Abstract
A typical breast MRI exam results in thousands of image slices from multiple sequences, collected over multiple time points and with different tissue contrasts. Computerized support systems help the radiologist to navigate through these images by detecting and characterizing parenchymal lesions. This chapter divides computerized systems for breast MRI into three main categories: computer-aided evaluation (CAE) systems that provide improved visualization of the image data and support the radiologists workflow; computer-aided diagnosis systems (CADx) that provide an estimate of the probability of a specific lesion being a cancer; and computer-aided detection and diagnosis (CADD) systems that first identify possible lesions and then classify them in terms of probability of being malignant or benign. Various steps of these automated systems are described such as lesion segmentation, feature extraction (including kinetic, morphological, and texture features), and lesion classification (by means of feature selection, training, and evaluation of classifiers). Moreover, systems for fully automated lesion detection as well as systems for motion correction (image registration) and breast segmentation are described. Finally, challenges that have hindered the widespread adoption of CAD systems for routine breast MRI clinical practice and opportunities for future research aimed at their improvement are illustrated.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
7.1 Introduction
Computerized support systems for mammography have been commercially available for many years and are used widely for diagnostic support and as a second reader. For mammographic systems, the term computer-aided detection (CADe) is typically used to denote a system that detects suspicious lesions while the term computer-aided diagnosis (CADx) is used to describe systems that provide an estimate of the probability that a detected lesion is cancer. The acronym “CAD” can indicate computed-aided detection, computed-aided diagnosis, or both. For breast MRI, the challenges are somewhat different. A typical MRI breast exam can result in thousands of image slices being acquired; images are volumetric and can be acquired in different planes; and there are multiple sequences, each of them resulting in a different tissue contrast, while dynamic contrast-enhanced (DCE) sequences provide additional temporal information. Computerized support systems are needed to help the radiologist to navigate through these images effectively. The high signal intensity in cancerous lesions that results from contrast enhancement provides excellent sensitivity, but the presence of many enhancing benign lesions and, in some cases, enhancing parenchymal tissue means that the differentiation between malignant and benign lesions is a difficult task. This chapter divides computerized decision support systems for breast MRI into three main categories: computer-aided evaluation (CAE) systems, which provide improved visualization of the image data and support the radiologists workflow; computer-aided diagnosis (CADx) systems, which provide an estimate of the probability of a specific lesion being a cancer; and computer-aided detection and diagnosis (CADD) systems, which first identify possible lesions and then classify them in terms of probability of being malignant or benign.
7.2 Computer-Aided Evaluation (CAE) Systems
There are several commercially available software packages that are designed to provide support to the breast radiologist evaluating a magnetic resonance imaging (MRI) examination of the breast.
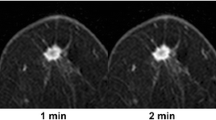
The main function of these packages is to provide a color-coded parametric map of the breast based on the Breast Imaging Reporting and Data System (BI-RADS) scheme [1] where enhancement kinetics are classified as persistent, plateau, or washout of contrast material. For each pixel in the image, a signal intensity curve is generated and the classification is performed as follows. First, an enhancement threshold is set based on the percentage increase in the signal in the first post-contrast image and only pixels that exceed this threshold are retained. Next, the software calculates the change in intensity in a delayed post-contrast image relative to the first post-contrast image. Finally, it determines whether each pixel intensity curve increases, decreases, or remains constant and assigns the corresponding color coding to an overlay map. The precise details of which time points are used, what the initial enhancement threshold should be, and which metric is used to distinguish between pixels that show washout, plateau, or continuous enhancement vary from platform to platform; but the essential principles are the same. In Fig. 7.1, signal enhancement ratio (SER) pixel-by-pixel maps are shown for a malignant and a benign lesion, together with the corresponding relative signal curves in Fig. 7.2. Here the signal enhancement ratio is defined as SER = (Sfirst − S0)/(Slast − S0) where S0, Sfirst, and Slast are the pre-contrast, first post-contrast, and last post-contrast signal intensities, respectively.
On the top row are images showing an invasive lobular carcinoma and on the bottom row images show a fibroadenoma. The left column shows the sagittal sections through the center of the lesions obtained as the first post-contrast frame using a fat-saturated gradient-echo sequence. On the right column, the signal enhancement ratio (SER) values are displayed as a color overlay on a pixel-by-pixel basis. Note the inhomogeneous color distribution with multiple yellow-red pixels in the malignant lesion and the homogeneous light blue color of the benign lesion
The signal enhancement curves corresponding to the lesions shown in Fig. 7.1. The malignant carcinoma (red) shows a more rapid initial enhancement followed by a slight late washout phase. The benign lesion has a slower initial enhancement without washout; i.e., it shows a continuous increase curve. These curves were obtained by considering the pixels showing the highest initial enhancement
Changes in magnet field strength, equipment vendor and model, software version, image acquisition protocols, contrast type, dose, and rate of administration, flushing with saline solution, and threshold values used to generate the color maps mean that overlay maps produced in one breast imaging center cannot be directly compared with those generated elsewhere. Pharmacokinetic (PK) models attempt to overcome this variability by estimating physiologically meaningful parameters such as the rate of exchange between capillaries and the extracellular space by fitting mathematical models to signal intensity curves [2]; however, these models require that images are acquired at a much higher temporal resolution than is common in clinical practice. The three-time-point (3TP) method [3] proposes a solution where just three images are acquired at specified time points and then a calibration scheme is used to estimate the pharmacokinetic parameters.
In addition to an overlay that color codes each pixel according to enhancement kinetics, some CAE systems provide tools for radiologists to identify and outline lesions of interest. The average signal intensity curve over the whole lesion can then be assessed, which reduces the effect of noise. Summary statistics that describe the area and extent of the lesion can also be produced from the segmented region of interest (ROI). Although CAE systems do provide some quantitative information, they are not designed to assign probabilities of malignancy to lesions in the image.
Several studies have evaluated commercially available CAE systems, including CADstream (Confirma, Bellevue, WA, USA) [4], Aegis (Sentinelle Medical, Toronto, Canada) [5], dynaCAD (Invivo, Pewaukee, WI, USA) [6], and 3TP [7]. Monique D. Dorrius and coworkers [8] carried out a systematic review and meta-analysis considering ten publications referencing commercially available systems. They reported that for experienced radiologists the sensitivity was unchanged by the use of a CAE system and there was a small but non-significant decrease in specificity. For residents with less than 6 months’ breast MRI experience, there was a significant improvement in sensitivity with CAE and no significant change in specificity. The authors concluded that CAE systems had little impact on accuracy overall and that inexperienced radiologists and residents benefitted the most from their use [8].
7.3 Computer-Aided Diagnosis (CADx) Systems
CAE systems can be very helpful in highlighting enhancing regions of the breast so that the radiologist can quickly direct their attention to these suspicious areas; however, they do not make use of other information such as the texture and the morphology of the lesion. CADx systems provide further support to the radiologist by combining kinetic, morphological, and textural information to predict whether a particular lesion is malignant or not. This is achieved by first delineating the suspicious lesion, then extracting multiple image features from the DCE sequence, and finally using a trained classifier to assign a probability of malignancy to the lesion. This is achieved using a machine learning algorithm, which is trained on previously labeled examples of malignant and benign lesions. Each of these components will be described in more detail below.
7.3.1 Lesion Segmentation
The accurate delineation of enhancing lesions is essential as it allows us to quantify the variation in contrast enhancement kinetics within the lesion and to extract morphological features that can represent its shape. Manual segmentation is a time-consuming and subjective process. Semi-automated methods have been shown to be faster and to reduce inter-observer variability [9]. Typically, such methods require the radiologist to mark a point at the center of the enhancing lesion or to draw a crude boundary or bounding box around the lesion. Upper and lower intensity thresholds are then set by the user, and pixels within the defined range, which are either connected to the seed point or lie within the bounding box, are defined as belonging to the lesion ROI. Usually the subtraction images or the enhancement maps are used to define the ROI as they have higher contrast between the lesion and the background.
More sophisticated approaches remove the need to manually define intensity threshold values and make use of all the DCE information available to improve the contrast between lesion and background. Weijie Chen and coworkers [10] developed an improved lesion segmentation algorithm based on fuzzy clustering that used the difference in contrast enhancement dynamics to identify pixels belonging to the foreground lesion. Yunfeng Cui and coworkers [11] used a Gaussian mixture model to automatically estimate threshold values that are used to identify pixels lying inside and outside of the lesion. A marker-controlled watershed method is then used to further refine the boundary. Other authors [12] described a system where the operator places two ellipses on the image, one identifying pixels inside the lesion the other containing background; and this information is used to classify all the remaining pixels. Alternative methods are those using a graph-cut-based algorithm that incorporates a spatial smoothness constraint [13].
Fully automated systems that carry out both detection and segmentation of lesions are discussed separately in Sect. 7.4.
7.3.2 Feature Extraction
Radiologists use well-defined descriptors [1] to characterize lesions, and these help to discriminate between malignant and benign lesions. Although there have been some attempts to build CADx systems based on categorical descriptors provided by radiologists [14], it is more common to extract continuous quantitative values that capture the same information. There are many papers describing different feature sets for use in CADx systems, and they can be grouped into three groups: kinetic (also called dynamic), morphological, and texture features.
7.3.2.1 Kinetic Features
There are many different ways of quantifying contrast enhancement in a lesion, but model-free methods, which attempt to characterize the shape of the signal enhancement curve, are the most commonly used. Features include the maximum enhancement, the time-to-peak enhancement, the rate of contrast uptake, and the rate of washout [15]. The normalized signal intensity values have been used directly [16]; however, when Jacob Levman and coworkers [17] compared several feature vectors, including one that used relative signal intensity alone and another that combined relative signal enhancement with the derivatives of the enhancement curve, they found that the more conventional feature vector based on the traditional parameters of maximum signal intensity enhancement, time of maximum enhancement, and maximum washout gave the most accurate results. Pharmacokinetic models require high temporal resolution and therefore are not suitable for most breast MRI exams, which typically only have 3–5 post-contrast images acquired at a lower temporal resolution, typically not lower than 60 s. Sanaz A. Jansen and coworkers [18] describe an empirical model that has just three parameters to fit and does not require an arterial input function. This approach may help to standardize kinetic parameters extracted from studies acquired at differing temporal resolutions, and features obtained using this model have been found to be relevant in lesion classification [19].
The contrast enhancement curve generated over an entire lesion will result in the averaging of pixel signal intensity curves. Several groups have attempted to cluster together pixels that show similar enhancement patterns in order to capture regions that show the greatest wash in and wash out of contrast. These include the mean-shift algorithm [20], vector quantization [21], and fuzzy c-means clustering [22]. Another approach is to differentiate between the signal enhancement in the center of the lesion and at the edge of the lesion [19].
7.3.2.2 Morphological Features
Radiologists use several morphological features such as the shape of the lesion, and the uniformity (i.e., pattern of internal distribution) of contrast enhancement to describe a lesion. Certain characteristics are associated with benign lesions while others tend to suggest a malignant lesion. For example, a stereotypical benign lesion may have a smooth margin, with an oval shape and internal septations, whereas a malignant lesion might have a speculated appearance with an irregular shape and rim enhancement. In order to use this information in a CADx system, it is necessary to quantify these findings. Various formula have been derived to capture information about circularity, convexity, irregularity, solidity, perimeter, compactness, etc. [9, 15, 23]. The sharpness of the lesion boundary, and the change in edge sharpness over the duration of the dynamic study are also useful morphological features [24, 25].
7.3.2.3 Texture Features
Texture features provide information about the heterogeneity of the contrast enhancement in the lesion. Since the mean signal intensity curve generated over the whole lesion region of interest does not reflect inhomogeneities within the lesion, many CADx algorithms also include the variance, skew, and kurtosis of each of the kinetic parameters measured from individual pixels within the ROI [15, 19]. However, features based purely on the statistical distribution of intensity values cannot capture spatial patterns. In 1973 Robert M. Haralick [26] introduced a method of mathematically describing textures in images that uses spatially dependent intensity information. Haralick features are based on a co-occurrence matrix Pij, which records the number of times that two pixels with values i and j occur in the region of interest separated by a distance d and an angle θ. Fourteen feature values can be derived from this matrix, including the angular second moment (ASM), energy, entropy, and contrast. Even more features can be obtained by varying values for d, θ, and the number of gray levels used to generate the matrix. Peter Gibbs and coworkers [27] showed that a combination of texture features could produce very accurate results, and Weijie Chen and coworkers [28] extended the method to three-dimensional (3D) volumetric regions of interest. Other texture features have also been used to discriminate between malignant and benign lesions, such as Gabor filters [13] or entropy of enhancement assessed by moving a 3 × 3 window over the lesion ROI [29].
7.3.3 Lesion Classification
Individual features rarely achieve high accuracy in isolation. However, when several features are combined, it is possible to achieve a better separation between malignant and benign lesions. Classification algorithms work by finding a boundary in multi-dimensional feature space that best separates two sets of labeled data points; once this boundary has been identified using training data, a new test case is projected into the feature space and, depending on which side of the decision boundary it falls, it is classified as malignant or benign. There are many different classifiers available that can take a set of features and return either a binary decision or a probability of malignancy.
7.3.3.1 Classifiers
Simple linear classifiers, such as linear discriminant analysis (LDA), have the advantage that they are easily understood and the contribution that each individual feature makes to the final decision can be calculated [27]. The disadvantage is that they cannot cope with data where the decision boundary is non-linear.
Support vector machines (SVMs) are more robust than LDA with small training datasets as they identify the decision boundary that maximizes the distance to the data points on either side. They can be extended to produce non-linear boundaries using different kernel functions and provide a mechanism for coping with misclassified points. The disadvantage of SVMs is that understanding the contribution of individual features to the classifier becomes much more difficult [17].
Decision trees are simple to understand, and Pascal Baltzer and coworkers achieved excellent results on a dataset of over 1,000 patients [14] using categorical features. The resulting tree could be represented by a series of simple decision rules; however, this approach is known to be prone to overfitting. Random forests , which are ensembles of many individual decision trees, are more robust [30] and have been used successfully to train breast CAD systems [19, 31]. It is possible to extract useful information about the importance of individual features using random forests, and methods of interpreting random forest models have also been explored [32].
Artificial neural networks (ANNs) attempt to mimic the way in which a human brain processes information. The features are connected to a layer of hidden nodes, and then these hidden nodes are connected to output nodes that represent the classes. A back-propagation method is used to learn the weights that connect the nodes together. Several groups have used neural networks for lesion characterization on breast MRI in the past [16, 33, 34], but the small size of the labeled training datasets that were available meant that these ANNs were restricted to a single hidden layer with just a few nodes. More recently there has been an explosion of interest in the use of deeper neural networks and more advanced networks that are specifically designed for images—these will be mentioned in later sections.
7.3.3.2 Feature Selection
Hundreds of quantitative features can be extracted from a DCE-MRI study, but using too many features increases computational complexity and may lead to overfitting. In practice, therefore, it is usually better to train a classification algorithm using a subset of the most discriminative features. Many methods of feature selection exist in the literature, and the simplest approach is to identify the top-ranking features individually. The discriminative power of a single feature can be quantified by using the receiver operating characteristic (ROC) analysis and calculating the area under the curve (AUC). Looking at one feature at a time, however, does not take into account the correlations between features, so methods that attempt to find the best combination of features have been proposed. Sequential forward search methods find the most discriminating feature first, and then search for a second feature that results in the greatest improvement in accuracy, and so on until the required number of features has been identified [34, 35]. Silvano Agliozzo and coworkers [23] used a genetic algorithm to identify the best subset of features. Some classification algorithms are able to automatically determine the relevance of features, for example, random forests [19] and Bayesian neural networks with automatic relevance determination [33].
7.3.3.3 Training and Evaluation of Classifiers
When a classifier is trained on labeled data, it is important to use a separate testing dataset to evaluate performance; otherwise the calculated accuracy will be overly optimistic. Overfitting of a classifier is said to occur when predictions made on the training set are very accurate but the performance on new unseen data is poor; i.e., the classification model fails to generalize. This can be avoided by careful attention to the way the labeled training data is used to create and test predictive models. Figure 7.3 illustrates a general framework for selecting the model parameters (for example, the number of trees in a random forest, hidden nodes in a neural network, or feature selection) using a labeled data set.
The best test of generalizability is obtained by using a completely independent testing dataset . This should be separated from the training data before any experiments are started to ensure that the choice of parameters or features is not biased. The remaining data is then split into a training set and a tuning set (often referred to as the validation set in the computer science literature). The training set is used to create the predictive model, and the tuning set is used to estimate performance. This process can then be repeated by switching cases in the training and tuning sets using a process known as cross-fold validation . The number of cases in the training, tuning, and testing datasets and the number of folds used will depend on the size of the available labeled dataset and the number of classes. For very small datasets, it is common to carry out a leave-one-out (LOO) experiment where all of the cases except one are used to train a classifier that is then used to predict the label on the remaining case. This process is repeated until each case has been held out and the reported accuracy is calculated. This procedure usually yields overly optimistic results. If the research then uses repeated LOO experiments to select model parameters and then reports on the most accurate configuration, then the results are also biased.
The independent testing set cannot be too small, or it will fail to capture the variability of the data and there will be a high variance in the error accuracy. In many cases, researchers will attempt to increase the number of labeled cases by using several lesions from a single patient. If this is done, then it is important to ensure that all of the lesions from a single patient are in the same dataset; i.e. it is incorrect to include lesions from the same patient in both the training set and the tuning or testing set.
Most classifiers return a numerical score between zero (definitely negative) and 1 (definitely positive). For a simple binary classifier, it is common to set a threshold of 0.5: everything with a higher score is considered to be positive and everything with a lower score is considered to be negative. Lowering this threshold results in a higher sensitivity (true positive fraction) and a lower specificity (true negative fraction) while raising the threshold produces a lower sensitivity and a higher specificity. The optimum setting for this threshold, which is also referred to as the decision point, will depend on the clinical context. The effect of changing the threshold can be visualized using the ROC curve, which plots sensitivity against (1-specificity) for different decision points. The AUC is often used to evaluate the performance of different CADx systems because it is independent of the single threshold.
7.4 Fully Automated Lesion Detection
In Sect. 7.3, it is assumed that a lesion has already been detected. A system that is capable of detecting suspicious lesions automatically, i.e., a CADe system, has the potential to speed up radiologists’ workflow and also to improve sensitivity by detecting otherwise overlooked cancers. In breast MRI, however, there are many regions of non-specific enhancement that may also be identified as lesions, and if a CADe system is to be useful, it is essential that the number of false positive detections is minimized. This makes the combined task of detection and diagnosis very challenging.
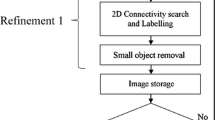
Several attempts to automatically detect lesions have been described in the literature. Mayer et al. [36] automatically segmented images into clusters of similar pixels using a hierarchical Gaussian pyramid and identified clusters with the highest local intensity values. This process led to the creation of about 2,500 objects for each breast exam from which morphological and dynamic features were extracted. After removing most of these objects using size and volume criteria, the remaining objects were classified as lesions or artifacts by a first ANN and then the lesions were classified as malignant or benign by a second ANN. Malignant lesions were detected with a sensitivity of 95% and a specificity of 92%. Diane M. Renz and coworkers [37] used the same approach on an independent dataset and reported a sensitivity of 97% and a specificity of 76%.
Anna Vignati and cowokers [38] proposed a lesion detection pipeline that included breast segmentation, image registration, and the normalization of contrast using the signal intensity in the blood vessels. In order to reduce the number of false-positive detections, they used a number of heuristically derived rules, which included a minimum size criteria of 20 mm3 and the rejection of any lesions where variation in signal intensity exceed a certain threshold. They were able to detect 89% of all lesions with a false detection rate of 4 false detections per breast. Most of the false-positive detections were due to blood vessels. The lesions detected were then classified as malignant or benign using a support vector machine [23].
Yan-Hao Huang and coworkers [39] used a thresholding method to isolate the enhancing tissue from background and then subdivided the enhancing regions into four groups using fuzzy clustering. This process still tended to identify background enhancement and vessels as suspicious, so a multi-scale Hessian filter was used to identify mass lesions. Morphological, texture, and enhancement features were extracted from the detected lesions; and logistic regression was used to classify malignant lesions. They reported a sensitivity of 92% with 4.6 false positives per case.
Albert Gubern-Merida and coworkers [31] used both Laplacian and Hessian filters to identify bright blob-like structures as potential lesions. Their patient population included women with both mass and non-mass malignant lesions and women with negative screening examinations and no breast cancer. Women with biopsy-proven benign lesions were not included in this study. They then compared several different classification methods and found that a random forest classifier gave the best performance with 7 false-positive lesions per patient at a sensitivity of 95%.
Hongbo Wu and coworkers [40] used an ANN with two hidden layers to classify small patches of the dynamic image as either lesion or non-lesion. In order to overcome the problem of insufficient labeled data to train a deep neural network, they used a denoising autoencoder [41], which allows features to be learned directly from unlabeled data. Once the network was pre-trained using unlabeled data, a smaller number of labeled patches were used to train the classifier to differentiate between lesions and non-lesions and a sensitivity of 92% with 17 false candidate lesion regions per volume was obtained. Once the lesions have been identified, it is possible to extract more conventional morphological and textural features and Hongbo Wu [42] found that adding a cascade of random forest classifiers, one to remove false-positive detections and one to differentiate between malignant and benign lesions, gave a final sensitivity of 94% at 0.12 false-positive detections per normal study. Figure 7.4 illustrates the work flow for the final classification algorithm.
Example of a processing pipeline for a breast MRI CADD system. (a) Preprocessing of the data typically involves motion correction. (b) An ANN is used to assign a lesion probability to each pixel in the image. (c) Once the lesion is identified, features can be extracted relating to enhancement, morphology, and texture. (d) A random forest classifier then uses these features to reduce false-positive detections. (e) A final classifier then differentiates between malignant and benign lesions
It is difficult to directly compare the results from these studies as the patient population differs in each case. In most studies, patients with biopsy-proven malignant or benign lesions are selected, but this does not assess the false-positive rate in examinations that do not contain any lesions at all. In the study by Albert Gubern-Mérida and coworkers [31], the false-positive rate is assessed on negative screening exams where 2 years’ follow-up confirmed that there was no breast cancer but no biopsied benign lesions were included in the study. In the master’s thesis by Hongbo Wu [42], the false-positive rate was also assessed on negative screening exams but benign lesions were present in the data used to train the classifiers.
7.5 Preprocessing: Motion Correction (Image Co-registration) and Breast Segmentation
In all CAD systems, features that quantify the change in intensity over time are used to differentiate between normal, benign, and malignant regions. Any motion between the pre- and post-contrast images will have an impact on these quantitative measures; therefore, motion correction , also referred to as image co-registration , is frequently carried out as a pre-processing step. The registration of contrast-enhanced breast MRI is challenging for two main reasons: the breast tissue is highly deformable and the changing intensity in enhancing regions can affect the accuracy of registration. Several methods have been evaluated motion correction for breast imaging [43,44,45,46]. Additional constraints on the deformable registration in order to prevent non-physiological changes in tumor volume have been described [46]. A framework for decoupling the effects of intensity changes due to motion and due to contrast enhancement has also been proposed [47]. Validation of motion correction is very difficult as the breast lacks anatomical landmarks that can be accurately localized in 3D images, and many landmarks are needed to assess a deformable registration algorithm. Some groups have used simulation studies based on finite element models of breast deformation [44, 48, 49] whilst others have attempted to carry out a subjective evaluation [43]. Albert Gubern-Mérida and coworkers [31] assessed the effect of motion correction on the final CAD outcome: the impact on overall accuracy was small but significant and that there was a greater improvement in accuracy for non-mass lesions. Figure 7.5 illustrates how motion correction can improve the quality of subtracted MRI images.
Motion correction. (a) Post-contrast image. (b) Result of subtracting the pre-contrast image from the post-contrast image. The effect of motion between the acquisition of these two images is well visible. (c) Subtracted image after motion correction [43]
Another useful preprocessing step for systems that perform both detection and diagnosis is breast segmentation . The areas of image artifact and high contrast enhancement in the chest can be misclassified as suspicious lesions, and, although radiologists are not affected by these errors, they do cause problems when evaluating automated systems. Processing time may also be affected as image features have to be calculated for every lesion identified by the system. Several breast segmentation algorithms have been developed for the purpose of assessing breast density with MRI [49,50,51,52,53,55], but Albert Gubern-Mérida and coworkers [31] noted that two lesions were missed due to segmentation errors, so there is still a need for improvements in this area.
7.6 Challenges
The use of CAD systems for mammography is widespread, but this is not true for MRI CAD despite over 15 years of research in this area. The computer-aided evaluation tools described in Sect. 7.2 are available on many commercial workstations, but these do not attempt to detect or diagnose lesions and cannot be used as a second reader.
One of the main differences between breast MRI and digital mammography is that there is much more variation in imaging protocols with MRI. Some differences—such as the use of 1.5 T or 3 T field strengths, the use of fat suppression, the type and dose contrast material and its injection protocol, or the timing of post-contrast images—will affect the relative signal intensity in the lesion compared with the background. Other differences—such as the choice of acquisition plane and the pixel size used—will affect the morphological features. A few studies have evaluated the accuracy of CAD systems that have been trained using data acquired using one protocol and then tested using data acquired using a different protocol, scanner or from a different institution. Weijie Chen and coworkers [33] compared datasets acquired on scanners from two different manufacturers and found that there was no significant difference in accuracy between a classifier trained on dataset 1 and tested on dataset 2 or vice versa. However, the protocols for the two datasets were very similar; both carried out acquisition in the coronal plane, no fat suppression was used, and the temporal resolution only differed by one second. Anna Vignati and coworkers [38] designed their lesion detection algorithm to work with both fat-saturated and non-fat saturated images, and their algorithm was trained and tested using images acquired using both protocols. Their two datasets also had very different temporal resolutions, and the effect of this was minimized by taking the mean signal intensity over the sequence of images and then normalizing intensity values using the intensity in the mammary blood vessels. These studies suggest that it is possible to design a CAD algorithm to work across different datasets from different institutions, but so far no authors have evaluated a fully automatic detection and classification algorithm in a clinically realistic scenario, where the software is tested on totally unseen images acquired using different protocols to those represented in the data used to train the algorithm.
The lack of standardization of imaging protocols is not the only reason that comparing the results of different breast MRI CAD studies is difficult. The patient populations also vary greatly from study to study, and this has an impact on the size and types of lesions used to train and test the algorithms. In most of the earlier studies, patients were undergoing MRI as a follow-up examination after mammography to either provide additional diagnostic information or to exclude the presence of additional lesions before surgery. In later studies, an increasing number of high-risk patients undergoing MRI screening have been included and, as a result, such studies may contain a greater proportion of very small lesions.
It is important to determine how best to incorporate CAD into breast radiologists’ workflow. For example, should the automated method be run before the radiologist reviews the images in order to speed up the work flow, or should it only be applied as a second look after the initial assessment has been made? The true impact of a breast MRI CAD system on sensitivity, specificity, and reporting times can only be evaluated in the context of the clinical workflow, which includes the breast radiologist; and only a few small studies [6, 56] have attempted this so far.
7.7 Opportunities
The use of additional MRI sequences to the DCE acquisition could improve discrimination between malignant and benign breast lesions. Although T2-weighted imaging is widely used in clinical breast MR, only a few studies have looked at the effect of adding T2 image features to classification [57, 58]. Diffusion-weighted imaging [59] and DCE-MRI with higher temporal resolution [60, 61] could also provide more discriminative features, but these sequences, especially the latter, are less commonly performed.
Incorporating information from previous MRI studies could also improve the accuracy of CAD systems. Women enrolled on MRI breast screening programs typically have annual examinations, and incorporating information from previous visits could improve specificity. Similarly, it could be advantageous to incorporate information derived from mammography.
Recent advances in machine learning have the potential to further improve the accuracy of breast MRI CAD. There has recently been a great deal of excitement over the use of convolutional neural networks (CNNs), which are capable of learning features directly from imaging data [62], and this approach has already been used to segment fibroglandular tissue in breast MRI images [53] and to classify lesions in mammography [63]. The performance of a CNN usually improves as the number of labeled training cases increases. Large databases of labeled images have been made available for several other CAD applications including mammography and nodule detection in chest computed tomography, and these have facilitated research and development in these areas. The creation of a large, publicly available, well-annotated, multi-institutional database for breast MRI would likely accelerate progress toward clinical CAD systems for breast MRI.
Significant progress has been made in the accuracy of breast MRI CAD, but in order to move this work into the clinical domain, it is essential that CAD platforms are tested in the context of the radiologist’s work flow and it is also essential that large-scale, multi-institutional studies are carried out to determine how robust these methods are when data is acquired on multiple scanners and with different protocols.
Abbreviations
- 3D:
-
Three-dimensional
- 3TP :
-
Three-time-point
- ANN:
-
Artificial neural network
- ASM :
-
Angular second moment
- AUC :
-
Area under the curve
- BI-RADS :
-
Breast Imaging Reporting and Data System
- CAD:
-
Computer-aided detection/diagnosis
- CADD :
-
Computer-aided detection and diagnosis
- CADe :
-
Computer-aided detection
- CADx :
-
Computer-aided diagnosis
- CAE :
-
Computer-aided evaluation
- CNN:
-
Convolutional neural networks
- DCE :
-
Dynamic contrast-enhanced
- LDA:
-
Linear discriminant analysis
- LOO :
-
Leave-one-out
- MRI :
-
Magnetic resonance imaging
- PK :
-
Pharmacokinetic
- ROC :
-
Receiver operating characteristic
- ROI :
-
Region of interest
- SER:
-
Signal enhancement ratio
- SVM:
-
Support vector machines
References
Morris EA, Comstock CE, Lee CH (2013) ACR BI-RADS® Magnetic Resonance Imaging. In: American College of Radiology. Breast Imaging Reporting and Data System® (BI-RADS®). 5th edition. American College of Radiology, Reston, VA, USA
Tofts PS, Brix G, Buckley DL et al (1999) Estimating kinetic parameters from dynamic contrast-enhanced T(1)-weighted MRI of a diffusable tracer: standardized quantities and symbols. J Magn Reson Imaging 10:223–232
Furman-Haran E, Degani H (2002) Parametric analysis of breast MRI. J Comput Assist Tomogr 26:376–386
Lehman CD, Peacock S, DeMartini WB, Chen X (2006) A new automated software system to evaluate breast MR examinations: improved specificity without decreased sensitivity. AJR Am J Roentgenol 187:51–56
Arazi-Kleinman T, Causer PA, Jong RA, Hill K, Warner E (2009) Can breast MRI computer-aided detection (CAD) improve radiologist accuracy for lesions detected at MRI screening and recommended for biopsy in a high-risk population? Clin Radiol 64:1166–1174
Baltzer PA, Freiberg C, Beger S et al (2009) Clinical MR-mammography: are computer-assisted methods superior to visual or manual measurements for curve type analysis? A systematic approach. Acad Radiol 16:1070–1076
Kelcz F, Furman-Haran E, Grobgeld D, Degani H (2002) Clinical testing of high-spatial-resolution parametric contrast-enhanced MR imaging of the breast. AJR Am J Roentgenol 179:1485–1492
Dorrius MD, Jansen-van der Weide MC, van Ooijen PM, Pijnappel RM, Oudkerk M (2011) Computer-aided detection in breast MRI: a systematic review and meta-analysis. Eur Radiol 21:1600–1608
Liney GP, Sreenivas M, Gibbs P, Garcia-Alvarez R, Turnbull LW (2006) Breast lesion analysis of shape technique: semiautomated vs. manual morphological description. J Magn Reson Imaging 23:493–498
Chen W, Giger ML, Bick U (2006) A fuzzy c-means (FCM)-based approach for computerized segmentation of breast lesions in dynamic contrast-enhanced MR images. Acad Radiol 13:63–72
Cui Y, Tan Y, Zhao B et al (2009) Malignant lesion segmentation in contrast-enhanced breast MR images based on the marker-controlled watershed. Med Phys 36:4359–4369
Levman J, Warner E, Causer P, Martel A (2014) Semi-automatic region-of-interest segmentation based computer-aided diagnosis of mass lesions from dynamic contrast-enhanced magnetic resonance imaging based breast cancer screening. J Digit Imaging 27:670–678
Zheng Y, Englander S, Baloch S et al (2009) STEP: spatiotemporal enhancement pattern for MR-based breast tumor diagnosis. Med Phys 36:3192–3204
Baltzer PAT, Dietzel M, Kaiser WA (2013) A simple and robust classification tree for differentiation between benign and malignant lesions in MR-mammography. Eur Radiol 23:2051–2060
Chen W, Giger ML, Lan L, Bick U (2004) Computerized interpretation of breast MRI: Investigation of enhancement-variance dynamics. Med Phys 31:1076–1082
Lucht RE, Knopp MV, Brix G (2001) Classification of signal-time curves from dynamic MR mammography by neural networks. Magn Reson Imaging 19:51–57
Levman J, Leung T, Causer P, Plewes D, Martel AL (2008) Classification of dynamic contrast-enhanced magnetic resonance breast lesions by support vector machines. IEEE Trans Med Imaging 27:688–696
Jansen SA, Fan X, Karczmar GS, Abe H, Schmidt RA, Newstead GM (2008) Differentiation between benign and malignant breast lesions detected by bilateral dynamic contrast-enhanced MRI: a sensitivity and specificity study. Magn Reson Med 59:747–754
Gallego-Ortiz C, Martel AL (2016) Improving the accuracy of computer-aided diagnosis for breast MR imaging by differentiating between mass and nonmass lesions. Radiology 278:679–688
Stoutjesdijk MJ, Veltman J, Huisman H et al (2007) Automated analysis of contrast enhancement in breast MRI lesions using mean shift clustering for ROI selection. J Magn Reson Imaging 26:606–614
Schlossbauer T, Leinsinger G, Wismuller A et al (2008) Classification of small contrast enhancing breast lesions in dynamic magnetic resonance imaging using a combination of morphological criteria and dynamic analysis based on unsupervised vector-quantization. Invest Radiol 43:56–64
Chen W, Giger ML, Bick U, Newstead GM (2006) Automatic identification and classification of characteristic kinetic curves of breast lesions on DCE-MRI. Med Phys 33:2878–2887
Agliozzo S, De Luca M, Bracco C et al (2012) Computer-aided diagnosis for dynamic contrast-enhanced breast MRI of mass-like lesions using a multiparametric model combining a selection of morphological, kinetic, and spatiotemporal features. Med Phys 39:1704–1715
Gilhuijs KG, Giger ML, Bick U (1998) Computerized analysis of breast lesions in three dimensions using dynamic magnetic-resonance imaging. Med Phys 25:1647–1654
Levman JE, Martel AL (2011) A margin sharpness measurement for the diagnosis of breast cancer from magnetic resonance imaging examinations. Acad Radiol 18:1577–1581
Haralick RM, Shanmugam K, Dinstein I (1973) Textural features for image classification. IEEE Trans Syst Man Cybern 6:610–621
Gibbs P, Turnbull LW (2003) Textural analysis of contrast-enhanced MR images of the breast. Magn Reson Med 50:92–98
Chen W, Giger ML, Li H, Bick U, Newstead GM (2007) Volumetric texture analysis of breast lesions on contrast-enhanced magnetic resonance images. Magn Reson Med 58:562–571
Ertaş G, Gülçür HO, Tunaci M (2007) Improved lesion detection in MR mammography: three-dimensional segmentation, moving voxel sampling, and normalized maximum intensity-time ratio entropy. Acad Radiol 14:151–161
Breiman L (2001) Random forests. Mach Learn 45:5–32
Gubern-Mérida A, Martí R, Melendez J et al (2015) Automated localization of breast cancer in DCE-MRI. Med Image Anal 20:265–274
Gallego-Ortiz C, Martel AL (2016) Interpreting extracted rules from ensemble of trees: application to computer-aided diagnosis of breast MRI. ICML workshop on human interpretability in machine learning (WHI 2016) arXiv:1606.08288. https://arxiv.org/abs/1606.08288. Accessed 30 Jun 2020
Chen W, Giger ML, Newstead GM, Bick U, Jansen SA, Li H, Lan L (2010) Computerized assessment of breast lesion malignancy using DCE-MRI robustness study on two independent clinical datasets from two manufacturers. Acad Radiol 17:822–829
Nie K, Chen J-H, Yu HJ, Chu Y, Nalcioglu O, Su M-Y (2008) Quantitative analysis of lesion morphology and texture features for diagnostic prediction in breast MRI. Acad Radiol 15:1513–1525
Rakoczy M, McGaughey D, Korenberg MJ, Levman J, Martel AL (2013) Feature selection in computer-aided breast cancer diagnosis via dynamic contrast-enhanced magnetic resonance images. J Digit Imaging 26:198–208
Mayer D, Vomweg TW, Faber H et al (2006) Fully automatic breast cancer diagnosis in contrast enhanced MRI. Int J CARS 1(Suppl 1):325–343
Renz DM, Böttcher J, Diekmann F et al (2012) Detection and classification of contrast-enhancing masses by a fully automatic computer-assisted diagnosis system for breast MRI. J Magn Reson Imaging 35:1077–1088
Vignati A, Giannini V, De Luca M et al (2011) Performance of a fully automatic lesion detection system for breast DCE-MRI. J Magn Reson Imaging 34:1341–1351
Huang YH, Chang YC, Huang CS, Chen JH, Chang RF (2014) Computerized breast mass detection using multi-scale Hessian-based analysis for dynamic contrastenhanced MRI. J Digit Imaging 27:649–660
Wu H, Gallego-Ortiz C, Martel A (2015) Deep artificial neural network approach to automated lesion segmentation in breast DCE-MRI. MICCAI-BIA 2015, Proceedings of the 3rd MICCAI workshop on breast image analysis, pp 73–80
Le QV (2013) Building high-level features using large scale unsupervised learning. 2013 IEEE international conference on acoustics, speech and signal processing: 8595–8598
Wu H (2016) Automatic computer aided diagnosis of breast cancer in dynamic contrast enhanced magnetic resonance images. Master’s thesis, University of Toronto. https://tspace.library.utoronto.ca/handle/1807/76158. Accessed 30 Jun 2020
Herrmann KH, Wurdinger S, Fischer DR et al (2007) Application and assessment of a robust elastic motion correction algorithm to dynamic MRI. Eur Radiol 17:259–264
Martel AL, Froh MS, Brock KK, Plewes DB, Barber DC (2007) Evaluating an optical-flow-based registration algorithm for contrast-enhanced magnetic resonance imaging of the breast. Phys Med Biol 52:3803–3816
Rueckert D, Sonoda LI, Hayes C, Hill DL, Leach MO, Hawkes DJ (1999) Nonrigid registration using free-form deformations: application to breast MR images. IEEE Trans Med Imaging 18:712–721
Rohlfing T, Maurer CR Jr, Bluemke DA, Jacobs MA (2003) Volume-preserving nonrigid registration of MR breast images using free-form deformation with an incompressibility constraint. IEEE Trans Med Imaging 22:730–741
Ebrahimi M, Martel AL (2009) A general PDE-framework for registration of contrast enhanced images. Med Image Comput Assist Interv 12:811–819
Schnabel JA, Tanner C, Castellano-Smith AD et al (2003) Validation of nonrigid image registration using finite-element methods: application to breast MR images. IEEE Trans Med Imaging 22:238–247
Mehrabian H, Richmond L, Lu Y, Martel AL (2018) Deformable registration for longitudinal breast MRI screening. J Digit Imaging 31(5):718–726
Nie K, Chen JH, Chan S et al (2008) Development of a quantitative method for analysis of breast density based on three-dimensional breast MRI. Med Phys 35:5253–5262
Martel AL, Gallego-Ortiz C, Lu Y (2016) Breast segmentation in MRI using Poisson surface reconstruction initialized with random forest edge detection. Proc. SPIE 9784, Medical Imaging 2016: Image Processing, 97841B. Accessed 27 August 2017
Ribes S, Didierlaurent D, Decoster N et al (2014) Automatic segmentation of breast MR images through a Markov random field statistical model. IEEE Trans Med Imaging 33:1986–1996
Dalmış MU, Litjens G, Holland K et al (2017) Using deep learning to segment breast and fibroglandular tissue in MRI volumes. Med Phys 44:533–546
Gubern-Mérida A, Kallenberg M, Mann RM, Marti R, Karssemeijer N (2015) Breast segmentation and density estimation in breast MRI: a fully automatic framework. IEEE J Biomed Heal Informatics 19:349–357
Fashandi H, Kuling G, Lu Y, Wu H, Martel AL (2019) An investigation of the effect of fat suppression and dimensionality on the accuracy of breast MRI segmentation using U-nets. Med Phys 46(3):1230–1244
Meinel LA, Stolpen AH, Berbaum KS, Fajardo LL, Reinhardt JM (2007) Breast MRI lesion classification: improved performance of human readers with a backpropagation neural network computer-aided diagnosis (CAD) system. J Magn Reson Imaging 25:89–95
Bhooshan N, Giger M, Lan L et al (2011) Combined use of T2-weighted MRI and T1-weighted dynamic contrast-enhanced MRI in the automated analysis of breast lesions. Magn Reson Med 66:555–564
Ballesio L, Savelli S, Angeletti M et al (2009) Breast MRI: Are T2 IR sequences useful in the evaluation of breast lesions? Eur J Radiol 71:96–101
Cai H, Liu L, Peng Y, Wu Y, Li L (2014) Diagnostic assessment by dynamic contrast-enhanced and diffusion-weighted magnetic resonance in differentiation of breast lesions under different imaging protocols. BMC Cancer 14:366
Platel B, Mus R, Welte T, Karssemeijer N, Mann R (2014) Automated characterization of breast lesions imaged with an ultrafast DCE-MR protocol. IEEE Trans Med Imaging 33:225–232
Abe H, Mori N, Tsuchiya K et al (2016) Kinetic analysis of benign and malignant breast lesions with ultrafast dynamic contrast-enhanced MRI: comparison with standard kinetic assessment. AJR Am J Roentgenol 207:1159–1166
Greenspan H, van Ginneken B, Summers RM (2016) Guest Editorial Deep Learning in Medical Imaging: Overview and future promise of an exciting new technique. IEEE Trans Med Imaging 35:1153–1159
Kooi T, Litjens G, van Ginneken B et al (2017) Large scale deep learning for computer aided detection of mammographic lesions. Med Image Anal 35:303–312
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Martel, A.L. (2020). CAD and Machine Learning for Breast MRI. In: Sardanelli, F., Podo, F. (eds) Breast MRI for High-risk Screening. Springer, Cham. https://doi.org/10.1007/978-3-030-41207-4_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-41207-4_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-41206-7
Online ISBN: 978-3-030-41207-4
eBook Packages: MedicineMedicine (R0)