Abstract
Reduced sulphur compounds such as sulphide, polysulphides, thiosulphate, and elemental sulphur are oxidized by a large and diverse group of prokaryotes. In many cases, intracellular globules of polymeric, water-insoluble sulphur are accumulated either as a transient product en route to sulphate or as the final product. Sulphur globule formation is especially widespread among sulphur-oxidizing Proteobacteria and occurs in purple sulphur bacteria of the family Chromatiaceae, in Beggiatoa species as well as in other “morphologically conspicuous” sulphur bacteria (e.g. Thioploca, Achromatium, Thiovulum). Sulphur globules are typically enclosed by a surface layer consisting of highly repetitive glycine-rich structural proteins (sulphur globule proteins, Sgps) and reside in the bacterial periplasm. Here, an overview of recent findings on the speciation of stored sulphur, the occurrence of Sgps and the enzymes involved in the formation and breakdown of bacterial sulphur globules is given.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Sulphur is the 16th element on the periodic table and the tenth most abundant element in the universe (Steudel and Chivers 2019). Sulphur serves essential functions in all living cells. In proteins it occurs not only in the form of cysteine and methionine, but also in iron-sulphur clusters, in several sulphur-containing cofactors like thiamine, biotin, coenzyme A and lipoic acid and is furthermore indispensable in tRNAs through a variety of modifications (Shigi 2014, 2018). Sulphur is a very versatile chemical element and undergoes permanent cycling in terrestrial as well as in marine environments. Dissimilatory sulphate reduction is the primary driver of the biogeochemical sulphur cycling. In this anaerobic respiratory process, sulphate is used as an electron acceptor instead of oxygen, nitrate or manganese [Mn(IV)] (Henkel et al 2019; Rabus et al 2015). In turn, hydrogen sulphide, polysulphides, thiosulphate, elemental sulphur and polythionates serve as electron donors for a huge array of chemo- and photolithotrophic bacteria and archaea such as Acidithiobacillus or Acidianus species (Dahl et al. 2008; Mangold et al. 2011; Kletzin et al. 2004; Frigaard and Dahl 2009; Dahl 2017). A large portion of these organisms forms sulphur globules both extracellularly and intracellularly (Dahl and Prange 2006; Dahl 2017; Maki 2013). Whether the sulphur accumulates as a transient or the final product varies depending on the species, the culture conditions and the reduced sulphur substrate.
Here, I attempt to give an update about the different sulphur-forming prokaryotes, the structure and chemical nature of bacterial sulphur inclusions and the metabolic pathways related to sulphur globule formation and degradation. An exclusive focus will be laid on sulphur globules deposited within the confines of the cell wall, i.e. sulphur present as a bacterial inclusion senso strictu. For further detailed information, the reader is referred to a number of reviews on oxidative sulphur metabolism (Frigaard and Dahl 2009; Dahl 2017; Wang et al. 2019; Friedrich et al. 2005; Dahl et al. 2008; Dahl and Prange 2006).
2 History
Internal sulphur globules are easily recognized even via light microscopy as they are highly light refractive (Fig. 1) and can reach diameters of several micrometres. Accordingly, the first mentioning of these conspicuous structures dates back to 1786, when Müller described intracellular spherical inclusions of unknown composition in ‘colourless’, egg-shaped algae (Müller 1786), later identified as Thiovulum majus (Rivière and Schmidt 2006). Over the following decades, several additional microorganisms were mentioned to contain similar inclusions (Ehrenberg 1838; Trevisan 1842; Perty 1852) and differentiated due to the presence of colour (the later Thiospirillum, Chromatium, Lamprocystis) and lack of colour (Beggiatoa, Thiothrix and Thiovulum). About a century after their discovery the cellular inclusions were proven to consist of elemental sulphur in Beggiatoa (C. Cramer in (Müller 1870); in Allochromatium vinosum (Cohn 1875); and in Thiovulum muelleri (Warming 1875)). A first systematic analysis of uncoloured and coloured ‘sulphobacteria’ and their sulphur globules was provided by Winogradsky (Winogradsky 1887), who also demonstrated the oxidation of hydrogen sulphide to stored sulphur under microaerophilic conditions in the chemotrophic Beggiatoa (Winogradsky 1889). Pioneering studies on the oxidation of sulphur in bacterial photosynthesis were done by van Niel whose classic studies about phototrophic sulphur bacteria and accumulation of elemental sulphur can be considered as milestones and provided the basis for further studies about sulphur compounds in photosynthesis (van Niel 1936; van Niel 1931). Those interested in the early research on sulphur bacteria are referred to discussions by (Waksman 1922; Waksman and Joffe 1922; Shively et al. 2006; Dahl and Prange 2006; Trüper 2008).
Allochromatium vinosum DSM 180T cells with sulphur globules visualized by light microscopy: (a) phase contrast (b) DNA stain with DAPI. In panel (c), staining with Nile red highlights sulphur globules. (d), DAPI and Nile red stain merged. Microscopy was carried out at room temperature using a Zeiss Axio Observer Z1 microscope (Zeiss, Jena, Germany) equipped with HXP 120 V light source and Axio Cam MR3 camera. Standard filter sets were used for DAPI (335–383 nm excitation and 420–470 nm emission) and Nile red (510–560 nm excitation and 590 nm long pass emission). Image acquisition and analysis were performed with Zen 2 software (Zeiss). Nile red is a useful probe of hydrophobic sites on proteins (Sackett and Wolff 1987) and probably interacts with the hydrophobic Sgps. The red halo in the Nile red image requires further investigation and currently remains unexplained. Photos courtesy of Fabian Grein
3 Elemental Sulphur
Naturally, sulphur occurs in a huge variety of environments (Cosmidis et al. 2019; Nims et al. 2019) such as volcanic areas including sulphidic springs (Macur et al. 2013; Kamyshny et al. 2014; Lau et al. 2017), deep-sea hydrothermal vents (Taylor et al. 1999), deep-sea hydrocarbon seeps (Eichinger et al. 2014) or marine sediments and salt marshes (Kamyshny and Ferdelman 2010; Zopfi et al. 2004; Jørgensen and Nelson 2004; Taylor and Wirsen 1997).
Elemental sulphur can be formed through abiogenic processes when sulphide is oxidized by molecular oxygen, possibly catalyzed by oxidized metals (Luther et al. 2011). Its presence in the environment is often associated with microbial oxidation of reduced sulphur compounds (Kleinjan et al. 2003).
Sulphur forms more than 30 solid allotropes, more than any other element. It exists in many forms, from cyclic octamers (S8 rings) that crystallize in different structures to sulphur chains with varying numbers of S-S bonds, and polysulphides (Sn2−) (Kamyshny and Ferdelman 2010; Meyer 1976; Trofimov et al. 2009). Only a few sulphur allotropes occur in biological systems. The thermodynamically most stable form at standard conditions is homocyclic, orthorhombic crystalline α-sulphur (α-S8) (cyclo-octasulphur) (Roy and Trudinger 1970; Steudel 1996a, b). S6, S7 and S12 rings have also been detected in samples of biological origin, while bigger rings up to S20 were made accessible by chemical synthesis (Steudel 1987, 2000). Commercially available sulphur consists mainly of S8 rings, traces of S7 rings that are responsible for the bright yellow colour (Steudel and Holz 1988) and polymeric sulphur. Polymeric sulphur consists of very long helically wound chains of almost all sizes (Steudel 2000). Regardless of the molecular size, all sulphur allotropes are hydrophobic, are not wetted by water and have very low solubilities in water (Steudel and Eckert 2003).
4 Organisms Forming Intracellular Sulphur Globules
Sulphur can accumulate in the form of water-insoluble globules as a transient or the final product during the oxidation of reduced sulphur compounds (sulphide, polysulphides, thiosulphate, polythionates and elemental sulphur). Accordingly, sulphur-forming bacteria share environments characterized by elevated levels of hydrogen sulphide mainly produced by bacterial sulphate reduction in anoxic sediments rich in organic nutrients or originating from hydrothermal vents or cold seeps.
Concerning their physiology, two large groups of sulphur-storing bacteria can be differentiated: The first are phototrophic prokaryotes that use sulphur compounds as electron donors for CO2 fixation in the light (Dahl 2017). Among these, purple sulphur bacteria of the family Chromatiaceae form intracellular sulphur deposits. On the other hand, chemotrophic (the classical “colourless”) sulphur-oxidizing prokaryotes use the energy derived from the oxidation of sulphur compounds with either oxygen, nitrate or Mn(IV) oxide as electron acceptors to fix carbon dioxide (Henkel et al. 2019; Dahl et al. 2008; Friedrich 1998; Kletzin et al. 2004; Wang et al. 2019). Sulphur compounds are also an important energy source for symbiotic associations of chemoautotrophic sulphur bacteria with marine organisms from unicellular protists (Ott et al. 2004) to metazoans, such as meduzoans (Abouna et al. 2015), bivalves (Frenkiel et al. 1996) and nematodes (Himmel et al. 2009). Although first discovered at hydrothermal vents, symbiosis with sulphur oxidizers is not limited to these highly specialized environments but has also been found in shallow subtidal sands, macrophyte debris, deep sea cold seeps, mangrove swamps, sea grass beds, anoxic marine basins, sewage outfalls and even rotting whale carcasses (Distel 1998; Kleiner et al. 2012; Petersen et al. 2016; Seah et al. 2019; Cavanaugh et al. 1981; Felbeck 1981; Nelson and Fisher 1995).
Concerning their systematic affiliation, the vast majority of organisms with reported capability for the formation of intracellular sulphur globules belong to the Proteobacteria (Table 1). Notable exceptions are the Gram-positives Thermoanaerobacter sulfurigignens and Thermoanaerobacterium thermosulfurigienes (Lee et al. 2007), which fall into the class Clostridia within the Firmicutes phylum. Sulphur globules within the confines of the cell have also been detected in Thermus scotoductus (Skirnisdottir et al. 2001) belonging to the class Deinococci within the phylum Thermus-Deinococcus.
Most reports about intracellular sulphur deposition are available for members of the α-, β-, γ- and ε-Proteobacteria (Table 1). The trait is widespread though not ubiquitous in alphaproteobacterial, microaerophilic, autotrophic magnetotactic bacteria which form the globules upon growth on sulphide and/or thiosulphate (Bazylinski et al. 2004, 2013; Bazylinski and Williams 2006; Keim et al. 2005; Williams et al. 2006; Lefevre et al. 2012; Spring and Bazylinski 2000). Another example among the Alphaproteobacteria is Azospirillum thiophilum for which intracellular sulphur globule formation has been described upon growth in the presence of sulphide (Lavrinenko et al. 2010). Within the β-Proteobacteria, we find the genus Macromonas (La Riviere and Schmidt 1999). The large cells of this genus are characterized by voluminous inclusions of calcium oxalate. In addition, sulphur globules may be present. Macromonas bipunctata can oxidize sulphide to sulphur by means of hydrogen peroxide; however, this process does not allow energy conservation (Willems 2014). Furthermore, the Betaproteobacterium Thermothrix azorensis, an aerobic, thermophilic, obligately chemolithoautotrophic sulphur oxidizer, appears to form inclusions of sulphur under certain growth conditions (incomplete thiosulphate oxidation, pH above 7.0) (Odintsova et al. 1996). Thiovulum is a spectacular genus belonging to the ε-branch of the Proteobacteria that has primarily been defined observationally by its large egg-shaped cells that can reach a length of 5–25μm. In the cells, sulphur globules are often concentrated at one cell pole (Marshall et al. 2012; La Riviere and Schmidt 1999). Thiovulum has so far evaded isolation in pure culture but appears to be a chemolithoautotrophic microaerophile. A single-cell genome is available (Marshall et al. 2012).
Among the Gammaproteobacteria, formation of intracellular sulphur globules is especially widespread (Table 1). Many of these bacteria belong to families within the order Chromatiales. Sulphur deposition is a characteristic trait of many purple sulphur bacteria of the family Chromatiaceae (Dahl 2017), while Thiorhodospira sibirica is the only phototrophic member of the family Ectothiorhodospiraceae that is capable of intracellular sulphur deposition (Bryantseva et al. 1999). In fact, this organism also forms extracellular sulphur deposits, the name-giving feature of the family. The Ectothiorhodospiraceae harbour additional species that store sulphur internally; these belong to the chemoautotrophic genus Thioalkalivibrio (Sorokin et al. 2003; Berben et al. 2015; Mu et al. 2016; Ahn et al. 2017). Recently, a member of the Thioalkalispiraceae, Endothiovibriovibrio diazotrophicus, was also described as containing intracellular sulphur globules (Bazylinski et al. 2017). The Thiotrichales are the second gammaproteobacterial order containing a variety of sulphur-storing chemotrophic sulphur oxidizers. Among these, the family Thiotrichaceae features some of the most conspicuous bacteria in nature. Species of the genera Thiomargerita and Achromatium as well as the filamentous sulphur-oxidizing bacteria of the genera Beggiatoa, Thiothrix and Thioploca are among the largest known prokaryotes (Mansor et al. 2015; Schulz and Jørgensen 2001; Schulz et al. 1999; Salman et al. 2016) and characterized by massive sulphur formation. The only representatives of the families Thiofilaceae and Thiolinaceae described so far also form intracellular sulphur globules (Boden and Scott 2018).
A vast majority of bacterial partners in thiotrophic symbioses with eukaryotes are taxonomically unclassified Gammaproteobacteria (Table 1). Regardless of whether the host is a protist or an invertebrate and whether the bacteria are associated as endo- or as ectosymbionts, formation of sulphur globules inside of their cells has often been noted (Grimonprez et al. 2018; Rinke et al. 2006, 2009; Bergin et al. 2018; Seah et al. 2019; Markert et al. 2011; Krieger et al. 2000; Frenkiel et al. 1996).
5 Subcellular Localization of Sulphur Globules
Internal sulphur globules are easily recognized even via light microscopy as they are highly light refractive (Fig. 1). Cultures and colonies of cells containing sulphur globules, therefore, exhibit a characteristic milky appearance (Fig. 2). Usually the diameter of sulphur globules is in the range of 1–3μm (Fig. 1), but sizes exceeding 15μm have also been reported (Williams et al. 1987; Head et al. 1996; Remsen 1978; Skirnisdottir et al. 2001). The sulphur can comprise 20–34% of the cell dry mass of Beggiatoa sp. and purple sulphur bacteria, respectively (Nelson and Castenholz 1981; Overmann 1997). While sulphur globules appear to be randomly localised in many bacterial species, specific cellular localizations have also been reported. In Thiovulum for example, the globules accumulate toward one cell pole (Marshall et al. 2012; La Riviere and Schmidt 1999).
Colonies Allochromatium vinosum DSM 180T grown (a) on malate in the absence of reduced sulphur compounds (b) grown in the presence of sulphide and thiosulphate. Colonies appear milky-white due to massive accumulation of sulphur globules inside of the cells. A. vinosum was cultivated for 10 days on plates solidified with 1% (w/v) phytagel as described by (Pattaragulwanit and Dahl 1995)
A series of technical problems must be considered when it comes to elucidating the subcellular localization of sulphur globules using microscopic techniques. Sulphur dissolves during the preparation of biological samples for electron microscopy, and in addition, any remaining sulphur is subject to thermal degradation under the electron beam. Therefore, sulphur deposits appear as a conspicuous, empty, and electron-lucent space in electron micrographs (Strohl et al. 1981; Remsen and Trüper 1973; Vetter 1985; Pasteris et al. 2001). Nevertheless, microscopic evaluation has resolved the periplasm as the intracellular compartment harbouring sulphur globules in many cases. In studies with free-living filamentous sulphur bacteria, including Thiothrix (Bland and Staley 1978; Larkin and Shinabarger 1983; Williams et al. 1987), Thioploca (Maier and Murray 1965), Thiofilum and Thiolinea (Boden and Scott 2018) as well as Beggiatoa (de Albuquerque et al. 2010; Maier and Murray 1965; Larkin and Strohl 1983), sulphur inclusions were found to be located within invaginated pockets of the cytoplasmic membrane. In some cases, the sulphur globules appeared as a membrane-bound inclusion in the cytoplasm with no apparent connection to the cytoplasmic membrane (Strohl et al. 1981), which may be an effect of the specific sectioning plane (Shively et al. 1989). Other examples for which a periplasmic localization of sulphur globules has been settled are species of the genera Thioalkalivibrio (Sorokin et al. 2001) and Thermus (Skirnisdottir et al. 2001).
In some cases, it has been problematic to distinguish putative sulphur vesicles from other vesicle-like storage structures such as polyhydroxyalkanoate bodies. This applies especially to chemoautotrophic sulphur–oxidizing endosymbionts that reside in animal organs, e.g., in specialized gills of Vesicomyid clams (Goffredi and Barry 2002) or in so-called trophosomes in Vestimeniferan worms like Riftia pachyptila (Felbeck 1981; Cavanaugh 1983). Inside these organs, the symbiontic bacterial cells exhibit roundish to polymorphic electron-translucent vesicles whose membranes are infoldings of the cytoplasmic membrane, and the enclosed spaces are contiguous with the periplasmic space. Although these vesicles obviously share common ultrastructural characteristics with sulphur-containing globules of other organisms, it has been debated whether these structures are indeed related to sulphur storage (Bright and Sorgo 2003; Maina and Maloyi 1998; Vetter 1985). On the other hand, electron spectroscopic imaging pictures clearly identified sulphur in the globules of gutless oligochaete worm endosymbionts. A cytoplasmic localization was inferred for the globules without analysis via cryo-EM (Krieger et al. 2000).
Interpretation of electron micrographs of phototrophic bacteria containing sulphur globules is complicated by the dense packing of these cells with intracytoplasmic membranes harbouring the photosynthetic apparatus. These so-called chromatophores are associated with sulphur globules in a highly organized manner. Careful inspection of electron micrographs revealed that some chromatophores, the insides of which are extracytoplasmic or periplasmic (depending on whether the insides are continuous with the periplasm or not), open into the space enclosing the sulphur globules, thus implying an extracytoplasmic location for the globules themselves (Pattaragulwanit et al. 1998).
It is obvious from the last paragraphs that high-resolution microscopy has so far not provided assignment of the correct subcellular compartment for sulphur deposition in all cases. Fortunately, another very valuable information resource is available. As early as 1963, an envelope was reported for the sulphur globules of the purple sulphur bacterium A. vinosum (Kran et al. 1963) that was soon identified as a protein envelope (Nicolson and Schmidt 1971; Schmidt and Kamen 1970). In A. vinosum, the envelope consists of four different proteins of 8.5–20.8 kDa named SgpA, SgpB, SgpC and SgpD (Brune 1995a; Pattaragulwanit et al. 1998; Weissgerber et al. 2014). Relative transcript abundances for all four of the corresponding genes strongly increase upon the exposure of the cells to sulphide, thiosulphate and elemental sulphur compared to photoorganoheterotrophic growth on malate in the absence of reduced sulphur compounds (Weissgerber et al. 2013; Weissgerber et al. 2014). All four proteins are synthesized as precursors carrying amino-terminal signal peptides mediating Sec-dependent transport across the cytoplasmic membrane (Weissgerber et al. 2014; Pattaragulwanit et al. 1998). The proposed targeting process was experimentally confirmed with a sgpA-phoA fusion in E. coli (Pattaragulwanit et al. 1998) which finally resolved the subcellular localization of the globules in purple sulphur bacteria of the family Chromatiaceae. Single-layered electron-dense envelopes of 2–5 nm have also been observed for the sulphur globules of Thioalkalivibrio paradoxus (Berben et al. 2015) as well as for Beggiatoa and Thiothrix species (Strohl et al. 1981; Williams et al. 1987). In Beggiatoa alba BL15D, the envelope is pentalaminar, 12–14 nm thick and consists of three electron dense layers of 3.5, 2.1 and 3.5 nm thickness (Strohl et al. 1982). Sulphur inclusion envelopes have been described as being fragile in fixatives used for transmission electron microscopy (Strohl et al. 1981) which may explain why they are not always visible in electron micrographs of sulphur-depositing bacteria. In fact, recently performed BLAST searches revealed the presence of genes encoding putative sulphur globule proteins targeted to the periplasm in almost all genome-sequenced, globule-forming Proteobacteria analyzed (unpublished) with Thiovulum and Macromonas as the only notable exceptions. Table 1 provides an overview of selected species. The number of predicted sgp genes in a given organism can vary from only one, e.g., in Beggiatoa alba or in Thiolinea disciformis to six in Thiothrix caldifontis (Table 1) and even 15 in Thiothrix lacustris (not shown). Taken together, these observations provide strong indication that the general target compartment for sulphur storage is not the bacterial cytoplasm, but that deposited sulphur is separated from the cytoplasm by a unit membrane which may be continuous with the cytoplasmic membrane, depending on the organism.
6 Properties and Function of Sulphur Globule Proteins
Brune already noted in 1995 that SgpA, SgpB and SgpC from the purple sulphur bacteria A. vinosum and Thiocapsa roseopersicina exhibit sequence similarity with structural proteins containing repetitive amino acid sequences rich in regularly spaced glycine-like cytoskeletal keratins, insect and blood fluke egg-shell proteins and plant cell wall proteins (Brune 1995a). SgpC shows some sequence similarity to Gly and Trp-rich regions of prion proteins (Brune 1995a). SgpD from A. vinosum appears to be a coiled-coil protein (Weissgerber et al. 2014). The coiled coil is a protein motif characterized by superhelical twisting of two or more alpha helices around one another. They can form rod-like tertiary structures and include the intermediate filaments of the metazoan cytoskeleton as well as bacteria specific cytoskeletal proteins that typically assemble into stable macromolecular scaffolds (Lin and Thanbichler 2013; Rose and Meier 2004). In general, sulphur globule proteins appear to be rich in glycine, alanine and asparagine. Tyrosine and glutamine and proline can also be major constituents, depending on the protein (Brune 1995a).
In plant cell walls, glycine-rich proteins form an important group of structural protein components (Ringli et al. 2001). The primary sequences of these proteins contain more than 60% glycine, which is considerably higher than the glycine content in bacterial Sgps (Brune 1995a). Just as in glycine-rich proteins from plants, the sequences of Sgps often follow the motif (Gly-X)n in the glycine rich regions. In the proteins forming sulphur globule envelopes, Ala, Pro, Ser and Tyr are common at the X position. In some cases the motif varies, e.g. (G-G-X)n, or is more complex. The structures proposed for the glycine-rich plant cell-wall proteins are antiparallel beta-pleated structures analogous to that of silk fibroin, in which the side chains of the X residues in the GXGX repeats all lie on one side of the sheet (Condit and Meagher 1986; Keller et al. 1988). If the sulphur globule proteins fold in a similar way, this may help to explain how they aid preserving the enclosed hydrophobic sulphur in a reactive state and at the same time impart hydrophilic properties to the globule surface (Steudel 1989). In the future, it will certainly be necessary to study secondary structure of Sgps in detail.
The presence of a protein envelope around the sulphur inclusions in sulphur-oxidizing bacteria suggests an important structure–function relationship. Indeed, mutants of A. vinosum lacking SgpB and SgpC are no longer able to oxidize sulphide and thiosulphate and to form sulphur inclusions from these sulphur compounds (Prange et al. 2004). In A. vinosum SgpA and SgpB can replace each other in the presence of SgpC. Still, SgpB and SgpA are not fully competent to replace each other as sulphur globule formation is not possible in mutants possessing soley SgpA or SgpB (Prange et al. 2004). A mutant containing SgpA and SgpB but lacking SgpC can grow on sulphide and thiosulphate. As this mutant forms significantly smaller sulphur globules, SgpC probably plays an important role in sulphur globule expansion. The construction of mutants lacking SgpA and SgpC or SgpA, SgpB and SgpC was not possible, leading to the conclusion that a basic level of Sgps is obligatory for cell survival even under conditions that do not allow sulphur globule formation (Prange et al. 2004). SgpD appears to be the most abundant of the A. vinosum sulphur globule proteins (Weissgerber et al. 2014). Genetic information about its role is not available because mutants lacking the respective gene have not yet been analyzed. The analysis of the A. vinsoum sulphur globule proteome revealed SgpB as the second most abundant sulphur globule protein in this organism while peptides originating from SgpA and SgpC were less frequently detected (Weissgerber et al. 2014). In A. vinosum, all four sgp genes form separate transcriptional units (Pattaragulwanit et al. 1998; Prange et al. 2004; Weissgerber et al. 2013). All four genes are constitutively expressed and their expression is significantly enhanced in the presence of sulphide and thiosulphate (Weissgerber et al. 2013; Prange et al. 2004).
Interestingly, cells of Beggiatoa alba grown in the absence of sulphur compounds apparently contained small rudimentary sulphur inclusion envelopes. It was hypothesized that these envelopes were present in collapsed form until a reduced sulphur source became available (Strohl et al. 1982). A direct/covalent attachment of chains of stored sulphur to the proteins enclosing the globules does not appear to occur as a vast majority of studied or predicted Sgps do not contain any cysteine residues (Brune 1995a; Weissgerber et al. 2014). It has been speculated that protein envelope may provide binding sites for sulphur-metabolizing enzymes (Schmidt et al. 1971). To elucidate the possibility that enzymes taking part in sulphur globule formation and/or oxidation are bound to or interact with the envelope proteins, similar to the situation found for polyhydroxyalkanoate (PHA) granules (Jendrossek 2009), the sulphur globule proteome of A. vinosum was enriched and analyzed (Weissgerber et al. 2014). While this approach identified 78 proteins that occur exclusively in the sulphur globule proteome and were not detected in the soluble and membrane fractions, none of the established components of the periplasmic sulphide- and thiosulphate-metabolizing enzymes appeared to be enriched with the globules.
7 Speciation of Sulphur
The chemical nature of the sulphur in the globules has been the subject of intensive controversy (Pickering et al. 1998; Prange et al. 1999b, 2002; George et al. 2002, 2008; Berg et al. 2014; Pasteris et al. 2001). It has been recognized several decades ago that this sulphur has several properties that do not go along with those of elemental sulphur outside of biological systems. The first discrepancy relates to its low density of 1.2 (Guerrero et al. 1984), compared to 2.1 for the common α-sulphur (Meyer 1976). Moreover, globule sulphur has been described as ‘liquid’ and ‘hydrophilic’ (Steudel 1989; Hageage et al. 1970), while all allotropes of sulphur are solid and virtually water-insoluble at room temperature. In fact, analysis of sulphur in biological systems is generally hampered by the variety of possible reactions, the high reactivity and short lifetime of sulphur compounds with intermediate oxidation states that may be formed during these reactions, the allotropic enantiotropy of S8 and its ability to catenate (Steudel 1982).
The first study focussing on the speciation of sulphur in sulphur globules dates back to 1970, applied polarizing microscopy and X-ray diffraction to a purple sulphur bacterium, A. vinosum (Hageage et al. 1970), and culminated in the conclusion that the sulphur is present in a ‘liquid’ or ‘liquid-like’ state. Later, this was questioned because Raman spectroscopy provided evidence for predominance of S8 sulphur in the globules from Beggiatoa and Thioploca (Pasteris et al. 2001). Several other studies applied synchrotron-based X-ray absorption near-edge structure spectroscopy (XANES) at the sulphur K-edge to investigate the nature of intracellular sulphur globules (George et al. 2008; Pickering et al. 2001; Prange et al. 1999a, 2002; Lee et al. 2007), however, with contradicting results. Our own work indicated different speciation depending on metabolic properties of the organisms and the environmental conditions: long sulphur chains very probably terminated by organic residues (mono- or bisorganyl polysulphanes) in purple sulphur bacteria, cyclo-octasulphur in chemotrophic sulphur oxidizers like Beggiatoa alba and Thiomargararita namibiensis and long chain polythionates in the aerobically grown acidophilic sulphur oxidizer Acidithiobacillus ferrooxidans (Prange et al. 1999a, 2002). Others pointed out shortcomings of the detection method applied that may suffer from spectroscopic distortions dependent upon particle size and compositions. Experimental data was provided suggesting that the spectral differences observed for the sulphur globules from different organisms are not due to differences in sulphur speciation but are solely due to differences in the particle sizes of the sulphur globules (George et al. 2008). More recently, Raman spectroscopy has been applied to various sulphur globule-forming bacteria. This non-destructive analytical technique circumvents many of the problems associated with other characterization methods, as measurements can be collected on solid, liquid and live samples at room temperature and atmospheric pressure. Characteristic internal vibrational (molecular) spectra make elemental sulphur easy to detect and characterize (Eichinger et al. 2014; Pasteris et al. 2001; Berg et al. 2014; Oren et al. 2015; Maurin et al. 2010; Himmel et al. 2009). Additionally, Raman mapping produces high spatial (~1μm) and spectral resolution. A first Raman study on the globules of Thioploca and Beggiatoa indicated the presence of S8 in a nano-crystalline form (Pasteris et al. 2001). Another study also identified S8 as the main form of elemental sulphur in sulphur globules produced in Beggiatoa filaments, but only in subpopulations located at the sulphide–oxygen interface of gradient tubes or in early growth stage cultures (Berg et al. 2014). Beggiatoa mats in a deeper sulphide-rich, anoxic zone, and freshwater gradient cultures gave rise to Raman signals suggesting a mixture of S8 rings and linear polysulphides (Sn 2−) within the globules. In vivo Raman spectra for Thiothrix presented the characteristic S8 structure previously described for the crystalline S8 standards and molten sulphur in the internal modes (Nims et al. 2019). The significance of the speciation of sulphur lies in the bioavailability of the different forms, i.e. amorphous polymeric or (nano)crystalline elemental sulphur. It has been shown, for example, that the purple sulphur bacterium Allochromatium vinosum, uses only the polymeric component and not the cyclo-octasulphur component when commercially available sulphur is provided as the substrate, i.e. the organism has difficulties to attack the comparatively stable S8 rings (Franz et al. 2007). It is possible that this also applies to sulphur stored inside of cells such that it is deposited in a chemical form that is readily available for further degradation.
8 Formation of Stored Sulphur
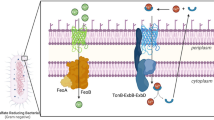
In general, sulphur stored in sulphur globules is formed by the oxidation of more reduced sulphur species. The location of the sulphur inclusions in the periplasmic space implies that globule formation from sulphide, polysulphides, thiosulphate, elemental sulphur and also the much less widely used substrate thiocyanate must occur in this cellular compartment (Fig. 3).
Pathways of sulphur globule formation and degradation. A given organism may contain all or only a subset of the pathways depicted (cf. Table 1). For clarity, reactions are not given with exact stoichiometries. TcDh, thiocyanate dehydrogenase. SQR, sulphide:quinone oxidoreductase, FccAB, flavoctochrome c sulphide dehydrogenase, rDsr, sulphur oxidizing Dsr system, sHdr, sulphur-oxidizing heterodisulphide reductase-like system, Apr, APS reductase, Sat, ATP sulfurylase. Question marks indicate that neither the pathways of sulphane sulphur transport into the cytoplasm nor sulphate export from the cytoplasm have been experimentally clarified
The main characterized enzymes for sulphide oxidation, FAD-containing flavocytochrome c (FccAB) and sulphide:quinone oxidoreductases (SQR) indeed reside in or are oriented towards the periplasm. All characterized SQRs are single-subunit flavoproteins asscociated with the cytoplasmic membrane. Based on protein structure, six distinct types of single subunit flavoprotein SQRs were identified (Marcia et al. 2009, 2010a, b; Shahak and Hauska 2008). Flavocytochrome is present as a soluble protein or as a membrane-bound enzyme and shows sulphide:cytochrome c oxidoreductase activity in vitro (Bosshard et al. 1986). Usually, the protein consists of a larger flavoprotein (FccB) and a smaller haemoprotein (FccA) (Castillo et al. 1994; Fukumori and Yamanaka 1979). All sulphur-globule-forming bacteria contain at least one and often several types of SQR. Flavocytochrome c can be present in addition (Table 1, Fig. 3). In A. vinosum, the gene for SqrF is followed in the same direction of transcription by two genes (Alvin_1196/97) each encoding a very short (32 amino acid) transmembrane protein. Deletion of the genes led to a ~ 60% reduced rate of sulphur formation from sulphide indicating a direct functional relation with SQR. Related genes occur in other purple sulphur bacteria, but nothing is known about their abundance and relevance in other sulphur-globule-forming bacteria (Weissgerber et al. 2013). In A. vinosum, mutants lacking flavocytochrome c sulphide oxidation proceeds with wild-type rates indicating that SQRs play a major role (Reinartz et al. 1998). Flavocytochrome c may represent a high affinity system for sulphide oxidation especially suited at very low sulphide concentration (Brune 1995b).
Polysulphides are the primary reaction products of SQR- and FccAB-catalyzed sulphide oxidation, and indeed, they are well-documented intermediates during the formation of sulphur globules from sulphide in A. vinosum (Prange et al. 2004). It is still unclear whether their conversion into sulphur stored in sulphur globules is a purely chemical process. Theoretically, this is possible because longer polysulphides are in equilibrium with elemental sulphur (Steudel et al. 1990). On the other hand, elevated protein and mRNA levels have been observed in A. vinosum for Alvin_1317–1319, constituting a putative sulphur or polysulphide reductase with highest similarity to archaeal SreABC (Laska et al. 2003). The active site molybdopterin-containing subunit PsrA is localized in the periplasm. This led to the proposal that this enzyme may be involved in the transformation of polysulphides to stored sulphur (Weissgerber et al. 2013; Weissgerber et al. 2014). However, only a minority of sulphur-globule-forming bacteria contains closely related genes shedding doubt on a general role of the encoded enzyme.
As apparent from Table 1, utilization of thiosulphate is very widespread among sulphur-globule-forming bacteria. When thiosulphate is oxidized, only its sulphane group is stored as sulphur; the sulphone sulphur is immediately excreted as sulphate. Thiosulphate oxidation is catalyzed by the periplasmic Sox system, consisting minimally of the proteins SoxXAK, SoxYZ and SoxB (Fig. 1). The heterodimeric SoxYZ protein acts as the central player and serves as a carrier of pathway intermediates (Sauvé et al. 2007). Recently, it has been shown that these intermediates are not simply bound to a cysteine residue located near the carboxy-terminus of the SoxY subunit as previously assumed but that the true carrier species is a SoxYZ-S-sulphane adduct (Grabarczyk and Berks 2017). The c-type cytochrome SoxXA(K) catalyzes the oxidative formation of a disulphide linkage between the sulphane sulphur of thiosulphate and the persulphurated active site cysteine residue of SoxY (Bamford et al. 2002; Ogawa et al. 2008; Grabarczyk and Berks 2017). Then, the sulphone group is hydrolytically released as sulphate. This reaction is catalyzed by SoxB (Grabarczyk et al. 2015; Sauvé et al. 2009) and leaves the original sulphane sulphur of thiosulphate bound to SoxY (Fig. 3). From here, the sulphur is transferred to the sulphur globules by an unknown mechanism, possibly involving the rhodoanese-like protein SoxL (Welte et al. 2009). It may be important to note in this regard that polysulphurated SoxY(S3–4)Z species occur as intermediates of thiosulphate oxidation catalyzed by a reconstituted Sox system in vitro (Grabarczyk and Berks 2017). Such polysulphurated species could serve as direct donors for sulphur globule formation. In organisms that do not form sulphur globules from thiosulphate, the Sox pathway involves one further crucial enzyme, SoxCD. This hemomolybdoprotein acts as a sulphane dehydrogenase and oxidizes the SoxY-bound sulphane sulphur stemming from thiosulphate to the level of a sulphone which is finally hydrolytically released as sulphate in a reaction catalyzed by SoxB. Among the sulphur-storing organisms tabulated in Table 1, Azospirillum thiophilum is the only one containing soxCD-homologous genes. Notably, this organism forms sulphur globules only in the presence of sulphide but not on thiosulphate (Kwak and Shin 2016; Lavrinenko et al. 2010).
Many phototrophic and also chemotrophic sulphur oxidizers use external elemental sulphur as a substrate and transform it into intracellular sulphur deposits before further oxidation (Franz et al. 2007). How external elemental sulphur is transformed into internally stored sulphur is currently completely unclear. A. vinosum needs direct cell–sulphur contact for the uptake of elemental sulphur (Franz et al. 2007). Further details remain to be investigated.
Some Thioalkalivibrio species are able to form sulphur globules from thiocyanate (SCN−) (Berben et al. 2017; Sorokin et al. 2002). Two different pathways for thiocyanate degradation have been described. In the first, a periplasmic cobalt-dependent enzyme, thiocyanate dehydrogenase, catalyzes direct oxidation of the sulphane atom, forming cyanate and sulphur (Berben et al. 2017; Tsallagov et al. 2019) (Fig. 3). The second pathway occurs in Thioalkalivibrio thiocyanodenitrificans (Berben et al. 2017) and involves hydrolysis of the C ≡ N bond by thiocyanate hydrolase to form carbonyl sulphide (COS) and ammonia. The carbonyl sulphide is further hydrolyzed to CO2 and sulphide by carbonyl sulphide hydrolase. T. thiocyanodenitrificans has not been reported to form sulphur globules from thiocyanate (Sorokin et al. 2004). In addition, none of the three genes for the subunits of its thiocyanate hydrolase encode signal peptides mediating transport into the periplasm (Berben et al. 2017). The carbonyl sulphide hydrolase from Thiobacillus thioparus also resides in the cytoplasm (Ogawa et al. 2013). It is therefore highly unlikely that the pathway is relevant for sulphur globule formation and it is not integrated into Fig. 3.
9 Degradation of Stored Sulphur
9.1 Oxidative Degradation
The majority of sulphur-storing organisms has the capacity to completely oxidize sulphur to sulphate (Table 1, Fig. 3). The enzyme systems involved reside in the cytoplasm necessitating sulphur transfer from the periplasm as the storage compartment to the cytoplasm as the compartment for further oxidation. How this transfer is achieved has not been not clarified. Low-molecular-weight organic persulphide such as glutathione amide persulphide has been proposed as a carrier molecule; however although potential transporters for such molecules are encoded in the genome of A. vinosum, they have not been genetically or biochemically characterized from this or any other sulphur-oxidizing prokaryote (Weissgerber et al. 2014).
Sulphur is never processed in a free form in the cytoplasm but rather in a protein-bound persulphidic state (Dahl 2015; Tanabe et al. 2019). A cascade of sulphur transfer reactions usually involving a rhodanese-like protein, a protein of the DsrE family and a TusA homolog delivers the sulphur to an oxidizing enzyme machinery that generates sulphite (Venceslau et al. 2014; Liu et al. 2014; Tanabe et al. 2019; Dahl 2015; Stockdreher et al. 2014). The sulphur carrier protein TusA has been recognized as a central element in these reactions (Tanabe et al. 2019). For better clarity, Fig. 3 shows only this central sulphur carrier protein instead of each single sulphur transferase. The pathway employed for the oxidation of protein-bound sulphane sulphur to sulphite can vary (Fig. 3).
The best-characterized cytoplasmic sulphite-generating pathway involves reverse-acting dissimilatory sulphite reductase rDsrAB as a central player (Pott and Dahl 1998; Dahl et al. 2005; Stockdreher et al. 2012). This pathway occurs in a majority of sulphur-globule-forming organisms (Table 1). The protein DsrC serves as the substrate-binding entity (Cort et al. 2008; Stockdreher et al. 2012). Presumably, the membrane-bound DsrMKJOP electron-transporting complex oxidizes persulphurated DsrC, thus generating a DsrC trisulphide, in which a sulphur atom is bridging two strictly conserved cysteine residues. As DsrC trisulphide has been identified as the reaction product of DsrAB in a sulphate reducer (Santos et al. 2015) and very probably serves as the substrate for oxidation catalysed by rDsrAB which releases sulphite and the reduced DsrC protein as products. The two released electrons are used to generate NADH. This reaction is catalyzed by the iron-sulphur flavoprotein DsrL, an intimate interaction partner of rDsrAB (Löffler et al. 2020).
The second sulphite-generating pathway, the so-called sulphur-oxidizing heterodisulphide reductase-like (sHdr) pathway (Cao et al. 2018; Koch and Dahl 2018), is much less studied and occurs in only a few organisms forming intracellular sulphur globules like Thiorhodospira sibirica (Table 1) and several Thioalkalivibrio species (Berben et al. 2019). The central element of this pathway is an enzyme complex resembling heterodisulphide reductase HdrABC from methanogenic archaea (Kaster et al. 2011; Wagner et al. 2017). The other crucial component of the pathway is a novel lipoate-binding protein (Cao et al. 2018). Both the sHdr complex and the lipoate-binding protein have been identified as indispensable for sulphur compound oxidation in the Alphaproteobacterium Hyphomicrobium denitrificans (Cao et al. 2018; Koch and Dahl 2018). The reaction mechanism of the Hdr-LbpA-based sulphur oxidation system is currently unclear although an experimentally testable hypothesis has recently been put forward (Tanabe et al. 2019).
Neither the Dsr nor the sHdr pathway is confined to sulphur oxidizers with the capacity for depositing intracellular sulphur globules. Both pathways also occur in sulphur oxidizers that do not form sulphur deposits, e.g. species of the genera Thiobacillus or Acidithiobacillus (Quatrini et al. 2009; Beller et al. 2006a, b). The Dsr and sHdr pathways occur virtually exclusively. Only very few organisms bear the genetic potential for both oxidation routes (Berben et al. 2019; Koch and Dahl 2018).
Sulphite is usually oxidized to the final product sulphate. All the sulphur-storing organisms tabulated in Table 1 that contain sulphite-generating enzyme systems in the cytoplasm also have the ability to oxidize sulphite in this compartment. Again, two pathways exist (Fig. 3) that can either occur individually or in parallel. The first pathway involves direct oxidation of sulphite to sulphate via the cytoplasm-oriented membrane-bound iron-sulphur molybdoenzyme SoeABC (Dahl et al. 2013). The second pathway proceeds via formation of the intermediate adenosine-5′-phosphosulphate (APS) and is catalyzed by APS reductase and ATP sulphurylase (Sat) (Dahl 1996; Parey et al. 2013). In A. vinosum, the periplasmic substrate-binding protein SoxYZ is needed in parallel to the cytoplasmic enzymes for effective sulphite oxidation (Dahl et al. 2013). Whether this also applies to other sulphur-oxidizing bacteria has not been elucidated.
9.2 Reductive Degradation
In purple sulphur bacteria, sulphur globules serve as an electron acceptor reserve that allow rudimentary anaerobic respiration under anoxic conditions leading to production of sulphide (van Gemerden 1968). Beggiatoa OH-75-2a used sulphur globules that were accumulated during aerobic thiosulphate oxidation to sustain anaerobic metabolism and several days of anoxia. Reduction of stored sulphur to sulphide with concomitant de novo synthesis of cell material was also found during anoxic incubation of Beggiatoa alba BL18LD. Furthermore, elemental sulphur stored as globules in thioautotrophic symbionts may serve as an electron sink, leading to production of sulphide during temporary anoxia (Gardebrecht et al. 2012; Arndt et al. 2001; Duplessis et al. 2004). Similar processes have been suggested for the sulphur globules in those organisms that lack enzymes to further oxidize stored sulphur, i.e. Dsr or sHdr systems, as is the case for Thiovulum for example (Table 1). Thiovulum may have to oscillate between an aerobic mode of energy conservation in which elemental sulphur accumulates in the cell and an anaerobic mode of energy conservation in which intracellular sulphur serves as an electron acceptor, perhaps with formate acting as an electron donor or via anaerobic sulphur disproportionation (Marshall et al. 2012). The mechanisms underlying reductive degradation of stored sulphur are unresolved.
10 Outlook
Much remains to be learned on bacterial sulphur globules. This especially applies to the abundance, function and structure of the proteins in the globule envelopes. As evident from Table 1, most—if not all—organisms depositing intracellular sulphur encode periplasmic sulphur globule proteins; however, the only organisms for which the proteins have been unambiguously identified are A. vinosum and Thiocapsa roseopersicina (Brune 1995a). None of the proteins have been structurally characterized nor have their interactions been analysed. Further research should also finally clarify the question whether any other proteins involved in formation or degradation of the globules may be specifically attached to the protein envelope.
References
Abouna S, Gonzalez-Rizzo S, Grimonprez A, Gros O (2015) First description of sulphur-oxidizing bacterial symbiosis in a cnidarian (Medusozoa) living in sulphidic shallow-water environments. PLoS One 10(5):e0127625. https://doi.org/10.1371/journal.pone.0127625
Ahn AC, Meier-Kolthoff JP, Overmars L, Richter M, Woyke T, Sorokin DY, Muyzer G (2017) Genomic diversity within the haloalkaliphilic genus Thioalkalivibrio. PLoS One 12(3):e0173517. https://doi.org/10.1371/journal.pone.0173517
Arndt C, Gaill F, Felbeck H (2001) Anaerobic sulfur metabolism in thiotrophic symbioses. J Exp Biol 204:741–750
Bamford VA, Bruno S, Rasmussen T, Appia-Ayme C, Cheesman MR, Berks BC, Hemmings AM (2002) Structural basis for the oxidation of thiosulfate by a sulfur cycle enzyme. EMBO J 21(21):5599–5610
Bazylinski DA, Williams TJ (2006) Ecophysiology of magnetotactic bacteria. In: Schüler D (ed) Magnetoreception and magnetosomes in bacteria, Microbiology monographs, vol 3. Springer, Berlin, pp 37–75
Bazylinski DA, Dean AJ, Williams TJ, Long LK, Middleton SL, Dubbels BL (2004) Chemolithoautotrophy in the marine, magnetotactic bacterial strains MV-1 and MV-2. Arch Microbiol 182(5):373–387
Bazylinski DA, Williams TJ, Lefevre CT, Trubitsyn D, Fang J, Beveridge TJ, Moskowitz BM, Ward B, Schubbe S, Dubbels BL, Simpson B (2013) Magnetovibrio blakemorei gen. nov., sp. nov., a magnetotactic bacterium (Alphaproteobacteria: Rhodospirillaceae) isolated from a salt marsh. Int J Syst Evol Microbiol 63(Pt 5):1824–1833. https://doi.org/10.1099/ijs.0.044453-0
Bazylinski DA, Morillo V, Lefevre CT, Viloria N, Dubbels BL, Williams TJ (2017) Endothiovibrio diazotrophicus gen. nov., sp. nov., a novel nitrogen-fixing, sulfur-oxidizing gammaproteobacterium isolated from a salt marsh. Int J Syst Evol Microbiol 67(5):1491–1498. https://doi.org/10.1099/ijsem.0.001743
Beller HR, Chai PSG, Letain TE, Chakicherla A, Larimer FW, Richardson PM, Coleman MA, Wood AP, Kelly DP (2006a) The genome sequence of the obligately chemolithoautotrophic, facultatively anaerobic bacterium Thiobacillus denitrificans. J Bacteriol 188(4):1473–1488
Beller HR, Letain TE, Chakicherla A, Kane SR, Legler TC, Coleman MA (2006b) Whole-genome transcriptional analysis of chemolithoautotrophic thiosulfate oxidation by Thiobacillus denitrificans under aerobic versus denitrifying conditions. J Bacteriol 188(19):7005–7015
Berben T, Sorokin DY, Ivanova N, Pati A, Kyrpides N, Goodwin LA, Woyke T, Muyzer G (2015) Complete genome sequence of Thioalkalivibrio paradoxus type strain ARh 1T, an obligately chemolithoautotrophic haloalkaliphilic sulfur-oxidizing bacterium isolated from a Kenyan soda lake. Stand Genomic Sci 10:105. https://doi.org/10.1186/s40793-015-0097-7
Berben T, Overmars L, Sorokin DY, Muyzer G (2017) Comparative genome analysis of three thiocyanate oxidizing Thioalkalivibrio species isolated from soda lakes. Front Microbiol 8:254. https://doi.org/10.3389/fmicb.2017.00254
Berben T, Overmars L, Sorokin DY, Muyzer G (2019) Diversity and distribution of sulfur oxidation-related genes in Thioalkalivibrio, a genus of chemolithoautotrophic and haloalkaliphilic sulfur-oxidizing bacteria. Front Microbiol 10:160. https://doi.org/10.3389/fmicb.2019.00160
Berg JS, Schwedt A, Kreutzmann AC, Kuypers MM, Milucka J (2014) Polysulfides as intermediates in the oxidation of sulfide to sulfate by Beggiatoa spp. Appl Environ Microbiol 80(2):629–636. https://doi.org/10.1128/AEM.02852-13
Bergin C, Wentrup C, Brewig N, Blazejak A, Erseus C, Giere O, Schmid M, De Wit P, Dubilier N (2018) Acquisition of a novel sulfur-oxidizing symbiont in the gutless marine worm Inanidrilus exumae. Appl Environ Microbiol 84(7):e02267–e02217. https://doi.org/10.1128/AEM.02267-17
Bland JA, Staley JT (1978) Observations on the biology of Thiothrix. Arch Microbiol 117:79–87
Boden R, Scott KM (2018) Evaluation of the genus Thiothrix Winogradsky 1888 (approved lists 1980) emend. Aruga et al. 2002: reclassification of Thiothrix disciformis to Thiolinea disciformis gen. nov., comb. nov., and of Thiothrix flexilis to Thiofilum flexile gen. nov., comb nov., with emended description of Thiothrix. Int J Syst Evol Microbiol 68(7):2226–2239. https://doi.org/10.1099/ijsem.0.002816
Bosshard HR, Davidson MW, Knaff DB, Millett F (1986) Complex formation and electron transfer between mitochondrial cytochrome c and flavocytochrome c 552 from Chromatium vinosum. J Biol Chem 261:190–193
Bright M, Sorgo A (2003) Ultrastructural reinvestigation of the trophosome in adults of Riftia pachyptila (Annelida, Siboglinidae). Invertebr Biol 122(4):345–366
Brune DC (1995a) Isolation and characterization of sulfur globule proteins from Chromatium vinosum and Thiocapsa roseopersicina. Arch Microbiol 163:391–399
Brune DC (1995b) Sulfur compounds as photosynthetic electron donors. In: Blankenship RE, Madigan MT, Bauer CE (eds) Anoxygenic photosynthetic bacteria, Advances in photosynthesis, vol 2. Kluwer Academic Publishers, Dordrecht, pp 847–870
Bryantseva IA, Gorlenko VM, Kompantseva EI, Imhoff JF, Sling J, Mityushina L (1999) Thiorhodospira sibirica gen. nov., sp. nov., a new alkaliphilic purple sulfur bacterium from a Siberian soda lake. Int J Syst Bacteriol 49:697–703. https://doi.org/10.1099/00207713-49-2-697
Cao X, Koch T, Steffens L, Finkensieper J, Zigann R, Cronan JE, Dahl C (2018) Lipoate-binding proteins and specific lipoate-protein ligases in microbial sulfur oxidation reveal an atpyical role for an old cofactor. eLife 7:e37439. https://doi.org/10.7554/eLife.37439
Castillo MCG, Lou BS, Ondrias MR, Robertson DE, Knaff DB (1994) Characterization of flavocytochrome c 552 from the thermophilic photosynthetic bacterium Chromatium tepidum. Arch Biochem Biophys 315:262–266
Cavanaugh CM (1983) Symbiontic chemoautotrophic bacteria in marine invertebrates from sulfide-rich habitats. Nature 302:58–61
Cavanaugh CM, Gardiner SL, Jones ML, Jannasch HW, Waterbury JB (1981) Prokaryotic cells in the hydrothermal vent tube worm Riftia pachyptila Jones: possible chemoautotrophic symbionts. Science 213:340–342
Chernousova E, Gridneva E, Grabovich M, Dubinina G, Akimov V, Rossetti S, Kuever J (2009) Thiothrix caldifontis sp. nov. and Thiothrix lacustris sp. nov., gammaproteobacteria isolated from sulfide springs. Int J Syst Evol Microbiol 59(12):3128–3135. https://doi.org/10.1099/ijs.0.009456-0
Cohn F (1875) Untersuchungen über Bakterien II. Beitr z Biol d Pflanzen 1:141–207
Condit CM, Meagher RB (1986) A gene encoding a novel glycine-rich structural protein of petunia. Nature 323:178–181. https://doi.org/10.1038/323178a0
Cort JR, Selan UM, Schulte A, Grimm F, Kennedy MA, Dahl C (2008) Allochromatium vinosum DsrC: solution-state NMR structure, redox properties and interaction with DsrEFH, a protein essential for purple sulfur bacterial sulfur oxidation. J Mol Biol 382:692–707
Cosmidis J, Nims CW, Diercks D, Templeton AS (2019) Formation and stabilization of elemental sulfur through organomineralization. Geochim Cosmochim Acta 247:59–82. https://doi.org/10.1016/j.gca.2018.12.025
Dahl C (1996) Insertional gene inactivation in a phototrophic sulphur bacterium: APS-reductase-deficient mutants of Chromatium vinosum. Microbiology 142:3363–3372. https://doi.org/10.1099/13500872-142-12-3363
Dahl C (2015) Cytoplasmic sulfur trafficking in sulfur-oxidizing prokaryotes. IUBMB Life 67(4):268–274. https://doi.org/10.1002/iub.1371
Dahl C (2017) Sulfur metabolism in phototrophic bacteria. In: Hallenbeck PC (ed) Modern topics in the phototrophic prokaryotes: metabolism, bioenergetics and omics. Springer International Publishing, Cham, pp 27–66. https://doi.org/10.1007/978-3-319-51365-2_2
Dahl C, Prange A (2006) Bacterial sulfur globules: occurrence, structure and metabolism. In: Shively JM (ed) Inclusions in prokaryotes, Microbiology monographs, vol 1. Springer, Berlin, Heidelberg, pp 21–51
Dahl C, Engels S, Pott-Sperling AS, Schulte A, Sander J, Lübbe Y, Deuster O, Brune DC (2005) Novel genes of the dsr gene cluster and evidence for close interaction of Dsr proteins during sulfur oxidation in the phototrophic sulfur bacterium Allochromatium vinosum. J Bacteriol 187(4):1392–1404
Dahl C, Friedrich CG, Kletzin A (2008) Sulfur oxidation in prokaryotes. In: Encyclopedia of life sciences (ELS). John Wiley & Sons, Chichester., http://www.els.net/. https://doi.org/10.1002/9780470015902.a9780470021155
Dahl C, Franz B, Hensen D, Kesselheim A, Zigann R (2013) Sulfite oxidation in the purple sulfur bacterium Allochromatium vinosum: identification of SoeABC as a major player and relevance of SoxYZ in the process. Microbiology 159:2626–2638
de Albuquerque JP, Keim CN, Lins U (2010) Comparative analysis of Beggiatoa from hypersaline and marine environments. Micron 41(5):507–517. https://doi.org/10.1016/j.micron.2010.01.009
Distel DL (1998) Evolution of chemoautotrophic endosymbioses in bivalves. Bioscience 48(4):277–286
Dubinina GA, Grabovich MY (1984) Isolation, cultivation and characteristics of Macromonas bipunctata. Mikrobiologiya 53:748–755
Dubinina G, Savvichev A, Orlova M, Gavrish E, Verbarg S, Grabovich M (2017) Beggiatoa leptomitoformis sp. nov., the first freshwater member of the genus capable of chemolithoautotrophic growth. Int J Syst Evol Microbiol 67(2):197–204. https://doi.org/10.1099/ijsem.0.001584
Duplessis MR, Ziebis W, Gros O, Caro A, Robidart J, Felbeck H (2004) Respiration strategies utilized by the gill endosymbiont from the host lucinid Codakia orbicularis (Bivalvia: Lucinidae). Appl Environ Microbiol 70(7):4144–4150
Ehrenberg CG (1838) Die Infusionsthierchen als vollkommene Organismen. Ein Blick in das tiefere organische Leben der Natur. Leopold Voss-Verlag, Leipzig
Eichinger I, Schmitz-Esser S, Schmid M, Fisher CR, Bright M (2014) Symbiont-driven sulfur crystal formation in a thiotrophic symbiosis from deep-sea hydrocarbon seeps. Environ Microbiol Rep 6(4):364–372. https://doi.org/10.1111/1758-2229.12149
Felbeck H (1981) Chemoautotrophic potential of the hydrothermal vent tube worm, Riftia pachyptila Jones (Vestimenifera). Science 213:336–338
Flood BE, Jones DS, Bailey JV (2015) Sedimenticola thiotaurini sp. nov., a sulfur-oxidizing bacterium isolated from salt marsh sediments, and emended descriptions of the genus Sedimenticola and Sedimenticola selenatireducens. Int J Syst Evol Microbiol 65(8):2522–2530. https://doi.org/10.1099/ijs.0.000295
Flood BE, Fliss P, Jones DS, Dick GJ, Jain S, Kaster AK, Winkel M, Mußmann M, Bailey JL (2016) Single-cell (meta-)genomics of a dimorphic Candidatus Thiomargarita nelsonii reveals genomic plasticity. Front Microbiol 3(7):602. https://doi.org/10.3389/fmicb.2016.00603
Franz B, Lichtenberg H, Hormes J, Modrow H, Dahl C, Prange A (2007) Utilization of solid “elemental” sulfur by the phototrophic purple sulfur bacterium Allochromatium vinosum: a sulfur K-edge XANES spectroscopy study. Microbiology 153:1268–1274
Frenkiel L, Gros O, Mouëza M (1996) Gill ultrastructure in Lucina pectinata (Bivalvia: Lucinidae) with reference to hemoglobin in bivalves with symbiotic sulphur-oxidizing bacteria. Mar Biol 125:511–524
Friedrich CG (1998) Physiology and genetics of sulfur-oxidizing bacteria. Adv Microb Physiol 39:235–289
Friedrich CG, Bardischewsky F, Rother D, Quentmeier A, Fischer J (2005) Prokaryotic sulfur oxidation. Curr Opin Microbiol 8(3):253–259
Frigaard NU, Dahl C (2009) Sulfur metabolism in phototrophic sulfur bacteria. Adv Microb Physiol 54:103–200
Frolov EN, Belousova EV, Lavrinenko K, Dubinina GA, Grabovich MY (2013) Capacity of Azospirillum thiophilum for lithotrophic growth coupled to oxidation of reduced sulfur compounds. Microbiology 82(3):271–279
Fukumori Y, Yamanaka T (1979) Flavocytochrome c of Chromatium vinosum. J Biochem 85:1405–1414
Gardebrecht A, Markert S, Sievert SM, Felbeck H, Thürmer A, Albrecht D, Wollherr A, Kabisch J, Le Bris N, Lehmann R, Daniel R, Liesegang H, Hecker M, Schweder T (2012) Physiological homogeneity among the endosymbionts of Riftia pachyptila and Tevnia jerichonana revealed by proteogenomics. ISME J 6(4):766–776. https://doi.org/10.1038/ismej.2011.137
George GN, Pickering IJ, Yu EY, Prince RC (2002) X-ray absorption spectroscopy of bacterial sulfur globules. Microbiology 148:2267–2268
George GN, Gnida M, Bazylinski DA, Prince RC, Pickering IJ (2008) X-ray absorption spectroscopy as a probe of microbial sulfur biochemistry: the nature of bacterial sulfur globules revisited. J Bacteriol 190(19):6376–6383. https://doi.org/10.1128/JB.00539-08
Goffredi SK, Barry JP (2002) Species-specific variation in sulfide physiology between closely related Vesicomyid clams. Mar Ecol Prog Ser 225:227–238
Grabarczyk DB, Berks BC (2017) Intermediates in the sox sulfur oxidation pathway are bound to a sulfane conjugate of the carrier protein SoxYZ. PLoS One 12(3):e0173395. https://doi.org/10.1371/journal.pone.0173395
Grabarczyk DB, Chappell PE, Johnson S, Stelzl LS, Lea SM, Berks BC (2015) Structural basis for specificity and promiscuity in a carrier protein/enzyme system from the sulfur cycle. Proc Natl Acad Sci U S A 112(52):E7166–E7175
Grimonprez A, Molza A, Laurent MCZ, Mansot JL, Gros O (2018) Thioautotrophic ectosymbiosis in Pseudovorticella sp., a peritrich ciliate species colonizing wood falls in marine mangrove. Eur J Protistol 62:43–55. https://doi.org/10.1016/j.ejop.2017.11.002
Guerrero R, Mas J, Pedros-Alio C (1984) Boyant density changes due to intracellular content of sulfur in Chromatium warmingii and Chromatium vinosum. Arch Microbiol 137:350–356
Hageage GJ Jr, Eanes ED, Gherna RL (1970) X-ray diffraction studies of the sulfur globules accumulated by Chromatium species. J Bacteriol 101:464–469
Head IM, Gray ND, Clarke KJ, Pickup RW, Jones JG (1996) The phylogenetic position and ultrastructure of the uncultured bacterium Achromatium oxaliferum. Microbiology 142:2341–2354
Henkel JV, Dellwig O, Pollehne F, Herlemann DPR, Leipe T, Schulz-Vogt HN (2019) A bacterial isolate from the Black Sea oxidizes sulfide with manganese(IV) oxide. Proc Natl Acad Sci U S A 116(25):12153–12155. https://doi.org/10.1073/pnas.1906000116
Himmel D, Maurin LC, Gros O, Mansot JL (2009) Raman microspectrometry sulfur detection and characterization in the marine ectosymbiotic nematode Eubostrichus dianae (Desmodoridae, Stilbonematidae). Biol Cell 101(1):43–54. https://doi.org/10.1042/BC20080051
Jendrossek D (2009) Polyhydroxyalkanoate granules are complex subcellular organelles (carbonosomes). J Bacteriol 191(10):3195–3202
Jørgensen BB, Nelson DC (2004) Sulfide oxidation in marine sediments: geochemistry meets microbiology. In: Sulfur biochemistry – past and present. Geological Society of America, Boulder, Colorado, pp 63–81
Kamyshny A, Ferdelman TG (2010) Dynamics of zero-valent sulfur species including polysulfides at seep sites on intertidal sand flats (Wadden Sea, North Sea). Mar Chem 121(1–4):17–26. https://doi.org/10.1016/j.marchem.2010.03.001
Kamyshny A, Druschel G, Mansaray ZF, Farquhar J (2014) Multiple sulfur isotopes fractionations associated with abiotic sulfur transformations in Yellowstone National Park geothermal springs. Geochem T 15(1):7. https://doi.org/10.1186/1467-4866-15-7
Kaster AK, Moll J, Parey K, Thauer RK (2011) Coupling of ferredoxin and heterodisulfide reduction via electron bifurcation in hydrogenotrophic methanogenic archaea. Proc Natl Acad Sci U S A 108(7):2981–2986
Keim CN, Solorzano G, Farina M, Lins U (2005) Intracellular inclusions of uncultured magnetotactic bacteria. Int Microbiol 8(2):111–117
Keller B, Sauer N, Lamb CJ (1988) Glycine-rich cell wall proteins in bean: gene structure and association of the protein with the vascular system. EMBO J 7:3625–3633. https://doi.org/10.1002/j.1460-2075.1988.tb03243.x
Kleiner M, Petersen JM, Dubilier N (2012) Convergent and divergent evolution of metabolism in sulfur-oxidizing symbionts and the role of horizontal gene transfer. Curr Opin Microbiol 15(5):621–631. https://doi.org/10.1016/j.mib.2012.09.003
Kleinjan WE, de Keizer A, Janssen AJH (2003) Biologically produced sulfur. In: Steudel R (ed) Elemental sulfur and sulfur-rich compounds I. Springer, Berlin, pp 167–187
Kletzin A, Urich T, Müller F, Bandeiras TM, Gomes CM (2004) Dissimilatory oxidation and reduction of elemental sulfur in thermophilic archaea. J Bioenerg Biomembr 36:77–91
Koch T, Dahl C (2018) A novel bacterial sulfur oxidation pathway provides a new link between the cycles of organic and inorganic sulfur compounds. ISME J 12(10):2479–2491. https://doi.org/10.1038/s41396-018-0209-7
Kran G, Schlote FW, Schlegel HG (1963) Cytologische Untersuchungen an Chromatium okenii Perty. Naturwissenschaften 50:728–730
Krieger J, Giere O, Dubilier N (2000) Localization of RubisCO and sulfur in endosymbiontic bacteria of the gutless marine oligochaete Inanidrilus leukodermatus (Annelida). Mar Biol 137:239–244
Kwak Y, Shin JH (2016) First Azospirillum genome from aquatic environments: whole-genome sequence of Azospirillum thiophilum BV-ST, a novel diazotroph harboring a capacity of sulfur-chemolithotrophy from a sulfide spring. Mar Genomics 25:21–24. https://doi.org/10.1016/j.margen.2015.11.001
La Riviere JWM, Schmidt K (1999) Morphologically conspicuous sulfur-oxidizing eubacteria. In: Dworkin M (ed) The prokaryotes: an evolving electronic resource for the microbiological community, vol 3. Springer, New York. http://link.springer-ny.com/link/service/books/10125/
Larkin JM, Shinabarger DL (1983) Characterization of Thiothrix nivea. Int J Syst Bacteriol 33:841–846
Larkin JM, Strohl WR (1983) Beggiatoa, Thiothrix, and Thioploca. Annu Rev Microbiol 37:341–367
Laska S, Lottspeich F, Kletzin A (2003) Membrane-bound hydrogenase and sulfur reductase of the hyperthermophilic and acidophilic archaeon Acidianus ambivalens. Microbiology 149:2357–2371
Lau GE, Cosmidis J, Grasby SE, Trivedi CB, Spear JR, Templeton AS (2017) Low-temperature formation and stabilization of rare allotropes of cyclooctasulfur (β-S8 and γ-S8) in the presence of organic carbon at a sulfur-rich glacial site in the Canadian high Arctic. Geochim Cosmochim Acta 200:218–231. https://doi.org/10.1016/j.gca.2016.11.036
Lavrinenko K, Chernousova E, Gridneva E, Dubinina G, Akimov V, Kuever J, Lysenko A, Grabovich M (2010) Azospirillum thiophilum sp. nov., a diazotrophic bacterium isolated from a sulfide spring. Int J Syst Evol Microbiol 60(Pt 12):2832–2837. https://doi.org/10.1099/ijs.0.018853-0
Lee YJ, Prange A, Lichtenberg H, Rohde M, Dashti M, Wiegel J (2007) In situ analysis of sulfur species in sulfur globules produced from thiosulfate by Thermoanaerobacter sulfurigignens and Thermoanaerobacterium thermosulfurigenes. J Bacteriol 189(20):7525–7529
Lefevre CT, Viloria N, Schmidt ML, Posfai M, Frankel RB, Bazylinski DA (2012) Novel magnetite-producing magnetotactic bacteria belonging to the Gammaproteobacteria. ISME J 6(2):440–450. https://doi.org/10.1038/ismej.2011.97
Lin L, Thanbichler M (2013) Nucleotide-independent cytoskeletal scaffolds in bacteria. Cytoskeleton 70(8):409–423
Liu LJ, Stockdreher Y, Koch T, Sun ST, Fan Z, Josten M, Sahl HG, Wang Q, Luo YM, Liu SJ, Dahl C, Jiang CY (2014) Thiosulfate transfer mediated by DsrE/TusA homologs from acidothermophilic sulfur-oxidizing archaeon Metallosphaera cuprina. J Biol Chem 289(39):26949–26959. https://doi.org/10.1074/jbc.M114.591669
Löffler M, Feldhues J, Venceslau SS, Kammler L, Grein F, IAC P, Dahl C (2020) DsrL mediates electron transfer between NADH and rDsrAB in Allochromatium vinosum. Environ Microbiol 22(2):783–795. https://doi.org/10.1111/1462-2920.14899
Luther GW 3rd, Findlay AJ, Macdonald DJ, Owings SM, Hanson TE, Beinart RA, Girguis PR (2011) Thermodynamics and kinetics of sulfide oxidation by oxygen: a look at inorganically controlled reactions and biologically mediated processes in the environment. Front Microbiol 2:62. https://doi.org/10.3389/fmicb.2011.00062
Macur RE, Jay ZJ, Taylor WP, Kozubal MA, Kocar BD, Inskeep WP (2013) Microbial community structure and sulfur biogeochemistry in mildly-acidic sulfidic geothermal springs in Yellowstone National Park. Geobiology 11(1):86–99. https://doi.org/10.1111/gbi.12015
Maier S, Murray RG (1965) The fine structure of Thioploca ingrica and a comparison with Beggiatoa. Can J Microbiol 11:645–655
Maina JN, Maloyi GMO (1998) Adaptations of a tropical swamp worm, Alma emini, for subsistence in a H2S-rich habitat: evolution of endosymbiontic bacteria, sulfide-metabolizing bodies, and a novel process of elimination of neutralized sulfide complexes. J Struct Biol 122(3):257–266
Maki JS (2013) Bacterial intracellular sulfur globules: structure and function. J Mol Microbiol Biotechnol 23(4–5):270–280
Mangold S, Valdés J, Holmes DS, Dopson M (2011) Sulfur metabolism in the extreme acidophile Acidithiobacillus caldus. Front Microbiol 2:17. https://doi.org/10.3389/fmcib.2011.00017
Mansor M, Hamilton TL, Fantle MS, Macalady JL (2015) Metabolic diversity and ecological niches of Achromatium populations revealed with single-cell genomic sequencing. Front Microbiol 6:822. https://doi.org/10.3389/fmicb.2015.00822
Marcia M, Ermler U, Peng GH, Michel H (2009) The structure of Aquifex aeolicus sulfide:quinone oxidoreductase, a basis to understand sulfide detoxification and respiration. Proc Natl Acad Sci U S A 106(24):9625–9630
Marcia M, Ermler U, Peng GH, Michel H (2010a) A new structure-based classification of sulfide:quinone oxidoreductases. Proteins: Struct Funct Bioinf 78(5):1073–1083
Marcia M, Langer JD, Parcej D, Vogel V, Peng GH, Michel H (2010b) Characterizing a monotopic membrane enzyme. Biochemical, enzymatic and crystallization studies on Aquifex aeolicus sulfide:quinone oxidoreductase. BBA-Biomembranes 1798(11):2114–2123
Markert S, Arndt C, Felbeck H, Becher D, Sievert SM, Hugler M, Albrecht D, Robidart J, Bench S, Feldman RA, Hecker M, Schweder T (2007) Physiological proteomics of the uncultured endosymbiont of Riftia pachyptila. Science 315(5809):247–250
Markert S, Gardebrecht A, Felbeck H, Sievert SM, Klose J, Becher D, Albrecht D, Thurmer A, Daniel R, Kleiner M, Hecker M, Schweder T (2011) Status quo in physiological proteomics of the uncultured Riftia pachyptila endosymbiont. Proteomics 11(15):3106–3117
Marshall IP, Blainey PC, Spormann AM, Quake SR (2012) A single-cell genome for Thiovulum sp. Appl Environ Microbiol 78(24):8555–8563. https://doi.org/10.1128/AEM.02314-12
Maurin LC, Himmel D, Mansot JL, Gros O (2010) Raman microspectrometry as a powerful tool for a quick screening of thiotrophy: an application on mangrove swamp meiofauna of Guadeloupe (F.W.I.). Mar Environ Res 69(5):382–389. https://doi.org/10.1016/j.marenvres.2010.02.001
Meyer B (1976) Elemental sulfur. Chem Rev 76(3):367–388. https://doi.org/10.1021/cr60301a003
Mu T, Zhou J, Yang M, Xing J (2016) Complete genome sequence of Thioalkalivibrio versutus D301 isolated from soda Lake in northern China, a typical strain with great ability to oxidize sulfide. J Biotechnol 227:21–22. https://doi.org/10.1016/j.jbiotec.2016.04.019
Müller C (1870) Chemisch-physikalische Beschreibung der Thermen von Baden in der Schweiz (Canton Aargau). Zehnder, Baden
Müller OF (1786) Animalcula infusoria fluviatilia et marina, quae detexit, systematice descripsit et ad vivum delineari curavit. Mülleri, Hauniae
Nelson DC, Castenholz RW (1981) Use of reduced sulfur compounds by Beggiatoa sp. J Bacteriol 147:140–154
Nelson DC, Fisher CR (1995) Chemoautotrophic and methanoautotrophic endosymbiontic bacteria at deep-sea vents and seeps. In: Karl DM (ed) Deep-sea hydrothermal vents. CRC Press, Boca Raton, FL, pp 125–167
Nicolson GL, Schmidt GL (1971) Structure of the Chromatium sulfur particle and its protein membrane. J Bacteriol 105:1142–1148
Nims C, Cron B, Wetherington M, Macalady J, Cosmidis J (2019) Low frequency Raman spectroscopy for micron-scale and in vivo characterization of elemental sulfur in microbial samples. Sci Rep 9(1):7971. https://doi.org/10.1038/s41598-019-44353-6
Nunoura T, Takaki Y, Kazama H, Kakuta J, Shimamura S, Makita H, Hirai M, Miyazaki M, Takai K (2014) Physiological and genomic features of a novel sulfur-oxidizing Gammaproteobacterium belonging to a previously uncultivated symbiotic lineage isolated from a hydrothermal vent. PLoS One 9(8):e104959. https://doi.org/10.1371/journal.pone.0104959
Odintsova EV, Jannasch H, Mamone JA, Langworthy TA (1996) Thermothrix azorensis sp. nov., an obligately chemolithoautotrophic, sulfur-oxidizing, thermophilic bacterium. Int J Syst Bacteriol 46(2):422–428
Ogawa T, Furusawa T, Nomura R, Seo D, Hosoya-Matsuda N, Sakurai H, Inoue K (2008) SoxAX binding protein, a novel component of the thiosulfate-oxidizing multienzyme system in the green sulfur bacterium Chlorobium tepidum. J Bacteriol 190(18):6097–6110
Ogawa T, Noguchi K, Saito M, Nagahata Y, Kato H, Ohtaki A, Nakayama H, Dohmae N, Matsushita Y, Odaka M, Yohda M, Nyunoya H, Katayama Y (2013) Carbonyl sulfide hydrolase from Thiobacillus thioparus strain THI115 is one of the beta-carbonic anhydrase family enzymes. J Am Chem Soc 135(10):3818–3825. https://doi.org/10.1021/ja307735e
Oren A, Mana L, Jehlicka J (2015) Probing single cells of purple sulfur bacteria with Raman spectroscopy: carotenoids and elemental sulfur. FEMS Microbiol Lett 362(6):fnv021. https://doi.org/10.1093/femsle/fnv021
Ott J, Bright M, Bulgheresi S (2004) Marine microbial thiotrophic ectosymbioses. In: Gibson RN, RJA A, JDM G (eds) Oceanography and marine biology – an annual review, vol 42. CRC Press, Boca Raton, pp 95–118
Overmann J (1997) Mahoney Lake: a case study of the ecological significance of phototrophic sulfur bacteria. Adv Microb Ecol 15:251–288
Parey K, Demmer U, Warkentin E, Wynen A, Ermler U, Dahl C (2013) Structural, biochemical and genetic characterization of ATP sulfurylase from Allochromatium vinosum. PLoS One 8(9):e74707
Pasteris JD, Freeman JJ, Goffredi SK, Buck KR (2001) Raman spectroscopic and laser confocal microscopic analysis of sulfur in living sulfur-precipitating marine bacteria. Chem Geol 180(1–4):3–18
Pattaragulwanit K, Dahl C (1995) Development of a genetic system for a purple sulfur bacterium: conjugative plasmid transfer in Chromatium vinosum. Arch Microbiol 164:217–222
Pattaragulwanit K, Brune DC, Trüper HG, Dahl C (1998) Molecular genetic evidence for extracytoplasmic localization of sulfur globules in Chromatium vinosum. Arch Microbiol 169:434–444
Perty M (1852) Zur Kenntnis kleinster Lebensformen nach Bau, Funktionen, Systematik, mit Spezialverzeichnis der in der Schweiz beobachteten. Jent und Reinert, Bern
Petersen JM, Kemper A, Gruber-Vodicka H, Cardini U, van der Geest M, Kleiner M, Bulgheresi S, Mussmann M, Herbold C, Seah BK, Antony CP, Liu D, Belitz A, Weber M (2016) Chemosynthetic symbionts of marine invertebrate animals are capable of nitrogen fixation. Nat Microbiol 2:16195. https://doi.org/10.1038/nmicrobiol.2016.195
Pfennig N, Trüper HG (1971) Type and neotype strains of the species of phototrophic bacteria maintained in pure culture. Int J Syst Bacteriol 21:19–24
Pickering TJ, Prince RC, Divers T, George GN (1998) Sulfur K-edge X-ray absorption spectroscopy for determining the chemical speciation of sulfur in biological systems. FEBS Lett 441:11–14
Pickering IJ, George GN, Yu EY, Brune DC, Tuschak C, Overmann J, Beatty JT, Prince RC (2001) Analysis of sulfur biochemistry of sulfur bacteria using x-ray absorption spectroscopy. Biochemistry 40:8138–8145
Pott AS, Dahl C (1998) Sirohaem-sulfite reductase and other proteins encoded in the dsr locus of Chromatium vinosum are involved in the oxidation of intracellular sulfur. Microbiology 144:1881–1894
Prange A, Arzberger I, Engemann C, Modrow H, Schumann O, Trüper HG, Steudel R, Dahl C, Hormes J (1999a) In situ analysis of sulfur in the sulfur globules of phototrophic sulfur bacteria by X-ray absorption near edge spectroscopy. Biochim Biophys Acta 1428:446–454
Prange A, Modrow H, Dahl C, Steudel R, Trüper HG, Hormes J (1999b) Structural analysis of sulfur in the sulfur globules of sulfur bacteria by X-Ray Near Edge Absorption Spectroscopy (XANES). Biospektrum Sonderausgabe zur Fr hjahrstagung der VAAM, Göttingen, 85
Prange A, Chauvistr R, Modrow H, Hormes J, Trüper HG, Dahl C (2002) Quantitative speciation of sulfur in bacterial sulfur globules: X-ray absorption spectroscopy reveals at least three different speciations of sulfur. Microbiology 148:267–276
Prange A, Engelhardt H, Trüper HG, Dahl C (2004) The role of the sulfur globule proteins of Allochromatium vinosum: mutagenesis of the sulfur globule protein genes and expression studies by real-time RT PCR. Arch Microbiol 182:165–174
Quatrini R, Appia-Ayme C, Denis Y, Jedlicki E, Holmes DS, Bonnefoy V (2009) Extending the models for iron and sulfur oxidation in the extreme acidophile Acidithiobacillus ferrooxidans. BMC Genomics 10:394. https://doi.org/10.1186/1471-2164-10-394
Rabus R, Venceslau SS, Wohlbrand L, Voordouw G, Wall JD, Pereira IA (2015) A post-genomic view of the ecophysiology, catabolism and biotechnological relevance of sulphate-reducing prokaryotes. Adv Microb Physiol 66:55–321
Reinartz M, Tschäpe J, Brüser T, Trüper HG, Dahl C (1998) Sulfide oxidation in the phototrophic sulfur bacterium Chromatium vinosum. Arch Microbiol 170:59–68
Remsen CC (1978) Comparative subcellular architecture of photosynthetic bacteria. In: Clayton RK, Sistrom WR (eds) The photosynthetic bacteria. Plenum Press, New York, pp 31–62
Remsen CC, Trüper HG (1973) The fine structure of Chromatium buderi. Arch Mikrobiol 90:269–280
Ringli C, Keller B, Ryser U (2001) Glycine-rich proteins as structural components of plant cell walls. Cell Mol Life Sci 58(10):1430–1441. https://doi.org/10.1007/PL00000786
Rinke C, Schmitz-Esser S, Stoecker K, Nussbaumer AD, Molnar DA, Vanura K, Wagner M, Horn M, Ott JA, Bright M (2006) “Candidatus Thiobios zoothamnicoli,” an ectosymbiotic bacterium covering the giant marine ciliate Zoothamnium niveum. Appl Environ Microbiol 72(3):2014–2021. https://doi.org/10.1128/AEM.72.3.2014-2021.2006
Rinke C, Schmitz-Esser S, Loy A, Horn M, Wagner M, Bright M (2009) High genetic similarity between two geographically distinct strains of the sulfur-oxidizing symbiont ‘Candidatus Thiobios zoothamnicoli’. FEMS Microbiol Ecol 67:229–241
Rivière JWM, Schmidt K (2006) Morphologically consicuous sulfur-oxidizing eubacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes, 3rd edn. Springer, New York, NY
Rose A, Meier I (2004) Scaffolds, levers, rods and springs: diverse cellular functions of long coiled-coil proteins. Cell Mol Life Sci 61(16):1996–2009
Roy AB, Trudinger PA (1970) The biochemistry of inorganic compounds of sulfur. Cambridge University Press, London
Russell SL, Corbett-Detig RB, Cavanaugh CM (2017) Mixed transmission modes and dynamic genome evolution in an obligate animal-bacterial symbiosis. ISME J 11(6):1359–1371. https://doi.org/10.1038/ismej.2017.10
Sackett DL, Wolff J (1987) Nile red as a polarity-sensitive fluorescent probe of hydrophobic protein surfaces. Anal Biochem 167:228–234
Salman V, Berben T, Bowers RM, Woyke T, Teske A, Angert ER (2016) Insights into the single cell draft genome of “Candidatus Achromatium palustre”. Stand Genomic Sci 11:28. https://doi.org/10.1186/s40793-016-0146-x
Santos AA, Venceslau SS, Grein F, Leavitt WD, Dahl C, Johnston DT, Pereira IA (2015) A protein trisulfide couples dissimilatory sulfate reduction to energy conservation. Science 350(6267):1541–1545
Sauvé V, Bruno S, Berks BC, Hemmings AM (2007) The SoxYZ complex carries sulfur cycle intermediates on a peptide swinging arm. J Biol Chem 282(32):23194–23204. https://doi.org/10.1074/jbc.M701602200
Sauvé V, Roversi P, Leath KJ, Garman EF, Antrobus R, Lea SM, Berks BC (2009) Mechanism for the hydrolysis of a sulfur-sulfur bond based on the crystal structure of the thiosulfohydrolase SoxB. J Biol Chem 284(32):21707–21718
Schmidt GL, Kamen MD (1970) Variable cellular composition of Chromatium in growing cultures. Arch Mikrobiol 73:1–18
Schmidt GL, Nicolson GL, Kamen MD (1971) Composition of the sulfur particle of Chromatium vinosum. J Bacteriol 105:1137–1141
Schulz HN, Jørgensen BB (2001) Big bacteria. Annu Rev Microbiol 55:105–137
Schulz HN, Brinkhoff T, Ferdelman TG, Hernndez Marin M, Teske A, Jørgensen BB (1999) Dense populations of a giant sulfur bacterium in namibian shelf sediments. Science 284:493–495
Seah BKB, Antony CP, Huettel B, Zarzycki J, Schada von Borzyskowski L, Erb TJ, Kouris A, Kleiner M, Liebeke M, Dubilier N, Gruber-Vodicka HR, Giovannoni SJ (2019) Sulfur-oxidizing symbionts without canonical genes for autotrophic CO2 fixation. mBio 10(3). https://doi.org/10.1128/mBio.01112-19
Shahak Y, Hauska G (2008) Sulfide oxidation from cyanobacteria to humans: sulfide-quinone oxidoreductase (SQR). In: Hell R, Dahl C, Knaff DB, Leustek T (eds) Sulfur metabolism in phototrophic organisms, Advances in photosynthesis and respiration, vol 27. Springer, Dordrecht, pp 319–335
Shigi N (2014) Biosynthesis and functions of sulfur modifications in tRNA. Front Genet 5:67. https://doi.org/10.3389/fgene.2014.00067
Shigi N (2018) Recent advances in our understanding of the biosynthesis of sulfur modifications in tRNAs. Front Microbiol 9:2679. https://doi.org/10.3389/fmicb.2018.02679
Shively JM, Bryant DA, Fuller RC, Konopka AE, Stevens JSE, Strohl WR (1989) Functional inclusions in prokaryotic cells. Int Rev Cytol 113:35–100
Shively JM, Cannon GC, Heinhorst S, Bryant DA, DasSarma S, Bazylinski D, Preiss J, Steinbüchel A, Docampo R, Dahl C (2006) Bacterial inclusions. In: Encyclopedia of life sciences. John Wiley & Sons, Chichester., http://www.els.net/. https://doi.org/10.1038/npg.els.0004268
Skirnisdottir S, Hreggvidsson GO, Holst O, Kristjansson JK (2001) Isolation and characterization of a mixotrophic sulfur-oxidizing Thermus scotoductus. Extremophiles 5(1):45–51. https://doi.org/10.1007/s007920000172
Sorokin DY, Lysenko AM, Mityushina LL, Tourova TP, Jones BE, Rainey FA, Robertson LA, Kuenen GJ (2001) Thioalkalimicrobium aerophilum gen. Nov., sp. nov. and Thioalkalimicrobium sibericum sp. nov., and Thioalkalivibrio versutus gen. nov., sp. nov., Thioalkalivibrio nitratis sp. nov. and Thioalkalivibrio denitrificans sp. nov., novel obligately alkaliphilic and obligately chemolithoautotrophic sulfur-oxidizing bacteria from soda lakes. Int J Syst Evol Microbiol 51:565–580
Sorokin DY, Tourova TP, Lysenko AM, Mityushina LL, Kuenen JG (2002) Thioalkalivibrio thiocyanoxidans sp. nov. and Thioalkalivibrio paradoxus sp. nov., novel alkaliphilic, obligately autotrophic, sulfur-oxidizing bacteria capable of growth on thiocyanate, from soda lakes. Int J Syst Evol Microbiol 52(Pt 2):657–664. https://doi.org/10.1099/00207713-52-2-657
Sorokin DY, Tourova TP, Sjollema KA, Kuenen JG (2003) Thialkalivibrio nitratireducens sp. nov., a nitrate-reducing member of an autotrophic denitrifying consortium from a soda lake. Int J Syst Evol Microbiol 53(Pt 6):1779–1783. https://doi.org/10.1099/ijs.0.02615-0
Sorokin DY, Tourova TP, Antipov AN, Muyzer G, Kuenen JG (2004) Anaerobic growth of the haloalkaliphilic denitrifying sulfur-oxidizing bacterium Thialkalivibrio thiocyanodenitrificans sp. nov. with thiocyanate. Microbiology 150(Pt 7):2435–2442. https://doi.org/10.1099/mic.0.27015-0
Spring S, Bazylinski DA (2000) Magnetotactic bacteria. In: Dworkin M (ed) The prokaryotes: an evolving electronic resource for the microbiological community, vol 3. Springer, New York. http://link.springer-ny.com/link/service/books/10125/
Steudel R (1982) Homocyclic sulfur molecules. Top Curr Chem 102:149–176
Steudel R (1987) Sulfur homocycles. In: Haiduc I, Sowerby DB (eds) The chemistry of inorganic homo- and heterocycles. Academic Press, London, pp 737–768
Steudel R (1989) On the nature of the “elemental sulfur” (S0) produced by sulfur-oxidizing bacteria- a model for S0 globules. In: Schlegel HG, Bowien B (eds) Autotrophic bacteria. Sciene Tech Publishers, Madison, WI, pp 289–303
Steudel R (1996a) Das gelbe Element und seine erstaunliche Vielseitigkeit. Chemie Unserer Zeit 30:226–234
Steudel R (1996b) Mechanism for the formation of elemental sulfur from aqueous sulfide in chemical and microbiological desulfurization processes. Ind Eng Chem Res 35:1417–1423
Steudel R (2000) The chemical sulfur cycle. In: Lens P, Hulshoff Pol W (eds) Environmental technologies to treat sulfur pollution. IWA Publishing, London, pp 1–31
Steudel R, Chivers T (2019) The role of polysulfide dianions and radical anions in the chemical, physical and biological sciences, including sulfur-based batteries. Chem Soc Rev 48(12):3279–3319. https://doi.org/10.1039/c8cs00826d
Steudel R, Eckert B (2003) Solid sulfur allotropes. In: Steudel R (ed) Elemental sulfur and sulfur-rich compounds. Springer, Berlin, pp 1–79
Steudel R, Holz B (1988) Detection of reactive sulfur molecules (S6, S7, S9, S∞) in commercial sulfur, in sulfur minerals, and in sulfur metals slowly cooled to 20°C. Z Naturforsch B43:581–589
Steudel R, Holdt G, Visscher PT, van Gemerden H (1990) Search for polythionates in cultures of Chromatium vinosum after sulfide incubation. Arch Microbiol 155:432–437
Stockdreher Y, Venceslau SS, Josten M, Sahl HG, Pereira IAC, Dahl C (2012) Cytoplasmic sulfurtransferases in the purple sulfur bacterium Allochromatium vinosum: evidence for sulfur transfer from DsrEFH to DsrC. PLoS One 7(7):e40785
Stockdreher Y, Sturm M, Josten M, Sahl HG, Dobler N, Zigann R, Dahl C (2014) New proteins involved in sulfur trafficking in the cytoplasm of Allochromatium vinosum. J Biol Chem 289(18):12390–12403. https://doi.org/10.1074/jbc.M113.536425
Strohl WR, Geffers I, Larkin JM (1981) Structure of the sulfur inclusion envelopes from four Beggiatoas. Curr Microbiol 6:75–79
Strohl WR, Howard KS, Larkin JM (1982) Ultrastructure of Beggiatoa alba strain B15LD. J Gen Microbiol 128:73–84
Tanabe TS, Leimkühler S, Dahl C (2019) The functional diversity of the prokaryotic sulfur carrier protein TusA. Adv Microb Physiol 75:233–277. https://doi.org/10.1016/bs.ampbs.2019.07.004
Taylor CD, Wirsen CO (1997) Microbiology and ecology of filamentous sulfur formation. Science 277(5331):1483–1485. https://doi.org/10.1126/science.277.5331.1483
Taylor CD, Wirsen CO, Gaill F (1999) Rapid microbial production of filamentous sulfur mats at hydrothermal vents. Appl Environ Microbiol 65(5):2235–2255
Trevisan V (1842) Prospetto della Flora Euganea. Coi Tipi del Seminario, Padua, pp 1–68
Trofimov BA, Sinegovskaya LM, Gusarova NK (2009) Vibrations of the S–S bond in elemental sulfur and organic polysulfides: a structural guide. J Sulfur Chem 30(5):518–554. https://doi.org/10.1080/17415990902998579
Trubitsyn D, Abreu F, Ward FB, Taylor T, Hattori M, Kondo S, Trivedi U, Staniland S, Lins U, Bazylinski DA (2016) Draft genome sequence of Magnetovibrio blakemorei strain MV-1, a marine vibrioid magnetotactic bacterium. Genome Announc 4(6). https://doi.org/10.1128/genomeA.01330-16
Trüper HG (2008) Sulfur and light? History and “thiology” of the phototrophic sulfur bacteria. In: Dahl C, Friedrich CG (eds) Microbial sulfur metabolism. Springer, Berlin, Heidelberg, pp 87–100
Tsallagov SI, Sorokin DY, Tikhonova TV, Popov VO, Muyzer G (2019) Comparative genomics of Thiohalobacter thiocyanaticus HRh1T and Guyparkeria sp. SCN-R1, halophilic chemolithoautotrophic sulfur-oxidizing Gammaproteobacteria capable of using thiocyanate as energy source. Front Microbiol 10:898. https://doi.org/10.3389/fmicb.2019.00898
van Gemerden H (1968) On the ATP generation by Chromatium in the dark. Arch Mikrobiol 64:118–124
van Niel CB (1931) On the morphology and physiology of the purple and green sulfur bacteria. Arch Mikrobiol 3:1–112
van Niel BC (1936) On the metabolism of the Thiorhodaceae. Arch Mikrobiol 7:323–358
Venceslau SS, Stockdreher Y, Dahl C, Pereira IAC (2014) The “bacterial heterodisulfide” DsrC is a key protein in dissimilatory sulfur metabolism. Biochim Biophys Acta 1837(7):1148–1164. https://doi.org/10.1016/j.bbabio.2014.03.007
Vetter RD (1985) Elemental sulfur in the gills of three species of clams containing chemoautotrophic symbiontic bacteria: a possible inorganic energy storage compound. Mar Biol 88:33–42
Wagner T, Koch J, Ermler U, Shima S (2017) Methanogenic heterodisulfide reductase (HdrABC-MvhAGD) uses two noncubane [4Fe-4S] clusters for reduction. Science 357(6352):699–703. https://doi.org/10.1126/science.aan0425
Waksman SA (1922) Microorganisms concerned in the oxidation of sulfur in the soil: I. Introductory. J Bacteriol 7(2):231–238
Waksman SA, Joffe JS (1922) Microorganisms concerned in the oxidation of sulfur in the soil: II. Thiobacillus thiooxidans, a new sulfur-oxidizing organism isolated from the soil. J Bacteriol 7(2):239–256
Wang R, Lin J-Q, Liu X-M, Pang X, Zhang C-J, Yang C-L, Gao X-Y, Lin C-M, Li Y-Q, Li Y, Lin J-Q, Chen L-X (2019) Sulfur oxidation in the acidophilic autotrophic Acidithiobacillus spp. Front Microbiol 9:3290. https://doi.org/10.3389/fmicb.2018.03290
Warming E (1875) Om nogle ved Danmarks kyster levede bakterier. Vidensk MeddDanNaturhistForen Khobenhavn 20–28:3–116
Weissgerber T, Zigann R, Bruce D, Chang YJ, Detter JC, Han C, Hauser L, Jeffries CD, Land M, Munk AC, Tapia R, Dahl C (2011) Complete genome sequence of Allochromatium vinosum DSM 180T. Stand Genomic Sci 5(3):311–330. https://doi.org/10.4056/sigs.2335270
Weissgerber T, Dobler N, Polen T, Latus J, Stockdreher Y, Dahl C (2013) Genome-wide transcriptional profiling of the purple sulfur bacterium Allochromatium vinosum DSM 180T during growth on different reduced sulfur compounds. J Bacteriol 195:4231–4245
Weissgerber T, Sylvester M, Kröninger L, Dahl C (2014) A comparative quantitative proteome study identifies new proteins relevant for sulfur oxidation in the purple sulfur bacterium Allochromatium vinosum. Appl Environ Microbiol 80(7):2279–2292. https://doi.org/10.1128/AEM.04182-13
Welte C, Hafner S, Krätzer C, Quentmeier AT, Friedrich CG, Dahl C (2009) Interaction between sox proteins of two physiologically distinct bacteria and a new protein involved in thiosulfate oxidation. FEBS Lett 583:1281–1286
Willems A (2014) The family Comamonadaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes – Alphaproteobacteria and Betaproteobacteria. Springer-Verlag, Berlin, Heidelberg, pp 777–851. https://doi.org/10.1007/978-3-642-30197-1_238
Williams TM, Unz RF, Doman T (1987) Ultrastructure of Thiothrix and “type 012N” bacteria. Appl Environ Microbiol 53(7):1560–1570
Williams TJ, Zhang CL, Scott JH, Bazylinski DA (2006) Evidence for autotrophy via the reverse tricarboxylic acid cycle in the marine magnetotactic coccus strain MC-1. Appl Environ Microbiol 72(2):1322–1329
Winogradsky SN (1887) Über Schwefelbakterien. Botanische Zeitung 45:489–508
Winogradsky SN (1889) Recherches physiologiques sur le sulfobactéries. Ann Inst Pasteur 3:49–60
Zopfi J, Ferdelman TG, Fossing H (2004) Distribution and fate of sulfur intermediates – sulfite, terathionate, thiosuflate and elemental sulfur – marine sediments. In: Sulfur biogeochemistry – past and present. Geological Society of America, Boulder, Colorado, pp 97–116
Acknowledgements
This work was supported by the Deutsche Forschungsgemeinschaft (grants Da 351/6-2 and Da 351/8-1).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Dahl, C. (2020). Bacterial Intracellular Sulphur Globules. In: Jendrossek, D. (eds) Bacterial Organelles and Organelle-like Inclusions. Microbiology Monographs, vol 34. Springer, Cham. https://doi.org/10.1007/978-3-030-60173-7_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-60173-7_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-60172-0
Online ISBN: 978-3-030-60173-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)