):463–471
Arrigo AP, Virot S, Chaufour S, Firdaus W, Kretz-Remy C, Diaz-Latoud C (2005) Hsp27 consolidates intracellular redox homeostasis by upholding glutathione in its reduced form and by decreasing iron intracellular levels. Antioxid Redox Signal 7:414–422
Asthana A, Bollapalli M, Tangirala R, Bakthisaran R, Mohan Rao C (2014) Hsp27 suppresses the Cu(2+)-induced amyloidogenicity, redox activity, and cytotoxicity of alpha-synuclein by metal ion stripping. Free Radic Biol Med 72:176–190
Azpurua J, Ke Z, Chen IX, Zhang Q, Ermolenko DN, Zhang ZD, Gorbunova V, Seluanov A (2013) Naked mole-rat has increased translational fidelity compared with the mouse, as well as a unique 28S ribosomal RNA cleavage. Proc Natl Acad Sci U S A 110(43):17350–17355
Bakthisaran R, Tangirala R, Rao Ch M (2015) Small heat shock proteins: role in cellular functions and pathology. Biochim Biophys Acta 1854(4):291–319
Barends TR, Werbeck ND, Reinstein J (2010) Disaggregases in 4 dimensions. Curr Opin Struct Biol 20(1):46–53
Benayoun BA, Pollina EA, Brunet A (2015) Epigenetic regulation of ageing: linking environmental inputs to genomic stability. Nat Rev Mol Cell Biol 16(10):593–610
Blackburn EH, Epel ES, Lin J (2015) Human telomere biology: a contributory and interactive factor in aging, disease risks, and protection. Science 350(6265):1193–1198
Book AJ, Yang P, Scalf M, Smith LM, Vierstra RD (2005) Tripeptidyl peptidase II. An oligomeric protease complex from Arabidopsis. Plant Physiol 138(2):1046–1057
Borbolis F, Syntichaki P (2015) Cytoplasmic mRNA turnover and ageing. Mech Ageing Dev 152:32–42
Brandman O, Hegde RS (2016) Ribosome-associated protein quality control. Nat Struct Mol Biol 23(1):7–15
Buffenstein R (2005) The naked mole-rat: a new long-living model for human aging research. J Gerontol A Biol Sci Med Sci 60(11):1369–1377
Buffenstein R (2008) Negligible senescence in the longest living rodent, the naked mole-rat: insights from a successfully aging species. J Comp Physiol B 178(4):439–445
Buffenstein R, Craft W (2021) The idiosyncratic physiological traits of the naked mole-rat; a resilient animal model of aging, longevity, and healthspan. In: Buffenstein R, Park TJ, Holmes MM (eds) The Extraordinary Biology of the Naked Mole-Rat. Springer, New York, pp 221–254
Buffenstein R, Yahav S (1991) Is the naked mole-rat Heterocephalus glaber an endothermic yet poikilothermic mammal? J Therm Biol 16(4):227–232
Bukau B, Horwich AL (1998) The Hsp70 and Hsp60 chaperone machines. Cell 92(3):351–366
Chandu D, Kumar A, Nandi D (2003) PepN, the major Suc-LLVY-AMC-hydrolyzing enzyme in Escherichia coli, displays functional similarity with downstream processing enzymes in Archaea and eukarya. Implications in cytosolic protein degradation. J Biol Chem 278(8):5548–5556
Chen HJ, Mitchell JC, Novoselov S, Miller J, Nishimura AL, Scotter EL, Vance CA, Cheetham ME, Shaw CE (2016) The heat shock response plays an important role in TDP-43 clearance: evidence for dysfunction in amyotrophic lateral sclerosis. Brain 139(Pt 5):1417–1432
Collins GA, Goldberg AL (2017) The logic of the 26S proteasome. Cell 169(5):792–806
Dabin J, Fortuny A, Polo SE (2016) Epigenome maintenance in response to DNA damage. Mol Cell 62(5):712–727
Dasgupta S, Castro LM, Dulman R, Yang C, Schmidt M, Ferro ES, Fricker LD (2014) Proteasome inhibitors alter levels of intracellular peptides in HEK293T and SH-SY5Y cells. PLoS One 9(7):e103604
de Lange T (2005) Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev 19(18):2100–2110
de Magalhaes JP, Costa J, Church GM (2007) An analysis of the relationship between metabolism, developmental schedules, and longevity using phylogenetic independent contrasts. J Gerontol A Biol Sci Med Sci 62(2):149–160
De AK, Kodys KM, Yeh BS, Miller-Graziano C (2000) Exaggerated human monocyte IL-10 concomitant to minimal TNF-alpha induction by heat-shock protein 27 (Hsp27) suggests Hsp27 is primarily an antiinflammatory stimulus. J Immunol 165(7):3951–3958
Delaney MA, Ward JM, Walsh TF, Chinnadurai SK, Kerns K, Kinsel MJ, Treuting PM (2016) Initial case reports of cancer in naked mole-rats (Heterocephalus glaber). Vet Pathol 53(3):691–696
Delaney M, Imai D, Buffenstein R (2021) Spontaneous disease and pathology of naked mole-rats. In: Buffenstein R, Park TJ, Holmes MM (eds) The Extraordinary Biology of the Naked mole-Rat. Springer, New York, pp 353–380
Denzel MS, Lapierre LR, Mack HID (2019) Emerging topics in C. elegans aging research: transcriptional regulation, stress response and epigenetics. Mech Ageing Dev 177:4–21
Deuker MM, Lewis KN, Ingaramo M, Kimmel J, Buffenstein R, Settleman J (2020) Unprovoked stabilization and nuclear accumulation of the naked mole-rat p53 protein. Sci Rep 10(1):6966
Doudna JA, Rath VL (2002) Structure and function of the eukaryotic ribosome: the next frontier. Cell 109(2):153–156
Esteller M (2011) Non-coding RNAs in human disease. Nat Rev Genet 12(12):861–874
Evdokimov A, Kutuzov M, Petruseva I, Lukjanchikova N, Kashina E, Kolova E, Zemerova T, Romanenko S, Perelman P, Prokopov D, Seluanov A, Gorbunova V, Graphodatsky A, Trifonov V, Khodyreva S, Lavrik O (2018) Naked mole rat cells display more efficient excision repair than mouse cells. Aging 10(6):1454–1473
Evfratov SA, Smekalova EM, Golovin AV, Logvina NA, Zvereva MI, Dontsova OA (2014) Structural features of the telomerase RNA gene in the naked mole rat Heterocephalus glaber. Acta Nat 6(2):41–47
Fabrizio P, Pozza F, Pletcher SD, Gendron CM, Longo VD (2001) Regulation of longevity and stress resistance by Sch9 in yeast. Science 292(5515):288–290
Fang X, Seim I, Huang Z, Gerashchenko MV, Xiong Z, Turanov AA, Zhu Y, Lobanov AV, Fan D, Yim SH, Yao X, Ma S, Yang L, Lee SG, Kim EB, Bronson RT, Sumbera R, Buffenstein R, Zhou X, Krogh A, Park TJ, Zhang G, Wang J, Gladyshev VN (2014) Adaptations to a subterranean environment and longevity revealed by the analysis of mole rat genomes. Cell Rep 8(5):1354–1364
Frye M, Harada BT, Behm M, He C (2018) RNA modifications modulate gene expression during development. Science 361(6409):1346–1349
Fukuda Y, Beck F, Plitzko JM, Baumeister W (2017) In situ structural studies of tripeptidyl peptidase II (TPPII) reveal spatial association with proteasomes. Proc Natl Acad Sci U S A 114(17):4412–4417
Geier E, Pfeifer G, Wilm M, Lucchiari-Hartz M, Baumeister W, Eichmann K, Niedermann G (1999) A giant protease with potential to substitute for some functions of the proteasome. Science 283(5404):978–981
George W (1979) Conservatism in the karyotypes of two African mole-rats (Rodentia, Bathyergidae). Zeitschrift für Säugetierkunde 44:278–285
Giguere CJ, Schnellmann RG (2008) Limitations of SLLVY-AMC in calpain and proteasome measurements. Biochem Biophys Res Commun 371(3):578–581
Glas R, Bogyo M, McMaster JS, Gaczynska M, Ploegh HL (1998) A proteolytic system that compensates for loss of proteasome function. Nature 392(6676):618–622
Glick D, Barth S, Macleod KF (2010) Autophagy: cellular and molecular mechanisms. J Pathol 221(1):3–12
Greer EL, Brunet A (2005) FOXO transcription factors at the interface between longevity and tumor suppression. Oncogene 24(50):7410–7425
Grimes KM, Reddy AK, Lindsey ML, Buffenstein R (2014) And the beat goes on: maintained cardiovascular function during aging in the longest-lived rodent, the naked mole-rat. Am J Physiol Heart Circ Physiol 307(3):H284–H291
Gutierrez-Vargas C (2020) Single-Particle Cryo-Electron Microscopy Studies of Ribosomes with Fragmented 28S rRNA. Ph.D. Thesis, Columbia University, New York, New York
Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, Klotzle B, Bibikova M, Fan JB, Gao Y, Deconde R, Chen M, Rajapakse I, Friend S, Ideker T, Zhang K (2013) Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell 49(2):359–367
Hansen M, Taubert S, Crawford D, Libina N, Lee SJ, Kenyon C (2007) Lifespan extension by conditions that inhibit translation in Caenorhabditis elegans. Aging Cell 6(1):95–110
Hansen M, Rubinsztein DC, Walker DW (2018) Autophagy as a promoter of longevity: insights from model organisms. Nat Rev Mol Cell Biol 19(9):579–593
Heinze I, Bens M, Calzia E, Holtze S, Dakhovnik O, Sahm A, Kirkpatrick JM, Szafranski K, Romanov N, Sama SN, Holzer K, Singer S, Ermolaeva M, Platzer M, Hildebrandt T, Ori A (2018) Species comparison of liver proteomes reveals links to naked mole-rat longevity and human aging. BMC Biol 16(1):82
Horvath S (2013) DNA methylation age of human tissues and cell types. Genome Biol 14(10):R115
Hulbert AJ, Faulks SC, Buffenstein R (2006) Oxidation-resistant membrane phospholipids can explain longevity differences among the longest-living rodents and similarly-sized mice. J Gerontol A Biol Sci Med Sci 61(10):1009–1018
Itoh K, Wakabayashi N, Katoh Y, Ishii T, Igarashi K, Engel JD, Yamamoto M (1999) Keap1 represses nuclear activation of antioxidant responsive elements by Nrf2 through binding to the amino-terminal Neh2 domain. Genes Dev 13(1):76–86
Jaiswal AK (2000) Regulation of genes encoding NAD(P)H:quinone oxidoreductases. Free Radic Biol Med 29(3–4):254–262
Jaiswal H, Conz C, Otto H, Wolfle T, Fitzke E, Mayer MP, Rospert S (2011) The chaperone network connected to human ribosome-associated complex. Mol Cell Biol 31(6):1160–1173
Jarvis JUM (1981) Eusociality in a mammal: cooperative breeding in naked mole-rat colonies. Science 212(4494):571–573
Jarvis JUM, O'Riain MJ, McDaid E (1991) Growth and factors affecting body size in naked mole-rats. In: Sherman PW, Jarvbis JUM, Alexander RD (eds) The Biology of the Naked Mole-Rat. Princeton University Press, Princeton, pp 358–383
Jayaraj GG, Hipp MS, Hartl FU (2020) Functional modules of the proteostasis network. Cold Spring Harb Perspect Biol 12(1):a033951
Jiang JJ, Cheng LH, Wu H, He YH, Kong QP (2016) Insights into long noncoding RNAs of naked mole-rat (Heterocephalus glaber) and their potential association with cancer resistance. Epigenetics Chromatin 9:51
Kampinga HH, Hageman J, Vos MJ, Kubota H, Tanguay RM, Bruford EA, Cheetham ME, Chen B, Hightower LE (2009) Guidelines for the nomenclature of the human heat shock proteins. Cell Stress Chaperones 14(1):105–111
Kapahi P, Zid BM, Harper T, Koslover D, Sapin V, Benzer S (2004) Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr Biol 14(10):885–890
Ke Z, Mallik P, Johnson AB, Luna F, Nevo E, Zhang ZD, Gladyshev VN, Seluanov A, Gorbunova V (2017) Translation fidelity coevolves with longevity. Aging Cell 16(5):988–993
Keane M, Craig T, Alfoldi J, Berlin AM, Johnson J, Seluanov A, Gorbunova V, Di Palma F, Lindblad-Toh K, Church GM, de Magalhaes JP (2014) The naked mole-rat genome resource: facilitating analyses of cancer and longevity-related adaptations. Bioinformatics 30(24):3558–3560
Kenyon CJ (2010) The genetics of ageing. Nature 464(7288):504–512
Kim EB, Fang X, Fushan AA, Huang Z, Lobanov AV, Han L, Marino SM, Sun X, Turanov AA, Yang P, Yim SH, Zhao X, Kasaikina MV, Stoletzki N, Peng C, Polak P, Xiong Z, Kiezun A, Zhu Y, Chen Y, Kryukov GV, Zhang Q, Peshkin L, Yang L, Bronson RT, Buffenstein R, Wang B, Han C, Li Q, Chen L, Zhao W, Sunyaev SR, Park TJ, Zhang G, Wang J, Gladyshev VN (2011) Genome sequencing reveals insights into physiology and longevity of the naked mole rat. Nature 479(7372):223–227
Kisselev AF, Akopian TN, Woo KM, Goldberg AL (1999) The sizes of peptides generated from protein by mammalian 26 and 20 S proteasomes. Implications for understanding the degradative mechanism and antigen presentation. J Biol Chem 274(6):3363–3371
Klaips CL, Jayaraj GG, Hartl FU (2018) Pathways of cellular proteostasis in aging and disease. J Cell Biol 217(1):51–63
Komoda T, Sato NS, Phelps SS, Namba N, Joseph S, Suzuki T (2006) The A-site finger in 23 S rRNA acts as a functional attenuator for translocation. J Biol Chem 281(43):32303–32309
Kruegel U, Robison B, Dange T, Kahlert G, Delaney JR, Kotireddy S, Tsuchiya M, Tsuchiyama S, Murakami CJ, Schleit J, Sutphin G, Carr D, Tar K, Dittmar G, Kaeberlein M, Kennedy BK, Schmidt M (2011) Elevated proteasome capacity extends replicative lifespan in Saccharomyces cerevisiae. PLoS Genet 7(9):e1002253
Labbadia J, Morimoto RI (2015) The biology of proteostasis in aging and disease. Annu Rev Biochem 84:435–464
Lee BP, Smith M, Buffenstein R, Harries LW (2020) Negligible senescence in naked mole rats may be a consequence of well-maintained splicing regulation. Geroscience 42(2):633–651
Leiser SF, Kaeberlein M (2010) The hypoxia-inducible factor HIF-1 functions as both a positive and negative modulator of aging. Biol Chem 391(10):1131–1137
Lewis KN, Mele J, Hornsby PJ, Buffenstein R (2012) Stress resistance in the naked mole-rat: the bare essentials – a mini-review. Gerontology 58(5):453–462
Lewis KN, Wason E, Edrey YH, Kristan DM, Nevo E, Buffenstein R (2015) Regulation of Nrf2 signaling and longevity in naturally long-lived rodents. Proc Natl Acad Sci U S A 112(12):3722–3727
Li J, Zhang Y, Li C, Xie J, Liu Y, Zhu W, Zhang X, Jiang S, Liu L, Ding Z (2013) HSPA12B attenuates cardiac dysfunction and remodelling after myocardial infarction through an eNOS-dependent mechanism. Cardiovasc Res 99(4):674–684
Li J, Labbadia J, Morimoto RI (2017) Rethinking HSF1 in stress, development, and organismal health. Trends Cell Biol 27(12):895–905
Liang S, Mele J, Wu Y, Buffenstein R, Hornsby PJ (2010) Resistance to experimental tumorigenesis in cells of a long-lived mammal, the naked mole-rat (Heterocephalus glaber). Aging Cell 9(4):626–635
Liggett A, Crawford LJ, Walker B, Morris TC, Irvine AE (2010) Methods for measuring proteasome activity: current limitations and future developments. Leuk Res 34(11):1403–1409
Lopez-Otin C, Blasco MA, Partridge L, Serrano M, Kroemer G (2013) The hallmarks of aging. Cell 153(6):1194–1217
Lowe R, Barton C, Jenkins CA, Ernst C, Forman O, Fernandez-Twinn DS, Bock C, Rossiter SJ, Faulkes CG, Ozanne SE, Walter L, Odom DT, Mellersh C, Rakyan VK (2018) Ageing-associated DNA methylation dynamics are a molecular readout of lifespan variation among mammalian species. Genome Biol 19(1):22
Lowe R, Danson AF, Rakyan VK, Yildizoglu S, Saldmann F, Viltard M, Friedlander G, Faulkes CG (2020) DNA methylation clocks as a predictor for ageing and age estimation in naked mole-rats, Heterocephalus glaber. Aging 12(5):4394–4406
Ma Y, Lu C, Li C, Li R, Zhang Y, Ma H, Zhang X, Ding Z, Liu L (2013) Overexpression of HSPA12B protects against cerebral ischemia/reperfusion injury via a PI3K/Akt-dependent mechanism. Biochim Biophys Acta 1832(1):57–66
MacRae SL, Croken MM, Calder RB, Aliper A, Milholland B, White RR, Zhavoronkov A, Gladyshev VN, Seluanov A, Gorbunova V, Zhang ZD, Vijg J (2015a) DNA repair in species with extreme lifespan differences. Aging 7(12):1171–1184
MacRae SL, Zhang Q, Lemetre C, Seim I, Calder RB, Hoeijmakers J, Suh Y, Gladyshev VN, Seluanov A, Gorbunova V, Vijg J, Zhang ZD (2015b) Comparative analysis of genome maintenance genes in naked mole rat, mouse, and human. Aging Cell 14(2):288–291
Madeo F, Zimmermann A, Maiuri MC, Kroemer G (2015) Essential role for autophagy in life span extension. J Clin Invest 125(1):85–93
Masuda K, Marasa B, Martindale JL, Halushka MK, Gorospe M (2009) Tissue- and age-dependent expression of RNA-binding proteins that influence mRNA turnover and translation. Aging 1(8):681–698
Mata J, Marguerat S, Bahler J (2005) Post-transcriptional control of gene expression: a genome-wide perspective. Trends Biochem Sci 30(9):506–514
McDonough H, Patterson C (2003) CHIP: a link between the chaperone and proteasome systems. Cell Stress Chaperones 8(4):303–308
Melen GJ, Pesce CG, Rossi MS, Kornblihtt AR (1999) Novel processing in a mammalian nuclear 28S pre-rRNA: tissue-specific elimination of an ‘intron’ bearing a hidden break site. EMBO J 18(11):3107–3118
Mitchell TW, Buffenstein R, Hulbert AJ (2007) Membrane phospholipid composition may contribute to exceptional longevity of the naked mole-rat (Heterocephalus glaber): a comparative study using shotgun lipidomics. Exp Gerontol 42(11):1053–1062
Miura K, Oiwa Y, Kawamura Y (2021) Induced pluripotent stem cells from cancer-resistant naked mole-rats. In: Buffenstein R, Park TJ, Holmes MM (eds) The Extraordinary Biology of the Naked Mole-Rat. Springer, New York, pp 329–339
Miyawaki S, Kawamura Y, Oiwa Y, Shimizu A, Hachiya T, Bono H, Koya I, Okada Y, Kimura T, Tsuchiya Y, Suzuki S, Onishi N, Kuzumaki N, Matsuzaki Y, Narita M, Ikeda E, Okanoya K, Seino K, Saya H, Okano H, Miura K (2016) Tumour resistance in induced pluripotent stem cells derived from naked mole-rats. Nat Commun 7:11471
Mordret E, Dahan O, Asraf O, Rak R, Yehonadav A, Barnabas GD, Cox J, Geiger T, Lindner AB, Pilpel Y (2019) Systematic detection of amino acid substitutions in proteomes reveals mechanistic basis of ribosome errors and selection for translation fidelity. Mol Cell 75(3):427–441
Natsidis P, Schiffer PH, Salvador-Martinez I, Telford MJ (2019) Computational discovery of hidden breaks in 28S ribosomal RNAs across eukaryotes and consequences for RNA Integrity Numbers. Sci Rep 9(1):19477
Neckers L, Workman P (2012) Hsp90 molecular chaperone inhibitors: are we there yet? Clin Cancer Res 18(1):64–76
Nguyen NN, Rana A, Goldman C, Moore R, Tai J, Hong Y, Shen J, Walker DW, Hur JH (2019) Proteasome beta5 subunit overexpression improves proteostasis during aging and extends lifespan in Drosophila melanogaster. Sci Rep 9(1):3170
Novoselov SS, Mustill WJ, Gray AL, Dick JR, Kanuga N, Kalmar B, Greensmith L, Cheetham ME (2013) Molecular chaperone mediated late-stage neuroprotection in the SOD1(G93A) mouse model of amyotrophic lateral sclerosis. PLoS One 8(8):e73944
O’Riain MJ, Jarvis JUM (1998) The dynamics of growth in naked mole-rats: the effects of litter order and changes in social structure. J Zool 246(1):49–60
Olzmann JA, Chin LS (2008) Parkin-mediated K63-linked polyubiquitination: a signal for targeting misfolded proteins to the aggresome-autophagy pathway. Autophagy 4(1):85–87
Orr ME, Garbarino VR, Salinas A, Buffenstein R (2016) Extended postnatal brain development in the longest-lived rodent: prolonged maintenance of neotenous traits in the naked mole-rat brain. Front Neurosci 10:504
Ott C, Konig J, Hohn A, Jung T, Grune T (2016) Macroautophagy is impaired in old murine brain tissue as well as in senescent human fibroblasts. Redox Biol 10:266–273
Pappireddi N, Martin L, Wuhr M (2019) A review on quantitative multiplexed proteomics. Chembiochem 20(10):1210–1224
Park TJ, Reznick J, Peterson BL, Blass G, Omerbasic D, Bennett NC, Kuich P, Zasada C, Browe BM, Hamann W, Applegate DT, Radke MH, Kosten T, Lutermann H, Gavaghan V, Eigenbrod O, Begay V, Amoroso VG, Govind V, Minshall RD, Smith ESJ, Larson J, Gotthardt M, Kempa S, Lewin GR (2017) Fructose-driven glycolysis supports anoxia resistance in the naked mole-rat. Science 356(6335):307–311
Pearl LH, Prodromou C (2006) Structure and mechanism of the Hsp90 molecular chaperone machinery. Annu Rev Biochem 75:271–294
Perez VI, Buffenstein R, Masamsetti V, Leonard S, Salmon AB, Mele J, Andziak B, Yang T, Edrey Y, Friguet B, Ward W, Richardson A, Chaudhuri A (2009) Protein stability and resistance to oxidative stress are determinants of longevity in the longest-living rodent, the naked mole-rat. Proc Natl Acad Sci 106(9):3059–3064
Perez FP, Moinuddin SS, ul ain Shamim Q, Joseph DJ, Morisaki J, Zhou X (2012) Longevity pathways: HSF1 and FoxO pathways, a new therapeutic target to prevent age-related diseases. Curr Aging Sci 5(2):87–95
Polanowski AM, Robbins J, Chandler D, Jarman SN (2014) Epigenetic estimation of age in humpback whales. Mol Ecol Resour 14(5):976–987
Powers RW 3rd, Kaeberlein M, Caldwell SD, Kennedy BK, Fields S (2006) Extension of chronological life span in yeast by decreased TOR pathway signaling. Genes Dev 20(2):174–184
Pride H, Yu Z, Sunchu B, Mochnick J, Coles A, Zhang Y, Buffenstein R, Hornsby PJ, Austad SN, Perez VI (2015) Long-lived species have improved proteostasis compared to phylogenetically-related shorter-lived species. Biochem Biophys Res Commun 457(4):669–675
Pyo JO, Yoo SM, Ahn HH, Nah J, Hong SH, Kam TI, Jung S, Jung YK (2013) Overexpression of Atg5 in mice activates autophagy and extends lifespan. Nat Commun 4:2300
Rangaraju S, Solis GM, Thompson RC, Gomez-Amaro RL, Kurian L, Encalada SE, Niculescu AB 3rd, Salomon DR, Petrascheck M (2015) Suppression of transcriptional drift extends C. elegans lifespan by postponing the onset of mortality. elife 4:e08833
Reddy NM, Kleeberger SR, Yamamoto M, Kensler TW, Scollick C, Biswal S, Reddy SP (2007) Genetic dissection of the Nrf2-dependent redox signaling-regulated transcriptional programs of cell proliferation and cytoprotection. Physiol Genomics 32(1):74–81
Richter K, Haslbeck M, Buchner J (2010) The heat shock response: life on the verge of death. Mol Cell 40(2):253–266
Ristow M, Schmeisser K (2014) Mitohormesis: promoting health and lifespan by increased levels of reactive oxygen species (ROS). Dose-response 12(2):288–341
Rivett AJ, Savory PJ, Djaballah H (1994) Multicatalytic endopeptidase complex: proteasome. Methods Enzymol 244:331–350
Rodriguez KA, Wywial E, Perez VI, Lambert AJ, Edrey YH, Lewis KN, Grimes K, Lindsey ML, Brand MD, Buffenstein R (2011) Walking the oxidative stress tightrope: a perspective from the naked mole-rat, the longest-living rodent. Curr Pharm Des 17(22):2290–2307
Rodriguez KA, Edrey YH, Osmulski P, Gaczynska M, Buffenstein R (2012) Altered composition of liver proteasome assemblies contributes to enhanced proteasome activity in the exceptionally long-lived naked mole-rat. PLoS One 7(5):e35890
Rodriguez KA, Osmulski PA, Pierce A, Weintraub ST, Gaczynska M, Buffenstein R (2014) A cytosolic protein factor from the naked mole-rat activates proteasomes of other species and protects these from inhibition. Biochim Biophys Acta 1842(11):2060–2072
Rodriguez KA, Valentine JM, Kramer DA, Gelfond JA, Kristan DM, Nevo E, Buffenstein R (2016) Determinants of rodent longevity in the chaperone-protein degradation network. Cell Stress Chaperones 21(3):453–466
Rose C, Vargas F, Facchinetti P, Bourgeat P, Bambal RB, Bishop PB, Chan SM, Moore AN, Ganellin CR, Schwartz JC (1996) Characterization and inhibition of a cholecystokinin-inactivating serine peptidase. Nature 380(6573):403–409
Rosenzweig R, Nillegoda NB, Mayer MP, Bukau B (2019) The Hsp70 chaperone network. Nat Rev Mol Cell Biol 20(11):665–680
Rothbard JB, Kurnellas MP, Brownell S, Adams CM, Su L, Axtell RC, Chen R, Fathman CG, Robinson WH, Steinman L (2012) Therapeutic effects of systemic administration of chaperone alphaB-crystallin associated with binding proinflammatory plasma proteins. J Biol Chem 287(13):9708–9721
Rubinsztein DC, Marino G, Kroemer G (2011) Autophagy and aging. Cell 146(5):682–695
Ruby JG, Smith M, Buffenstein R (2018) Naked mole-rat mortality rates defy Gompertzian laws by not increasing with age. elife 7:e31157
Salway KD, Gallagher EJ, Page MM, Stuart JA (2011) Higher levels of heat shock proteins in longer-lived mammals and birds. Mech Ageing Dev 132(6-7):287–297
Selman C, Tullet JM, Wieser D, Irvine E, Lingard SJ, Choudhury AI, Claret M, Al-Qassab H, Carmignac D, Ramadani F, Woods A, Robinson IC, Schuster E, Batterham RL, Kozma SC, Thomas G, Carling D, Okkenhaug K, Thornton JM, Partridge L, Gems D, Withers DJ (2009) Ribosomal protein S6 kinase 1 signaling regulates mammalian life span. Science 326(5949):140–144
Seluanov A, Chen Z, Hine C, Sasahara TH, Ribeiro AA, Catania KC, Presgraves DC, Gorbunova V (2007) Telomerase activity coevolves with body mass not lifespan. Aging Cell 6(1):45–52
Seluanov A, Hine C, Bozzella M, Hall A, Sasahara TH, Ribeiro AA, Catania KC, Presgraves DC, Gorbunova V (2008) Distinct tumor suppressor mechanisms evolve in rodent species that differ in size and lifespan. Aging Cell 7(6):813–823
Seluanov A, Hine C, Azpurua J, Feigenson M, Bozzella M, Mao Z, Catania KC, Gorbunova V (2009) Hypersensitivity to contact inhibition provides a clue to cancer resistance of naked mole-rat. Proc Natl Acad Sci U S A 106(46):19352–19357
Seluanov A, Gladyshev VN, Vijg J, Gorbunova V (2018) Mechanisms of cancer resistance in long-lived mammals. Nat Rev Cancer 18(7):433–441
Simonsen A, Cumming RC, Brech A, Isakson P, Schubert DR, Finley KD (2008) Promoting basal levels of autophagy in the nervous system enhances longevity and oxidant resistance in adult Drosophila. Autophagy 4(2):176–184
Smith-Vikos T, Slack FJ (2012) MicroRNAs and their roles in aging. J Cell Sci 125(Pt 1):7–17
Southworth LK, Owen AB, Kim SK (2009) Aging mice show a decreasing correlation of gene expression within genetic modules. PLoS Genet 5(12):e1000776
Steffen KK, Dillin A (2016) A ribosomal perspective on proteostasis and aging. Cell Metab 23(6):1004–1012
Stubbs TM, Bonder MJ, Stark AK, Krueger F, Team BIAC, von Meyenn F, Stegle O, Reik W (2017) Multi-tissue DNA methylation age predictor in mouse. Genome Biol 18(1):68
Suarez-Cabrera C, de la Pena B, Gonzalez LL, Page A, Martinez-Fernandez M, Casanova ML, Paramio JM, Rojo-Sebastian A, Moreno-Bueno G, Maroto A, Ramirez A, Navarro M (2018) The Ras-related gene ERAS is involved in human and murine breast cancer. Sci Rep 8(1):13038
Swovick K, Welle KA, Hryhorenko JR, Seluanov A, Gorbunova V, Ghaemmaghami S (2018) Cross-species comparison of proteome turnover kinetics. Mol Cell Proteomics 17(4):580–591
Tacutu R, Thornton D, Johnson E, Budovsky A, Barardo D, Craig T, Diana E, Lehmann G, Toren D, Wang J, Fraifeld VE, de Magalhaes JP (2018) Human ageing genomic resources: new and updated databases. Nucleic Acids Res 46(D1):D1083–D1090
Tan L, Ke Z, Tombline G, Macoretta N, Hayes K, Tian X, Lv R, Ablaeva J, Gilbert M, Bhanu NV, Yuan ZF, Garcia BA, Shi YG, Shi Y, Seluanov A, Gorbunova V (2017) Naked mole rat cells have a stable epigenome that resists iPSC reprogramming. Stem Cell Rep 9(5):1721–1734
Taylor RC, Dillin A (2011) Aging as an event of proteostasis collapse. Cold Spring Harb Perspect Biol 3(5):a004440
Taylor KR, Milone NA, Rodriguez CE (2017) Four cases of spontaneous neoplasia in the naked mole-rat (Heterocephalus glaber), a putative cancer-resistant species. J Gerontol A Biol Sci Med Sci 72(1):38–43
Thompson A, Schafer J, Kuhn K, Kienle S, Schwarz J, Schmidt G, Neumann T, Johnstone R, Mohammed AK, Hamon C (2003) Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal Chem 75(8):1895–1904
Thompson MJ, vonHoldt B, Horvath S, Pellegrini M (2017) An epigenetic aging clock for dogs and wolves. Aging (Albany NY) 9(3):1055–1068
Thomson E, Ferreira-Cerca S, Hurt E (2013) Eukaryotic ribosome biogenesis at a glance. J Cell Sci 126(21):4815–4821
Tian X, Azpurua J, Hine C, Vaidya A, Myakishev-Rempel M, Ablaeva J, Mao Z, Nevo E, Gorbunova V, Seluanov A (2013) High-molecular-mass hyaluronan mediates the cancer resistance of the naked mole rat. Nature 499(7458):346–349
Tian X, Azpurua J, Ke Z, Augereau A, Zhang ZD, Vijg J, Gladyshev VN, Gorbunova V, Seluanov A (2015) INK4 locus of the tumor-resistant rodent, the naked mole rat, expresses a functional p15/p16 hybrid isoform. Proc Natl Acad Sci U S A 112(4):1053–1058
Tong KI, Kobayashi A, Katsuoka F, Yamamoto M (2006) Two-site substrate recognition model for the Keap1-Nrf2 system: a hinge and latch mechanism. Biol Chem 387(10–11):1311–1320
Triplett JC, Tramutola A, Swomley A, Kirk J, Grimes K, Lewis K, Orr M, Rodriguez K, Cai J, Klein JB, Perluigi M, Buffenstein R, Butterfield DA (2015) Age-related changes in the proteostasis network in the brain of the naked mole-rat: implications promoting healthy longevity. Biochim Biophys Acta 1852(10A):2213–2224
Vilchez D, Morantte I, Liu Z, Douglas PM, Merkwirth C, Rodrigues AP, Manning G, Dillin A (2012) RPN-6 determines C. elegans longevity under proteotoxic stress conditions. Nature 489(7415):263–268
Vilchez D, Saez I, Dillin A (2014) The role of protein clearance mechanisms in organismal ageing and age-related diseases. Nat Commun 5:5659
Wagner A (2005) Energy constraints on the evolution of gene expression. Mol Biol Evol 22(6):1365–1374
Walerych D, Kudla G, Gutkowska M, Wawrzynow B, Muller L, King FW, Helwak A, Boros J, Zylicz A, Zylicz M (2004) Hsp90 chaperones wild-type p53 tumor suppressor protein. J Biol Chem 279(47):48836–48845
Webster JM, Darling AL, Uversky VN, Blair LJ (2019) Small heat shock proteins, big impact on protein aggregation in neurodegenerative disease. Front Pharmacol 10:1047
Wei YN, Hu HY, Xie GC, Fu N, Ning ZB, Zeng R, Khaitovich P (2015) Transcript and protein expression decoupling reveals RNA binding proteins and miRNAs as potential modulators of human aging. Genome Biol 16:41
Williams AB, Schumacher B (2016) p53 in the DNA-damage-repair process. Cold Spring Harb Perspect Med 6(5):a026070
Wilson DN, Doudna Cate JH (2012) The structure and function of the eukaryotic ribosome. Cold Spring Harb Perspect Biol 4(5):4(5):a011536
Xiao B, Wang S, Yang G, Sun X, Zhao S, Lin L, Cheng J, Yang W, Cong W, Sun W, Kan G, Cui S (2017a) HIF-1alpha contributes to hypoxia adaptation of the naked mole rat. Oncotarget 8(66):109941–109951
Xiao B, Li L, Xu C, Zhao S, Lin L, Cheng J, Yang W, Cong W, Kan G, Cui S (2017b) Transcriptome sequencing of the naked mole rat (Heterocephalus glaber) and identification of hypoxia tolerance genes. Biol Open 6(12):1904–1912
Xie J, de Souza Alves V, von der Haar T, O’Keefe L, Lenchine RV, Jensen KB, Liu R, Coldwell MJ, Wang X, Proud CG (2019) Regulation of the elongation phase of protein synthesis enhances translation accuracy and modulates lifespan. Curr Biol 29(5):737–749
Yang SB, Tien AC, Boddupalli G, Xu AW, Jan YN, Jan LY (2012) Rapamycin ameliorates age-dependent obesity associated with increased mTOR signaling in hypothalamic POMC neurons. Neuron 75(3):425–436
Yoon MS (2017) The role of mammalian target of rapamycin (mTOR) in insulin signaling. Nutrients 9(11):1176
Yuno A, Lee MJ, Lee S, Tomita Y, Rekhtman D, Moore B, Trepel JB (2018) Clinical evaluation and biomarker profiling of Hsp90 inhibitors. Methods Mol Biol 1709:423–441
Yusupova G, Yusupov M (2014) High-resolution structure of the eukaryotic 80S ribosome. Annu Rev Biochem 83:467–486
Zaher HS, Green R (2009) Fidelity at the molecular level: lessons from protein synthesis. Cell 136(4):746–762
Zhang Y, Bokov A, Gelfond J, Soto V, Ikeno Y, Hubbard G, Diaz V, Sloane L, Maslin K, Treaster S, Rendon S, van Remmen H, Ward W, Javors M, Richardson A, Austad SN, Fischer K (2014) Rapamycin extends life and health in C57BL/6 mice. J Gerontol A Biol Sci Med Sci 69(2):119–130
Zhang MW, Zhao P, Yung WH, Sheng Y, Ke Y, Qian ZM (2018) Tissue iron is negatively correlated with TERC or TERT mRNA expression: a heterochronic parabiosis study in mice. Aging 10(12):3834–3850
Zhang P, Wu W, Chen Q, Chen M (2019) Non-coding RNAs and their integrated networks. J Integr Bioinform 16(3):20190027
Zhao S, Lin L, Kan G, Xu C, Tang Q, Yu C, Sun W, Cai L, Xu C, Cui S (2014) High autophagy in the naked mole rat may play a significant role in maintaining good health. Cell Physiol Biochem 33(2):321–332
Acknowledgements
We gratefully acknowledge Calico Life Sciences LLC for funding this work. We also thank David Botstein, Yao Wong, Ewan St. John Smith and Thomas Park for their constructive critique of this manuscript.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Narayan, V., McMahon, M., O’Brien, J.J., McAllister, F., Buffenstein, R. (2021). Insights into the Molecular Basis of Genome Stability and Pristine Proteostasis in Naked Mole-Rats. In: Buffenstein, R., Park, T.J., Holmes, M.M. (eds) The Extraordinary Biology of the Naked Mole-Rat. Advances in Experimental Medicine and Biology, vol 1319. Springer, Cham. https://doi.org/10.1007/978-3-030-65943-1_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-65943-1_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-65942-4
Online ISBN: 978-3-030-65943-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)