Abstract
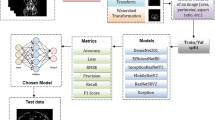
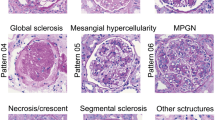
Recent breakthroughs in the classification of thousands of images into defined categories by deep learning has led to the application of these algorithms and models to the field of medicine, particularly in histology and pathology. The evaluation of histological images obtained by a kidney biopsy is a key step in the diagnosis and assessment of disease status in the practice of clinical nephrology. Many studies have investigated how deep learning can be applied to nephropathology to evaluate complex structures present inside the kidneys. In this chapter, we provide a brief discussion on nephropathology and introduce some of the most recent studies on artificial intelligence involving this field. Additionally, we also discuss about the implications of and challenges associated with the application of AI to nephropathology. The studies are divided into three subcategories based on the key objective of the presented study. These include (i) identifying the position of specific structures, especially the glomerulus (detection), (ii) segmentation and identification of various structures present in histological images (segmentation), and (iii) classification of specific images into clinically and pathologically defined categories (classification).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Jayapandian CP, Chen Y, Janowczyk AR, Palmer MB, Cassol CA, Sekulic M, et al. Development and evaluation of deep learning-based segmentation of histologic structures in the kidney cortex with multiple histologic stains. Kidney Int. 2021;99(1):86–101. https://doi.org/10.1016/j.kint.2020.07.044.
Bellur SS, Roberts ISD, Troyanov S, Royal V, Coppo R, Cook HT, et al. Reproducibility of the Oxford classification of immunoglobulin A nephropathy, impact of biopsy scoring on treatment allocation and clinical relevance of disagreements: evidence from the VALidation of IGA study cohort. Nephrol Dial Transplant. 2019;34(10):1681–90. https://doi.org/10.1093/ndt/gfy337.
Barisoni L, Troost JP, Nast C, Bagnasco S, Avila-Casado C, Hodgin J, et al. Reproducibility of the NEPTUNE descriptor-based scoring system on whole-slide images and histologic and ultrastructural digital images. Mod Pathol. 2016;29(7):671–84. https://doi.org/10.1038/modpathol.2016.58.
Loupy A, Haas M, Roufosse C, Naesens M, Adam B, Afrouzian M, et al. The Banff 2019 Kidney Meeting Report (I): Updates on and clarification of criteria for T cell- and antibody-mediated rejection. Am J Transplant. 2020;20(9):2318–31. https://doi.org/10.1111/ajt.15898.
Samsi S, Jarjour WN, Krishnamurthy A. Glomeruli segmentation in H&E stained tissue using perceptual organization. In: 2012 IEEE Signal Processing in Medicine and Biology Symposium (SPMB). 2012. p. 1–5. https://doi.org/10.1109/SPMB.2012.6469464.
Kakimoto T, Okada K, Hirohashi Y, Relator R, Kawai M, Iguchi T, et al. Automated image analysis of a glomerular injury marker desmin in spontaneously diabetic Torii rats treated with losartan. J Endocrinol. 2014;222(1):43–51. https://doi.org/10.1530/JOE-14-0164.
Kato T, Relator R, Ngouv H, Hirohashi Y, Takaki O, Kakimoto T, et al. Segmental HOG: new descriptor for glomerulus detection in kidney microscopy image. BMC Bioinformatics. 2015;16:316. https://doi.org/10.1186/s12859-015-0739-1.
Simon O, Yacoub R, Jain S, Tomaszewski JE, Sarder P. Multi-radial LBP Features as a Tool for Rapid Glomerular Detection and Assessment in Whole Slide Histopathology Images. Sci Rep. 2018;8(1):2032. https://doi.org/10.1038/s41598-018-20453-7.
Marée R, Dallongeville S, Olivo-Marin J, Meas-Yedid V. An approach for detection of glomeruli in multisite digital pathology. In: 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI). 2016. p. 1033–6. https://doi.org/10.1109/ISBI.2016.7493442.
Temerinac-Ott M, Forestier G, Schmitz J, Hermsen M, Bräsen JH, Feuerhake F, et al. Detection of glomeruli in renal pathology by mutual comparison of multiple staining modalities. In: Proceedings of the 10th International Symposium on Image and Signal Processing and Analysis. 2017. p. 19–24. https://doi.org/10.1109/ISPA.2017.8073562.
Bukowy JD, Dayton A, Cloutier D, Manis AD, Staruschenko A, Lombard JH, et al. Region-based convolutional neural nets for localization of glomeruli in trichrome-stained whole kidney sections. J Am Soc Nephrol. 2018;29(8):2081–8. https://doi.org/10.1681/ASN.2017111210.
Girshick R, Donahue J, Darrell T, Malik J. Rich feature hierarchies for accurate object detection and semantic segmentation. In: 2014 IEEE Conference on Computer Vision and Pattern Recognition. 2014. p. 580–7. https://doi.org/10.1109/CVPR.2014.81.
Gadermayr M, Klinkhammer BM, Boor P, Merhof D. Do we need large annotated training data for detection applications in biomedical imaging? A case study in renal glomeruli detection. in: machine learning in medical imaging. Springer International Publishing; 2016. p. 18–26. https://doi.org/10.1007/978-3-319-47157-0_3.
Gadermayr M, Eschweiler D, Jeevanesan A, Klinkhammer BM, Boor P, Merhof D. Segmenting renal whole slide images virtually without training data. Comput Biol Med. 2017;90:88–97. https://doi.org/10.1016/j.compbiomed.2017.09.014.
Gadermayr M, Dombrowski A-K, Klinkhammer BM, Boor P, Merhof D. CNN cascades for segmenting sparse objects in gigapixel whole slide images. Comput Med Imaging Graph. 2019;71:40–8. https://doi.org/10.1016/j.compmedimag.2018.11.002.
Gadermayr M, Gupta L, Appel V, Boor P, Klinkhammer BM, Merhof D. Generative adversarial networks for facilitating stain-independent supervised and unsupervised segmentation: a study on kidney histology. IEEE Trans Med Imaging. 2019;38(10):2293–302. https://doi.org/10.1109/TMI.2019.2899364.
Bueno G, Fernandez-Carrobles MM, Gonzalez-Lopez L, Deniz O. Glomerulosclerosis identification in whole slide images using semantic segmentation. Comput Methods Programs Biomed. 2020;184:105273. https://doi.org/10.1016/j.cmpb.2019.105273.
Ronneberger O, Fischer P, Brox T. U-Net: convolutional networks for biomedical image segmentation. In: Medical image computing and computer-assisted intervention – MICCAI 2015. Springer International Publishing; 2015. p. 234–41. https://doi.org/10.1007/978-3-319-24574-4_28.
Badrinarayanan V, Kendall A, Cipolla R. SegNet: a deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans Pattern Anal Mach Intell. 2017;39(12):2481–95. https://doi.org/10.1109/TPAMI.2016.2644615.
Bueno G, Gonzalez-Lopez L, Garcia-Rojo M, Laurinavicius A, Deniz O. Data for glomeruli characterization in histopathological images. Data Brief. 2020;29:105314. https://doi.org/10.1016/j.dib.2020.105314.
Bouteldja N, Klinkhammer BM, Bülow RD, Droste P, Otten SW, von Stillfried SF, et al. Deep learning–based segmentation and quantification in experimental kidney histopathology. J Am Soc Nephrol. 2021;32(1):52–68. https://doi.org/10.1681/ASN.2020050597.
Ginley B, Jen K-Y, Han SS, Rodrigues L, Jain S, Fogo AB, et al. Automated computational detection of interstitial fibrosis, tubular atrophy, and glomerulosclerosis. J Am Soc Nephrol. 2021;32(4):837-50. https://doi.org/10.1681/ASN.2020050652.
Barros GO, Navarro B, Duarte A, Dos-Santos WLC. PathoSpotter-K: a computational tool for the automatic identification of glomerular lesions in histological images of kidneys. Sci Rep. 2017;7:46769. https://doi.org/10.1038/srep46769.
Sheehan SM, Korstanje R. Automatic glomerular identification and quantification of histological phenotypes using image analysis and machine learning. Am J Physiol Renal Physiol. 2018;315(6):F1644–51. https://doi.org/10.1152/ajprenal.00629.2017.
Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, et al. ImageNet large scale visual recognition challenge. Int J Comput Vis. 2015;115(3):211–52. https://doi.org/10.1007/s11263-015-0816-y.
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z. Rethinking the inception architecture for computer vision. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR). 2016. p. 2818–26. https://doi.org/10.1109/CVPR.2016.308.
Kannan S, Morgan LA, Liang B, Cheung MG, Lin CQ, Mun D, et al. Segmentation of glomeruli within trichrome images using deep learning. Kidney Int Rep. 2019;4(7):955–62. https://doi.org/10.1016/j.ekir.2019.04.008.
Marsh JN, Matlock MK, Kudose S, Liu T-C, Stappenbeck TS, Gaut JP, et al. Deep learning global glomerulosclerosis in transplant kidney frozen sections. IEEE Trans Med Imaging. 2018;37(12):2718–28. https://doi.org/10.1109/TMI.2018.2851150.
Wu B, Zhang X, Zhao S, Xie L, Zeng C, Liu Z, et al. G2C: a generator-to-classifier framework integrating multi-stained visual cues for pathological glomerulus classification. Proceedings of the AAAI Conference on Artificial Intelligence. 2019;33(01):1214–21. https://doi.org/10.1609/aaai.v33i01.33011214.
Uchino E, Suzuki K, Sato N, Kojima R, Tamada Y, Hiragi S, et al. Classification of glomerular pathological findings using deep learning and nephrologist-AI collective intelligence approach. Int J Med Inform. 2020;141:104231. https://doi.org/10.1016/j.ijmedinf.2020.104231.
Yamaguchi R, Kawazoe Y, Shimamoto K, Shinohara E, Tsukamoto T, Shintani-Domoto Y, et al. Glomerular classification using convolutional neural networks based on defined annotation criteria and concordance evaluation among clinicians. Kidney Int Rep. 2021;6(3):716–26. https://doi.org/10.1016/j.ekir.2020.11.037.
Chagas P, Souza L, Araújo I, Aldeman N, Duarte A, Angelo M, et al. Classification of glomerular hypercellularity using convolutional features and support vector machine. Artif Intell Med. 2020;103:101808. https://doi.org/10.1016/j.artmed.2020.101808.
Zeng C, Nan Y, Xu F, Lei Q, Li F, Chen T, et al. Identification of glomerular lesions and intrinsic glomerular cell types in kidney diseases via deep learning. J Pathol. 2020;252(1):53-64. https://doi.org/10.1002/path.5491.
Chen Y, Li M, Hao F, Han W, Niu D, Wang C. Classification of glomerular spikes using Convolutional Neural Network. In: Proceedings of the 2020 Conference on Artificial Intelligence and Healthcare. New York, NY, USA: Association for Computing Machinery; 2020. p. 254–8. https://doi.org/10.1145/3433996.3434043.
Dietterich TG, Lathrop RH, Lozano-Pérez T. Solving the multiple instance problem with axis-parallel rectangles. Artif Intell. 1997;89(1):31–71. https://doi.org/10.1016/S0004-3702(96)00034-3.
Ginley B, Lutnick B, Jen K-Y, Fogo AB, Jain S, Rosenberg A, et al. Computational segmentation and classification of diabetic glomerulosclerosis. J Am Soc Nephrol. 2019 Oct;30(10):1953–67. https://doi.org/10.1681/ASN.2018121259.
Ginley B, Jen K-Y, Rosenberg A, Rossi GM, Jain S, Sarder P. Fully automated classification of glomerular lesions in lupus nephritis. In: Medical Imaging 2020: Digital Pathology. International Society for Optics and Photonics; 2020. p. 113200Y. https://doi.org/10.1117/12.2548528.
Ligabue G, Pollastri F, Fontana F, Leonelli M, Furci L, Giovanella S, et al. Evaluation of the classification accuracy of the kidney biopsy direct immunofluorescence through convolutional neural networks. Clin J Am Soc Nephrol. 2020;15(10):1445–54. https://doi.org/10.2215/CJN.03210320.
Choi G, Kim Y-G, Cho H, Kim N, Lee H, Moon KC, et al. Automated detection algorithm for C4d immunostaining showed comparable diagnostic performance to pathologists in renal allograft biopsy. Mod Pathol. 2020;33(8):1626–34. https://doi.org/10.1038/s41379-020-0529-9.
Kolachalama VB, Singh P, Lin CQ, Mun D, Belghasem ME, Henderson JM, et al. Association of pathological fibrosis with renal survival using deep neural networks. Kidney Int Rep. 2018;3(2):464–75. https://doi.org/10.1016/j.ekir.2017.11.002.
Sheehan S, Mawe S, Cianciolo RE, Korstanje R, Mahoney JM. Detection and classification of novel renal histologic phenotypes using deep neural networks. Am J Pathol. 2019;189(9):1786–96. https://doi.org/10.1016/j.ajpath.2019.05.019.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this entry
Cite this entry
Noriaki, S., Eiichiro, U., Yasushi, O. (2022). Artificial Intelligence in Kidney Pathology. In: Lidströmer, N., Ashrafian, H. (eds) Artificial Intelligence in Medicine. Springer, Cham. https://doi.org/10.1007/978-3-030-64573-1_181
Download citation
DOI: https://doi.org/10.1007/978-3-030-64573-1_181
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-64572-4
Online ISBN: 978-3-030-64573-1
eBook Packages: MedicineReference Module Medicine