Abstract
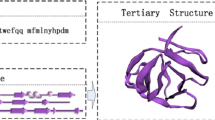
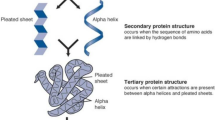
The significance of determining the structure of proteins comes up from the importance of their role in the body, making proteins the primary target of drugs and key to developing new drugs. There are several experimental methods to determine PS ‘Protein structure’, but a reliable and direct prediction method to determine PS is not yet available. Hence, it was necessary to use intelligent systems that predicted PS based on the sequence of amino acids. Predicting the SSP “secondary structure of protein” is a very important step. It has increasingly become the basis for a number of ways to predict protein structure and thus know its function.
The aim of this research is to design and implement a high-performance method for predicting SSP from the AAS “amino acid sequence” and calculating prediction accuracy based on recorded results. To achieve this goal, the work was done in two phases: first, to construct an SSP system, based solely on the AAS for the protein using ANFIS “Adaptive Neuro-Fuzzy Inference Systems” This system achieves accuracy of up to 65% and is a good accuracy compared to many SSP systems.
The second stage was the construction of two SSP systems to study the effect of coding the data used on the system’s input on its accuracy. Both systems had same designs and structures using artificial neural network. The input in the second is coded using Profiles, which showed a significant improvement in system accuracy exceeded 10%, bringing the accuracy of total prediction up to 70.5%.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aliev, R.A., Fazlollahi, B., Aliev, R.R.: Soft Computing and its Application in Business and Economics, vol. 157, p. 110. Springer, Heidelberg (2004). https://doi.org/10.1007/978-3-540-44429-9
Zadeh, L.A.: Fuzzy set. Inform. Control 8(3), 338–353 (1965). https://doi.org/10.2307/2272014
Maind, S.B., Wankar, P.: Research paper on basic of artificial neural network. Int. J. Recent Innov. Trends Comput. Commun. 2(1), 96–100 (2014). https://doi.org/10.17762/ijritcc.v2i1.2920
Bibel, W.: On a scientific discipline (once) named AI. In: IJCAI, pp. 5143–5149 (2018)
Maass, W.: Networks of spiking neurons: the third generation of neural network models. Neural Netw. 10(9), 1659–1671 (1997). https://doi.org/10.1016/S0893-6080(97)00011-7
Ni, X.: Research of data mining based on neural networks. World Acad. Sci. Eng. Technol. 39, 381–384 (2008)
Zadeh, L.A.: Fuzzy logic. Computer 21(4), 83–93 (1988). https://doi.org/10.1109/2.53
Yardimci, A.: Application of soft computing to medical problems. In: International CONFERENCE 2009 on Intelligent System Design and Applications, pp. 614–619 (2009). https://doi.org/10.1109/ISDA.2009.168
Hunter, L.: Molecular biology for computer scientists, pp. 18–24 (2014)
Clark, D.P., Pazdernik, N.J., McGehee, M.: Molecular Biology. Elsevier Science, Amsterdam (2018)
Lodish, H., Berk, A.: Molecular Cell Biology. WH Freeman and Company, New York (2004)
Voet, D., Voet, J., Pratt, C.: Principles of Biochemistry. 3rd International student edition edn. Wiley, New York (2009)
Miller, N.: The misfolding diseases unfold. Beremans, UK (2011)
Chaudhuri, T.K., Paul, S.: Protein-misfolding diseases and chaperone-based therapeutic approaches. FEBS J. 273(7), 1331–1349 (2006). https://doi.org/10.1111/j.1742-4658.2006.05181.x
Khan, S., Vihinen, M.: Spectrum of disease-causing mutations in protein secondary structures. BMC Struct. Biol. 7(1) (2007). https://doi.org/10.1186/1472-6807-7-56
Imanov, E., Anwar, A.: Artificial neural network for the left ventricle detection. In: 10th International Conference 2019 on Theory and Application of Soft Computing, Computing with Words and Perceptions (2019)
Sharma, V., Rai, S., Dev, A.: A comprehensive study of artificial neural networks. Int. J. Adv. Res. Comput. Sci. Softw. Eng. 2(10), 278–284 (2012)
Wei, M., Bai, B., Sung, A.H., Liu, Q., Wang, J., Cather, M.E.: Predicting injection profiles using ANFIS. Inform. Sci. 177(20), 4445–4461 (2007). https://doi.org/10.1016/j.ins.2007.03.021
Pollastri, G., Przybylski, D., Rost, B., Baldi, P.: Improving the prediction of protein secondary structure in three and eight classes using recurrent neural networks and profiles. Proteins: Struct. Funct. Bioinf. 47(2), 228–235 (2002)
Rost, B., Sander, C.: Prediction of protein secondary structure at better than 70% accuracy. J. Mol. Biol. 232(2), 548–599 (1993)
Gribskov, M., Mclachlant, A.D., Eisenberg, D.: Profile analysis: detection of distantly related proteins. Proc. Natl. Acad. Sci. 84(13), 4355–4358 (1987). https://doi.org/10.1073/pnas.84.13.4355
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Imanov, E., Shaheen, R. (2021). Soft Computing for Prediction of Secondary Structure of the Protein. In: Aliev, R.A., Kacprzyk, J., Pedrycz, W., Jamshidi, M., Babanli, M., Sadikoglu, F.M. (eds) 14th International Conference on Theory and Application of Fuzzy Systems and Soft Computing – ICAFS-2020 . ICAFS 2020. Advances in Intelligent Systems and Computing, vol 1306. Springer, Cham. https://doi.org/10.1007/978-3-030-64058-3_56
Download citation
DOI: https://doi.org/10.1007/978-3-030-64058-3_56
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-64057-6
Online ISBN: 978-3-030-64058-3
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)