Abstract
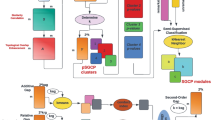
A semi-supervised gene co-expressed pattern finding method, PatGeneClus is presented in this paper. PatGeneClus attempts to find all possible biologically relevant gene coherent patterns from any microarray dataset by exploiting both gene expression similarity as well as GO-similarity. PatGeneClus uses a graph-based clustering algorithm called DClique to generate a set of clusters of high biological relevance. We establish the effectiveness of PatGeneClus over several benchmark datasets using well-known validity measures. The clusters obtained by PatGeneClus have been found to be biologically significant due to their high p-values, Q-values and clustalW scores.
The supplementary materials are available at http://agnigarh.tezu.ernet.in/~rosy8/shared.html
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lagreid, A., Hvidsten, T.R., Midelfart, H., Komorowski, J., Sandvik, A.K.: Predicting gene ontology biological process from temporal gene expression patterns. Genome Res. 13(5), 965–979 (2003)
Harris, M.A., et al.: The gene ontology (GO) database and informatics resource. Nucleic Acids Res. 32(Database issue), D258–D261 (2004)
Macintyre, G., Bailey, J., Gustafsson, D., Haviv, I., Kowalczyk, A.: Using gene ontology annotations in exploratory microarray clustering to understand cancer etiology. Pattern Recogn. Lett. 31(14), 2138–2146 (2010)
Verbanck, M., Le, S., Pages, J.: A new unsupervised gene clustering algorithm based on the integration of biological knowledge into expression data. BMC Bioinform. 14, 42 (2013)
Pesquita, C., Faria, D., Falcao, A.O., Lord, P., Couto, F.M.: Semantic similarity in biomedical ontologies. PLoS Comput. Biol. 5(7), e1000443 (2009)
Pesquita, C., Faria, D., Bastos, H., Falcao, A.O., Couto, F.: Evaluating GO based semantic similarity measures. In: ISMB/ECCB 2007 SIG Meeting Program Materials, International Society for Computational Biology (2007)
Gentleman, R.: Visualizing and distances using GO (2005)
Ovaska, K., Laakso, M., Hautaniemi, S.: Fast gene ontology based clustering for microarray experiments. Bio Data Min. 1(1) (2008)
Mandal, K., Sarmah, R.: A Density-Based Clustering for Gene Expression Data Using Gene Ontology, Lecture Notes in Networks and Systems (2018)
Brionne, A., Juanchich, A., Hennequet-Antier, C.: ViSEAGO: a Bioconductor package for clustering biological functions using gene ontology and semantic similarity. BioData Min. 12(1), 16 (2019)
Paul, A.K., Shill, P.C.: Incorporating gene ontology into fuzzy relational clustering of microarray gene expression data. Biosystems 163, 1–10 (2018)
Baishya, R.C., Sarmah, R., Bhattacharyya, D.K., Dutta, M.: A similarity measure for clustering gene expression data. In: Proceedings of International Conference on Applied Algorithms, Kolkata, India, pp. 245–256 (2014)
Cho, R.J., Campbell, M.J., Winzeler, E.A., Steinmetz, L.: A genome-wide transcriptional analysis of the mitotic cell cycle. Mol. Cell 2(1), 65–73 (1998)
DeRisi, J.L., Iyer, V.R., Brown, P.O.: Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686 (1997)
Nymark, P., Lindholm, P.M., Korpela, M.V., Lahti, L., Ruosaari, S., Kaski, S., Hollmen, J., Anttila, S., Kinnula, V.L., Knuutila, S.: Gene expression profiles in asbestos-exposed epithelial and mesothelial lung cell lines. BMC Genom. 8, 62 (2007)
Berriz, F.G., et al.: Characterizing gene sets with funcassociate. Bioinformatics 19, 2502–2504 (2003)
Warde-Farley, D., Donaldson, S.L., Comes, O., Zuberi, K., Badrawi, R., Chao, P., Franz, M., Grouios, C., Kazi, F., Lopes, C.T., Maitland, A., Mostafavi, S., Montojo, J., Shao, Q., Wright, G., Bader, G.D., Morris, Q.: The geneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 38, W214–W220 (2010)
Larkin, M.A., Blackshields, G., Brown, N.P., Chenna, R., McGettigan, H., McWilliam, H., Valentin, F., Wallace, I.M., Wilm, A., Lopez, R., Thompson, J.D., Gibson, T.J., Higgins, D.G.: Clustalw and clustalx version 2. Bioinformatics 23(21), 2947–2948 (2007)
Cheng, Y., Church, G.M.: Biclustering of expression data. In: Proceedings of ISMB 2000, pp. 93–103. AAAI Press (2000)
Sharan, R., Shamir, R.: Click: a clustering algorithm with applications to gene expression analysis. In: Proceedings of 8th International Conference on Intelligent Systems for Molecular Biology. AAAI Press (2000)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Baishya, R.C., Sarmah, R., Bhattacharyya, D.K. (2020). Improving Co-expressed Gene Pattern Finding Using Gene Ontology. In: Dehuri, S., Mishra, B., Mallick, P., Cho, SB., Favorskaya, M. (eds) Biologically Inspired Techniques in Many-Criteria Decision Making. BITMDM 2019. Learning and Analytics in Intelligent Systems, vol 10. Springer, Cham. https://doi.org/10.1007/978-3-030-39033-4_20
Download citation
DOI: https://doi.org/10.1007/978-3-030-39033-4_20
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-39032-7
Online ISBN: 978-3-030-39033-4
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)