Abstract
Extracellular vesicles (EV) are small membrane-coated structures secreted by all cells of the body and can be detected in all bodily fluids, including urine. EV contents (e.g. proteins and distinct RNA classes) reflect the physiological state of their cells of origin, offering a new source of biomarkers. Accordingly, urinary Extracellular Vesicles (uEVs) are emerging as a source for early biomarkers of kidney damage and beyond, holding the potential to replace the conventional invasive techniques including kidney biopsy. However, the lack of standardization and sample collection and isolation methods, and the influence of factors such as inter- and intra-individual variability create difficulties in interpreting current results. Here we review recent results and reported uses of especially urinary EVs and also pinpoint approaches to be considered when designing experiments.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction
While the history of urinary Extracellular Vesicles (uEV) is relatively young, knowledge of their biologic roles appear to grossly follow those of vesicles found in other bodily fluids.
With the already impressive impact list of EVs, it is fair to state that these abundant vesicular structures once considered as handy waste packages to be excreted out of the body, now appear to open new understanding to many key biological phenomena. These include, but not limited to, general cell-to-cell communication and regulation of immune reaction and extends to roles in spread of cancer cells to precise modulation of tissue -and cell-type specific functions.
Detailed information of molecular mechanisms mediated by EVs and their “holistic” understanding in distinct tissue functions is rapidly emerging. Notably, many of these mechanisms have now been established and appear to parallel generic EV functions in different biological fluids. Interestingly, many recent reports have highlighted the potential of customized EVs as targeted future “magic bullets” in precision medicine to enable advanced personalized and tissue targeted treatments. EVs are lucrative sources of personalized disease biomarkers, snapshots of cellular and tissue pathophysiology. These snapshots provide unforeseen accuracy to monitor distinct functions and, more importantly, to highlight new understanding of basic cellular biological events.
Reproducible use of extracellular vesicles in academic or applied research for clinically relevant problems still needs stringent standardization at many levels including sample collection, storage, EV analytics and downstream applications. In spite of valuable standardization guidelines already achieved, with particular and continued efforts by the International Society for Extracellular Vesicles [1], many important steps remain to be achieved and strict adherence to published guidelines enforced. During the rapid expansion of EV applications in a multitude of areas and from an ever increasing number of laboratories, the sheer credibility and reproducibility of results will need much stronger attention. Before EVs can be adopted in wide general use, especially in the biomarker applications, several fundamental limitations and controversies still exist.
Here we review recent results and reported uses of especially urinary EVs and also pinpoint approaches to be considered when designing experiments.
3.2 Urinary EV Biology
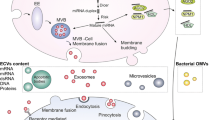
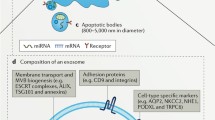
Extracellular vesicles are well characterized to their main categories of exosomes, microvesicles and apoptotic bodies (Fig. 3.1). This division is somewhat arbitrary and mostly considers the biogenesis and size of vesicles, without going into other physicochemical features nor into their precise biochemical characteristics. These basic EV properties have been well described several recent excellent reviews [2,3,4,5,6].
The milestone report of urinary EVs by Pisitkun et al. 2004 translated the physicochemical features of EVs also into urine, including their size, shape and contents. Thus, it appears that uEVs show the similar variety of protein, RNA and lipid content as EVs from other biological sources. This suggests but does not prove that EV functions in the distinct sources are identical.
Traditional view tells that urine is the vehicle to excrete metabolic waste products from not only the kidney tissue but, via glomerular filtration and tubular secretion, from the whole body. It is becoming increasingly evident, mostly by uEV studies, that urine is far beyond simple means to excrete waste from the body but indeed is a complex, poorly understood mixture of different biologically active molecular species. Notably, urinary contents dynamically reflect metabolic extremes within an individual, e.g. after strong physical exercise leading to dehydration [7]. These extremes result in well-established changes in urinary osmolality as well as in some key urinary “normal contents”. While uEV changes could reflect individual physiological adaptation to stress situations at the organ level, it is also evident that such changes are similarly seen following shift in e.g. dietary patterns, disease states and upon introduction of new medication. This means that uEV content reflects more the total body reaction and implies that all uEV clinical studies should also contain comprehensive reporting of clinical chemistry values as well as activity, diet, and, particularly, use of medications.
With the uEV contents of specific proteins beyond the kidney, it is conceivable that uEVs indeed reflect systemic rather than merely local tissue level physiology. How EVs from circulation get access to urine remains an open question but could involve transglomerular passage through the glomerular filtration barrier (GFB) or active secretion by tubular epithelial cells.
Normal urine contains cell type-specific proteins from the glomerular filtration barrier, particularly the glomerular visceral epithelial cells (podocytes) [8], and electron microscopy of glomeruli also show abundance of vesicles in this site [9]. Final proof visualizing EVs passaging the GFB remain to be achieved.
EV contents in general include specific functional, structural and metabolic proteins, lipids and, particularly, a rich content of RNA species (mi/messenger/lncRNA) and DNA [4, 10]. uEVs show a closely similar repertoire of molecules [6]. While their respective physiologic significance remains still poorly studied, it is tempting to speculate that these active molecules continue to have their respective functions also in urine. In this respect, EV surface molecules could serve as address codes to target their active contents to specific downstream sites [8]. However, further consensus needs to be reached on the methods to isolate and characterize the EV surface contents as passive adsorption of a variety of molecules in this site is possible [11, 12]. After targeting and adhesion, EVs may be taken up by a variety of well described mechanisms [13,14,15] to subsequently induce functional changes [5].
It is interesting to note that diet [16], use of medication, exercise and other still poorly understood factors [17,18,19] may cause intra-individual changes in vesicle content as detected by proteomics or RNomics. Our own results indicate significant intra-individual variation in first morning urine uEV size, class and especially contents distribution during daily monitoring over 3 months of first morning urine voids (Holthofer et al., unpublished).
Our recent results have shown distinct gender and ethnic differences in uEV contents of healthy subjects, which may be only partly explained by dietary differences (Xu et al., unpublished). These results emphasize the crucial importance of recording comprehensive phenotypic data of subjects under study to understand the dynamic changes in uEV signatures and factors behind. With this complexity in mind, our approach to understand biological roles of uEVs has consisted of using three pronged approach (see Fig. 3.2) and, especially, established animal models of diseases under study. Notably, this approach as a first step eliminates differences based on e.g. gender, genetic heterogeneity, diet, medication, and exercise (Xu et al., unpublished). It is also interesting to note that our preliminary studies with human subjects show rapid changes in distinct uEV categories upon medication: when type II (adult onset) diabetics started with SGLT1 inhibitor or miconatsole, distinct intra-individual changes in miRNA were observed (Barreiro et al., unpublished). This shows the rapid, dynamic changes in cellular physiology that can be recapitulated as cellular content snapshots in the form of uEV changes.
Taken together, there are currently many unknown individual factors which may explain the observed complexity of uEV patterns (as well as EVs from other bodily sources). It is also evident that significant advances need to be achieved to understand the abundance of factors influencing uEV contents before they can be fully introduced as biomarkers or tools to precision medicine.
3.3 Urinary Extracellular Vesicle Isolation
Currently, numerous methods are used for extracellular vesicle isolation e.g. ultracentrifugation [6], filtration [20], precipitation [21] and hydrostatic filtration dialysis [22] based techniques, or combination of these. Isolation methods make use of several of the extracellular vesicle properties e.g. size, density and solubility and this is why different subpopulations of extracellular vesicles are enriched by different methods. The method or combination of methods to be applied usually depends on the starting material and the volume [23]. However, there is presently no solid standardization approaches of protocols and, in many cases, methods applied are reported loosely to confound result interpretation and repeatability. Details on the isolation methods, advantages and disadvantages such as processing times, costs, ease of implementation, and co-isolation of contaminants are well described in recent detailed reviews [8, 24,25,26,27]. For reporting a new method, a modification of an existing method or, when applying a method for the first time, we strongly recommend strict adherence to MISEV2018 guideline details [1].
3.4 Inter- and Intra- Individual Variability in EVs Samples
Few studies have assessed systematically the intra- and inter- individual variation of EV contents, and most of them focus on results based on proteomics findings.
Accordingly, in many proteomics based studies, it has been reported that inter- and intra- individual variation is lower in EVs compared to crude urine [28]. In addition, inter-individual variation was described to be higher than intra- individual variation in urinary EVs [29]. In contrast, both inter- and intra- variation for cell culture EVs proteins were reported to be low [30].
Interestingly, the opposite was described for miRNA urinary EV content (inter-individual variation was lower than intra- individual variation) [31] and less variability for miRNA isolated from plasma derived EVs [32] was observed.
The differences regarding inter and intra- individual variation in these reports may be due to the use of different biological fluids, differences in pre-analytical variables and to the diversity of EV isolation methods used. Thus, for biomarkers research it is of key importance to determine the variation for each sample type, downstream application and, vesicle isolation method in order to calculate a solid sample number [33].
Age, gender, and race differences have also been reported to affect the size of the vesicles and the protein content [34, 35]. Thus, these factors should be considered for a well-balanced experimental design.
3.5 Example of Experimental Setups Relevant in uEV Studies
To avoid oversimplification and misinterpretation of uEV results seen especially in urines from diabetic patients, our experimental setup has built on utilization of three levels of meticulously monitored EV parameters as follows:
-
1.
EVs from in vitro cultured diabetes target cells
-
2.
uEVs from established experimental model of diabetes in the rat
-
3.
uEVs from human diabetic (type 1 diabetes) patient urines
This comprehensive approach to study uEVs in Diabetic Kidney Disease (DKD), starting from in vitro and in vivo models to human samples, aims to identify better tools and predictive marker candidates for early DKD diagnostics and disease management.
For the in vitro studies (Figs. 3.3 and 3.4) our hypothesis was that the use of known diabetes target cells and their EV secretion patterns upon diabetic conditions gives the most simple and targeted information of respective EV secretion responses. This approach allows to study effects of a variety of modifications, e.g. variations of insulin, glucose or established pharmacologic manipulation with standardized harvesting of EV response, respectively. Thus, we used conditioned culture media from all three resident glomerular cell types, the kidney targets of diabetes: podocytes, mesangial and endothelial cells with the adjacent proximal tubular cells. According to the study protocol, cells were grown with or without diabetes -mimicking conditions (collaboration with Prof Richard Coward’s lab, Bristol University, UK). EVs from cell culture media were isolated by Hydrostatic Filtration dialysis [36]. Rigorous quality control of the isolated vesicles was done by Western blot, nanoparticle analysis and electron microscopy. Samples were processed for small and long RNA sequencing, and quantitative proteomics. With this data, we seek to define a panel of dysregulated miRNAs, mRNAs and proteins (by cell type) in these in vitro conditions. All data layers are being integrated with multiomics approach to reveal novel candidate pathways involved in insulin resistance and other parameters. Preliminary results of the study from podocytes are shown.
3.6 uEV Omics Studies for Biomarker Research
The application of omics techniques to EV samples has increased remarkably as a consequence of general interest into EV for biomarker research. This is clearly reflected in the amount of publications involving EV proteomics, transcriptomics, lipidomics, or metabolomics, which are thoroughly documented in recent reviews [37,38,39,40,41]. Even if these techniques offer exciting possibilities to profile EV contents precisely, it is important to consider the technical challenges that accompany them in order to design experiments accordingly [42]. Table 3.1 shows examples of recent studies involving specifically urinary extracellular vesicles. As evident from Table 3.1, the use of different approaches to isolate EV, workflows for isolation of features under study (e.g. RNA, proteins), and normalization protocols vary from study to study. The lack of standardization in general and only a few pilot studies available to compare particular omics approaches using same sample sets affects interpretation on what is the preferential approach to study uEV.
All omics approaches can be considered ultrasensitive. This fact, especially in transcriptomics approaches emphasizes the crucial importance of selecting optimized uEV isolation methods and their critical application at all steps. Poor early quality measures will automatically result in poor data quality. Reports have shown that different uEV isolation methods result in dissimilar miRNA containing EV populations and/or even co-isolate non-EV miRNA [43,44,45]. However, most of the comparisons have thus far focused in uEV miRNA analysis. Thus, more studies are needed to reach a clear picture on the effect of early EV isolation steps on RNA type outcomes.
Furthermore, the library preparation protocol applied has major effects on sequencing results as e.g. (i) small vs total RNA library approaches affect RNA biotype distribution [40], (ii) poly(A) library preparation approach does not perform well with partially degraded RNAs. Thus, integrity of EV samples RNA has to be evaluated before choosing this kind of library, (iii) Library Kits that employ universal adapters with fixed sequences generate biases in small RNA sequencing. As reported by Srinivasan et al. [46], the kits that use adapters with degenerate bases reduce biases for small RNA sequencing.
Normalization of quantitative data is also a possible confounding factor. Whether it is better to normalize per starting volume of urine, creatinine or EV related values (e.g. particle number, presence of CD9) is not clear and remains an open debate [47]. MISEV 2018 guidelines acknowledge the lack of agreement on normalization strategy and calls for more studies on the topic.
Validation of sequencing results is usually done through qPCR. However, for example the lack of a reference miRNA and biases introduced by the library preparation itself could difficult validation of the respective miRNA targets [48]. As housekeeping genes used normally may not work properly for EV sequencing data, ISEV recommends to use multiple reference genes to normalize qPCR data [49]. For more information on EV RNA analysis, methodologies and biases, we recommend an ISEV position review by Mateescu et al. [50].
3.7 Future of uEV Studies
Urine remains an underutilized diagnostic matrix, which can be easily and painlessly collected in large quantities, repeatedly and processed easily for further analytics. Factors limiting its diagnostic consist of dynamic contents together with constituents causing serious artifacts in downstream analytics. For these reasons, rigorous standardization of urine is needed and methods are now available for successful use of urine. Notably, to solve these challenges we recently developed a method combining equalization of urinary electrolytes while neutralizing effects of pigments and manage Tamm-Horsfall [51]. Interestingly, the method developed simultaneously suits perfectly for unbiased collection of uEVs to release the full analytical power of the uEVs. It is notable, that uEVs contain 5–20 times the contents of proteins, RNA and lipids as compared to crude urine and, furthermore, without many known artifacts associated with the use of crude urine.
With the wealth of data being increasingly published on the biology, derivation, association to disease and future therapeutic aspects of EVs, it is now reasonable to expect that the uEVs will show their power as future analytic and source for biomarkers valuable for precision medicine.
To process the large number of samples needed to define reliable biomarkers, automated methods to isolate efficiently extracellular vesicles are needed. This is especially important to reduce the processing time; to reduce the number of steps needed to process samples (to reduce batch effects). Reducing these factors to essential ones, addition of technical replicates and follow up samples in large studies would be achievable in reasonable times. However, performance of commercially available kits (better for high-throughput) is variable when compared to more traditional and well characterized isolation methods [52, 53]. More studies are needed to compare isolation methods including control and disease samples and their use in downstream applications. In addition to the commercially available kits new approaches are emerging to automate and standardize the isolation process [54].
With the increasing interest in EVs as source of novel biomarkers, the need to develop devices that could both isolate and detect markers (Lab-on-a-chip) has grown substantially. Promising devices have already been designed, based e.g. in diverse microfluidic strategies to isolate successfully extracellular vesicles as well as detection of their RNA and protein contents and the combination of both on single device [55, 56]. These approaches provide an interesting future solution for quick EV biomarker detection in clinical settings.
Together with the innovation in isolation methods, exiting advances were made in single EV detection in e.g. flow cytometry, which could be applied for accurate biomarker detection [57].
References
Thery, C., Witwer, K.W., Aikawa, E., Alcaraz, M.J., Anderson, J.D., Andriantsitohaina, R., et al.: Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles. 7(1), 1535750 (2018)
Kalluri, R., LeBleu, V.S.: The biology, function, and biomedical applications of exosomes. Science (New York, NY). 367(6478) (2020)
van Niel, G., D’Angelo, G., Raposo, G.: Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19(4), 213–228 (2018)
Yanez-Mo, M., Siljander, P.R., Andreu, Z., Zavec, A.B., Borras, F.E., Buzas, E.I., et al.: Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles. 4, 27066 (2015)
Maas, S.L.N., Breakefield, X.O., Weaver, A.M.: Extracellular vesicles: unique intercellular delivery vehicles. Trends Cell. Biol. 27(3), 172–188 (2017)
Pisitkun, T., Shen, R.F., Knepper, M.A.: Identification and proteomic profiling of exosomes in human urine. Proc. Natl. Acad. Sci. U. S. A. 101(36), 13368–13373 (2004)
Maughan, R.J., Shirreffs, S.M.: Dehydration and rehydration in competative sport. Scand. J. Med. Sci. Sports. 20(Suppl 3), 40–47 (2010)
Musante, L., Tataruch, D.E., Holthofer, H.: Use and isolation of urinary exosomes as biomarkers for diabetic nephropathy. Front. Endocrinol. 5, 149 (2014)
Rastaldi, M.P., Armelloni, S., Berra, S., Li, M., Pesaresi, M., Poczewski, H., et al.: Glomerular podocytes possess the synaptic vesicle molecule Rab3A and its specific effector rabphilin-3a. Am. J. Pathol. 163(3), 889–899 (2003)
Huttenhofer, A., Mayer, G.: Circulating miRNAs as biomarkers of kidney disease. Clin. Kidney J. 10(1), 27–29 (2017)
Miranda, K.C., Bond, D.T., McKee, M., Skog, J., Paunescu, T.G., Da Silva, N., et al.: Nucleic acids within urinary exosomes/microvesicles are potential biomarkers for renal disease. Kidney Int. 78(2), 191–199 (2010)
Bryzgunova, O.E., Zaripov, M.M., Skvortsova, T.E., Lekchnov, E.A., Grigor’eva, A.E., Zaporozhchenko, I.A., et al.: Comparative study of extracellular vesicles from the urine of healthy individuals and prostate cancer patients. PLoS One. 11(6), e0157566 (2016)
Horibe, S., Tanahashi, T., Kawauchi, S., Murakami, Y., Rikitake, Y.: Mechanism of recipient cell-dependent differences in exosome uptake. BMC Cancer. 18(1), 47 (2018)
Mathieu, M., Martin-Jaular, L., Lavieu, G., Thery, C.: Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat. Cell Biol. 21(1), 9–17 (2019)
Mulcahy, L.A., Pink, R.C., Carter, D.R.: Routes and mechanisms of extracellular vesicle uptake. J. Extracell. Vesicles. 3 (2014)
Eitan, E., Tosti, V., Suire, C.N., Cava, E., Berkowitz, S., Bertozzi, B., et al.: In a randomized trial in prostate cancer patients, dietary protein restriction modifies markers of leptin and insulin signaling in plasma extracellular vesicles. Aging Cell. 16(6), 1430–1433 (2017)
Rigamonti, A.E., Bollati, V., Pergoli, L., Iodice, S., De Col, A., Tamini, S., et al.: Effects of an acute bout of exercise on circulating extracellular vesicles: tissue-, sex-, and BMI-related differences. Int. J. Obes. 2019 (2005)
Fruhbeis, C., Helmig, S., Tug, S., Simon, P., Kramer-Albers, E.M.: Physical exercise induces rapid release of small extracellular vesicles into the circulation. J. Extracell. Vesicles. 4, 28239 (2015)
Zachar, R., Jensen, B.L., Svenningsen, P.: Dietary Na(+) intake in healthy humans changes the urine extracellular vesicle prostasin abundance while the vesicle excretion rate, NCC, and ENaC are not altered. Am. J. Physiol. Renal Physiol. 317(6), F1612–F1f22 (2019)
Cheruvanky, A., Zhou, H., Pisitkun, T., Kopp, J.B., Knepper, M.A., Yuen, P.S., et al.: Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am. J. Physiol. Renal Physiol. 292(5), F1657–F1661 (2007)
Rider, M.A., Hurwitz, S.N., Meckes Jr., D.G.: ExtraPEG: a polyethylene glycol-based method for enrichment of extracellular vesicles. Sci. Rep. 6, 23978 (2016)
Musante, L., Tataruch, D., Gu, D., Benito-Martin, A., Calzaferri, G., Aherne, S., et al.: A simplified method to recover urinary vesicles for clinical applications, and sample banking. Sci. Rep. 4, 7532 (2014)
Gardiner, C., Di Vizio, D., Sahoo, S., Théry, C., Witwer, K.W., Wauben, M., et al.: Techniques used for the isolation and characterization of extracellular vesicles: results of a worldwide survey. J. Extracell. Vesicles. 5, 32945 (2016)
Konoshenko, M.Y., Lekchnov, E.A., Vlassov, A.V., Laktionov, P.P.: Isolation of extracellular vesicles: general methodologies and latest trends. Biomed. Res. Int. 2018, 8545347 (2018)
Barreiro, K., Holthofer, H.: Urinary extracellular vesicles. A promising shortcut to novel biomarker discoveries. Cell Tissue Res. 369(1), 217–227 (2017)
Merchant, M.L., Rood, I.M., Deegens, J.K.J., Klein, J.B.: Isolation and characterization of urinary extracellular vesicles: implications for biomarker discovery. Nat. Rev. Nephrol. 13(12), 731–749 (2017)
van der Pol, E., Boing, A.N., Gool, E.L., Nieuwland, R.: Recent developments in the nomenclature, presence, isolation, detection and clinical impact of extracellular vesicles. J. Throm. Haem. 14(1), 48–56 (2016)
Lee, J., McKinney, K.Q., Pavlopoulos, A.J., Niu, M., Kang, J.W., Oh, J.W., et al.: Altered proteome of extracellular vesicles derived from bladder cancer patients urine. Mol. Cells. 41(3), 179–187 (2018)
Wang, S., Kojima, K., Mobley, J.A., West, A.B.: Proteomic analysis of urinary extracellular vesicles reveal biomarkers for neurologic disease. EBioMedicine. 45, 351–361 (2019)
Tiruvayipati, S., Wolfgeher, D., Yue, M., Duan, F., Andrade, J., Jiang, H., et al.: Variability in protein cargo detection in technical and biological replicates of exosome-enriched extracellular vesicles. PLoS One. 15(3), e0228871 (2020)
Ben-Dov, I.Z., Whalen, V.M., Goilav, B., Max, K.E., Tuschl, T.: Cell and microvesicle urine microRNA deep sequencing profiles from healthy individuals: observations with potential impact on biomarker studies. PLoS One. 11(1), e0147249 (2016)
Sanz-Rubio, D., Martin-Burriel, I., Gil, A., Cubero, P., Forner, M., Khalyfa, A., et al.: Stability of circulating Exosomal miRNAs in healthy subjects. Sci. Rep. 8(1), 10306 (2018)
Oeyen, E., Willems, H., Kindt, R., Sandra, K., Boonen, K., Hoekx, L., et al.: Determination of variability due to biological and technical variation in urinary extracellular vesicles as a crucial step in biomarker discovery studies. J. Extracell. Vesicles. 8(1), 1676035 (2019)
Noren Hooten, N., McFarland, M.H., Freeman, D.W., Mode, N.A., Ezike, N., Zonderman, A.B., et al.: Association of extracellular vesicle protein cargo with race and clinical markers of mortality. Sci. Rep. 9(1), 17582 (2019)
Gustafson, C.M., Shepherd, A.J., Miller, V.M., Jayachandran, M.: Age- and sex-specific differences in blood-borne microvesicles from apparently healthy humans. Biol. Sex Differ. 6, 10 (2015)
Barreiro, K., Huber, T.B., Holthofer, H.: Isolating urinary extracellular vesicles as biomarkers for diabetic disease. Meth. Mol. Biol. (Clifton, NJ). 2067, 175–188 (2020)
Zebrowska, A., Skowronek, A., Wojakowska, A., Widlak, P., Pietrowska, M.: Metabolome of exosomes: focus on vesicles released by cancer cells and present in human body fluids. Int. J. Mol. Sci. 20(14) (2019)
Williams, C., Palviainen, M., Reichardt, N.C., Siljander, P.R., Falcon-Perez, J.M.: Metabolomics applied to the study of extracellular vesicles. Metabolites. 9(11) (2019)
Erozenci, L.A., Bottger, F., Bijnsdorp, I.V., Jimenez, C.R.: Urinary exosomal proteins as (pan-)cancer biomarkers: insights from the proteome. FEBS Lett. 593(13), 1580–1597 (2019)
Turchinovich, A., Drapkina, O., Tonevitsky, A.: Transcriptome of extracellular vesicles: state-of-the-art. Front. Immunol. 10, 202 (2019)
Gezsi, A., Kovacs, A., Visnovitz, T., Buzas, E.I.: Systems biology approaches to investigating the roles of extracellular vesicles in human diseases. Exp. Mol. Med. 51(3), 1–11 (2019)
Ramirez, M.I., Amorim, M.G., Gadelha, C., Milic, I., Welsh, J.A., Freitas, V.M., et al.: Technical challenges of working with extracellular vesicles. Nanoscale. 10(3), 881–906 (2018)
Srinivasan, S., Yeri, A., Cheah, P.S., Chung, A., Danielson, K., De Hoff, P., et al.: Small RNA sequencing across diverse biofluids identifies optimal methods for exRNA isolation. Cell. 177(2), 446-62.e16 (2019)
Mussack, V., Wittmann, G., Pfaffl, M.W.: Comparing small urinary extracellular vesicle purification methods with a view to RNA sequencing-enabling robust and non-invasive biomarker research. Biomol. Detect. Quant. 17, 100089 (2019)
Cheng, L., Sun, X., Scicluna, B.J., Coleman, B.M., Hill, A.F.: Characterization and deep sequencing analysis of exosomal and non-exosomal miRNA in human urine. Kidney Int. 86(2), 433–444 (2014)
Srinivasan, S., Duval, M.X., Kaimal, V., Cuff, C., Clarke, S.H.: Assessment of methods for serum extracellular vesicle small RNA sequencing to support biomarker development. J. Extracell. Vesicles. 8(1), 1684425 (2019)
Gunasekaran, P.M., Luther, J.M., Byrd, J.B.: For what factors should we normalize urinary extracellular mRNA biomarkers? Biomol. Detect. Quant. 17, 100090 (2019)
Git, A., Dvinge, H., Salmon-Divon, M., Osborne, M., Kutter, C., Hadfield, J., et al.: Systematic comparison of microarray profiling, real-time PCR, and next-generation sequencing technologies for measuring differential microRNA expression. RNA (New York, NY). 16(5), 991–1006 (2010)
Hill, A.F., Pegtel, D.M., Lambertz, U., Leonardi, T., O’Driscoll, L., Pluchino, S., et al.: ISEV position paper: extracellular vesicle RNA analysis and bioinformatics. J. Extracell. Vesicles. 2 (2013)
Mateescu, B., Kowal, E.J., van Balkom, B.W., Bartel, S., Bhattacharyya, S.N., Buzas, E.I., et al.: Obstacles and opportunities in the functional analysis of extracellular vesicle RNA – An ISEV position paper. J. Extracell. Vesicles. 6(1), 1286095 (2017)
Xu, X., Barreiro, K., Musante, L., Kretz, O., Lin, H., Zou, H., et al.: Management of Tamm-Horsfall protein for reliable urinary analytics. Proteomics Clin. Appl. 13(6), e1900018 (2019)
Tian, Y., Gong, M., Hu, Y., Liu, H., Zhang, W., Zhang, M., et al.: Quality and efficiency assessment of six extracellular vesicle isolation methods by nano-flow cytometry. J. Extracell. Vesicles. 9(1), 1697028 (2020)
Patel, G.K., Khan, M.A., Zubair, H., Srivastava, S.K., Khushman, M., Singh, S., et al.: Comparative analysis of exosome isolation methods using culture supernatant for optimum yield, purity and downstream applications. Sci. Rep. 9(1), 5335 (2019)
Sunkara, V., Kim, C.J., Park, J., Woo, H.K., Kim, D., Ha, H.K., et al.: Fully automated, label-free isolation of extracellular vesicles from whole blood for cancer diagnosis and monitoring. Theranostics. 9(7), 1851–1863 (2019)
Chiriaco, M.S., Bianco, M., Nigro, A., Primiceri, E., Ferrara, F., Romano, A., et al.: Lab-on-chip for exosomes and microvesicles detection and characterization. Sensors (Basel, Switzerland). 18(10) (2018)
Ramshani, Z., Zhang, C., Richards, K., Chen, L., Xu, G., Stiles, B.L., et al.: Extracellular vesicle microRNA quantification from plasma using an integrated microfluidic device. Comm. Biol. 2, 189 (2019)
Wang, S., Khan, A., Huang, R., Ye, S., Di, K., Xiong, T., et al.: Recent advances in single extracellular vesicle detection methods. Biosens. Bioelectron. 154, 112056 (2020)
Babicki, S., Arndt, D., Marcu, A., Liang, Y., Grant, J.R., Maciejewski, A., et al.: Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res. 44(W1), W147–W153 (2016)
Zhou, G., Xia, J.: Using OmicsNet for network integration and 3D visualization. Curr. Protoc. Bioinformatics. 65(1), e69 (2019)
Ghai, V., Wu, X., Bheda-Malge, A., Argyropoulos, C.P., Bernardo, J.F., Orchard, T., et al.: Genome-wide profiling of urinary extracellular vesicle microRNAs associated with diabetic nephropathy in type 1 diabetes. Kidney Int. Rep. 3(3), 555–572 (2018)
Khurana, R., Ranches, G., Schafferer, S., Lukasser, M., Rudnicki, M., Mayer, G., et al.: Identification of urinary exosomal noncoding RNAs as novel biomarkers in chronic kidney disease. RNA (New York, NY). 23(2), 142–152 (2017)
Salih, M., Demmers, J.A., Bezstarosti, K., Leonhard, W.N., Losekoot, M., van Kooten, C., et al.: Proteomics of urinary vesicles links plakins and complement to polycystic kidney disease. J Am Soc Nephrol. 27(10), 3079–3092 (2016)
Puhka, M., Takatalo, M., Nordberg, M.E., Valkonen, S., Nandania, J., Aatonen, M., et al.: Metabolomic profiling of extracellular vesicles and alternative normalization methods reveal enriched metabolites and strategies to study prostate cancer-related changes. Theranostics. 7(16), 3824–3841 (2017)
Clos-Garcia, M., Loizaga-Iriarte, A., Zuniga-Garcia, P., Sanchez-Mosquera, P., Rosa Cortazar, A., Gonzalez, E., et al.: Metabolic alterations in urine extracellular vesicles are associated to prostate cancer pathogenesis and progression. J. Extracell. Vesicles. 7(1), 1470442 (2018)
Skotland, T., Ekroos, K., Kauhanen, D., Simolin, H., Seierstad, T., Berge, V., et al.: Molecular lipid species in urinary exosomes as potential prostate cancer biomarkers. Eur. J. Cancer (Oxford, England: 1990). 70, 122–132 (2017)
Acknowledgements
Research lines reported here are based on fruitful collaborative efforts and are gratefully acknowledged as follows:
-
Prof Richard Coward’s team (Bristol University, UK, in-vitro models);
-
Prof Leif Groop’s team (University of Helsinki, Finland, Clinical samples);
-
Dr. Carol Forsblom (The Finnish Diabetic Nephropathy Study, Finland, Clinical Samples;);
-
Prof Riitta Lassila (University of Helsinki, Finland, in vivo experimental model);
-
And Dr. Denis Delic, Dr. German Leparc, Marcel Rosler (Boehringer Ingelheim Pharma GmbH & Co. KG, Germany, short and long RNA sequencing)
This study was supported by grants from the Paulo Foundation of Finland and NovoNordisk Foundation.
TBH was supported by the DFG (CRC1192, HU 1016/8-2, HU 1016/11-1, HU 1016/12-1), by the BMBF (STOP-FSGS-01GM1518C), and by the European Research Council-ERC (grant 616891).
This project has received funding from the Innovative Medicines Initiative 2 Joint Undertaking under grant agreement no. 115974 (BEAt-DKD; T.B.H., H.H.). This Joint Undertaking receives support from the European Union’s Horizon 2020 research and innovation programme and EFPIA and by the H2020-IMI2 consortium BEAt-DKD.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Barreiro, K., Huber, T.B., Holthofer, H. (2021). Urinary Extracellular Vesicles Magic Particles for Biomarker Discovery. In: Baptista Carreira dos Santos, H.M. (eds) Translational Urinomics. Advances in Experimental Medicine and Biology(), vol 1306. Springer, Cham. https://doi.org/10.1007/978-3-030-63908-2_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-63908-2_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-63907-5
Online ISBN: 978-3-030-63908-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)