Abstract
Epithelial ion channels and transporters play critical roles in salt and water homeostasis. The expression, assembly, and trafficking of these proteins are tightly regulated at both the transcriptional and posttranscriptional levels. For example, ion channel residence at the plasma membrane is regulated in response to intra- and extracellular signals that can alter anterograde trafficking, endocytosis, and/or degradation. While regulation at the plasma membrane has been thoroughly characterized for a number of ion channels and transporters, the mechanisms underlying early events in the endoplasmic reticulum (ER) have only more recently been defined. Notably, all epithelial ion channels and transporters are synthesized at and insert into the ER membrane, at which time posttranslational modifications are added, the protein begins to fold, and subunits of oligomeric proteins assemble. However, before epithelial ion channels and transporters can traffic from the ER, they are subject to protein quality control “decisions,” which ensure that only properly modified, folded, and assembled subunits are directed to the Golgi apparatus and ultimately to the plasma membrane via vesicle intermediates. Here, we first describe the general steps to which ion channels and transporters are subject in the ER and highlight the factors that facilitate protein folding. We will also discuss the events that lead to the degradation of improperly modified, misfolded, and misassembled proteins in the ER. This pathway is known as ER-associated degradation, or ERAD. We will then provide specific examples of critical epithelial proteins that undergo these events in ER protein quality control.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Endoplasmic reticulum (ER)
- ER-associated degradation (ERAD)
- Kidney
- Ion channels
- Ion transporters
- Molecular chaperones
- ENaC
- Na,K-ATPase
- ROMK
- V2R
- NCC
- Aquaporin-2
7.1 Introduction
Approximately one-third of all cellular proteins, including all membrane proteins, transit through the secretory pathway (Ghaemmaghami et al. 2003). The translation of these secretory proteins begins in the cytosol. Signal recognition particle (SRP) binds to either an N-terminal signal sequence or the first transmembrane domain (TMD) of a nascent protein as it emerges from the ribosome and stalls translation (Walter and Blobel 1981; Halic et al. 2004, 2006). The SRP–ribosome nascent chain complex is transferred to the ER membrane by virtue of SRP binding to the SRP receptor, which facilitates ribosome docking to the Sec61 translocation channel in the ER membrane (Meyer and Dobberstein 1980; Gilmore et al. 1982). After the ribosome docks onto Sec61, translation resumes and secretory proteins are cotranslationally translocated into the ER lumen through Sec61 (Fig. 7.1) (Gorlich et al. 1992). For secretory proteins, the signal sequence may be cleaved by signal peptidase (Fig. 7.1a). For membrane proteins, such as epithelial ion channels and transporters, the TMDs move laterally through a Sec61 “gate” and are thus integrated into the lipid bilayer. Alternating TMDs act as either signal sequences or stop transfer sequences (Fig. 7.1b–c) (Egea and Stroud 2010). After membrane integration is complete, downstream events—such as protein folding, oligomeric protein assembly, and posttranslational modifications—occur. These events precede a quality control checkpoint which ensures that only properly folded proteins, and especially membrane proteins, can leave the ER and traffic to the plasma membrane (Barlowe and Helenius 2016).
Protein translocation. The ribosome nascent chain complex docks at the Sec61 channel. For soluble proteins (a) a signal sequence orients the polypeptide with positively charged amino acids facing the cytosol. The signal sequence is cleaved by signal peptidase prior to translocation of the nascent polypeptide chain into the ER lumen. For membrane proteins (b and c) the first transmembrane domain acts as a signal sequence to orient the N-terminus either toward the cytosol (b) or toward the ER lumen (extracellular) (c). Transmembrane domains move through a lateral gate in the Sec61 channel into the lipid bilayer. Subsequent transmembrane domains act as either stop transfers (b) or signal anchors (c) to position the more C-terminal transmembrane domains appropriately. Figure republished with permission of Rapoport et al. (2017). Permission conveyed through Copyright Clearance Center, Inc
7.2 Protein Folding in the Endoplasmic Reticulum
The integration of multi-pass ion-conducting membrane proteins is particularly challenging for the following reasons. First, ion-conducting pores are lined with polar residues that are embedded in the lipid bilayer. In fact, an analysis of all known TMDs predicts that ~30% are only marginally hydrophobic and may not integrate efficiently into the ER membrane (Hessa et al. 2007). Therefore, TMDs either from within the protein or from another subunit in an oligomeric complex are often required to stabilize polar TMDs (Beguin et al. 1998, 2000; Lu et al. 2000; Buck et al. 2007). Second, the final structure of a membrane protein requires the assembly of TMDs that might be synthesized early (i.e., at the N-terminus) and later (i.e., at the C-terminus). Thus, a TMD at the N-terminus might be orphaned for an extended time until it can assemble with a partner TMD. Third, the folding of a membrane protein takes place in three chemically distinct compartments: The ER lumen, which will ultimately be extracellular, the ER lipid bilayer, and the cytoplasm (Fig. 7.1). With few exceptions, it is generally unclear how the folding of a protein in these environments is coordinated. Finally, ribosomes synthesizing subunits of an oligomeric protein may not utilize adjacent Sec61 pores, so after membrane insertion each subunit will need to diffuse in the plane of the ER to find one another. To overcome some of these hurdles, protein folding is aided by the contributions of other factors.
7.2.1 The Role of Molecular Chaperones in Protein Folding
Molecular chaperones bind nascent polypeptides chains as they emerge from the ribosome and facilitate folding, prevent aggregation, promote proper oligomerization, and/or trafficking from the ER. Chaperones continue to act often through cycles of binding and release until a mature protein conformation is reached. Chaperones are a structurally diverse group of proteins that include Hsp70s, Hsp90s, lectins, thiol oxidoreductases, and small heat shock proteins (Buchner 2019; Genest et al. 2019; Lang et al. 2017), each of which are required for the quality control of ion channels and transporters.
One of the most abundant and promiscuous chaperones are the Hsp70s (Fig. 7.2). Hsp70s are located in both the cytosol and the ER and participate in ATP-dependent protein folding. Hsp70s are composed of an N-terminal nucleotide-binding domain (NBD) and a substrate-binding domain (SBD) composed of eight β strands and C-terminal helical “lid domain” that is connected to the SBD via a linker (Fig. 7.2b) (Pobre et al. 2019; Alderson et al. 2016). In the ATP bound state, Hsp70 binds weakly to substrates. ATP hydrolysis, stimulated by the J-domain containing Hsp40s (Fig. 7.2a) (Misselwitz et al. 1998; Kityk et al. 2018), results in a conformational change whereby the “lid” closes over the SBD, which results in a tightly bound state (Fig. 7.2c) (Mayer and Kityk 2015; Mayer 2018). In addition to stimulating Hsp70 ATPase activity, Hsp40s deliver substrate to the Hsp70 and may provide specificity (Fig. 7.2a). In humans, there are 13 Hsp70s and ~50 Hsp40s (Kampinga et al. 2009). After ATP hydrolysis, a second class of co-chaperones, the nucleotide exchange factors (NEFs), stimulate the release of both the hydrolyzed nucleotide (i.e., ADP) and substrate (Fig. 7.2b) (Bracher and Verghese 2015). If the protein fails to reach a mature confirmation, Hsp70 may reengage the substrate to support additional rounds of folding. Alternatively, Hsp70s can target misfolded proteins in the ER for degradation, which is described below.
Hsp70 cycle, cellular roles, and conformational changes. (a) Hsp40 J-domain proteins (JDPs) provide specificity for the diverse cellular roles and various substrates of Hsp70s. (b) Hsp70 ATPase/substrate binding cycle as described in the text. Substrate (dark red) is targeted to Hps70 by JDPs. Structures of bacterial DnaK in the ATP bound open conformation (PDB 4B9Q) (Kityk et al. 2012) and the ADP-bound closed conformation (PDB 2KHO) (Bertelsen et al. 2009) bound to substrate (Zhu et al. 1996) are shown. Substrate-binding domain (SBD) is green. The nucleotide-binding domain (dark (lobe I) and light blue (lobe II)) linker between SBD and NBD is yellow. Nucleotide exchange factors (NEFs) promote release of ADP and ATP rebinding. (c) Model of Hsp70 conformational changes induced by ATP-binding and hydrolysis. Figure republished with permission of Mayer and Gierasch (2019). Permission conveyed through Copyright Clearance Center, Inc
7.2.2 The Role of the Chaperone-Like Lectins in Protein Folding
Another class of ER chaperones, the glycan-binding lectins, recognize proteins containing N-linked glycans. Glycans are appended cotranslationally to polypeptides by the oligosaccharyl-transferase, which appends a core-glycan (GlcNAc2Man9Glc3) to an Asn in the sequence Asn-X-Ser/Thr (where X can be any amino acid except Pro) (Fig. 7.3a). The glycan is then trimmed by glucosidases to yield a mono-glucosylated glycan (GlcNAc2Man9Glc), which enables the binding of two lectins, calnexin (CNX) and calreticulin (CRT) (Fig. 7.3b). CNX is a type I membrane protein, and CRT is a soluble paralog. While CNX/CRT binding was initially thought to be entirely glycan dependent, later evidence indicated that CNX/CRT can bind polypeptides in the absence of glycans (Danilczyk and Williams 2001; Sandhu et al. 2007; Brockmeier et al. 2009). CNX/CRT can also recruit a thioxidoreductase as well as a peptidyl-prolyl isomerase to respectively catalyze the formation of disulfide bonds in the ER and the interconversion of trans–cis Pro, both of which augment protein folding. In most cases, these events are sufficient to permit the mature protein to exit the ER after the action of another glucosidase. However, if folding is problematic, the UDP-glucose glycoprotein glucosyltransferase (UGGT1) acts as a protein folding sensor. This enzyme adds a single glucose back to the core glycan, preventing ER exit and allowing for CNX/CRT reassociation (Fig. 7.3b) (Cherepanova et al. 2016). In principle, the cycle of glucose re-addition by the glucosyltransferase, favoring CNX/CRT binding, and glucose removal could continue indefinitely if the protein has failed to mature. However, proteins that exit the CNX/CRT cycle prior to reaching a mature state can be modified by EDEM (ER degradation enhancing α-mannosidase-like protein) (Ninagawa et al. 2014), thereby generating a substrate that is targeted for ERAD (Figs. 7.3b–c and 7.4, step 1–2). Although this cycle has been established for several model proteins, it remains unclear whether all proteins, especially membrane proteins, are subject to this cycle. In addition, some proteins are not glycosylated or the glycans may not be accessible to the enzymes that regulate protein retention and ERAD. Recent evidence indicates that other lectins, XTP-3B and Os9, help “decode” substrates for ERAD (van der Goot et al. 2018).
N-linked glycosylation and the role of chaperone-like lectins in protein folding and degradation. (a) The core N-linked glycan unit is cotranslationally attached to nascent polypeptide chains as they enter the ER. (b) As described in the text, glycans are trimmed and glycoproteins undergo cycles of lectin binding and release prior to either degradation or reaching a mature confirmation. (c) Yos9 delivers the glycoprotein substrate to the Hrd1 complex by association with Hrd3. Figure republished with permission of Smith et al. (2011). Permission conveyed through Copyright Clearance Center, Inc
ERAD-L/M pathway. Step 1: N-linked glycans are trimmed from 8 to 7 mannose residues (depicted as green circles). Step 2: The lectin Yos9 next binds to the terminal α1,6-linked mannose residues and the misfolded protein is inserted into Hrd1 channel with assistance from Hrd3 and Der1 (see discussion of retrotranslocation for alternative scenarios). Step 3: A single spanning membrane protein (ERAD-M substrate) interacts with Hrd1 through misfolded TMDs. Step 4: Substrate is ubiquitinated by the E3 ligase Hrd1. Step 5: Cdc48 (p97 in mammals) binds Ubx2 and the Cdc48 cofactors, Ufd1/Npl4 (UN) bind to the ubiquitinated substrate. Step 6: The substrate is retrotranslocated from the ER to the cytosol using the energy generated by Cdc48 ATP-hydrolysis. A deubiquitinating enzyme (DUB) trims the ubiquitin chain triggering substrate release from the Cdc48 complex. Step 7: The substrate is degraded by the 26S proteasome. Figure republished with permission of Wu and Rapoport (2018). Permission conveyed through Copyright Clearance Center, Inc
7.3 Endoplasmic Reticulum-Associated Degradation
During ERAD, substrates are first recognized by molecular chaperones (Ismail and Ng 2006; Meusser et al. 2005; Vembar and Brodsky 2008; Hebert and Molinari 2012; Rutkevich and Williams 2011; Young 2014). Following recognition, an integral membrane ERAD substrate—such as an ion channel or transporter—is next ubiquitinated by a series of ER-associated ubiquitin ligases (Fig. 7.4, step 3–5) (Preston and Brodsky 2017). The protein is then “retrotranslocated” or removed from the ER and transported to the cytosol through the action of an energy requiring complex that associates with ubiquitinated proteins (Fig. 7.4, step 6) (Bodnar and Rapoport 2017a). Finally, the ubiquitinated substrate is delivered to and degraded by the cytosolic 26S proteasome (Fig. 7.4, step 7) (Hiller et al. 1996; Hampton et al. 1996; Raasi and Wolf 2007).
7.3.1 Recognition of ERAD Substrates
Chaperones recognize folding lesions that include exposed hydrophobic patches, unpaired charged residues within membrane domains, or discrete sequences termed “degrons” that target proteins for degradation. Extensive experiments to characterize the degradation pathway of a diverse group of ERAD substrates have led to their placement in three categories based on the site of the folding lesion: ERAD-L (ER lumen), ERAD-M (membrane), or ERAD-C (cytosolic) (Vashist and Ng 2004; Taxis et al. 2003; Carvalho et al. 2006). When examined using model substrates, each class requires a unique set of factors for ERAD targeting. We will briefly discuss these requirements here, which were developed using yeast, but we note that the rules positioning substrates into these unique groups do not necessarily hold up for membrane proteins in mammalian cells (Bernasconi et al. 2010).
ERAD-L substrates are soluble proteins and are recognized by ER lumenal chaperones, including an Hsp70, BiP, along with Hsp40 and NEF cochaperones, select lectins, and thiol oxidoreductases. In addition to its role in protein folding, BiP identifies and targets ERAD-L substrates for ERAD and acts in conjunction with a subset of the seven Hsp40s (ERdj1-7) in the ER lumen (Pobre et al. 2019). For example, ERdj3 binds transiently to substrate through conserved, hydrophobic residues before delivering the substrate to BiP. Like most Hsp40s, ERdj3 is a dimer and interacts with BiP through the 70 amino acid J-domain. Once identified, BiP directs misfolded proteins to the mannose trimming EDEM proteins (Oda et al. 2003) and then on to a membrane complex for retrotranslocation and ubiquitination.
Due to the fact that the folding lesion resides within the membrane, ERAD-M substrates generally require fewer chaperones for substrate recognition. BiP, Hsp40s, and lectins are often dispensable to target ERAD-M substrates for degradation (Gardner et al. 2000; Sato et al. 2009). ERAD-M substrates rely instead on the Hrd1 complex to recognize misfolded lesions within the membrane (Figs. 7.4 and 7.5) (Sato et al. 2009). In one model, the exposure of hydrophilic or charged residues within the membrane may act as a folding lesion, which is in contrast to the exposure of hydrophobic residues in soluble proteins (e.g., ERAD-L substrates) or the cytoplasmic domain in ERAD-C proteins. To date, the mechanisms that lead to the selection of ion channels and transporters that are targeted for ERAD have not been defined.
Hrd1 E3 Ligase complex. Model of the Saccharomyces cerevisiae Hrd1 complex required for the degradation of ERAD-L substrates (black). Mammalian homologs are indicated in pink text. Key factors include the putative retrotranslocation channel, Hrd1, the adapter proteins, Usa1, Hrd3, and Der1, that complex with Hrd1, the Cdc48-Ufd1 complex (Npl4 is not shown) that retrotranslocates substrate, components of the ubiquitination machinery (Ubx2, Ubc7, Cue1) and chaperones, Kar2 and Yos9. Figure republished with permission of (Nguyen et al. 2009). Permission conveyed through Copyright Clearance Center, Inc
These substrates are integral membrane proteins with folding lesions facing the cytoplasm and are recognized and targeted for degradation by cytosolic chaperones. For example, a well-characterized ion channel target of the ERAD pathway is the Cystic Fibrosis Transmembrane Conductance Regulator, CFTR. Mutations in CFTR are responsible for Cystic Fibrosis and the most prevalent CFTR mutant, F508del, results in ER retention and premature degradation by the ERAD pathway (Cheng et al. 1990). The bulk of the CFTR protein resides in the cytosol and consequently cytosolic chaperones are required to target CFTR for ERAD, whereas the ER lumenal chaperones are dispensable (Zhang et al. 2001). For example, cytosolic Hsp70 and sHsps are required for CFTR degradation (Rubenstein and Zeitlin 2000; Ahner et al. 2007) (also see Chap. 6 of this volume and Volume 3, Chaps. 15 and 16).
7.3.2 Ubiquitination of ERAD Substrates
The ubiquitination of an ERAD substrate occurs through a series of three enzymatic reactions carried out by an E1 ubiquitin-activating enzyme, an E2 ubiquitin-conjugating enzyme and an E3 ubiquitin ligase. The ubiquitin-activating enzyme catalyzes the formation of a high-energy thioester bond between the C-terminus of ubiquitin and an active site cysteine residue in an ATP-dependent reaction. The ubiquitin is then transferred to the active-site cysteine in an ubiquitin-conjugating enzyme. An E3 enzyme next facilitates the interaction between an E2 and the substrate and the transfer of ubiquitin to, most commonly, a Lys side chain. Ubiquitin conjugation onto Cys, Ser, Thr, or the N-terminus has also been observed (Shimizu et al. 2010; Swatek and Komander 2016). Polyubiquitin chains are formed when ubiquitin is appended to one of the seven Lys residues in ubiquitin (K6, K11, K27, K29, K33, K48, and K63). K48 polyubiquitin chains are the most common ubiquitin linkage for proteasomal degradation, and specificity of the linkage is provided by the E2. At least 4 ubiquitins in series are required for proteasome-dependent degradation. There are ~100 ubiquitin ligases in yeast and ~600 in humans, which reflects the role of these enzymes in substrate recognition, whereas there are only ~38 E2s and 2 E1s in humans (Mehrtash and Hochstrasser 2018). Hrd1 and Doa10 are the dedicated E3s for the ERAD pathway in yeast, and conserved homologs (HRD1 and gp78, and TEB4/MARCH6, respectively) function similarly in human cells. Each of these ligases is an ER membrane protein with the catalytic “RING” domains deposited in the cytosol. Other mammalian E3s associated with the ERAD pathway include TRC8, THEM129, Rfp2, RMA1, RNF145, RNF170, and RNF185 (Stagg et al. 2009; van den Boomen et al. 2014; Younger et al. 2006; Lu et al. 2011; El Khouri et al. 2013; Jiang et al. 2018; Lerner et al. 2007).
To facilitate substrate ubiquitination, each ligase requires somewhat different ubiquitin-conjugating enzymes. Hrd1 acts with Ubc7 in yeast to ubiquitinate ERAD-L or ERAD-M substrates. Ubc7 is tethered to the ER membrane by a membrane protein, Cue1, which directly activates conjugating activity (Fig. 7.5) (Metzger et al. 2013). Hrd1 also forms oligomers that are stabilized by another membrane protein, Usa1, which enhances ubiquitination activity (Horn et al. 2009; Carvalho et al. 2010). In addition, oligomerization increases the activity of gp78, the mammalian homolog of Hrd1, and gp78 can build polyubiquitin chains on the E2, Ube2g2, which in vitro can be transferred en bloc to an ERAD substrate, thereby increasing ubiquitination efficiency (Li et al. 2007). It is not clear whether this occurs in vivo. In contrast, Doa10 functions with both Ubc6 and Ubc7. Unlike Ubc7, Ubc6 is a tail-anchored protein in the ER membrane. Doa10 ubiquitinates ERAD-C substrates but exceptions to this rule have been noted (Habeck et al. 2015). Recently, the role of a conserved 16 amino acid, C-terminal element (CTE) in Doa10 was identified. Mutations in the CTE block the ubiquitination and degradation of select Doa10-dependent substrates. Therefore, the CTE may be important for substrate recognition. Mutations in the CTE of the mammalian homolog, MARCH6/TEB4, resulted in a phenotype similar to the Doa10 mutant (Zattas et al. 2016). Even though the action of these enzymes on the degradation of several ERAD substrates has been noted, the rules that define mechanisms underlying E3-dependent substrate selection are ill defined.
7.3.3 Retrotranslocation of ERAD Substrates
The driving force for the extraction or retrotranslocation of proteins from the ER into the cytosol is provided by the p97 complex (or Cdc48 complex in yeast) (Bays et al. 2001; Jarosch et al. 2002; Rabinovich et al. 2002; Ye et al. 2001). p97/Cdc48 is a hexameric AAA+-ATPase and hydrolyzes ATP to spool proteins through a central aperture (Bodnar and Rapoport 2017b; Ripstein et al. 2017). Each p97/Cdc48 monomer contains two ATPase domains (D1 and D2) and an N-terminal domain (N). The two ATPase domains are stacked on top of one another. The N-terminal domains extend from the D1 ring, and movement of the N domains are coupled to ATP hydrolysis (Tang et al. 2010; Bodnar et al. 2018). p97/Cdc48 is involved in a variety of cellular activities that, in addition to ERAD, include mitochondrial associated degradation, removal of proteins from chromatin, membrane fusion, and vesicular trafficking (Xia et al. 2016). To date, ~38 p97 binding partners and cofactors have been identified, which dictate the cellular activity of p97 (Hanzelmann and Schindelin 2017).
During ERAD, the p97/Cdc48 cofactors—Ufd1 and Npl4—recognize polyubiquitinated proteins (Meyer et al. 2000, 2002). In addition to providing the driving force for retrotranslocation, the enzyme acts as an “unfoldase” (Bodnar et al. 2018). After recognition of a ubiquitinated substrate, ATP binds the D1 ring and the enzyme unfolds the protein using energy provided by ATP hydrolysis in the D2 ring. Partial trimming of the ubiquitin chain by a deubiquitinating enzyme may facilitate entry into the p97/Cdc48 central pore (Bodnar and Rapoport 2017b). ERAD remains proficient in a yeast strain lacking the deubiquitinating enzyme, suggesting that functional analogs are involved in retrotranslocation (Stein et al. 2014).
Does the retrotranslocation of an ERAD substrate require a dedicated channel? Early studies suggested that Sec61 (see, Fig. 7.1) is also required for the retrotranslocation and degradation of soluble ERAD substrates (Pilon et al. 1997, 1998; Wiertz et al. 1996; Plemper et al. 1997; Willer et al. 2008). However, later studies determined that Sec61 is dispensable for their retrotranslocation (Sato and Hampton 2006; Wahlman et al. 2007; Garza et al. 2009). In yeast, recent data strongly suggest that the E3 ubiquitin ligase, Hrd1, can also serve as a retrotranslocation channel for soluble substrates. For example, Hrd1 is necessary and sufficient for the retrotranslocation of an ER lumenal protein in vitro (Carvalho et al. 2010), and the structure of Hrd1 displays channel-like properties (Schoebel et al. 2017). In fact, Hrd1 links ERAD-requiring proteins both in the ER lumen and in the cytoplasm (Nakatsukasa et al. 2013). Nevertheless, the solubilization and extraction of membrane proteins require associated proteins, such as Der1 and Dfm1 (Gauss et al. 2006; Stein et al. 2014; Willer et al. 2008; Garza et al. 2009; Mehnert et al. 2014; Neal et al. 2018).
Yeast Der1 (Derlin in mammals) is an ER resident transmembrane protein required for the degradation of both soluble and membrane proteins, i.e., ERAD-L and ERAD-M substrates (Knop et al. 1996; Vashist et al. 2001; Neal et al. 2018; Hitt and Wolf 2004; Vashist and Ng 2004; Baldridge and Rapoport 2016). Der1 coprecipitates with other proteins involved in the ERAD pathway, and the association between Hrd1 and Der1 is mediated by Usa1 (Fig. 7.5) (Horn et al. 2009; Mehnert et al. 2014). p97 also binds to Der1 in an Hrd1-dependent manner (Gauss et al. 2006), consistent with Der1’s role as an accessory during retrotranslocation (Fig. 7.5). The relationship between Der1 was further delineated via a site-specific cross-linking study in which it was established that the transmembrane domains of Der1 are in direct contact with both Hrd1and Hrd3 and the lumenal domains and the transmembrane domains bind an ERAD substrate (Mehnert et al. 2014). Consistent with a direct role for Der1 in retrotranslocation, antibodies against Der1 inhibited the retrotranslocation of a soluble ERAD substrate that could be observed in real time (Wahlman et al. 2007). Together, Hrd3, which binds BiP (Mehnert et al. 2015), may recruit ERAD clients to Der1 (Fig. 7.5), and as the substrate emerges on the cytosolic face of the ER through the action of Hrd1, it is ubiquitinated by the cytoplasmic RING domain in Hrd1 (Mehnert et al. 2014).
Both Der1/Derlin and Dfm1 are members of the rhomboid protease family, which clip TMDs within membrane-spanning domains. However, both proteins lack critical catalytic residues and are therefore considered inactive rhomboids. A recent study indicated that rhomboids can distort and thin a lipid bilayer, suggesting that these proteins are unable to clip integral membrane ERAD substrates but instead help solubilize them prior to retrotranslocation (Kreutzberger et al. 2019). Consistent with this view, a genetic screen for ERAD effectors in yeast identified Dfm1 as being required for the degradation of integral membrane but not soluble ERAD substrates (Neal et al. 2018). Interestingly, yeast lacking Dfm1 rapidly acquire suppressor mutations that duplicate a region of the yeast chromosome that harbors the Hrd1 gene. Indeed, Hrd1 overexpression compensates for the deletion of Dfm1 (Neal et al. 2018). Together these data position a membrane complex, comprised of Hrd1 and inactive rhomboids, as playing a central role in retrotranslocation, assisted by the action of the p97/Cdc48 ATPase (Fig. 7.5). It is currently unknown whether these factors are involved in the degradation of misfolded epithelial ion channels and transporters.
7.3.4 Degradation by the 26S Proteasome
Prior to proteasome-dependent degradation, accessory factors facilitate the processing and targeting of substrates to this multi-catalytic protease. First, N-linked glycans may be removed by the cytosolic N-glycanase, Png1. Png1 is recruited to the ER membrane by Cdc48 (Li et al. 2006; Suzuki et al. 2000), suggesting that it acts immediately after or with retrotranslocation. Second, Dsk2 and Rad23 harbor both ubiquitin association domains as well as ubiquitin-like domains, which allows them to shuttle substrates to the proteasome (Richly et al. 2005; Verma et al. 2004; Bard et al. 2018). In yeast, deletion of the genes encoding these factors has a minimal effect on ERAD, suggesting that they function with only a subset of substrates. Consistent with this view, mutations in the human Png1 homolog result in a catastrophic disease, which may arise from impaired mitochondrial function (Suzuki 2015; Kong et al. 2018).
The 26S proteasome is composed of the 20S core particle (CP) (Finley et al. 2016) and the 19S regulatory particle (RP). The 20S CP contains four stacked heteroheptameric rings. Two rings of β subunits are flanked by two rings of α subunits. Three out of the seven β subunits contain proteolytic activity (six total active sites). The 20S CP contains three protease activities, chymotrypsin-like, trypsin-like, and caspase-like that cleave after hydrophobic, basic, and acidic residues, respectively. The 20S core of the proteasome forms a chamber that reduces nonspecific degradation of cellular proteins (Bard et al. 2018).
The 19S RP is responsible for unfolding and translocating polypeptide substrates into the 20S proteolytic chamber. The 19S RP is made of a base and a lid. The base consists of a six subunit AAA-ATPase and non-ATPase subunits that have multiple sites for binding ubiquitin and ubiquitin-like proteins, such as Rad23 and Dsk2 (see above). The hexameric AAA-ATPase ring is at the center of the base and provides the “motor” necessary to translocate proteins into the 20S proteolytic chamber. The lid also contains deubiquitinating enzymes that remove ubiquitin chains prior to polypeptide translocation into the 20S CP.
Recently, two cryo-EM structures of a substrate-engaged human 26S proteasome were published (Dong et al. 2019; de la Pena et al. 2018). Based on data from these studies, updated molecular models for substrate processing by the proteasome were proposed. It was reported that the 19S RP can be found in up to seven unique allosteric conformations (Dong et al. 2019; de la Pena et al. 2018; Bard et al. 2018). These include conformational rearrangements of both the base and the lid domain of 19S RP in response to the nucleotide-bound state, the presence of substrate, occupancy of the deubiquitinating enzyme, and the occupancy of ubiquitin receptors. Therefore, substrates are initially targeted through ubiquitin chains by ubiquitin receptors prior to an unstructured region of the substrate initiating ATP-dependent translocation into the CP. Substrate engagement triggers major structural and conformational changes to augment 19S RP-dependent peptide translocation (Bard et al. 2018).
7.4 Epithelial Ion Channels and Transporters Subject to ER Protein Quality Control
7.4.1 The Na,K-ATPase
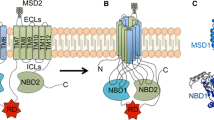
The Na,K-ATPase pump is ubiquitously expressed in epithelia and can reside on either the apical or basolateral membrane. The Na,K-ATPase facilitates the transport of sodium and potassium across the membrane and generates ion gradients to maintain membrane potential and drive vectorial sodium-dependent transepithelial transport by pumping three Na+ ions into the extracellular space in exchange for two K+ ions (Jorgensen 1986). The Na,K-ATPase pump is a heterodimer composed of an ⍺-subunit that contains the ion-transport activity, and a β-subunit that is essential for channel maturation and trafficking (Geering 2008). The ⍺-subunit is a ten TMD-containing protein with N- and C-termini facing the cytosol (Blanco 2005). The β-subunit is a single TMD-spanning glycoprotein with a short cytosolic tail and large extracellular domain (Geering 2001; Blanco 2005). While some studies suggested alternative subunit ratios (Yoshimura et al. 2008; Clifford and Kaplan 2008) analysis of the high-resolution structures of the Na,K-ATPase confirms a 1:1 ⍺:β stoichiometry (Morth et al. 2007; Shinoda et al. 2009). There are four isomers of the ⍺-subunit (⍺1, ⍺2, ⍺3, and ⍺4) and three isoforms of the β-subunit (β1, β2, and β3), and each show both tissue-specific and developmental differences in their expression patterns (Geering 2001; Blanco 2005). Each isoform also has unique kinetic properties and responds differently to substrate, ATP, and/or inhibitors (Blanco and Mercer 1998; Crambert et al. 2000). There is also evidence that the Na,K-ATPase is involved in cell adhesion and the maintenance of adherens junctions as the cytosolic domain of the pump binds E-cadherin and the ankyrin and spectrin cytoskeleton (Kizhatil et al. 2007; Nelson et al. 1990; Nelson and Veshnock 1987). The β-subunit dimerizes with β-subunits on opposing cells to maintain adherens junctions (Tokhtaeva et al. 2011; Padilla-Benavides et al. 2010). A more thorough discussion of the physiological roles and function of the Na,K-ATPase are provided in Volume 3, Chap. 1, of this series.
7.4.1.1 Assembly and ER-Associated Degradation of the Na,K-ATPase
In the absence of the β-subunit, the ⍺-subunit is retained in the ER and targeted for ERAD (Beguin et al. 1998, 2000; Beggah et al. 1996, 1999). Several studies have provided insight into the mechanism by which the ⍺-subunit is degraded (Beguin et al. 1998, 2000; Hasler et al. 2001). First, a series of truncated ⍺-subunit TMD constructs were expressed in oocytes. Data indicated that the first four TMDs act alternatively as efficient “signal-anchor sequences” and “stop-transfer sequences” (see Fig. 7.1), thereby integrating TMD1-4 in the predicted topology (Beguin et al. 1998). However, efficient membrane integration of TMD6-10 required both C-terminal sequence information and the presence of the β-subunit. In the absence of the β-subunit, TMD7 was unable to act as a signal anchor sequence but remained in the cytosol. Therefore, the extracellular loop between TMD7-8, which contains a degradation signal (Beguin et al. 2000), is exposed to the cytosol, creating an unstable, immature protein (Beguin et al. 1998). Coexpressing the β-subunit promoted membrane integration of TMD7 and masked the degradation signal (Beguin et al. 2000). Membrane integration of the TMD7-8 pair resulted in proper localization of the TMD7-8 loop within the ER lumen and stabilized not only the full-length ⍺-subunit but any construct containing at least TMD1-8. In the presence of the β-subunit, exposure of the degradation signal is only transient (Beguin et al. 1998, 2000). Thus, the β-subunit exhibits chaperone-like activity.
Although channel assembly is required for ER exit of the ⍺1-subunit (Beguin et al. 1998, 2000; Beggah et al. 1996, 1999), whether oligomerization is a prerequisite for trafficking of the β-subunit is less clear (Noguchi et al. 1990). However, later studies established that all mature β-subunits were bound to ⍺-subunits (Clifford and Kaplan 2008), and Xenopus oocyte β-subunits were ER retained in the absence of ⍺-subunits (Ackermann and Geering 1990; Jaunin et al. 1992; Beggah et al. 1996). The ⍺-subunit also appears to be the limiting factor for ⍺-β assembly (Tokhtaeva et al. 2009). As with ⍺-subunits, ER retained unassembled β subunits are rapidly degraded, and mutations in the β-subunit TMD at residues predicted to interact with ⍺-subunit TMD7 or the β-subunit extracellular domain results in ER-retention of the β-subunit and reduced colocalization with the ⍺-subunit (Shinoda et al. 2009). Corresponding mutations in the ⍺-subunit similarly resulted in impaired assembly and ER retention of the subunits (Tokhtaeva et al. 2009). These studies are all consistent with an inability of the β-subunit to traffic to the Golgi in the absence of the ⍺-subunit.
7.4.1.2 The Roles of Chaperones in Na,K-ATPase Regulation
The role of molecular chaperones in Na,K-ATPase folding, assembly, quality control, and trafficking is not well characterized, but some evidence exists that BiP and calnexin regulate Na,K-ATPase biogenesis (Beggah et al. 1996; Tokhtaeva et al. 2010). Moreover, the least stable isoforms of the α- and β-subunits associated strongly with BiP, as did ER-retained subunits, whereas subunits with complex (Golgi appended) glycans did not coprecipitate with BiP. Consistent with these observations, coexpressing the α- and β-subunits reduced BiP association, presumably by promoting assembly, maturation, and trafficking of the Na,K-ATPase heterodimer (Beggah et al. 1996). These data strongly suggest that BiP is important either for Na,K-ATPase folding and assembly and/or for preferentially binding and targeting misfolded subunits for ERAD. These data are consistent with earlier data on yeast BiP, which is required for the folding and secretion of a wild-type version of a yeast glycoprotein; when mutated, BiP instead targets the substrate for ERAD (Simons et al. 1995; Plemper et al. 1997).
As described above, calnexin recognizes N-linked glycans and retains immature proteins in the ER. The three isoforms of the Na/K-ATPase β-subunit, β1, β2, and β3, possess 3, 8, and 2 glycosylation sites, respectively. Surprisingly, β1 and β3 subunit functions are not impaired when glycosylation is blocked (Takeda et al. 1988; Lian et al. 2006; Beggah et al. 1997; Zamofing et al. 1989). However, when glycosylation of the β2 isomer is blocked, the protein is retained in the ER and fails to assemble with the α-subunit. All eight glycosylation sites are not required for trafficking of β2, but glycosylation sites two and seven are critical (Tokhtaeva et al. 2010). These data are consistent with studies in yeast that established a hierarchy of glycosylation sites in secreted proteins: Those that reside in hard-to-fold regions appear to be most vital for selecting the protein for ER retention and ERAD (Spear and Ng 2005). Consistent with these data, hydrophobic amino acids near glycosylation sites are required for calnexin binding and calnexin-mediated maturation of the β2-subunit (Tokhtaeva et al. 2010).
7.4.2 The Epithelial Sodium Channel
Epithelial Sodium Channel (ENaC) is formed by homologous α-, β-, and γ-subunits, and each subunit contains two transmembrane domains (TMDs), a large extracellular loop (ECL) and cytosolic N- and C-termini (Canessa et al. 1994a). ENaC is highly expressed in the lung, colon, and kidney epithelia where it reabsorbs sodium. In the kidney, ENaC is expressed on the apical surface of the distal convoluted tubule, connecting tubule, and collecting duct, where ENaC is responsible for Na+ reabsorption from urine and regulates salt and water homeostasis and therefore blood pressure (Rossier 2014; Bhalla and Hallows 2008; Snyder 2005; Palmer 2015). ENaC also provides the driving force for K+ secretion by acting in conjunction with the ROMK potassium channel. Gain of function mutations in ENaC lead to hypertension and Liddle’s syndrome (Staub et al. 1996), and loss of function mutations lead to hypotension, salt-wasting and Pseudohypoaldosteronism Type 1 (PHA1) (Grunder et al. 1997; Bhalla and Hallows 2008). In the lung, ENaC is negatively regulated by CFTR. In the absence of CFTR, ENaC hyperactively reabsorbs Na+ which further dehydrates the airway mucosa and exacerbates cystic fibrosis symptoms (Mall et al. 2004; Hobbs et al. 2013; Gentzsch et al. 2010; Ismailov et al. 1996; Yan et al. 2004). ENaC trafficking and surface expression are highly regulated and have been extensively studied, which is discussed in Volume 3, Chap. 18 of this series.
7.4.2.1 Posttranslational Modifications of ENaC
The ENaC subunits are modified by N-linked glycosylation, the formation of disulfide bonds, palmitoylation, acetylation, and proteolytic cleavage in the Golgi (Buck and Brodsky 2018). Interestingly, the α-, β-, and γENaC subunits are subject to different posttranslational modifications, even though they are ~30–40% identical (Bhalla and Hallows 2008; Canessa et al. 1994a; Mueller et al. 2010; Valentijn et al. 1998; Weisz et al. 2000; Snyder et al. 1994). For example, the α-, β-, and γENaC subunits contain 6, 13, and 7 sites for N-linked glycosylation, respectively (Buck and Brodsky 2018), and blocking glycosylation in any subunit altered channel conductance and Na+-dependent inhibition (Kashlan et al. 2018). Expression of β-subunits lacking glycans led to the most dramatic effect, with an overall reduction in Na+ current of ~50–80%. Interestingly, the α-subunit was more sensitive to protease under these conditions. Although glycosylation of the α-subunit was previously shown to be dispensable (Canessa et al. 1994a; Snyder et al. 1994), other studies established that tunicamycin treatment, which blocks the addition of N-linked sugars, inhibited ENaC current (Kieber-Emmons et al. 1999). ENaC trafficking is also subject to Na+-dependent regulation: low Na+ favors the maturation of glycans, consistent with trafficking to the later secretory pathway, whereas high Na+ results in the acquisition of immature glycans (Heidrich et al. 2015). Because glycan maturation occurs in the Golgi under low Na+ conditions (0 mM Na+ in media), ENaC may follow an alternative trafficking pathway that bypasses the Golgi under conditions of high Na+ (130 mM Na+ in media) (Heidrich et al. 2015).
Members of the protein disulfide isomerase (PDI) family oxidize Cys residues within the ER to form disulfide bonds in nascent proteins (Galligan and Petersen 2012). The formation of disulfide bonds requires a thioredoxin motif (Cys-X-X-Cys), and to date 21 human PDIs are known (Galligan and Petersen 2012). Each ENaC subunit contains 16 conserved Cys residues, so perhaps not surprisingly each subunit in the channel forms several intrasubunit disulfide bonds, which are important for function (Buck and Brodsky 2018). For example, mutating the first or sixth Cys in α- or βENaC reduces surface expression, but mutating any of the other Cys residues results in an increase in sodium current. Disulfide bonds are also required for self-inhibition in response to Na+ (Sheng et al. 2007; Firsov et al. 1999).
In contrast to most PDIs that harbor two thioredoxin motifs, some family members contain only a single Cys-X-X-Cys motif. These PDIs are unable to catalyze the formation of disulfide bonds in substrates but can form covalent bonds to single Cys residues and thus rearrange existing disulfide bonds. One of these family members is ERp29, which promotes ENaC trafficking from the ER to the Golgi (Grumbach et al. 2014). Specifically, ERp29 promotes the association of βENaC with a cargo recognition component of the coat complex II (COPII), which regulates ER exit and thus channel maturation.
A third posttranslational modification to which ENaC subunits are subject is the conjugation of palmitate, which is transferred to a Cys, Ser, or Thr near a hydrophobic amino acid patch by a palmitoyl transferase (De and Sadhukhan 2018; Mueller et al. 2010; Mukherjee et al. 2014). Palmitoylation increases the net hydrophobicity of a substrate, and ~50% of all human ion channels are palmitoylated. The addition of palmitate can inhibit protein–protein interactions, act as a signal for endocytosis and block recycling to the plasma membrane (Shipston 2011; Nadolski and Linder 2007; Linder and Deschenes 2007; Fukata et al. 2016). Both the βENaC (Cys43 and Cys557) and the γENaC (Cys33 and Cys41) subunits are modified (Mueller et al. 2010; Mukherjee et al. 2014). Treatment of cortical collecting duct cells with a palmitoyl transferase inhibitor reduced ENaC-specific Na+ current (Mukherjee et al. 2017), and mutating the palmitoylation sites decreased ENaC channel open probability (Po) and therefore reduced Na+ current (Mueller et al. 2010; Mukherjee et al. 2014). Among the 23 mammalian palmitoyl transferases, coexpression of five (DHCC1, 2, 3, 7, and 14) increased Na+ current, and ENaC coprecipitated with each of these enzymes (Mukherjee et al. 2017). All of the transferases are expressed in either the ER and/or the Golgi (Ohno et al. 2006), which raises the possibility that this modification affects early events during ENaC quality control.
Finally, each of the ENaC subunits is acetylated (Butler et al. 2015). Although protein acetylation is usually associated with histones, many other proteins are subject to this modification (Sadoul et al. 2008; Drazic et al. 2016). ENaC subunit acetylation increases both the cellular levels of the channel and the amount of ENaC present at the plasma membrane (Butler et al. 2015). Acetylation inhibitors increase ENaC current and expression, perhaps by inhibiting ubiquitination. Because ubiquitination of ENaC at the cell surface triggers endocytosis, these data suggest that acetylation and ubiquitination of Lys residues on the cytoplasmic tails of ENaC are antagonistic (Zhou et al. 2013; Oberfeld et al. 2011; Butler et al. 2015). In contrast, HDAC7 deacetylates ENaC early in the secretory pathway, because inhibition of HDAC7 increases the total ENaC pool, i.e., not just the population of surface-expressed ENaC (Butler et al. 2015). In the future, it will be important to determine if HDAC7-mediated deacetylation alters the targeting of ENaC for ERAD.
7.4.2.2 Regulation of ENaC by ERAD
Expression of all three ENaC subunits is required for efficient channel assembly and trafficking to the cell surface, and when the subunits are expressed individually they are rapidly targeted for ERAD (Snyder 2005; Staub et al. 1997; Valentijn et al. 1998; Asher et al. 1996; Masilamani et al. 1999). Moreover, even when all three subunits are present a significant percentage of the ENaC subunits continue to be degraded (Staub et al. 1997; Valentijn et al. 1998; Malik et al. 2001; Kashlan et al. 2007). To identify factors required for ENaC degradation, a yeast ENaC expression system was developed (Mueller et al. 2007; Buck et al. 2010, 2017). Initial studies using the yeast model confirmed that ENaC is targeted for ERAD, and more specifically that ENaC degradation requires the proteasome and the Cdc48 complex (Kashlan et al. 2007) (see above). The sHsps, which prevent protein aggregation in an ATP-independent manner, are also required for ⍺ENaC degradation (Kashlan et al. 2007). Importantly, overexpression of a mammalian sHsp in MDCK cells decreased ENaC expression, and co-expression of the sHsp with ENaC decreased Na+ current and ENaC surface expression, consistent with results from the yeast model (Kashlan et al. 2007). Additionally, deletion of the ER lumenal Hsp40s, Jem1, and Scj1, which normally interact with BiP, stabilized ENaC in yeast, and co-expression of the human homologs with ENaC in Xenopus oocytes also reduced ENaC current and surface expression (Buck et al. 2010). Surprisingly, ENaC degradation showed no dependence on the yeast BiP homolog, even though the Hsp40s played an important role in degradation (Buck et al. 2010). Because ENaC ubiquitination was reduced in the absence of the Hsp40s, the Hsp40s—but not BiP—appear to be required to recognize or target ENaC for degradation.
Although BiP does not regulate ENaC degradation in yeast, another Hsp70-like molecule, Lhs1, is present in the ER. Lhs1 is a member of the Hsp110/GRP170 family of proteins, which act as a NEF for Hsp70 (Baxter et al. 1996; Craven et al. 1996; Steel et al. 2004; Tyson and Stirling 2000). Similar to Hsp70, mammalian Lhs1 can also bind misfolded proteins (Behnke and Hendershot 2014) and acts as a “holdase” to prevent misfolded protein aggregation (de Keyzer et al. 2009). In the yeast model, we found that ⍺ENaC degradation required this Hsp70-like NEF, and an unglycosylated species of ⍺ENaC was preferentially stabilized (Buck et al. 2013). Consistent with these results, the mammalian Lhs1 homolog, GRP170, also promoted ⍺ENaC degradation and preferentially bound to unglycosylated ⍺ENaC in HEK293 cells (Buck et al. 2013). In contrast to strains lacking the Hsp40s, ubiquitinated ⍺ENaC accumulated in the ER membrane when Lhs1 was deleted (Buck et al. 2017). These data suggest that Lhs1/GRP170 acts at a post-ubiquitination step and are consistent with this factor playing a role in ⍺ENaC retrotranslocation. Similarly, a role for GRP170 in the retrotranslocation of a viral protein and a misfolded glycoprotein, NHK, from the ER was observed, but retrotranslocation was BiP-dependent in both of these cases (Inoue and Tsai 2015, 2016).
Even though BiP is dispensable for ENaC biogenesis or quality control, the cytosolic Hsp70 promotes ENaC trafficking to the cell surface by enhancing ENaC association with COPII (Chanoux et al. 2012). Surprisingly, the closely related, constitutively expressed homolog of Hsp70, Hsc70, instead promotes ENaC degradation (Chanoux et al. 2013; Goldfarb et al. 2006). Although a mechanism for Hsc70-mediated degradation is unknown, the chaperone probably targets ENaC to the ERAD pathway. Consistent with these results, some stabilization of ENaC was noted when degradation was examined in yeast expressing a temperature-sensitive allele of a cytosolic Hsp70, Ssa1 (Buck et al. 2010).
Other modulators of ENaC stability and trafficking include PON2 (Paraoxonase-2) and Derlin-1 (Shi et al. 2017; You et al. 2017). PON2 is an ER-associated lactonase that is expressed in the ENaC-expressing principal cells of the distal nephron (Shi et al. 2017). The Caenorhabditis elegans PON2 homolog, MEC-6, promotes trafficking of an ENaC family member, the mechanosensitive degenerin, MEC-4 (Harel et al. 2004; Chelur et al. 2002). In transfected HEK293 cells, PON2 coimmunoprecipitates with ENaC, but in contrast to MEC-6, PON2 overexpression lowers ENaC currents and reduces ENaC surface expression (Shi et al. 2017). The reason that PON2 and MEC-6 appear to function antagonistically toward their respective sodium channels is unclear, but more work on these ill-defined chaperone-like proteins is essential. Similarly, overexpression of Derlin-1 promotes ENaC degradation in mammalian cells and associates with the channel (You et al. 2017). However, whether or not Derlin-1 serves as the ENaC retrotranslocation channel alone on in conjunction with other factors, perhaps gp78 or HRD1, must also be defined.
In yeast, ENaC degradation is dependent on both the ERAD-L/M E3 ubiquitin ligase, Hrd1, as well as the ERAD-C ubiquitin ligase, Doa10 (Buck et al. 2010). Whereas βENaC is nearly completely stabilized in a yeast strain lacking both Hrd1 and Doa10, a significant portion of both ⍺- and γENaC continues to be degraded. This result suggests that a third unidentified ubiquitin ligase targets ⍺- and γENaC for degradation. Because Hrd1 and Doa10 play a role in ENaC degradation, ENaC may contain folding lesions in multiple cellular compartments or Hrd1 and Doa10 may compensate for one another. To date, it remains mysterious which E3 ubiquitin ligases act on misfolded forms of ENaC in the mammalian ER.
7.4.2.3 ENaC Channel Assembly and ER Exit
Assembly of the α-, β-, and γENaC subunits likely occurs in the ER, and indirect evidence supports this assumption. First, in yeast, where nearly all ENaC is retained in the ER, the ENaC subunits coprecipitate (Buck et al. 2010, 2017). Second, in mammalian systems, immature channels lacking complex glycans or Golgi-associated protease activation also associate with one another, suggesting assembly occurs in a pre-Golgi compartment (Hughey et al. 2003). Third, when constructs containing only single ⍺ENaC domains were coexpressed with wild-type ENaC, channel trafficking was blocked (Bruns et al. 2003). These data imply that the single domain constructs assembled with wild-type ENaC subunits and prevented ENaC trafficking. While some heteromeric membrane proteins assemble in a defined order (Wanamaker et al. 2003; Adams et al. 1997; Feige et al. 2015), there is no evidence that ENaC assembly is also ordered. Nevertheless, intersubunit contacts between each of the ENaC domains is required for channel assembly, which was highlighted in the recent cryo-EM structure of the trimeric channel (Noreng et al. 2018). Assembly of ENaC TMDs is particularly important to reach a mature conformation. Evidence to support this view is derived from the expression of a series of chimeric ⍺-βENaC constructs, which were then coexpressed with wild-type ⍺ENaC and γENaC. It was established that interactions between the TMDs alone, and not interactions between the large ER loops, are required to evade chaperone-dependent degradation (Buck et al. 2017).
What is the signal for the assembled channel to exit the ER after assembly is complete? Early data indicated that the ⍺ENaC subunit can form a homotrimer that inefficiently traffics to the plasma membrane in the absence of the β- and γENaC subunits, whereas the β- and γENaC subunits require the ⍺-subunit for ER exit (Canessa et al. 1994b). Therefore, it was hypothesized that ⍺ENaC possesses an ER exit signal (Canessa et al. 1994b; Mueller et al. 2007). Indeed, using a reporter protein fused to cytoplasmic segments of ENaC, a five amino acid motif was identified that acted as a trafficking signal (Mueller et al. 2007). The mechanism by which these residues promote ER exit, or whether a competing ER retention sequence is also present, is unknown.
7.4.3 Other Epithelial Ion Channels and Transporters Regulated by ERAD
7.4.3.1 Renal Outer Medullary Potassium Channel
Renal Outer Medullary Potassium Channel (ROMK) (Kir1.1/KCNJ1), is a member of the inwardly rectifying potassium channel family. ROMK is a homotetramer, and each ROMK subunit contains two TMDs, a potassium selectivity filter, and large cytosolic N- and C-terminal tails (Ho et al. 1993). Like ENaC, ROMK is expressed on the apical surface of epithelial cells in the distal nephron of the kidney. ROMK is responsible for potassium secretion and also regulates salt-water homeostasis. ROMK physiology, trafficking, and regulation are discussed in depth in Volume 3, Chap. 19, of this series.
To date, over 60 mutations have been identified in ROMK that result in Type II Bartter syndrome, a disease that is associated with early developmental defects and lifelong hypokalemic alkalosis (Welling 2016; Ji et al. 2008). N-terminal frameshifts result in a nonfunctional channel, but some mutations alter ion selectivity, gating, or ligand binding (Fang et al. 2010; Peters et al. 2003). Although it was unknown whether other mutations instead altered protein folding, potentially targeting the channel for ERAD, four mutations that cluster in a C-terminal immunoglobulin-like domain result in trafficking defects in oocytes and/or HEK293 cells (Fallen et al. 2009; Peters et al. 2003). Later, via the development of a yeast model, the same mutants were found to be selected for proteasome-dependent degradation in a p97/Cdc48- and Hsp70-dependent manner (O’Donnell et al. 2017; Mackie et al. 2018). The same mutants were also degraded in a proteasome- and p97/Cdc48-dependent manner in mammalian cells. It will be important to next define which of the other uncharacterized disease-causing mutants are also targeted for ERAD.
7.4.3.2 Thiazide-Sensitive Sodium Chloride Cotransporter
Sodium Chloride Cotransporter (NCC) is a member of the SLC12 cation chloride cotransporter family of electroneutral transporters and is a 12 TMD protein with a glycosylated extracellular loop and large cytosolic N- and C-termini (Castaneda-Bueno et al. 2010). NCC is expressed on the apical surface of the distal convoluted tubule and also regulates sodium reabsorption, thereby playing a major role in salt-water homeostasis and blood pressure regulation. The transporter is also a target for the thiazide diuretics. Mutations in NCC result in Gitelman Syndrome, which is typified by salt-wasting, hypokalemic alkalosis, hypomagnesemia, and hypocalciura (Fulchiero and Seo-Mayer 2019). NCC is discussed in depth in Volume 3, Chap. 3, of this series.
Several characterized Gitelman Syndrome-causing mutations in NCC result in the production of an immature protein that contains only core N-linked glycans (Kunchaparty et al. 1999; De Jong et al. 2002; Sabath et al. 2004). To determine whether NCC is an ERAD substrate, and whether these mutations accelerated ERAD, a yeast expression system for NCC was developed (Needham et al. 2011). As anticipated, NCC was primarily ER localized, and degradation was slowed in the presence of a proteasome inhibitor. Further, NCC was ubiquitinated primarily by Hrd1 in an Hsp70-dependent manner. In mammalian cells, immature NCC was again ubiquitinated and targeted for proteasome-dependent degradation. Consistent with the data in yeast, Hsp70 overexpression increased the proportion of immature NCC and increased the degradation rate (Needham et al. 2011). A role for accelerated chaperone-dependent ERAD in Gitelman Syndrome was then verified via a mass spectrometry analysis. Major hits from this screen included Hsp90 and Hsp90 cochaperones, CHIP—an E3 ubiquitin ligase previously implicated in ERAD (McDonough and Patterson 2003)—an Hsp70/Hsp90 organizer protein, an Hsp40, as well as the known NCC effector, Hsp70 (Donnelly et al. 2013). Through the application of an Hsp90 inhibitor, it was established that NCC maturation depended on Hsp90 function (Donnelly et al. 2013). Notably, Gitelman syndrome mutants associated more strongly with Hsp70, Hsp40, and CHIP than wild-type NCC (Donnelly et al. 2013). The data are consistent with sequential NCC quality control in the ER, whereby the Hsp70 and Hsp90 chaperone complex acts sequentially to monitor NCC folding and ultimately promote channel trafficking (Donnelly et al. 2013).
7.4.3.3 V2 Vasopressin Receptor
The V2 vasopressin receptor (V2R) is a G-protein coupled receptor (GPCR) stimulated by arginine-vasopressin (AVP) (Birnbaumer et al. 1992). AVP responds to either an increase in NaCl or a decrease in blood pressure and binds V2R, which resides in the basolateral membrane of principal cells in the distal nephron. Activation of V2R by AVP triggers AQP2 insertion in the plasma membrane and allows for water reabsorption from urine. In the absence of functional V2R, AQP2 remains embedded in intracellular vesicles. Mutations in the V2R (AVPR2 gene) cause X-linked nephrogenic diabetes insipidus (NDI) (Hoekstra et al. 1996), and >270 NDI-causing mutations have been identified (Milano et al. 2017). These mutations are responsible for ~90% of the inherited cases of NDI, while the other ~10% are caused by mutations in the AQP2 gene, as described below. Patients with NDI are unable to respond to AVP and produce copious volumes of urine (polyuria) and exhibit excessive thirst (polydipsia).
NDI-causing V2R mutants are either retained in the ER or Golgi, or they traffic to the plasma membrane and are defective (Milano et al. 2017). Among nine NDI-causing mutants, most are ER-retained and subject to ERAD (Robben et al. 2005). Later, three V2R mutants with alterations in the cytosolic portion of the protein (L62P), within the membrane (V226E), and within the lumen (G201D) were characterized (Schwieger et al. 2008). G201D localized throughout the secretory pathway, whereas the L62P mutant was found exclusively in the ER. The V226E mutant was visualized in both the ER and ER-Golgi intermediate compartment. Proteasome inhibition stabilized the unglycosylated and core glycosylated species of both the wild type and the mutant V2Rs, and all of the proteins were ubiquitinated to varying degrees. Consistent with V2R being a substrate for ERAD, the unglycosylated and core glycosylated V2R mutants coimmunoprecipitated with p97 and Derlin-1. Although the chaperone and ubiquitin ligase requirements for degradation were not addressed, this will be an important future area of research.
7.4.3.4 Aquaporin-2
Aquaporin-2 (AQP2) is responsible for water reabsorption in the distal nephron and is expressed on the apical surface of principal cells, where ENaC, V2R, and ROMK also reside. AQP2 is a six TMD homotetramer (Bai et al. 1996). Over 60 NDI-causing mutations in AQP2 have been described (Milano et al. 2017), many of which lead to protein retention in the ER (Lin et al. 2002; Lloyd et al. 2005; Tamarappoo and Verkman 1998; Marr et al. 2002; Iolascon et al. 2007). Some of these mutants function if they reach the cell surface, suggesting the folding defect is minor (Leduc-Nadeau et al. 2010; Marr et al. 2002). Interestingly, 11 of the 65 mutants are dominant (Kuwahara et al. 2001; Sohara et al. 2006; Asai et al. 2003; Mulders et al. 1998) and form heterotrimers with wild-type subunits that are either misrouted to the lysosome (Marr et al. 2002) or the basolateral membrane (Kamsteeg et al. 2003) or are retained in the Golgi (Mulders et al. 1998; Procino et al. 2003).
Perhaps not surprisingly, several of the AQP2 NDI-associated mutants are ERAD substrates, but there is a dearth of information on the factors that recognize and target AQP2 mutants for degradation. One study observed that, in contrast to wild-type AQP2, the glycosylated form of four AQP2 mutants was significantly more stable than the unglycosylated species. Therefore, lectins may be involved in AQP2 folding and quality control, but this has not been investigated (Buck et al. 2004). There is also some evidence that the ERAD-associated E3 ubiquitin ligase, CHIP (see above), regulates AQP2 levels (Wu et al. 2018). AQP2 levels are higher in cells lacking CHIP, and CHIP seems to be required for both proteasomal and lysosomal AQP2 degradation. In an in vitro assay CHIP ubiquitinates AQP2, but Hsc70 blocks ubiquitination. AQP2 expression is also increased in a CHIP knockout mouse and water intake is lower, which is consistent with increased AQP2 activity (Wu et al. 2018). A more thorough discussion of AQPs is provided in Volume 3, Chap. 30, of this series.
7.4.3.5 Polycystin-2
Polycystin-2 (PC2) is a six TMD nonselective cation channel with cytosolic N- and C-termini (Mochizuki et al. 1996). The transmembrane domain of PC2 is homologous to polycystin-1 (PC1), which is a PC2 partner, and the TRP channels (Somlo and Ehrlich 2001). PC2 function is dependent on cellular residence. In the ER, PC2 acts as a Ca2+ release channel (Clapham 2003; Koulen et al. 2002), whereas at the plasma membrane PC2 may facilitate Ca2+ entry (Luo et al. 2003). In embryonic nodal cells and the primary cilia of renal tubular cells, PC2 acts as a flow sensor (McGrath et al. 2003), and when PC2 is inhibited the Ca2+ response to flow stimulation is absent (Nauli et al. 2003). Mutations in PC2—as well as PC1—are responsible for autosomal dominant polycystic kidney disease (ADPKD) (Torres and Harris 2006), which is the fourth leading cause of kidney failure.
Mounting evidence suggests that PC2 is targeted for ERAD. First, PC2 is ubiquitinated, and PC2 degradation is proteasome dependent (Liang et al. 2008). Second, PC2 degradation is dependent on p97, and PC2-GFP localizes to the ER and coprecipitates with p97 in MDCK cells. Liang et al. (2008) also determined that Herp, which may function similarly to the Hrd1-Der1 assembly factor, Usa1 (see above) and recruits substrates to this complex (Leitman et al. 2014), contributes to PC2 ERAD. It will be interesting to determine whether disease-causing mutations in PC2 enhance ERAD, as seen for several of the substrates discussed above, and which chaperones select PC2 for degradation.
7.4.3.6 The Sodium-Potassium Chloride Cotransporter-2
The sodium-potassium chloride cotransporter-2, NKCC2, is another renal protein that, when mutated, gives rise to a disease associated with altered salt-water balance (type I Bartter Syndrome). NKCC2, which is homologous to NCC (see above), resides in the thick ascending limb of the loop of Henle and plays a pivotal role in NaCl reabsorption. Prior work established that the protein is targeted for ERAD, which is consistent with its slow rate of ER exit (Zaarour et al. 2012). More recent evidence indicates that immature, ER glycosylated forms of the substrate are selected for degradation through the action of the ER lectin Os9 (Seaayfan et al. 2016). As in many of the examples outlined in this section, ongoing work on NKCC2 will better define how the transporter is targeted for ERAD. A more thorough discussion of NKCC2 function is provided in Volume 3, Chap. 2, of this series.
7.5 Conclusions and Future Directions
In this chapter, we have provided an overview of the many hurdles that a nascent protein in the ER must overcome. We then outlined the chaperones, chaperone-like factors, and enzymes that impact protein folding and degradation, and focused on the mechanisms that lead to the selection, ubiquitination, and delivery of an ERAD substrate to the proteasome. As described in the preceding section, a growing body of literature indicates that TMD-containing proteins, many of which reside in epithelia, are targeted to this pathway. While disease-causing mutations may slow ER exit and hasten ERAD, even wild-type versions of these proteins fold inefficiently and are targeted for ERAD. Therefore, much has been learned from studying the biogenesis of both wild type and disease-associated mutants. As highlighted throughout the chapter the use of model systems has led to significant insights on ER quality control processes.
Although the field is in its infancy, there is growing interest in harnessing the knowledge obtained from studies on ER protein quality toward the development of therapeutics that correct misfolded epithelial proteins. Therefore, a better definition of how these varied ion channels and transporters assemble in the ER and traffic to the plasma membrane is imperative. Small molecules that target any of these steps might lead to the development of new therapeutic options.
For example, pharmacological chaperones are under development that target ER retained V2R mutants (Los et al. 2010). Like molecular chaperones, pharmacological chaperones are designed to assist protein folding. One class of these molecules are the nonpeptide vasopressin receptor antagonists, or vaptans (Francis and Tang 2004). Vaptans mimic the structure of AVP (Mouillac et al. 1995), are cell permeable, and thus bind the AVP binding site on V2R in the ER. Vaptans stabilize the mutant V2R allowing for maturation and trafficking while avoiding ERAD (Milano et al. 2017). Six vaptans can promote V2R trafficking, maturation, plasma membrane expression, and the AVP response (Robben et al. 2007; Morello et al. 2000; Wuller et al. 2004; Bernier et al. 2004). In small scale clinical trials for a vaptan, SR49059, patient urine volume was decreased while urine osmolality was significantly increased.
Perhaps the best example in which pharmacological approaches to target ERAD-associated diseases has been successful is Cystic Fibrosis. The F508del CFTR mutation, which targets this unfolded, ER-retained protein for ERAD, is significantly stabilized, can traffic to the apical membrane, and function in the presence of an FDA-approved drug combination (Lukacs and Verkman 2012). Current knowledge suggests these compounds bind directly to the mutated channel. One compound helps link two domains during channel synthesis, while the other compound binds to the cell surface population of F508del CFTR and increases its open probability (Clancy 2018). While improvements in the efficacy of this treatment are actively being pursued, these data provide hope that other drugs that target misfolded ion channels and transporters can be identified. In the case of Cystic Fibrosis, a molecular definition of the cause of the disease was instrumental in developing this therapy. For the other diseases discussed in this chapter, we similarly propose that basic research will pave the way for future cures.
References
Ackermann U, Geering K (1990) Mutual dependence of Na,K-ATPase alpha- and beta-subunits for correct posttranslational processing and intracellular transport. FEBS Lett 269(1):105–108
Adams CM, Snyder PM, Welsh MJ (1997) Interactions between subunits of the human epithelial sodium channel. J Biol Chem 272(43):27295–27300
Ahner A, Nakatsukasa K, Zhang H, Frizzell RA, Brodsky JL (2007) Small heat-shock proteins select {delta}F508-CFTR for endoplasmic reticulum-associated degradation. Mol Biol Cell 18(3):806–814
Alderson TR, Kim JH, Markley JL (2016) Dynamical structures of Hsp70 and Hsp70-Hsp40 complexes. Structure 24(7):1014–1030. https://doi.org/10.1016/j.str.2016.05.011
Asai T, Kuwahara M, Kurihara H, Sakai T, Terada Y, Marumo F, Sasaki S (2003) Pathogenesis of nephrogenic diabetes insipidus by aquaporin-2 C-terminus mutations. Kidney Int 64(1):2–10. https://doi.org/10.1046/j.1523-1755.2003.00049.x
Asher C, Wald H, Rossier BC, Garty H (1996) Aldosterone-induced increase in the abundance of Na+ channel subunits. Am J Phys 271(2 Pt 1):C605–C611
Bai L, Fushimi K, Sasaki S, Marumo F (1996) Structure of aquaporin-2 vasopressin water channel. J Biol Chem 271(9):5171–5176
Baldridge RD, Rapoport TA (2016) Autoubiquitination of the Hrd1 ligase triggers protein retrotranslocation in ERAD. Cell 166(2):394–407. https://doi.org/10.1016/j.cell.2016.05.048
Bard JAM, Goodall EA, Greene ER, Jonsson E, Dong KC, Martin A (2018) Structure and function of the 26S proteasome. Annu Rev Biochem 87:697–724. https://doi.org/10.1146/annurev-biochem-062917-011931
Barlowe C, Helenius A (2016) Cargo capture and bulk flow in the early secretory pathway. Annu Rev Cell Dev Biol 32:197–222. https://doi.org/10.1146/annurev-cellbio-111315-125016
Baxter BK, James P, Evans T, Craig EA (1996) SSI1 encodes a novel Hsp70 of the Saccharomyces cerevisiae endoplasmic reticulum. Mol Cell Biol 16(11):6444–6456
Bays NW, Wilhovsky SK, Goradia A, Hodgkiss-Harlow K, Hampton RY (2001) HRD4/NPL4 is required for the proteasomal processing of ubiquitinated ER proteins. Mol Biol Cell 12(12):4114–4128
Beggah A, Mathews P, Beguin P, Geering K (1996) Degradation and endoplasmic reticulum retention of unassembled alpha- and beta-subunits of Na,K-ATPase correlate with interaction of BiP. J Biol Chem 271(34):20895–20902
Beggah AT, Jaunin P, Geering K (1997) Role of glycosylation and disulfide bond formation in the beta subunit in the folding and functional expression of Na,K-ATPase. J Biol Chem 272(15):10318–10326
Beggah AT, Beguin P, Bamberg K, Sachs G, Geering K (1999) beta-subunit assembly is essential for the correct packing and the stable membrane insertion of the H,K-ATPase alpha-subunit. J Biol Chem 274(12):8217–8223
Beguin P, Hasler U, Beggah A, Horisberger JD, Geering K (1998) Membrane integration of Na,K-ATPase alpha-subunits and beta-subunit assembly. J Biol Chem 273(38):24921–24931
Beguin P, Hasler U, Staub O, Geering K (2000) Endoplasmic reticulum quality control of oligomeric membrane proteins: topogenic determinants involved in the degradation of the unassembled Na,K-ATPase alpha subunit and in its stabilization by beta subunit assembly. Mol Biol Cell 11(5):1657–1672
Behnke J, Hendershot LM (2014) The large Hsp70 Grp170 binds to unfolded protein substrates in vivo with a regulation distinct from conventional Hsp70s. J Biol Chem 289(5):2899–2907. https://doi.org/10.1074/jbc.M113.507491
Bernasconi R, Galli C, Calanca V, Nakajima T, Molinari M (2010) Stringent requirement for HRD1, SEL1L, and OS-9/XTP3-B for disposal of ERAD-LS substrates. J Cell Biol 188(2):223–235. https://doi.org/10.1083/jcb.200910042
Bernier V, Lagace M, Lonergan M, Arthus MF, Bichet DG, Bouvier M (2004) Functional rescue of the constitutively internalized V2 vasopressin receptor mutant R137H by the pharmacological chaperone action of SR49059. Mol Endocrinol 18(8):2074–2084. https://doi.org/10.1210/me.2004-0080
Bertelsen EB, Chang L, Gestwicki JE, Zuiderweg ER (2009) Solution conformation of wild-type E. coli Hsp70 (DnaK) chaperone complexed with ADP and substrate. Proc Natl Acad Sci U S A 106(21):8471–8476. https://doi.org/10.1073/pnas.0903503106
Bhalla V, Hallows KR (2008) Mechanisms of ENaC regulation and clinical implications. J Am Soc Nephrol 19(10):1845–1854
Birnbaumer M, Seibold A, Gilbert S, Ishido M, Barberis C, Antaramian A, Brabet P, Rosenthal W (1992) Molecular cloning of the receptor for human antidiuretic hormone. Nature 357(6376):333–335. https://doi.org/10.1038/357333a0
Blanco G (2005) Na,K-ATPase subunit heterogeneity as a mechanism for tissue-specific ion regulation. Semin Nephrol 25(5):292–303. https://doi.org/10.1016/j.semnephrol.2005.03.004
Blanco G, Mercer RW (1998) Isozymes of the Na-K-ATPase: heterogeneity in structure, diversity in function. Am J Phys 275(5 Pt 2):F633–F650
Bodnar N, Rapoport T (2017a) Toward an understanding of the Cdc48/p97 ATPase. F1000Res 6:1318. https://doi.org/10.12688/f1000research.11683.1
Bodnar NO, Rapoport TA (2017b) Molecular mechanism of substrate processing by the Cdc48 ATPase complex. Cell 169(4):722–735.e729. https://doi.org/10.1016/j.cell.2017.04.020
Bodnar NO, Kim KH, Ji Z, Wales TE, Svetlov V, Nudler E, Engen JR, Walz T, Rapoport TA (2018) Structure of the Cdc48 ATPase with its ubiquitin-binding cofactor Ufd1-Npl4. Nat Struct Mol Biol 25(7):616–622. https://doi.org/10.1038/s41594-018-0085-x
Bracher A, Verghese J (2015) The nucleotide exchange factors of Hsp70 molecular chaperones. Front Mol Biosci 2:10. https://doi.org/10.3389/fmolb.2015.00010
Brockmeier A, Brockmeier U, Williams DB (2009) Distinct contributions of the lectin and arm domains of calnexin to its molecular chaperone function. J Biol Chem 284(6):3433–3444. https://doi.org/10.1074/jbc.M804866200
Bruns JB, Hu B, Ahn YJ, Sheng S, Hughey RP, Kleyman TR (2003) Multiple epithelial Na+ channel domains participate in subunit assembly. Am J Physiol Renal Physiol 285(4):F600–F609
Buchner J (2019) Molecular chaperones and protein quality control: an introduction to the JBC Reviews thematic series. J Biol Chem 294(6):2074–2075. https://doi.org/10.1074/jbc.REV118.006739
Buck TM, Brodsky JL (2018) Epithelial sodium channel biogenesis and quality control in the early secretory pathway. Curr Opin Nephrol Hypertens 27(5):364–372. https://doi.org/10.1097/MNH.0000000000000438
Buck TM, Eledge J, Skach WR (2004) Evidence for stabilization of aquaporin-2 folding mutants by N-linked glycosylation in endoplasmic reticulum. Am J Physiol Cell Physiol 287(5):C1292–C1299. https://doi.org/10.1152/ajpcell.00561.2003
Buck TM, Wagner J, Grund S, Skach WR (2007) A novel tripartite motif involved in aquaporin topogenesis, monomer folding and tetramerization. Nat Struct Mol Biol 14(8):762–769
Buck TM, Kolb AR, Boyd C, Kleyman TR, Brodsky JL (2010) The ER associated degradation of the epithelial sodium channel requires a unique complement of molecular chaperones. Mol Biol Cell 21(6):1047–1058
Buck TM, Plavchak L, Roy A, Donnelly BF, Kashlan OB, Kleyman TR, Subramanya AR, Brodsky JL (2013) The Lhs1/GRP170 chaperones facilitate the endoplasmic reticulum-associated degradation of the epithelial sodium channel. J Biol Chem 288(25):18366–18380. https://doi.org/10.1074/jbc.M113.469882
Buck TM, Jordahl AS, Yates ME, Preston GM, Cook E, Kleyman TR, Brodsky JL (2017) Interactions between intersubunit transmembrane domains regulate the chaperone-dependent degradation of an oligomeric membrane protein. Biochem J 474(3):357–376. https://doi.org/10.1042/BCJ20160760
Butler PL, Staruschenko A, Snyder PM (2015) Acetylation stimulates the epithelial sodium channel by reducing its ubiquitination and degradation. J Biol Chem 290(20):12497–12503. https://doi.org/10.1074/jbc.M114.635540
Canessa CM, Merillat AM, Rossier BC (1994a) Membrane topology of the epithelial sodium channel in intact cells. Am J Phys 267(6 Pt 1):C1682–C1690
Canessa CM, Schild L, Buell G, Thorens B, Gautschi I, Horisberger JD, Rossier BC (1994b) Amiloride-sensitive epithelial Na+ channel is made of three homologous subunits. Nature 367(6462):463–467
Carvalho P, Goder V, Rapoport TA (2006) Distinct ubiquitin-ligase complexes define convergent pathways for the degradation of ER proteins. Cell 126(2):361–373
Carvalho P, Stanley AM, Rapoport TA (2010) Retrotranslocation of a misfolded luminal ER protein by the ubiquitin-ligase Hrd1p. Cell 143(4):579–591. https://doi.org/10.1016/j.cell.2010.10.028
Castaneda-Bueno M, Vazquez N, Bustos-Jaimes I, Hernandez D, Rodriguez-Lobato E, Pacheco-Alvarez D, Carino-Cortes R, Moreno E, Bobadilla NA, Gamba G (2010) A single residue in transmembrane domain 11 defines the different affinity for thiazides between the mammalian and flounder NaCl transporters. Am J Physiol Renal Physiol 299(5):F1111–F1119. https://doi.org/10.1152/ajprenal.00412.2010
Chanoux RA, Robay A, Shubin CB, Kebler C, Suaud L, Rubenstein RC (2012) Hsp70 promotes epithelial sodium channel functional expression by increasing its association with coat complex II and its exit from endoplasmic reticulum. J Biol Chem 287(23):19255–19265. https://doi.org/10.1074/jbc.M112.357756
Chanoux RA, Shubin CB, Robay A, Suaud L, Rubenstein RC (2013) Hsc70 negatively regulates epithelial sodium channel trafficking at multiple sites in epithelial cells. Am J Physiol Cell Physiol 305(7):C776–C787. https://doi.org/10.1152/ajpcell.00059.2013
Chelur DS, Ernstrom GG, Goodman MB, Yao CA, Chen L, Hagan RO’, Chalfie M (2002) The mechanosensory protein MEC-6 is a subunit of the C. elegans touch-cell degenerin channel. Nature 420(6916):669–673. https://doi.org/10.1038/nature01205
Cheng SH, Gregory RJ, Marshall J, Paul S, Souza DW, White GA, O’Riordan CR, Smith AE (1990) Defective intracellular transport and processing of CFTR is the molecular basis of most cystic fibrosis. Cell 63(4):827–834
Cherepanova N, Shrimal S, Gilmore R (2016) N-linked glycosylation and homeostasis of the endoplasmic reticulum. Curr Opin Cell Biol 41:57–65. https://doi.org/10.1016/j.ceb.2016.03.021
Clancy JP (2018) Rapid therapeutic advances in CFTR modulator science. Pediatr Pulmonol 53(S3):S4–S11. https://doi.org/10.1002/ppul.24157
Clapham DE (2003) TRP channels as cellular sensors. Nature 426(6966):517–524. https://doi.org/10.1038/nature02196
Clifford RJ, Kaplan JH (2008) beta-Subunit overexpression alters the stoicheometry of assembled Na-K-ATPase subunits in MDCK cells. Am J Physiol Renal Physiol 295(5):F1314–F1323. https://doi.org/10.1152/ajprenal.90406.2008
Crambert G, Hasler U, Beggah AT, Yu C, Modyanov NN, Horisberger JD, Lelievre L, Geering K (2000) Transport and pharmacological properties of nine different human Na, K-ATPase isozymes. J Biol Chem 275(3):1976–1986
Craven RA, Egerton M, Stirling CJ (1996) A novel Hsp70 of the yeast ER lumen is required for the efficient translocation of a number of protein precursors. EMBO J 15(11):2640–2650
Danilczyk UG, Williams DB (2001) The lectin chaperone calnexin utilizes polypeptide-based interactions to associate with many of its substrates in vivo. J Biol Chem 276(27):25532–25540. https://doi.org/10.1074/jbc.M100270200
De Jong JC, Van Der Vliet WA, Van Den Heuvel LP, Willems PH, Knoers NV, Bindels RJ (2002) Functional expression of mutations in the human NaCl cotransporter: evidence for impaired routing mechanisms in Gitelman’s syndrome. J Am Soc Nephrol 13(6):1442–1448
de Keyzer J, Steel GJ, Hale SJ, Humphries D, Stirling CJ (2009) Nucleotide binding by Lhs1p is essential for its nucleotide exchange activity and for function in vivo. J Biol Chem 284(46):31564–31571. https://doi.org/10.1074/jbc.M109.055160
de la Pena AH, Goodall EA, Gates SN, Lander GC, Martin A (2018) Substrate-engaged 26S proteasome structures reveal mechanisms for ATP-hydrolysis-driven translocation. Science 362(6418):eaav0725. https://doi.org/10.1126/science.aav0725
De I, Sadhukhan S (2018) Emerging roles of DHHC-mediated protein S-palmitoylation in physiological and pathophysiological context. Eur J Cell Biol 97(5):319–338. https://doi.org/10.1016/j.ejcb.2018.03.005
Dong Y, Zhang S, Wu Z, Li X, Wang WL, Zhu Y, Stoilova-McPhie S, Lu Y, Finley D, Mao Y (2019) Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature 565(7737):49–55. https://doi.org/10.1038/s41586-018-0736-4
Donnelly BF, Needham PG, Snyder AC, Roy A, Khadem S, Brodsky JL, Subramanya AR (2013) Hsp70 and Hsp90 multichaperone complexes sequentially regulate thiazide-sensitive cotransporter endoplasmic reticulum-associated degradation and biogenesis. J Biol Chem 288(18):13124–13135. https://doi.org/10.1074/jbc.M113.455394
Drazic A, Myklebust LM, Ree R, Arnesen T (2016) The world of protein acetylation. Biochim Biophys Acta 1864(10):1372–1401. https://doi.org/10.1016/j.bbapap.2016.06.007
Egea PF, Stroud RM (2010) Lateral opening of a translocon upon entry of protein suggests the mechanism of insertion into membranes. Proc Natl Acad Sci U S A 107(40):17182–17187. https://doi.org/10.1073/pnas.1012556107
El Khouri E, Le Pavec G, Toledano MB, Delaunay-Moisan A (2013) RNF185 is a novel E3 ligase of endoplasmic reticulum-associated degradation (ERAD) that targets cystic fibrosis transmembrane conductance regulator (CFTR). J Biol Chem 288(43):31177–31191. https://doi.org/10.1074/jbc.M113.470500
Fallen K, Banerjee S, Sheehan J, Addison D, Lewis LM, Meiler J, Denton JS (2009) The Kir channel immunoglobulin domain is essential for Kir1.1 (ROMK) thermodynamic stability, trafficking and gating. Channels (Austin) 3(1):57–68
Fang L, Li D, Welling PA (2010) Hypertension resistance polymorphisms in ROMK (Kir1.1) alter channel function by different mechanisms. Am J Physiol Renal Physiol 299(6):F1359–F1364. https://doi.org/10.1152/ajprenal.00257.2010
Feige MJ, Behnke J, Mittag T, Hendershot LM (2015) Dimerization-dependent folding underlies assembly control of the clonotypic alphabetaT cell receptor chains. J Biol Chem 290(44):26821–26831. https://doi.org/10.1074/jbc.M115.689471
Finley D, Chen X, Walters KJ (2016) Gates, channels, and switches: elements of the proteasome machine. Trends Biochem Sci 41(1):77–93. https://doi.org/10.1016/j.tibs.2015.10.009
Firsov D, Robert-Nicoud M, Gruender S, Schild L, Rossier BC (1999) Mutational analysis of cysteine-rich domains of the epithelium sodium channel (ENaC). Identification of cysteines essential for channel expression at the cell surface. J Biol Chem 274(5):2743–2749
Francis GS, Tang WH (2004) Vasopressin receptor antagonists: will the “vaptans” fulfill their promise? JAMA 291(16):2017–2018. https://doi.org/10.1001/jama.291.16.2017
Fukata Y, Murakami T, Yokoi N, Fukata M (2016) Local palmitoylation cycles and specialized membrane domain organization. Curr Top Membr 77:97–141. https://doi.org/10.1016/bs.ctm.2015.10.003
Fulchiero R, Seo-Mayer P (2019) Bartter syndrome and Gitelman syndrome. Pediatr Clin N Am 66(1):121–134. https://doi.org/10.1016/j.pcl.2018.08.010
Galligan JJ, Petersen DR (2012) The human protein disulfide isomerase gene family. Hum Genomics 6:6. https://doi.org/10.1186/1479-7364-6-6
Gardner RG, Swarbrick GM, Bays NW, Cronin SR, Wilhovsky S, Seelig L, Kim C, Hampton RY (2000) Endoplasmic reticulum degradation requires lumen to cytosol signaling. Transmembrane control of Hrd1p by Hrd3p. J Cell Biol 151(1):69–82
Garza RM, Sato BK, Hampton RY (2009) In vitro analysis of Hrd1p-mediated retrotranslocation of its multispanning membrane substrate 3-hydroxy-3-methylglutaryl (HMG)-CoA reductase. J Biol Chem 284(22):14710–14722. https://doi.org/10.1074/jbc.M809607200
Gauss R, Sommer T, Jarosch E (2006) The Hrd1p ligase complex forms a linchpin between ER-lumenal substrate selection and Cdc48p recruitment. EMBO J 25(9):1827–1835. https://doi.org/10.1038/sj.emboj.7601088
Geering K (2001) The functional role of beta subunits in oligomeric P-type ATPases. J Bioenerg Biomembr 33(5):425–438
Geering K (2008) Functional roles of Na,K-ATPase subunits. Curr Opin Nephrol Hypertens 17(5):526–532. https://doi.org/10.1097/MNH.0b013e3283036cbf
Genest O, Wickner S, Doyle SM (2019) Hsp90 and Hsp70 chaperones: collaborators in protein remodeling. J Biol Chem 294(6):2109–2120. https://doi.org/10.1074/jbc.REV118.002806
Gentzsch M, Dang H, Dang Y, Garcia-Caballero A, Suchindran H, Boucher RC, Stutts MJ (2010) The cystic fibrosis transmembrane conductance regulator impedes proteolytic stimulation of the epithelial Na+ channel. J Biol Chem 285(42):32227–32232. https://doi.org/10.1074/jbc.M110.155259
Ghaemmaghami S, Huh WK, Bower K, Howson RW, Belle A, Dephoure N, O’Shea EK, Weissman JS (2003) Global analysis of protein expression in yeast. Nature 425(6959):737–741
Gilmore R, Blobel G, Walter P (1982) Protein translocation across the endoplasmic reticulum. I. Detection in the microsomal membrane of a receptor for the signal recognition particle. J Cell Biol 95(2 Pt 1):463–469
Goldfarb SB, Kashlan OB, Watkins JN, Suaud L, Yan W, Kleyman TR, Rubenstein RC (2006) Differential effects of Hsc70 and Hsp70 on the intracellular trafficking and functional expression of epithelial sodium channels. Proc Natl Acad Sci U S A 103(15):5817–5822
Gorlich D, Prehn S, Hartmann E, Kalies KU, Rapoport TA (1992) A mammalian homolog of SEC61p and SECYp is associated with ribosomes and nascent polypeptides during translocation. Cell 71(3):489–503
Grumbach Y, Bikard Y, Suaud L, Chanoux RA, Rubenstein RC (2014) ERp29 regulates epithelial sodium channel functional expression by promoting channel cleavage. Am J Physiol Cell Physiol 307(8):C701–C709. https://doi.org/10.1152/ajpcell.00134.2014
Grunder S, Firsov D, Chang SS, Jaeger NF, Gautschi I, Schild L, Lifton RP, Rossier BC (1997) A mutation causing pseudohypoaldosteronism type 1 identifies a conserved glycine that is involved in the gating of the epithelial sodium channel. EMBO J 16(5):899–907. https://doi.org/10.1093/emboj/16.5.899
Habeck G, Ebner FA, Shimada-Kreft H, Kreft SG (2015) The yeast ERAD-C ubiquitin ligase Doa10 recognizes an intramembrane degron. J Cell Biol 209(2):261–273. https://doi.org/10.1083/jcb.201408088
Halic M, Becker T, Pool MR, Spahn CM, Grassucci RA, Frank J, Beckmann R (2004) Structure of the signal recognition particle interacting with the elongation-arrested ribosome. Nature 427(6977):808–814. https://doi.org/10.1038/nature02342
Halic M, Blau M, Becker T, Mielke T, Pool MR, Wild K, Sinning I, Beckmann R (2006) Following the signal sequence from ribosomal tunnel exit to signal recognition particle. Nature 444(7118):507–511. https://doi.org/10.1038/nature05326
Hampton RY, Gardner RG, Rine J (1996) Role of 26S proteasome and HRD genes in the degradation of 3-hydroxy-3-methylglutaryl-CoA reductase, an integral endoplasmic reticulum membrane protein. Mol Biol Cell 7(12):2029–2044
Hanzelmann P, Schindelin H (2017) The interplay of cofactor interactions and post-translational modifications in the regulation of the AAA+ ATPase p97. Front Mol Biosci 4:21. https://doi.org/10.3389/fmolb.2017.00021
Harel M, Aharoni A, Gaidukov L, Brumshtein B, Khersonsky O, Meged R, Dvir H, Ravelli RB, McCarthy A, Toker L, Silman I, Sussman JL, Tawfik DS (2004) Structure and evolution of the serum paraoxonase family of detoxifying and anti-atherosclerotic enzymes. Nat Struct Mol Biol 11(5):412–419. https://doi.org/10.1038/nsmb767
Hasler U, Crambert G, Horisberger JD, Geering K (2001) Structural and functional features of the transmembrane domain of the Na,K-ATPase beta subunit revealed by tryptophan scanning. J Biol Chem 276(19):16356–16364. https://doi.org/10.1074/jbc.M008778200
Hebert DN, Molinari M (2012) Flagging and docking: dual roles for N-glycans in protein quality control and cellular proteostasis. Trends Biochem Sci 37(10):404–410. https://doi.org/10.1016/j.tibs.2012.07.005
Heidrich E, Carattino MD, Hughey RP, Pilewski JM, Kleyman TR, Myerburg MM (2015) Intracellular Na+ regulates epithelial Na+ channel maturation. J Biol Chem 290(18):11569–11577. https://doi.org/10.1074/jbc.M115.640763
Hessa T, Meindl-Beinker NM, Bernsel A, Kim H, Sato Y, Lerch-Bader M, Nilsson I, White SH, von Heijne G (2007) Molecular code for transmembrane-helix recognition by the Sec61 translocon. Nature 450(7172):1026–1030. https://doi.org/10.1038/nature06387
Hiller MM, Finger A, Schweiger M, Wolf DH (1996) ER degradation of a misfolded luminal protein by the cytosolic ubiquitin-proteasome pathway. Science 273(5282):1725–1728
Hitt R, Wolf DH (2004) Der1p, a protein required for degradation of malfolded soluble proteins of the endoplasmic reticulum: topology and Der1-like proteins. FEMS Yeast Res 4(7):721–729
Ho K, Nichols CG, Lederer WJ, Lytton J, Vassilev PM, Kanazirska MV, Hebert SC (1993) Cloning and expression of an inwardly rectifying ATP-regulated potassium channel. Nature 362(6415):31–38. https://doi.org/10.1038/362031a0
Hobbs CA, Da Tan C, Tarran R (2013) Does epithelial sodium channel hyperactivity contribute to cystic fibrosis lung disease? J Physiol 591(18):4377–4387. https://doi.org/10.1113/jphysiol.2012.240861
Hoekstra JA, van Lieburg AF, Monnens LA, Hulstijn-Dirkmaat GM, Knoers VV (1996) Cognitive and psychosocial functioning of patients with congenital nephrogenic diabetes insipidus. Am J Med Genet 61(1):81–88. https://doi.org/10.1002/(SICI)1096-8628(19960102)61:1<81::AID-AJMG17>3.0.CO;2-S
Horn SC, Hanna J, Hirsch C, Volkwein C, Schutz A, Heinemann U, Sommer T, Jarosch E (2009) Usa1 functions as a scaffold of the HRD-ubiquitin ligase. Mol Cell 36(5):782–793. https://doi.org/10.1016/j.molcel.2009.10.015
Hughey RP, Mueller GM, Bruns JB, Kinlough CL, Poland PA, Harkleroad KL, Carattino MD, Kleyman TR (2003) Maturation of the epithelial Na+ channel involves proteolytic processing of the alpha- and gamma-subunits. J Biol Chem 278(39):37073–37082
Inoue T, Tsai B (2015) A nucleotide exchange factor promotes endoplasmic reticulum-to-cytosol membrane penetration of the nonenveloped virus simian virus 40. J Virol 89(8):4069–4079. https://doi.org/10.1128/JVI.03552-14
Inoue T, Tsai B (2016) The Grp170 nucleotide exchange factor executes a key role during ERAD of cellular misfolded clients. Mol Biol Cell 27(10):1650–1662. https://doi.org/10.1091/mbc.E16-01-0033
Iolascon A, Aglio V, Tamma G, D’Apolito M, Addabbo F, Procino G, Simonetti MC, Montini G, Gesualdo L, Debler EW, Svelto M, Valenti G (2007) Characterization of two novel missense mutations in the AQP2 gene causing nephrogenic diabetes insipidus. Nephron Physiol 105(3):p33–p41. https://doi.org/10.1159/000098136
Ismail N, Ng DT (2006) Have you HRD? Understanding ERAD is DOAble! Cell 126(2):237–239
Ismailov II, Awayda MS, Jovov B, Berdiev BK, Fuller CM, Dedman JR, Kaetzel M, Benos DJ (1996) Regulation of epithelial sodium channels by the cystic fibrosis transmembrane conductance regulator. J Biol Chem 271(9):4725–4732
Jarosch E, Taxis C, Volkwein C, Bordallo J, Finley D, Wolf DH, Sommer T (2002) Protein dislocation from the ER requires polyubiquitination and the AAA-ATPase Cdc48. Nat Cell Biol 4(2):134–139
Jaunin P, Horisberger JD, Richter K, Good PJ, Rossier BC, Geering K (1992) Processing, intracellular transport, and functional expression of endogenous and exogenous alpha-beta 3 Na,K-ATPase complexes in Xenopus oocytes. J Biol Chem 267(1):577–585
Ji W, Foo JN, O’Roak BJ, Zhao H, Larson MG, Simon DB, Newton-Cheh C, State MW, Levy D, Lifton RP (2008) Rare independent mutations in renal salt handling genes contribute to blood pressure variation. Nat Genet 40(5):592–599. https://doi.org/10.1038/ng.118
Jiang LY, Jiang W, Tian N, Xiong YN, Liu J, Wei J, Wu KY, Luo J, Shi XJ, Song BL (2018) Ring finger protein 145 (RNF145) is a ubiquitin ligase for sterol-induced degradation of HMG-CoA reductase. J Biol Chem 293(11):4047–4055. https://doi.org/10.1074/jbc.RA117.001260
Jorgensen PL (1986) Structure, function and regulation of Na,K-ATPase in the kidney. Kidney Int 29(1):10–20
Kampinga HH, Hageman J, Vos MJ, Kubota H, Tanguay RM, Bruford EA, Cheetham ME, Chen B, Hightower LE (2009) Guidelines for the nomenclature of the human heat shock proteins. Cell Stress Chaperones 14(1):105–111. https://doi.org/10.1007/s12192-008-0068-7
Kamsteeg EJ, Bichet DG, Konings IB, Nivet H, Lonergan M, Arthus MF, van Os CH, Deen PM (2003) Reversed polarized delivery of an aquaporin-2 mutant causes dominant nephrogenic diabetes insipidus. J Cell Biol 163(5):1099–1109
Kashlan OB, Mueller GM, Qamar MZ, Poland PA, Ahner A, Rubenstein RC, Hughey RP, Brodsky JL, Kleyman TR (2007) Small heat shock protein alphaA-crystallin regulates epithelial sodium channel expression. J Biol Chem 282(38):28149–28156
Kashlan OB, Kinlough CL, Myerburg MM, Shi S, Chen J, Blobner BM, Buck TM, Brodsky JL, Hughey RP, Kleyman TR (2018) N-linked glycans are required on epithelial Na+ channel subunits for maturation and surface expression. Am J Physiol Renal Physiol 314(3):F483–F492. https://doi.org/10.1152/ajprenal.00195.2017
Kieber-Emmons T, Lin C, Foster MH, Kleyman TR (1999) Antiidiotypic antibody recognizes an amiloride binding domain within the alpha subunit of the epithelial Na+ channel. J Biol Chem 274(14):9648–9655
Kityk R, Kopp J, Sinning I, Mayer MP (2012) Structure and dynamics of the ATP-bound open conformation of Hsp70 chaperones. Mol Cell 48(6):863–874. https://doi.org/10.1016/j.molcel.2012.09.023
Kityk R, Kopp J, Mayer MP (2018) Molecular mechanism of J-domain-TRIGGERED ATP hydrolysis by Hsp70 chaperones. Mol Cell 69(2):227–237.e224. https://doi.org/10.1016/j.molcel.2017.12.003
Kizhatil K, Davis JQ, Davis L, Hoffman J, Hogan BL, Bennett V (2007) Ankyrin-G is a molecular partner of E-cadherin in epithelial cells and early embryos. J Biol Chem 282(36):26552–26561. https://doi.org/10.1074/jbc.M703158200
Knop M, Finger A, Braun T, Hellmuth K, Wolf DH (1996) Der1, a novel protein specifically required for endoplasmic reticulum degradation in yeast. EMBO J 15(4):753–763
Kong J, Peng M, Ostrovsky J, Kwon YJ, Oretsky O, McCormick EM, He M, Argon Y, Falk MJ (2018) Mitochondrial function requires NGLY1. Mitochondrion 38:6–16. https://doi.org/10.1016/j.mito.2017.07.008
Koulen P, Cai Y, Geng L, Maeda Y, Nishimura S, Witzgall R, Ehrlich BE, Somlo S (2002) Polycystin-2 is an intracellular calcium release channel. Nat Cell Biol 4(3):191–197. https://doi.org/10.1038/ncb754
Kreutzberger AJB, Ji M, Aaron J, Mihaljevic L, Urban S (2019) Rhomboid distorts lipids to break the viscosity-imposed speed limit of membrane diffusion. Science 363(6426):eaao0076. https://doi.org/10.1126/science.aao0076
Kunchaparty S, Palcso M, Berkman J, Velazquez H, Desir GV, Bernstein P, Reilly RF, Ellison DH (1999) Defective processing and expression of thiazide-sensitive Na-Cl cotransporter as a cause of Gitelman’s syndrome. Am J Phys 277(4):F643–F649. https://doi.org/10.1152/ajprenal.1999.277.4.F643
Kuwahara M, Iwai K, Ooeda T, Igarashi T, Ogawa E, Katsushima Y, Shinbo I, Uchida S, Terada Y, Arthus MF, Lonergan M, Fujiwara TM, Bichet DG, Marumo F, Sasaki S (2001) Three families with autosomal dominant nephrogenic diabetes insipidus caused by aquaporin-2 mutations in the C-terminus. Am J Hum Genet 69(4):738–748. https://doi.org/10.1086/323643
Lang S, Pfeffer S, Lee PH, Cavalie A, Helms V, Forster F, Zimmermann R (2017) An update on Sec61 channel functions, mechanisms, and related diseases. Front Physiol 8:887. https://doi.org/10.3389/fphys.2017.00887
Leduc-Nadeau A, Lussier Y, Arthus MF, Lonergan M, Martinez-Aguayo A, Riveira-Munoz E, Devuyst O, Bissonnette P, Bichet DG (2010) New autosomal recessive mutations in aquaporin-2 causing nephrogenic diabetes insipidus through deficient targeting display normal expression in Xenopus oocytes. J Physiol 588(Pt 12):2205–2218. https://doi.org/10.1113/jphysiol.2010.187674
Leitman J, Shenkman M, Gofman Y, Shtern NO, Ben-Tal N, Hendershot LM, Lederkremer GZ (2014) Herp coordinates compartmentalization and recruitment of HRD1 and misfolded proteins for ERAD. Mol Biol Cell 25(7):1050–1060. https://doi.org/10.1091/mbc.E13-06-0350
Lerner M, Corcoran M, Cepeda D, Nielsen ML, Zubarev R, Ponten F, Uhlen M, Hober S, Grander D, Sangfelt O (2007) The RBCC gene RFP2 (Leu5) encodes a novel transmembrane E3 ubiquitin ligase involved in ERAD. Mol Biol Cell 18(5):1670–1682. https://doi.org/10.1091/mbc.e06-03-0248
Li G, Zhao G, Zhou X, Schindelin H, Lennarz WJ (2006) The AAA ATPase p97 links peptide N-glycanase to the endoplasmic reticulum-associated E3 ligase autocrine motility factor receptor. Proc Natl Acad Sci U S A 103(22):8348–8353. https://doi.org/10.1073/pnas.0602747103
Li W, Tu D, Brunger AT, Ye Y (2007) A ubiquitin ligase transfers preformed polyubiquitin chains from a conjugating enzyme to a substrate. Nature 446(7133):333–337. https://doi.org/10.1038/nature05542
Lian WN, Wu TW, Dao RL, Chen YJ, Lin CH (2006) Deglycosylation of Na+/K+-ATPase causes the basolateral protein to undergo apical targeting in polarized hepatic cells. J Cell Sci 119(Pt 1):11–22. https://doi.org/10.1242/jcs.02706
Liang G, Li Q, Tang Y, Kokame K, Kikuchi T, Wu G, Chen XZ (2008) Polycystin-2 is regulated by endoplasmic reticulum-associated degradation. Hum Mol Genet 17(8):1109–1119. https://doi.org/10.1093/hmg/ddm383
Lin SH, Bichet DG, Sasaki S, Kuwahara M, Arthus MF, Lonergan M, Lin YF (2002) Two novel aquaporin-2 mutations responsible for congenital nephrogenic diabetes insipidus in Chinese families. J Clin Endocrinol Metab 87(6):2694–2700. https://doi.org/10.1210/jcem.87.6.8617
Linder ME, Deschenes RJ (2007) Palmitoylation: policing protein stability and traffic. Nat Rev Mol Cell Biol 8(1):74–84. https://doi.org/10.1038/nrm2084
Lloyd DJ, Hall FW, Tarantino LM, Gekakis N (2005) Diabetes insipidus in mice with a mutation in aquaporin-2. PLoS Genet 1(2):e20. https://doi.org/10.1371/journal.pgen.0010020
Los EL, Deen PM, Robben JH (2010) Potential of nonpeptide (ant)agonists to rescue vasopressin V2 receptor mutants for the treatment of X-linked nephrogenic diabetes insipidus. J Neuroendocrinol 22(5):393–399. https://doi.org/10.1111/j.1365-2826.2010.01983.x
Lu Y, Turnbull IR, Bragin A, Carveth K, Verkman AS, Skach WR (2000) Reorientation of aquaporin-1 topology during maturation in the endoplasmic reticulum. Mol Biol Cell 11(9):2973–2985
Lu JP, Wang Y, Sliter DA, Pearce MM, Wojcikiewicz RJ (2011) RNF170 protein, an endoplasmic reticulum membrane ubiquitin ligase, mediates inositol 1,4,5-trisphosphate receptor ubiquitination and degradation. J Biol Chem 286(27):24426–24433. https://doi.org/10.1074/jbc.M111.251983
Lukacs GL, Verkman AS (2012) CFTR: folding, misfolding and correcting the DeltaF508 conformational defect. Trends Mol Med 18(2):81–91. https://doi.org/10.1016/j.molmed.2011.144003
Luo Y, Vassilev PM, Li X, Kawanabe Y, Zhou J (2003) Native polycystin 2 functions as a plasma membrane Ca2+-permeable cation channel in renal epithelia. Mol Cell Biol 23(7):2600–2607
Mackie TD, Kim BY, Subramanya AR, Bain DJ, O’Donnell AF, Welling PA, Brodsky JL (2018) The endosomal trafficking factors CORVET and ESCRT suppress plasma membrane residence of the renal outer medullary potassium channel (ROMK). J Biol Chem 293(9):3201–3217. https://doi.org/10.1074/jbc.M117.819086
Malik B, Schlanger L, Al-Khalili O, Bao HF, Yue G, Price SR, Mitch WE, Eaton DC (2001) Enac degradation in A6 cells by the ubiquitin-proteosome proteolytic pathway. J Biol Chem 276(16):12903–12910
Mall M, Grubb BR, Harkema JR, O’Neal WK, Boucher RC (2004) Increased airway epithelial Na+ absorption produces cystic fibrosis-like lung disease in mice. Nat Med 10(5):487–493
Marr N, Bichet DG, Hoefs S, Savelkoul PJ, Konings IB, De Mattia F, Graat MP, Arthus MF, Lonergan M, Fujiwara TM, Knoers NV, Landau D, Balfe WJ, Oksche A, Rosenthal W, Muller D, Van Os CH, Deen PM (2002) Cell-biologic and functional analyses of five new Aquaporin-2 missense mutations that cause recessive nephrogenic diabetes insipidus. J Am Soc Nephrol 13(9):2267–2277
Masilamani S, Kim GH, Mitchell C, Wade JB, Knepper MA (1999) Aldosterone-mediated regulation of ENaC alpha, beta, and gamma subunit proteins in rat kidney. J Clin Invest 104(7):R19–R23
Mayer MP (2018) Intra-molecular pathways of allosteric control in Hsp70s. Philos Trans R Soc Lond Ser B Biol Sci 373(1749):20170183. https://doi.org/10.1098/rstb.2017.0183
Mayer MP, Gierasch LM (2019) Recent advances in the structural and mechanistic aspects of Hsp70 molecular chaperones. J Biol Chem 294(6):2085–2097. https://doi.org/10.1074/jbc.REV118.002810
Mayer MP, Kityk R (2015) Insights into the molecular mechanism of allostery in Hsp70s. Front Mol Biosci 2:58. https://doi.org/10.3389/fmolb.2015.00058
McDonough H, Patterson C (2003) CHIP: a link between the chaperone and proteasome systems. Cell Stress Chaperones 8(4):303–308
McGrath J, Somlo S, Makova S, Tian X, Brueckner M (2003) Two populations of node monocilia initiate left-right asymmetry in the mouse. Cell 114(1):61–73
Mehnert M, Sommer T, Jarosch E (2014) Der1 promotes movement of misfolded proteins through the endoplasmic reticulum membrane. Nat Cell Biol 16(1):77–86. https://doi.org/10.1038/ncb2882
Mehnert M, Sommermeyer F, Berger M, Kumar Lakshmipathy S, Gauss R, Aebi M, Jarosch E, Sommer T (2015) The interplay of Hrd3 and the molecular chaperone system ensures efficient degradation of malfolded secretory proteins. Mol Biol Cell 26(2):185–194. https://doi.org/10.1091/mbc.E14-07-1202
Mehrtash AB, Hochstrasser M (2018) Ubiquitin-dependent protein degradation at the endoplasmic reticulum and nuclear envelope. Semin Cell Dev Biol 93:111–124. https://doi.org/10.1016/j.semcdb.2018.09.013
Metzger MB, Liang YH, Das R, Mariano J, Li S, Li J, Kostova Z, Byrd RA, Ji X, Weissman AM (2013) A structurally unique E2-binding domain activates ubiquitination by the ERAD E2, Ubc7p, through multiple mechanisms. Mol Cell 50(4):516–527. https://doi.org/10.1016/j.molcel.2013.04.004
Meusser B, Hirsch C, Jarosch E, Sommer T (2005) ERAD: the long road to destruction. Nat Cell Biol 7(8):766–772
Meyer DI, Dobberstein B (1980) A membrane component essential for vectorial translocation of nascent proteins across the endoplasmic reticulum: requirements for its extraction and reassociation with the membrane. J Cell Biol 87(2 Pt 1):498–502
Meyer HH, Shorter JG, Seemann J, Pappin D, Warren G (2000) A complex of mammalian ufd1 and npl4 links the AAA-ATPase, p97, to ubiquitin and nuclear transport pathways. EMBO J 19(10):2181–2192
Meyer HH, Wang Y, Warren G (2002) Direct binding of ubiquitin conjugates by the mammalian p97 adaptor complexes, p47 and Ufd1-Npl4. EMBO J 21(21):5645–5652
Milano S, Carmosino M, Gerbino A, Svelto M, Procino G (2017) Hereditary nephrogenic diabetes insipidus: pathophysiology and possible treatment. An update. Int J Mol Sci 18(11):2385. https://doi.org/10.3390/ijms18112385
Misselwitz B, Staeck O, Rapoport TA (1998) J proteins catalytically activate Hsp70 molecules to trap a wide range of peptide sequences. Mol Cell 2(5):593–603
Mochizuki T, Wu G, Hayashi T, Xenophontos SL, Veldhuisen B, Saris JJ, Reynolds DM, Cai Y, Gabow PA, Pierides A, Kimberling WJ, Breuning MH, Deltas CC, Peters DJ, Somlo S (1996) PKD2, a gene for polycystic kidney disease that encodes an integral membrane protein. Science 272(5266):1339–1342
Morello JP, Salahpour A, Laperriere A, Bernier V, Arthus MF, Lonergan M, Petaja-Repo U, Angers S, Morin D, Bichet DG, Bouvier M (2000) Pharmacological chaperones rescue cell-surface expression and function of misfolded V2 vasopressin receptor mutants. J Clin Invest 105(7):887–895. https://doi.org/10.1172/JCI8688
Morth JP, Pedersen BP, Toustrup-Jensen MS, Sorensen TL, Petersen J, Andersen JP, Vilsen B, Nissen P (2007) Crystal structure of the sodium-potassium pump. Nature 450(7172):1043–1049. https://doi.org/10.1038/nature06419
Mouillac B, Chini B, Balestre MN, Elands J, Trumpp-Kallmeyer S, Hoflack J, Hibert M, Jard S, Barberis C (1995) The binding site of neuropeptide vasopressin V1a receptor. Evidence for a major localization within transmembrane regions. J Biol Chem 270(43):25771–25777
Mueller GM, Kashlan OB, Bruns JB, Maarouf AB, Aridor M, Kleyman TR, Hughey RP (2007) Epithelial sodium channel exit from the endoplasmic reticulum is regulated by a signal within the carboxyl cytoplasmic domain of the alpha subunit. J Biol Chem 282(46):33475–33483. https://doi.org/10.1074/jbc.M707339200
Mueller GM, Maarouf AB, Kinlough CL, Sheng N, Kashlan OB, Okumura S, Luthy S, Kleyman TR, Hughey RP (2010) Cys palmitoylation of the beta subunit modulates gating of the epithelial sodium channel. J Biol Chem 285(40):30453–30462. https://doi.org/10.1074/jbc.M110.151845
Mukherjee A, Mueller GM, Kinlough CL, Sheng N, Wang Z, Mustafa SA, Kashlan OB, Kleyman TR, Hughey RP (2014) Cysteine palmitoylation of the gamma subunit has a dominant role in modulating activity of the epithelial sodium channel. J Biol Chem 289(20):14351–14359. https://doi.org/10.1074/jbc.M113.526020
Mukherjee A, Wang Z, Kinlough CL, Poland PA, Marciszyn AL, Montalbetti N, Carattino MD, Butterworth MB, Kleyman TR, Hughey RP (2017) Specific palmitoyltransferases associate with and activate the epithelial sodium channel. J Biol Chem 292(10):4152–4163. https://doi.org/10.1074/jbc.M117.776146
Mulders SM, Bichet DG, Rijss JP, Kamsteeg EJ, Arthus MF, Lonergan M, Fujiwara M, Morgan K, Leijendekker R, van der Sluijs P, van Os CH, Deen PM (1998) An aquaporin-2 water channel mutant which causes autosomal dominant nephrogenic diabetes insipidus is retained in the Golgi complex. J Clin Invest 102(1):57–66
Nadolski MJ, Linder ME (2007) Protein lipidation. FEBS J 274(20):5202–5210. https://doi.org/10.1111/j.1742-4658.2007.06056.x
Nakatsukasa K, Brodsky JL, Kamura T (2013) A stalled retrotranslocation complex reveals physical linkage between substrate recognition and proteasomal degradation during ER-associated degradation. Mol Biol Cell 24(11):1765–1775, S1761–1768. https://doi.org/10.1091/mbc.E12-12-0907
Nauli SM, Alenghat FJ, Luo Y, Williams E, Vassilev P, Li X, Elia AE, Lu W, Brown EM, Quinn SJ, Ingber DE, Zhou J (2003) Polycystins 1 and 2 mediate mechanosensation in the primary cilium of kidney cells. Nat Genet 33(2):129–137. https://doi.org/10.1038/ng1076
Neal S, Jaeger PA, Duttke SH, Benner C, Glass CK, Ideker T, Hampton RY (2018) The Dfm1 Derlin is required for ERAD retrotranslocation of integral membrane proteins. Mol Cell 69(2):306–320.e304. https://doi.org/10.1016/j.molcel.2017.12.012
Needham PG, Mikoluk K, Dhakarwal P, Khadem S, Snyder AC, Subramanya AR, Brodsky JL (2011) The thiazide-sensitive NaCl cotransporter is targeted for chaperone-dependent endoplasmic reticulum-associated degradation. J Biol Chem 286(51):43611–43621. https://doi.org/10.1074/jbc.M111.288928
Nelson WJ, Veshnock PJ (1987) Ankyrin binding to (Na+ + K+)ATPase and implications for the organization of membrane domains in polarized cells. Nature 328(6130):533–536. https://doi.org/10.1038/328533a0
Nelson WJ, Shore EM, Wang AZ, Hammerton RW (1990) Identification of a membrane-cytoskeletal complex containing the cell adhesion molecule uvomorulin (E-cadherin), ankyrin, and fodrin in Madin-Darby canine kidney epithelial cells. J Cell Biol 110(2):349–357
Nguyen AD, Lee SH, DeBose-Boyd RA (2009) Insig-mediated, sterol-accelerated degradation of the membrane domain of hamster 3-hydroxy-3-methylglutaryl-coenzyme A reductase in insect cells. J Biol Chem 284(39):26778–26788. https://doi.org/10.1074/jbc.M109.032342
Ninagawa S, Okada T, Sumitomo Y, Kamiya Y, Kato K, Horimoto S, Ishikawa T, Takeda S, Sakuma T, Yamamoto T, Mori K (2014) EDEM2 initiates mammalian glycoprotein ERAD by catalyzing the first mannose trimming step. J Cell Biol 206(3):347–356. https://doi.org/10.1083/jcb.201404075
Noguchi S, Higashi K, Kawamura M (1990) A possible role of the beta-subunit of (Na,K)-ATPase in facilitating correct assembly of the alpha-subunit into the membrane. J Biol Chem 265(26):15991–15995
Noreng S, Bharadwaj A, Posert R, Yoshioka C, Baconguis I (2018) Structure of the human epithelial sodium channel by cryo-electron microscopy. elife 7:e39340. https://doi.org/10.7554/eLife.39340
O’Donnell BM, Mackie TD, Subramanya AR, Brodsky JL (2017) Endoplasmic reticulum-associated degradation of the renal potassium channel, ROMK, leads to type II Bartter syndrome. J Biol Chem 292(31):12813–12827. https://doi.org/10.1074/jbc.M117.786376
Oberfeld B, Ruffieux-Daidie D, Vitagliano JJ, Pos KM, Verrey F, Staub O (2011) Ubiquitin-specific protease 2-45 (Usp2-45) binds to epithelial Na+ channel (ENaC)-ubiquitylating enzyme Nedd4-2. Am J Physiol Renal Physiol 301(1):F189–F196. https://doi.org/10.1152/ajprenal.00487.2010
Oda Y, Hosokawa N, Wada I, Nagata K (2003) EDEM as an acceptor of terminally misfolded glycoproteins released from calnexin.[comment]. Science 299(5611):1394–1397
Ohno Y, Kihara A, Sano T, Igarashi Y (2006) Intracellular localization and tissue-specific distribution of human and yeast DHHC cysteine-rich domain-containing proteins. Biochim Biophys Acta 1761(4):474–483. https://doi.org/10.1016/j.bbalip.2006.03.010
Padilla-Benavides T, Roldan ML, Larre I, Flores-Benitez D, Villegas-Sepulveda N, Contreras RG, Cereijido M, Shoshani L (2010) The polarized distribution of Na+,K+-ATPase: role of the interaction between {beta} subunits. Mol Biol Cell 21(13):2217–2225. https://doi.org/10.1091/mbc.E10-01-0081
Palmer BF (2015) Regulation of potassium homeostasis. Clin J Am Soc Nephrol 10(6):1050–1060. https://doi.org/10.2215/CJN.08580813
Peters M, Ermert S, Jeck N, Derst C, Pechmann U, Weber S, Schlingmann KP, Seyberth HW, Waldegger S, Konrad M (2003) Classification and rescue of ROMK mutations underlying hyperprostaglandin E syndrome/antenatal Bartter syndrome. Kidney Int 64(3):923–932. https://doi.org/10.1046/j.1523-1755.2003.00153.x
Pilon M, Schekman R, Romisch K (1997) Sec61p mediates export of a misfolded secretory protein from the endoplasmic reticulum to the cytosol for degradation. EMBO J 16(15):4540–4548
Pilon M, Romisch K, Quach D, Schekman R (1998) Sec61p serves multiple roles in secretory precursor binding and translocation into the endoplasmic reticulum membrane. Mol Biol Cell 9(12):3455–3473
Plemper RK, Bohmler S, Bordallo J, Sommer T, Wolf DH (1997) Mutant analysis links the translocon and BiP to retrograde protein transport for ER degradation. Nature 388(6645):891–895
Pobre KFR, Poet GJ, Hendershot LM (2019) The endoplasmic reticulum (ER) chaperone BiP is a master regulator of ER functions: getting by with a little help from ERdj friends. J Biol Chem 294(6):2098–2108. https://doi.org/10.1074/jbc.REV118.002804
Preston GM, Brodsky JL (2017) The evolving role of ubiquitin modification in endoplasmic reticulum-associated degradation. Biochem J 474(4):445–469. https://doi.org/10.1042/BCJ20160582
Procino G, Carmosino M, Marin O, Brunati AM, Contri A, Pinna LA, Mannucci R, Nielsen S, Kwon TH, Svelto M, Valenti G (2003) Ser-256 phosphorylation dynamics of Aquaporin 2 during maturation from the ER to the vesicular compartment in renal cells. FASEB J 17(13):1886–1888. https://doi.org/10.1096/fj.02-0870fje
Raasi S, Wolf DH (2007) Ubiquitin receptors and ERAD: a network of pathways to the proteasome. Semin Cell Dev Biol 18(6):780–791. https://doi.org/10.1016/j.semcdb.2007.09.008
Rabinovich E, Kerem A, Frohlich KU, Diamant N, Bar-Nun S (2002) AAA-ATPase p97/Cdc48p, a cytosolic chaperone required for endoplasmic reticulum-associated protein degradation. Mol Cell Biol 22(2):626–634
Rapoport TA, Li L, Park E (2017) Structural and mechanistic insights into protein translocation. Annu Rev Cell Dev Biol 33:369–390. https://doi.org/10.1146/annurev-cellbio-100616-060439
Richly H, Rape M, Braun S, Rumpf S, Hoege C, Jentsch S (2005) A series of ubiquitin binding factors connects CDC48/p97 to substrate multiubiquitylation and proteasomal targeting. Cell 120(1):73–84
Ripstein ZA, Huang R, Augustyniak R, Kay LE, Rubinstein JL (2017) Structure of a AAA+ unfoldase in the process of unfolding substrate. elife 6:e25754. https://doi.org/10.7554/eLife.25754
Robben JH, Knoers NV, Deen PM (2005) Characterization of vasopressin V2 receptor mutants in nephrogenic diabetes insipidus in a polarized cell model. Am J Physiol Renal Physiol 289(2):F265–F272. https://doi.org/10.1152/ajprenal.00404.2004
Robben JH, Sze M, Knoers NV, Deen PM (2007) Functional rescue of vasopressin V2 receptor mutants in MDCK cells by pharmacochaperones: relevance to therapy of nephrogenic diabetes insipidus. Am J Physiol Renal Physiol 292(1):F253–F260. https://doi.org/10.1152/ajprenal.00247.2006
Rossier BC (2014) Epithelial sodium channel (ENaC) and the control of blood pressure. Curr Opin Pharmacol 15:33–46. https://doi.org/10.1016/j.coph.2013.11.010
Rubenstein RC, Zeitlin PL (2000) Sodium 4-phenylbutyrate downregulates Hsc70: implications for intracellular trafficking of DeltaF508-CFTR. Am J Physiol Cell Physiol 278(2):C259–C267
Rutkevich LA, Williams DB (2011) Participation of lectin chaperones and thiol oxidoreductases in protein folding within the endoplasmic reticulum. Curr Opin Cell Biol 23(2):157–166. https://doi.org/10.1016/j.ceb.2010.10.011
Sabath E, Meade P, Berkman J, de los Heros P, Moreno E, Bobadilla NA, Vazquez N, Ellison DH, Gamba G (2004) Pathophysiology of functional mutations of the thiazide-sensitive Na-Cl cotransporter in Gitelman disease. Am J Physiol Renal Physiol 287(2):F195–F203. https://doi.org/10.1152/ajprenal.00044.2004
Sadoul K, Boyault C, Pabion M, Khochbin S (2008) Regulation of protein turnover by acetyltransferases and deacetylases. Biochimie 90(2):306–312. https://doi.org/10.1016/j.biochi.2007.06.009
Sandhu N, Duus K, Jorgensen CS, Hansen PR, Bruun SW, Pedersen LO, Hojrup P, Houen G (2007) Peptide binding specificity of the chaperone calreticulin. Biochim Biophys Acta 1774(6):701–713. https://doi.org/10.1016/j.bbapap.2007.03.019
Sato BK, Hampton RY (2006) Yeast Derlin Dfm1 interacts with Cdc48 and functions in ER homeostasis. Yeast 23(14-15):1053–1064. https://doi.org/10.1002/yea.1407
Sato BK, Schulz D, Do PH, Hampton RY (2009) Misfolded membrane proteins are specifically recognized by the transmembrane domain of the Hrd1p ubiquitin ligase. Mol Cell 34(2):212–222
Schoebel S, Mi W, Stein A, Ovchinnikov S, Pavlovicz R, DiMaio F, Baker D, Chambers MG, Su H, Li D, Rapoport TA, Liao M (2017) Cryo-EM structure of the protein-conducting ERAD channel Hrd1 in complex with Hrd3. Nature 548(7667):352–355. https://doi.org/10.1038/nature23314
Schwieger I, Lautz K, Krause E, Rosenthal W, Wiesner B, Hermosilla R (2008) Derlin-1 and p97/valosin-containing protein mediate the endoplasmic reticulum-associated degradation of human V2 vasopressin receptors. Mol Pharmacol 73(3):697–708. https://doi.org/10.1124/mol.107.040931
Seaayfan E, Defontaine N, Demaretz S, Zaarour N, Laghmani K (2016) OS9 protein interacts with Na-K-2Cl co-transporter (NKCC2) and targets its immature form for the endoplasmic reticulum-associated degradation pathway. J Biol Chem 291(9):4487–4502. https://doi.org/10.1074/jbc.M115.702514
Sheng S, Maarouf AB, Bruns JB, Hughey RP, Kleyman TR (2007) Functional role of extracellular loop cysteine residues of the epithelial Na+ channel in Na+ self-inhibition. J Biol Chem 282(28):20180–20190. https://doi.org/10.1074/jbc.M611761200
Shi S, Buck TM, Kinlough CL, Marciszyn AL, Hughey RP, Chalfie M, Brodsky JL, Kleyman TR (2017) Regulation of the epithelial Na+ channel by paraoxonase-2. J Biol Chem 292(38):15927–15938. https://doi.org/10.1074/jbc.M117.785253
Shimizu Y, Okuda-Shimizu Y, Hendershot LM (2010) Ubiquitylation of an ERAD substrate occurs on multiple types of amino acids. Mol Cell 40(6):917–926. https://doi.org/10.1016/j.molcel.2010.11.033
Shinoda T, Ogawa H, Cornelius F, Toyoshima C (2009) Crystal structure of the sodium-potassium pump at 2.4 A resolution. Nature 459(7245):446–450. https://doi.org/10.1038/nature07939
Shipston MJ (2011) Ion channel regulation by protein palmitoylation. J Biol Chem 286(11):8709–8716. https://doi.org/10.1074/jbc.R110.210005
Simons JF, Ferro-Novick S, Rose MD, Helenius A (1995) BiP/Kar2p serves as a molecular chaperone during carboxypeptidase Y folding in yeast. J Cell Biol 130(1):41–49
Smith MH, Ploegh HL, Weissman JS (2011) Road to ruin: targeting proteins for degradation in the endoplasmic reticulum. Science 334(6059):1086–1090. https://doi.org/10.1126/science.1209235
Snyder PM (2005) Minireview: regulation of epithelial Na+ channel trafficking. Endocrinology 146(12):5079–5085
Snyder PM, McDonald FJ, Stokes JB, Welsh MJ (1994) Membrane topology of the amiloride-sensitive epithelial sodium channel. J Biol Chem 269(39):24379–24383
Sohara E, Rai T, Yang SS, Uchida K, Nitta K, Horita S, Ohno M, Harada A, Sasaki S, Uchida S (2006) Pathogenesis and treatment of autosomal-dominant nephrogenic diabetes insipidus caused by an aquaporin 2 mutation. Proc Natl Acad Sci U S A 103(38):14217–14222. https://doi.org/10.1073/pnas.0602331103
Somlo S, Ehrlich B (2001) Human disease: calcium signaling in polycystic kidney disease. Curr Biol 11(9):R356–R360
Spear ED, Ng DT (2005) Single, context-specific glycans can target misfolded glycoproteins for ER-associated degradation. J Cell Biol 169(1):73–82. https://doi.org/10.1083/jcb.200411136
Stagg HR, Thomas M, van den Boomen D, Wiertz EJ, Drabkin HA, Gemmill RM, Lehner PJ (2009) The TRC8 E3 ligase ubiquitinates MHC class I molecules before dislocation from the ER. J Cell Biol 186(5):685–692. https://doi.org/10.1083/jcb.200906110
Staub O, Dho S, Henry P, Correa J, Ishikawa T, McGlade J, Rotin D (1996) WW domains of Nedd4 bind to the proline-rich PY motifs in the epithelial Na+ channel deleted in Liddle’s syndrome. EMBO J 15(10):2371–2380
Staub O, Gautschi I, Ishikawa T, Breitschopf K, Ciechanover A, Schild L, Rotin D (1997) Regulation of stability and function of the epithelial Na+ channel (ENaC) by ubiquitination. EMBO J 16(21):6325–6336
Steel GJ, Fullerton DM, Tyson JR, Stirling CJ (2004) Coordinated activation of Hsp70 chaperones. Science 303(5654):98–101
Stein A, Ruggiano A, Carvalho P, Rapoport TA (2014) Key steps in ERAD of luminal ER proteins reconstituted with purified components. Cell 158(6):1375–1388. https://doi.org/10.1016/j.cell.2014.07.050
Suzuki T (2015) The cytoplasmic peptide:N-glycanase (Ngly1)-basic science encounters a human genetic disorder. J Biochem 157(1):23–34. https://doi.org/10.1093/jb/mvu068
Suzuki T, Park H, Hollingsworth NM, Sternglanz R, Lennarz WJ (2000) PNG1, a yeast gene encoding a highly conserved peptide:N-glycanase. J Cell Biol 149(5):1039–1052
Swatek KN, Komander D (2016) Ubiquitin modifications. Cell Res 26(4):399–422. https://doi.org/10.1038/cr.2016.39
Takeda K, Noguchi S, Sugino A, Kawamura M (1988) Functional activity of oligosaccharide-deficient (Na,K)ATPase expressed in Xenopus oocytes. FEBS Lett 238(1):201–204
Tamarappoo BK, Verkman AS (1998) Defective aquaporin-2 trafficking in nephrogenic diabetes insipidus and correction by chemical chaperones. J Clin Investig 101(10):2257–2267
Tang WK, Li D, Li CC, Esser L, Dai R, Guo L, Xia D (2010) A novel ATP-dependent conformation in p97 N-D1 fragment revealed by crystal structures of disease-related mutants. EMBO J 29(13):2217–2229. https://doi.org/10.1038/emboj.2010.104
Taxis C, Hitt R, Park SH, Deak PM, Kostova Z, Wolf DH (2003) Use of modular substrates demonstrates mechanistic diversity and reveals differences in chaperone requirement of ERAD. J Biol Chem 278(38):35903–35913
Tokhtaeva E, Sachs G, Vagin O (2009) Assembly with the Na,K-ATPase alpha(1) subunit is required for export of beta(1) and beta(2) subunits from the endoplasmic reticulum. Biochemistry 48(48):11421–11431. https://doi.org/10.1021/bi901438z
Tokhtaeva E, Munson K, Sachs G, Vagin O (2010) N-glycan-dependent quality control of the Na,K-ATPase beta(2) subunit. Biochemistry 49(14):3116–3128. https://doi.org/10.1021/bi100115a
Tokhtaeva E, Sachs G, Souda P, Bassilian S, Whitelegge JP, Shoshani L, Vagin O (2011) Epithelial junctions depend on intercellular trans-interactions between the Na,K-ATPase beta(1) subunits. J Biol Chem 286(29):25801–25812. https://doi.org/10.1074/jbc.M111.252247
Torres VE, Harris PC (2006) Mechanisms of disease: autosomal dominant and recessive polycystic kidney diseases. Nat Clin Pract Nephrol 2(1):40–55. https://doi.org/10.1038/ncpneph0070
Tyson JR, Stirling CJ (2000) LHS1 and SIL1 provide a lumenal function that is essential for protein translocation into the endoplasmic reticulum. EMBO J 19(23):6440–6452. https://doi.org/10.1093/emboj/19.23.6440
Valentijn JA, Fyfe GK, Canessa CM (1998) Biosynthesis and processing of epithelial sodium channels in Xenopus oocytes. J Biol Chem 273(46):30344–30351
van den Boomen DJ, Timms RT, Grice GL, Stagg HR, Skodt K, Dougan G, Nathan JA, Lehner PJ (2014) TMEM129 is a Derlin-1 associated ERAD E3 ligase essential for virus-induced degradation of MHC-I. Proc Natl Acad Sci U S A 111(31):11425–11430. https://doi.org/10.1073/pnas.1409099111
van der Goot AT, Pearce MMP, Leto DE, Shaler TA, Kopito RR (2018) Redundant and antagonistic roles of XTP3B and OS9 in decoding glycan and non-glycan degrons in ER-associated degradation. Mol Cell 70(3):516–530.e516. https://doi.org/10.1016/j.molcel.2018.03.026
Vashist S, Ng DT (2004) Misfolded proteins are sorted by a sequential checkpoint mechanism of ER quality control. J Cell Biol 165(1):41–52
Vashist S, Kim W, Belden WJ, Spear ED, Barlowe C, Ng DT (2001) Distinct retrieval and retention mechanisms are required for the quality control of endoplasmic reticulum protein folding. J Cell Biol 155(3):355–368. https://doi.org/10.1083/jcb.200106123
Vembar SS, Brodsky JL (2008) One step at a time: endoplasmic reticulum-associated degradation. Nat Rev Mol Cell Biol 9(12):944–957
Verma R, Oania R, Graumann J, Deshaies RJ (2004) Multiubiquitin chain receptors define a layer of substrate selectivity in the ubiquitin-proteasome system. Cell 118(1):99–110
Wahlman J, DeMartino GN, Skach W, Bulleid NJ, Brodsky JL, Johnson AE (2007) Real-time fluorescence detection of ERAD substrate retro-translocation in a mammalian in vitro system. Cell 129(5):943–955
Walter P, Blobel G (1981) Translocation of proteins across the endoplasmic reticulum III. Signal recognition protein (SRP) causes signal sequence-dependent and site-specific arrest of chain elongation that is released by microsomal membranes. J Cell Biol 91(2 Pt 1):557–561
Wanamaker CP, Christianson JC, Green WN (2003) Regulation of nicotinic acetylcholine receptor assembly. Ann N Y Acad Sci 998:66–80
Weisz OA, Wang JM, Edinger RS, Johnson JP (2000) Non-coordinate regulation of endogenous epithelial sodium channel (ENaC) subunit expression at the apical membrane of A6 cells in response to various transporting conditions. J Biol Chem 275(51):39886–39893
Welling PA (2016) ROMK and Bartter syndrome type 2. Ion channels and transporters of epithelia in health and disease. Springer, New York
Wiertz EJ, Tortorella D, Bogyo M, Yu J, Mothes W, Jones TR, Rapoport TA, Ploegh HL (1996) Sec61-mediated transfer of a membrane protein from the endoplasmic reticulum to the proteasome for destruction. Nature 384(6608):432–438
Willer M, Forte GM, Stirling CJ (2008) Sec61p is required for ERAD-L: genetic dissection of the translocation and ERAD-L functions of Sec61P using novel derivatives of CPY. J Biol Chem 283(49):33883–33888. https://doi.org/10.1074/jbc.M803054200
Wu X, Rapoport TA (2018) Mechanistic insights into ER-associated protein degradation. Curr Opin Cell Biol 53:22–28. https://doi.org/10.1016/j.ceb.2018.04.004
Wu Q, Moeller HB, Stevens DA, Sanchez-Hodge R, Childers G, Kortenoeven MLA, Cheng L, Rosenbaek LL, Rubel C, Patterson C, Pisitkun T, Schisler JC, Fenton RA (2018) CHIP regulates aquaporin-2 quality control and body water homeostasis. J Am Soc Nephrol 29(3):936–948. https://doi.org/10.1681/ASN.2017050526
Wuller S, Wiesner B, Loffler A, Furkert J, Krause G, Hermosilla R, Schaefer M, Schulein R, Rosenthal W, Oksche A (2004) Pharmacochaperones post-translationally enhance cell surface expression by increasing conformational stability of wild-type and mutant vasopressin V2 receptors. J Biol Chem 279(45):47254–47263. https://doi.org/10.1074/jbc.M408154200
Xia D, Tang WK, Ye Y (2016) Structure and function of the AAA+ ATPase p97/Cdc48p. Gene 583(1):64–77. https://doi.org/10.1016/j.gene.2016.02.042
Yan W, Samaha FF, Ramkumar M, Kleyman TR, Rubenstein RC (2004) Cystic fibrosis transmembrane conductance regulator differentially regulates human and mouse epithelial sodium channels in Xenopus oocytes. J Biol Chem 279(22):23183–23192. https://doi.org/10.1074/jbc.M402373200
Ye Y, Meyer HH, Rapoport TA (2001) The AAA ATPase Cdc48/p97 and its partners transport proteins from the ER into the cytosol. Nature 414(6864):652–656
Yoshimura SH, Iwasaka S, Schwarz W, Takeyasu K (2008) Fast degradation of the auxiliary subunit of Na+/K+-ATPase in the plasma membrane of HeLa cells. J Cell Sci 121(Pt 13):2159–2168. https://doi.org/10.1242/jcs.022905
You H, Ge Y, Zhang J, Cao Y, Xing J, Su D, Huang Y, Li M, Qu S, Sun F, Liang X (2017) Derlin-1 promotes ubiquitylation and degradation of the epithelial Na+ channel, ENaC. J Cell Sci 130(6):1027–1036. https://doi.org/10.1242/jcs.198242
Young JC (2014) The role of the cytosolic HSP70 chaperone system in diseases caused by misfolding and aberrant trafficking of ion channels. Dis Model Mech 7(3):319–329. https://doi.org/10.1242/dmm.014001
Younger JM, Chen L, Ren HY, Rosser MF, Turnbull EL, Fan CY, Patterson C, Cyr DM (2006) Sequential quality-control checkpoints triage misfolded cystic fibrosis transmembrane conductance regulator. Cell 126(3):571–582. https://doi.org/10.1016/j.cell.2006.06.041
Zaarour N, Demaretz S, Defontaine N, Zhu Y, Laghmani K (2012) Multiple evolutionarily conserved Di-leucine like motifs in the carboxyl terminus control the anterograde trafficking of NKCC2. J Biol Chem 287(51):42642–42653. https://doi.org/10.1074/jbc.M112.399162
Zamofing D, Rossier BC, Geering K (1989) Inhibition of N-glycosylation affects transepithelial Na+ but not Na+-K+-ATPase transport. Am J Phys 256(5 Pt 1):C958–C966. https://doi.org/10.1152/ajpcell.1989.256.5.C958
Zattas D, Berk JM, Kreft SG, Hochstrasser M (2016) A conserved C-terminal element in the yeast Doa10 and human MARCH6 ubiquitin ligases required for selective substrate degradation. J Biol Chem 291(23):12105–12118. https://doi.org/10.1074/jbc.M116.726877
Zhang Y, Nijbroek G, Sullivan ML, McCracken AA, Watkins SC, Michaelis S, Brodsky JL (2001) Hsp70 molecular chaperone facilitates endoplasmic reticulum-associated protein degradation of cystic fibrosis transmembrane conductance regulator in yeast. Mol Biol Cell 12(5):1303–1314
Zhou R, Tomkovicz VR, Butler PL, Ochoa LA, Peterson ZJ, Snyder PM (2013) Ubiquitin-specific peptidase 8 (USP8) regulates endosomal trafficking of the epithelial Na+ channel. J Biol Chem 288(8):5389–5397. https://doi.org/10.1074/jbc.M112.425272
Zhu X, Zhao X, Burkholder WF, Gragerov A, Ogata CM, Gottesman ME, Hendrickson WA (1996) Structural analysis of substrate binding by the molecular chaperone DnaK. Science 272(5268):1606–1614
Acknowledgments
The authors would like to thank the editors, Kirk L. Hamilton and Daniel C. Devor, for the invitation to contribute to this edition. This work was supported by NIDDK DK109024 and DK117126 to T.M.B. and GM075061, GM131732 and DK079307 to J.L.B.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 The American Physiological Society
About this chapter
Cite this chapter
Buck, T.M., Brodsky, J.L. (2020). Epithelial Ion Channel Folding and ER-Associated Degradation (ERAD). In: Hamilton, K.L., Devor, D.C. (eds) Basic Epithelial Ion Transport Principles and Function. Physiology in Health and Disease. Springer, Cham. https://doi.org/10.1007/978-3-030-52780-8_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-52780-8_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-52779-2
Online ISBN: 978-3-030-52780-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)