Abstract
Class XVIII myosins represent a branch of the myosin family tree characterized by the presence of large N- and C-terminal extensions flanking a generic myosin core. These myosins display the highest sequence similarity to conventional class II muscle myosins and are compatible with but not restricted to myosin-2 contractile structures. Instead, they fulfill their functions at diverse localities, such as lamella, actomyosin bundles, the Golgi apparatus, focal adhesions, the cell membrane, and within sarcomeres. Sequence comparison of active-site residues and biochemical data available thus far indicate that this myosin class lacks active ATPase-driven motor activity, suggesting that its members function as structural myosins. An emerging body of evidence indicates that this structural capability is essential for the organization, maturation, and regulation of the contractile machinery in both muscle and nonmuscle cells. This is supported by the clear association of myosin-18A (Myo18A) and myosin-18B (Myo18B) dysregulation with diseases such as cancer and various myopathies.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Myosin-18
- Golgi
- GOLPH3
- Transport
- Surfactant protein A receptor
- Cancer
- Tumor suppressor
- Sarcomere
- Myofibrillogenesis
19.1 Introduction
The class XVIII myosin family encompasses two distinct myosins known as Myo18A and Myo18B, which are synthesized from two different genes. These are amongst the most recently identified myosins with the mouse Myo18A gene discovered as recently as the year 2000 through a gene expression screen of bone marrow stromal cells (Furusawa et al. 2000). Originally named MysPDZ due to the presence of a PDZ domain within its unique ~400 amino acid N-terminal extension, this protein was later renamed Myo18A following the identification of other PDZ-myosins in vertebrates and Drosophila. Together, these myosins constituted the new myosin XVIII class (Yamashita et al. 2000; Berg et al. 2001). The second family member, Myo18B, was first reported in 2001 following partial annotation of the human MYO18B gene (Berg et al. 2001). Phylogenetic analysis suggests that MYO18B, which is exclusively found in vertebrates, is likely derived from duplication of the MYO18A gene, which is found in both invertebrates and vertebrates (Salamon et al. 2003). At the protein level, Myo18A and Myo18B share ~40% identity and are distinguished from other classes by their large N-terminal and C-terminal extensions, which flank either side of a generic myosin core structure. While the literature demonstrates unique roles for these two proteins at specific subcellular localities, it remains unclear whether they have distinct or overlapping functions at sites where they both localize, such as stress fibers in nonmuscle cells and sarcomeric structures in striated muscle.
19.2 Myosin-18A
19.2.1 MYO18A Gene Architecture and RNA Expression Across Cell Lines and Tissues
Myo18A, located on mouse chromosome 11 and arm q11.2 of chromosome 17 in humans (MYO18A), was first discovered in 2000 in a gene expression screen of bone marrow stromal cells, where it was found to be upregulated in cell lines with increased hematopoietic supportive ability (Furusawa et al. 2000). In mouse, the ORF encodes for two splice isoforms that produce long (Myo18Aα) and short (Myo18Aβ) versions of the protein, which consist of 2035 amino acids (230 kDa) and 1719 amino acids (196 kDa), respectively. The human homologues display similar dimensions with Myo18Aα comprising 2054 amino acids (~233 kDa) and Myo18Aβ comprising 1723 amino acids (~196 kDa) (Fig. 19.1). The central part of Myo18Aα (residues 405–1972) exhibits homology to generic myosins and displays the highest similarity with conventional muscle myosin-2, as both have extended coiled-coil regions in the C-terminal half of the protein. Further examination of the domain structure and SMART-based sequence analysis reveals the existence of a region rich in lysine and glutamate residues (KE motif) and a PDZ domain inside the long N-terminal extension (Fig. 19.1). Whilst a few other myosin classes described thus far also contain additional domains at the N- or C-terminus (e.g., myosin class III, IX, XV, and XVI (Odronitz and Kollmar 2007)), the presence of the PDZ domain is unique and prompted Furusawa et al. to give this novel myosin the name MysPDZ, which denoted “Myosin containing a PDZ domain” (Furusawa et al. 2000). An independent study also identified Myo18A in a human peripheral blood cDNA library during a screen for novel interaction partners of the cytosolic tyrosine kinase JAK3. Accordingly, this protein was designated as MAJN (molecule associated with Jak3 N-terminal) (Ji et al. 2000).
Schematic representation of the domain structure of human (Hs, Homo sapiens) and mouse (Mm, Mus musculus) Myosin-18A splice isoforms. These construct diagrams illustrate the large N-terminal and C-terminal extensions characteristic of class XVIII myosins, which flank the generic myosin core containing a motor domain, neck region, and coiled-coil tail domain. Specifically for Myo18A, splice isoforms are distinguished by the presence of a KE-rich region, PDZ domain, or PPII (polyproline II helix) domain within the N-terminal extension. Like myosin-2 motors, this protein forms homotypic dimers via the tail domain. The precise structure and organization of the N-terminal and C-terminal extensions remain to be resolved
When mRNA expression levels were examined, ubiquitous expression of Myo18A was detected in all tissues and cell lines tested (Furusawa et al. 2000). However, a longer transcript with a size of 10 kb was enriched in heart and skeletal muscle. The normal 7.5-kb Myo18Aα transcript was found in most tissues (brain, thymus, lung, liver, etc.) and a shorter transcript of 7 kb, later identified as Myo18Aβ, was observed in hematopoietic cell lines (Furusawa et al. 2000; Mori et al. 2003). The long transcript has not been explicitly studied further; however, the larger size and muscle expression profile indicate that it could in fact be the recently identified Myo18Aγ splice isoform (Horsthemke et al. 2019). In addition to these splice isoforms, which have been verified at both the RNA and protein levels, a short version of Myo18A has also been identified in a macrophage cell line, where it is tyrosine-phosphorylated after CSF-1 stimulation (Cross et al. 2004). Intriguingly, this protein had a molecular weight of only 110 kDa (p110 myosin 18A), and peptide analysis suggests that the protein boundaries span from the middle of the motor domain to the end of the coiled-coil region. The lack of evidence for an alternative translational start within the Myo18A mRNA suggests that this protein results from post-translational cleavage of one of the higher molecular mass isoforms.
19.2.2 Myo18A Biochemistry: Protein Domain Architecture and Functional Properties
Myo18A has a complex domain architecture that comprises a number of different elements with various functions. In the Myo18Aα splice isoform, the N-terminal extension has thus far been shown to contain two distinct functional regions: a KE motif (containing 11 lysine and 3 glutamate residues within a stretch of twenty amino acids) located at the far N-terminus and a PDZ domain that is found in the middle of the N-terminal extension (Fig. 19.1) (Furusawa et al. 2000). Myo18Aγ comprises a polyproline II helix (PPII) of yet unknown function in its N-terminal extension whereas Myo18Aβ completely lacks this domain. The molecular function of the Myo18Aα N-terminal extension has been investigated in two studies to date. Isogawa et al. utilized GFP-tagged deletion constructs of human Myo18A to characterize actin-binding properties in lysates of HeLa cells that overexpress the respective proteins (Isogawa et al. 2005). The N-terminal extension construct displayed a strong, ATP-independent interaction with actin in cosedimentation assays, and this result was validated with bacterially-expressed deletion constructs. Analysis of a selection of purified truncation and mutation constructs in these assays localized the actin-binding interface to the middle of the N-terminal extension, outside of the KE motif and PDZ domain. Through a series of cosedimentation affinity assays, it was shown that a recombinant human Myo18A protein construct spanning from the N-terminus to the start of the PDZ domain weakly interacted with actin (KA of 6.5 μM), whilst a construct encompassing the whole N-terminal extension had a six-fold enhanced actin affinity with a KA of 1 μM (Taft et al. 2013). Actin affinity is further enhanced five-fold by the direct binding to GOLPH3, a phosphoprotein of the Golgi membrane (cf. 19.2.3.). These data demonstrate that the N-terminal extension combines at least two functional modules that mediate actin binding, are dependent from one another, and can be regulated by binding of an interaction partner, GOLPH3.
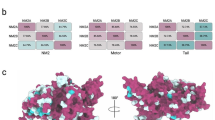
The adjacent structural element in the domain architecture of Myo18A is its motor domain. A fundamental feature of the myosin family is the inherent ability of their motor domains to hydrolyze ATP and to convert the resulting energy into force and directed movement along actin filaments (Geeves et al. 2005; Bloemink and Geeves 2011). The basic structural elements in the motor domain known to be essential for the enzymatic activity of myosins are the P-loop, switch-1, and switch-2 (Preller and Manstein 2013; Geeves 2016). Sequence alignments with other myosin motor domains show that these three aforementioned motifs are modified in Myo18A (Fig. 19.2). Most strikingly, two highly conserved serine residues in switch-1, which usually coordinate the γ-phosphate of ATP and Mg2+ for efficient catalysis, are mutated to alanine and threonine. In addition, the salt bridge between an arginine in switch-1 and a glutamate in switch-2 cannot be formed due to a glutamate-to-glutamine exchange in Myo18A. As the salt bridge has been shown to be essential for ATPase activity and motor function of myosins (Onishi et al. 1998; Furch et al. 1999), the sequence variations found in Myo18A raise the question of whether this myosin can function as an active motor. To date, three studies have addressed this question, none of which could demonstrate motor function or ATPase activity for class XVIII myosins. The first study by Guzik-Lendrum et al. assessed the biochemical properties of the sole Drosophila melanogaster (Dm) class XVIII myosin (Myo18), which has the highest homology to mammalian Myo18A (Guzik-Lendrum et al. 2011). They purified motor domain constructs with and without the PDZ domain using the baculovirus/Sf9 insect cell system and found that neither of the constructs exhibits ATPase activity or can even bind ATP. Actin-binding affinity of the Myo18 motor domain was demonstrated and quantified with cosedimentation assays (KD ~1 μM) and optical trap studies showed the existence of short-lived actin interactions. The functional properties of mouse Myo18A were subsequently studied by the same group with purified S1 constructs, which display weak ATP and ADP affinities (~80 and ~100 μM) (Guzik-Lendrum et al. 2013). Furthermore, the long (Myo18Aα) and short (Myo18Aβ) isoforms exhibit actin-binding affinities of ~5 and ~50 μM, and no regulation of the actin interaction by nucleotide binding or light chain phosphorylation was found. In line with the previous study, no ATPase activity or motor function was detected for mouse Myo18A, and negative-stain electron microscopy revealed no structural changes of the S1 conformation associated with changes in the nucleotide state (ATP, ADP, or rigor). The third study sought to understand the molecular function of human Myo18A and its interaction with nucleotides, actin, and GOLPH3 (Taft et al. 2013). Nucleotide and actin affinities of motor domain constructs produced with the baculovirus/Sf9 insect cell system were found to be 10–100 times higher for the human protein compared to its mouse homologue. In line with the mouse protein, human Myo18A does not exhibit ATPase activity. Intriguingly, in the ADP state, the human Myo18A motor domain displays the highest actin affinity and the most complete binding to actin. Negative-stain electron microscopy revealed the typical arrowhead pattern of myosin bound to actin. In contrast to generic myosins, the decoration was not only complete and well-ordered in the absence of nucleotide and in the presence of ADP but also in the presence of excess ATP.
Alignment of Hs Myosin-18A and Hs Myosin-18B P-loop, switch-1, and switch-2 sequences with Dictyostelium discoideum (Dd) myosin-2, Hs skeletal muscle myosin-2, and Hs β-cardiac muscle myosin-2. Conserved sequence motifs are defined in dark grey regions, while deviations in Myo18A and Myo18A are shown in bold
The myosin motor domain adjoins to the light chain-binding domain, which binds the essential light chain (ELC) and regulatory light chain (RLC) of conventional class II muscle myosins. In the case of unconventional myosins, which contain one or more IQ motifs, this region mediates the interaction with calmodulin (CaM) or CaM-like light chains (Odronitz and Kollmar 2007). Initial examination of the Myo18A domain structure suggested the presence of one IQ motif following the motor domain (Furusawa et al. 2000; Mori et al. 2003). However, subsequent analysis proposed the existence of two IQ motifs based on comprehensive alignments of Myo18A sequences across subclasses and phyla with myosin-2 (Guzik-Lendrum et al. 2013). This was experimentally confirmed by the co-purification of S1 and HMM mouse Myo18A constructs along with both ELC and RLC. Whilst it had already previously been shown that overexpressed Myo18A binds to and co-purifies with RLC in COS7 cells (Tan et al. 2008), the ability of Myo18A to bind both ELC and RLC of the nonmuscle myosin-2 (NM2) system suggests a role for Myo18A in NM2-dependent cellular structures and processes.
The C-terminal end of the light chain-binding domain leads over to the “bend” motif (WPWW/WQWW/WPWM), which is followed by a highly conserved tryptophan residue located eight residues further, where the coiled-coil domain of class II myosins starts. According to sequence analyses, the coiled-coil domain in human Myo18A encompasses amino acids 1253-1966 (Fig. 19.1). This strongly suggests that the protein exists and functions inside cells as a dimer of two heavy chains, which has been confirmed for human and mouse Myo18A by chemical crosslinking of HMM constructs and by co-immunoprecipitation (Mori et al. 2005; Isogawa et al. 2005), and for mouse Myo18A HMM by negative-stain electron microscopy (Guzik-Lendrum et al. 2013). When purified untruncated mouse Myo18Aβ protein was subjected to electron microscopy, the formation of unusual short and condensed antiparallel dimers was observed (Billington et al. 2015). In addition, Myo18Aβ has been shown to co-assemble with NM2A to form mixed bipolar filaments, where its incorporation reduces the average filament length (from 310 to 220 nm).
So far, the C-terminal extension is the least-characterized domain of Myo18A. Extending from residue 1972 to the C-terminus, this domain of 83 amino acids is predicted not to form coiled-coil structures. It appears reasonable that this domain provides Myo18A with additional binding motifs for protein-protein interactions. In line with this, it has been shown that the C-terminal extension of Myo18A directly interacts with the C-terminal last 8 residues of the Rho GTPase activator βPIX (PAK-interacting exchange factor-β) (cf. 19.2.4) (Hsu et al. 2010). No additional direct binding partner of the C-terminal extension has been described yet.
19.2.3 The Role of Myo18A at the Golgi
The Golgi apparatus is a membranous system essential for the intracellular sorting and transport of proteins and is known to connect to both the microtubule and the actin cytoskeletons (reviewed in (Gurel et al. 2014)). The phospholipid phosphatidylinositol-4-phosphate (PI4P) is enriched in Golgi membranes and directly binds to a few proteins containing pleckstrin homology (PH) domains (Dowler et al. 2000; Levine and Munro 2002). An in vitro lipid-binding screen identified the Golgi phosphoprotein GOLPH3 as a direct binding partner for PI4P, which was subsequently shown to localize to the Golgi membrane (Dippold et al. 2009). It was observed that latrunculin B-induced actin depolymerization and GOLPH3 knockdown both result in rapid Golgi compaction. In search of an interaction partner that can mediate the connection between GOLPH3 and the actin cytoskeleton, immunoprecipitation studies using HeLa cell lysates and mass spectrometry identified a ~250-kDa putative interaction partner to be Myo18A. Whilst a direct interaction between Myo18A and GOLPH3 was validated by reciprocal co-immunoprecipitation of both proteins from cell lysates by specific antibodies, other Golgi-localized myosins (NM2B, myosin-6) were shown not to interact with GOLPH3 (Dippold et al. 2009). Further mapping the interaction region, pulldown experiments with bacterially-expressed domains of Myo18A showed direct binding between GOLPH3 and the N-terminal extension and motor/IQ domains of Myo18A. As both domains interact with GOLPH3 independently of each other, bipartite binding can be assumed. It should be noted that the expression of myosin motor domains in bacteria does not produce functional protein as a certain setup of chaperones is needed for proper folding (McNally et al. 1991). In cellular studies utilizing siRNA-mediated knockdown of GOLPH3, Myo18A localization to the Golgi was abolished. Further, a functional Myo18A/GOLPH3 complex is needed for normal trans-Golgi appearance and efficient tubule and vesicle budding, as well as directed vesicle trafficking away from the Golgi ribbon (Dippold et al. 2009). To understand whether active force generation by Myo18A is required for its cellular function, a Myo18A construct with ATP-binding pocket mutations (G520S/K521A in the P-loop) was utilized to rescue the Myo18A-knockdown phenotype of condensed Golgi. The fact that this rescue failed prompted the authors to suggest that force generation via the Myo18A motor is needed to stretch and shape the Golgi. This interpretation is additionally backed up by a study by Tan et al., where overexpression of a similar ATP-binding pocket mutation (G520D/K521E) in Myo18A generates a dominant-negative phenotype similar to the depletion of Myo18A (cf. 19.2.4) (Tan et al. 2008). However, the fact that Myo18A function can be altered by mutations in the active site is in contrast with the aforementioned biochemical studies that could not detect any ATPase or motor function for Myo18A from different species (cf. 19.2.2). There are at least three possible explanations for this obvious contradiction. Firstly, it is possible that inside the cell, Myo18A actually functions as a molecular motor either activated by thus far unidentified binding partners or by other mechanisms such as posttranslational modifications, the binding of metabolites, or the performance of specific chaperones. Secondly, the mutations that have been introduced in the center of the Myo18A globular motor domain could impair its proper folding and as such, lead to decreased protein stability and the observed phenotypes. Thirdly, the mutations may not deactivate motor function but rather interfere with the proper discrimination of the nucleotide state, which is shown to regulate the actin-binding properties of human Myo18A (Taft et al. 2013). Additionally, this could also affect the functional competence of mixed bipolar filaments of Myo18A and NM2A, which have been detected inside living cells (Billington et al. 2015), where NM2A provides the motor activity for the mixed myosin filaments.
Independent of the exact mechanism of how Myo18A performs its function at the Golgi, further studies by the same group support its interaction with GOLPH3 and its participation in maintaining Golgi morphology and function. In secretory tissues, a GOLPH3 paralogue, GOLPH3L, was identified that antagonizes GOLPH3 function as it binds to the Golgi membrane via PI4P interaction and supports anterograde trafficking without binding to Myo18A (Ng et al. 2013). Accordingly, GOLPH3L blocks PI4P for GOLPH3 binding and thereby suppresses Myo18A-mediated anchoring of the Golgi to the actin cytoskeleton. In a subsequent study it was shown that the nuclear DNA-PK protein kinase is activated by doxorubicin-induced DNA damage to phosphorylate GOLPH3, which leads to an increased interaction with Myo18A (Farber-Katz et al. 2014). The enhanced interaction impairs vesicle trafficking from the Golgi membrane to the plasma membrane and thus induces fragmentation of the Golgi as a result of DNA damage. It has previously been described that GOLPH3 is an oncogene that is frequently overexpressed and amplified in human cancers (Scott et al. 2009; Buschman et al. 2015; Buschman and Field 2018). GOLPH3 acts as an oncogenic factor by increasing cell migration. Overexpression of GOLPH3 and siRNA-mediated knockdown of both GOLPH3 and Myo18A show that the entire PI4P/GOLPH3/Myo18A/F-actin complex is necessary and sufficient to promote Golgi reorientation towards the leading edge via the actin cytoskeleton and as such is able to enhance directional cell migration (Xing et al. 2016).
Although a plethora of experimental evidence speaks in favor of a direct role of the Myo18A/GOLPH3 complex in the maintenance of Golgi morphology and function, a recent study provides data that question the previous results (Bruun et al. 2017). Here, Hammer and coworkers used immunofluorescence studies with specific antibodies and overexpression of GFP-tagged proteins and found no proof of Myo18A localization to the Golgi or an impact of reduced Myo18A levels on Golgi morphology. As their study focuses on Myo18Aα, the authors consider that unknown splice variants of Myo18A may facilitate the Golgi-shaping via GOLPH3. However, based on their experimental results, they assume that global F-actin disassembly is the major reason for the observed Golgi morphology.
19.2.4 Myo18A Has Multiple Roles in Focal Adhesions, Stress Fibers and Lamellipodia
Myo18A has biochemically been defined as an actin-binding protein that comprises four actin-binding sites per heavy chain dimer (Taft et al. 2013). On the cellular level, a few studies have analyzed its localization and role in the context of different actin structures, specifically focal adhesions, actin stress fibers and lamellar actomyosin bundles. Characterization of the subcellular distribution of overexpressed EYFP-tagged Myo18A protein and deletion constructs in fibroblasts revealed that the KE motif mediates actin fiber co-distribution, whilst the PDZ domain controls the localization of Myo18A to the inner surface of the cell membrane (Mori et al. 2005). FRAP experiments revealed random and rapid short-range movement of punctate structures in the cytoplasm. The cellular function of Myo18A has also been assessed in the context of lamellar actomyosin bundles (Tan et al. 2008). Here, a tripartite complex of Myo18A with the myosin light chain kinase MRCK (myotonic dystrophy-related Cdc42-binding kinase) and the leucine-rich adapter protein LRAP35a was shown to co-localize with lamellar actomyosin filaments. In this protein complex, LRAP35a binds and activates MRCK and simultaneously binds to Myo18A, which directs the complex to actomyosin bundles. The activated MRCK subsequently phosphorylates the RLC, which is bound to Myo18A. In parallel, MRCK also phosphorylates NM2A and thereby initiates actomyosin assembly in the lamella to enable cell protrusion and cell migration. The results of this study are in line with the findings of Billington et al., who applied TIRF-SIM microscopy of fluorescently-tagged Myo18Aα and Myo18Aβ in combination with NM2A and found that these myosins co-assemble in living cells (Billington et al. 2015). These results were further confirmed with antibodies specific for Myo18A and NM2A. Thus, Myo18A can incorporate into NM2 filaments and might regulate the assembly and functional properties of structures like actomyosin bundles and stress fibers. In addition, a direct interaction of Myo18A with the PAK2/βPIX/GIT1 (p21-activated kinase 2/PAK-interacting exchange factor-β/G protein-coupled receptor kinase-interactor 1) complex was established in a proteomics approach that sought to identify new proteins that interact with PAK2 in human epidermoid carcinoma cells (Hsu et al. 2010). The interaction of Myo18A with PAK2 is indirect and mediated via βPIX/GIT1 to direct the whole complex to lamellipodia and membrane ruffles. Myo18A-depleted cells display decreased cell migration, an enlarged morphology, and also increased actin stress fiber density and focal adhesion formation. It appears that Myo18A is essential for lamellipodia targeting of the complex, as Myo18A depletion and disruption of its interaction with βPIX both induce accumulation of the complex at focal adhesions and decrease cell migration (Hsu et al. 2010, 2014).
19.2.5 Membrane Proteins Identified as Myo18A
A striking feature of Myo18A is its implication in binding of surfactant protein A (SPA) at the cell surface. Chroneos and coworkers identified Myo18A as SPA-receptor 210 (SP-R210) using mass spectrometry and antibodies specific for the C-terminus of Myo18A (Yang et al. 2005). They showed that Myo18A overexpression in COS-1 cells enables SPA binding and that SP-R210-directed antibodies detect bacterially-expressed Myo18A domains that correspond to the region encompassing the IQ motifs and the first half of the coiled-coil domain. Intriguingly, these antibodies block the binding of SPA to macrophages (Yang et al. 2005) as well as T-cell-mediated immune responses (Samten et al. 2008). Further, the stable expression of the C-terminal domain of Myo18A in macrophages is also shown to induce a dominant-negative inhibition of SP-R210 protein expression (Szeliga et al. 2005; Yang et al. 2015). In addition, a single study by a different group applied mass spectrometry supported by immunoblotting with a Myo18A-specific antibody and identified CD245, a surface antigen of peripheral blood lymphocytes, as Myo18A (De Masson et al. 2016). In line with the aforementioned studies, both the long and the short isoforms (230 and 190 kDa) were detected, and the authors suggest a mechanism for Myo18A recruitment-mediated activation of NK cell cytotoxicity. Although the results of these studies were substantiated with numerous independent experiments, these findings raise a number of obvious questions. Myo18A does not contain transmembrane domains and has exclusively been detected in the cytosolic compartment of cells in all other studies. Further, it is completely unclear how Myo18A is externalized to the cell surface as a Golgi/ER-mediated process should have been detected in other studies. Future research may shed light on this unique role for Myo18A and unravel the underlying cellular and functional mechanisms.
19.2.6 Participation of Myo18A in Muscle Development and Integrity
A central feature of the other class XVIII myosin, Myo18B, is its implication in the process of striated muscle maturation (cf. 19.3.2). Initial evidence indicating that Myo18A likewise contributes to muscle development stems from studies in Drosophila and zebrafish. The sole class XVIII myosin in Drosophila, Myo18, was shown to localize to cell-cell contacts in myocytes during myoblast fusion (Bonn et al. 2013). The protein Rolling Pebbles 7, which is an integral part of a protein complex at myoblast fusion pores, was identified as a direct interaction partner essential for Myo18 localization to these sites. A possible role of Myo18 at these sites is to organize the actin cytoskeleton to widen the fusion pore. As deficiency of Myo18 does not impact muscle development, it can be speculated that other myosins have redundant functions in Drosophila. In mature muscles, Drosophila Myo18 localizes adjacent to the Z-disc but does not directly co-localize with Z-disc markers, implying an additional role not connected to myoblast development. Two studies shed light on the participation of Myo18A in myofiber integrity in zebrafish. It was found that both Myo18Aα and Myo18Aβ are expressed in somite borders during zebrafish development (Cao et al. 2014). Knockdown of Myo18A or overexpression of the PDZ module was shown to disrupt myofiber integrity. Accordingly, it was suggested that Myo18A mediates adhesion of myofibers to the cell membrane, permitting their stable attachment to the extracellular matrix. In a follow-up study by the same group, yeast two-hybrid screening and co-immunoprecipitation identified p190Rho-guanine nucleotide exchange factor (p190RhoGEF) and Golgin45 as novel binding partners for the PDZ module of Myo18A during zebrafish development (Cao et al. 2016). These experiments also confirmed the direct interaction of Myo18A with LURAP (LRAP35A, (Tan et al. 2008)). Knockdown of Myo18A alone or in combination with the aforementioned three binding partners severely disrupts myofiber integrity during development supposedly via a combined effect on the localization of dystrophin via a Myo18A-mediated impact on the actin cytoskeleton and/or via Myo18A-mediated support of Golgi morphology and trafficking (Cao et al. 2016).
A recent study identified a new class XVIII myosin isoform in mouse myocytes and human heart, termed Myo18Aγ (Horsthemke et al. 2019). Myo18A-directed antibodies were used to define the Myo18A isoform expression pattern in mouse ventricular myocytes and detected a protein of a molecular mass of ~267 kDa, which is not compatible with either Myo18Aα or Myo18Aβ. RNA-Seq confirmed that this protein constitutes a novel isoform of Myo18A, which has not been described previously. This isoform comprises an alternate N-terminal extension, as well as an extended C-terminus (Fig. 19.1). Overexpression of EYFP-tagged Myo18Aγ led to its accumulation within the A-band of sarcomeres in mouse ventricular myocytes whilst Myo18A-knockout mice displayed disorganized sarcomeres.
19.2.7 Disease Implications of Myo18A in Cancer and Virus Transport
In terms of disease, Myo18A has predominantly been investigated in cancer, where it is described to be part of fusion genes. However, it is also proposed to play a role in virus assembly and secretion.
An exon array analysis study identified alternative splicing of MYO18A in non-small cell lung cancer, which caused in-frame variations in the protein sequence (Langer et al. 2010). Further, the MYO18A gene was identified as a partner in the three-way chromosomal translocation of stem cell leukemia-lymphoma syndrome (Walz et al. 2005). Here, exon 32 of MYO18A is fused with exon 9 of FGFR1 (fibroblast growth factor receptor 1), which creates a constitutively-active tyrosine kinase that stimulates cell growth and proliferation. This leads to thrombocytopenia with decreased number and size of megakaryocytes and increased monocyte, eosinophil, and basophil cell numbers, establishing an unusual myelodysplastic/myeloproliferative disease (Walz et al. 2005). An additional fusion gene, MYO18A-PDGFRB (platelet-derived growth factor receptor beta), was detected by long-distance inverse PCR in eosinophilia-associated atypical myeloproliferative neoplasms (Walz et al. 2009). Fusion of exon 40 of MYO18A with exon 10 of PDGFRB creates a large protein (2661 aa) that retains most of Myo18A (98%) and contains the C-terminal 59% of PDGFRB. The proposed activation mechanism is based on the dimerization or oligomerization of the PDGFRB part by the Myo18A coiled-coil domain. In addition, a three-way translocation of the highly promiscuous oncogene MLL, a histone methyltransferase, and the reciprocal partner gene MYO18A was described in acute myeloid leukemia (Ussowicz et al. 2012). As the fusion gene is in frame, it translates into a functional fusion protein that encompasses the N-terminal 362 amino acids of Myo18A and the C-terminal 2607 amino acids (of 3969) of MLL, producing a chimeric protein that preserves methyltransferase activity and contains the N-terminal PDZ domain of Myo18A. However, the consequence of this fusion on the localization, regulation, or activity of the resulting protein is not yet known. An additional cancer-related role for Myo18A, not linked to gene fusion events, was proposed for its abundance in prostate cancer cell lines (Peckham 2016). Here, increased levels of four unconventional myosins, Myo1b, Myo9b, Myo10, and Myo18A, were reported (Makowska et al. 2015). Conversely, knockdown of Myo18A results in re-organization of long NM2A-rich stress fibers. This is in line with the previously reported ability of Myo18A to form mixed filaments with NM2A (Billington et al. 2015). The re-organized stress fibers, depleted of Myo18A, locate along the length of prostate cancer cells, which results in reduced directionality persistence in cell migration (Makowska et al. 2015). These findings suggest a contribution of increased Myo18A levels to prostate cancer metastasis.
Using a systems biology approach to follow the fate of organelles during human cytomegalovirus (HCMV) infection, Myo18A was found to localize to the plasma membrane in primary human fibroblasts (Jean Beltran et al. 2016). Following viral infection, Myo18A is translocated to the viral assembly complex and supports virus production by an as of yet unknown mechanism. The authors speculate that a Myo18A-mediated tethering of virus-loaded vesicles to NM2-containing myosin filaments provides the molecular basis for vesicle movement. A role for Myo18A in the virus secretion process was described for the hepatitis C virus (Bishé et al. 2012). Silencing of GOLPH3 or Myo18A inhibits hepatitis C virus secretion and leads to virion accumulation inside the cell. These findings are in line with a role of Myo18A in Golgi budding via GOLPH3 interaction (Dippold et al. 2009) and in virus secretion pathways (Jean Beltran et al. 2016).
19.3 Myosin-18B
19.3.1 Molecular and Cell Biology
The existence of a second class XVIII myosin family member, Myo18B, was first reported by Berg et al. in 2001 following partial annotation of the human MYO18B gene (Berg et al. 2001). MYO18B maps on human chromosome 22q12.1 between the GRK3 and SEZ6L genes, in an area of ~322 kb (25,742,133 – 26,063,847) and produces an mRNA transcript of 8577 bp from 43 coding exons and a 5′ non-coding exon (NCBI, NM_032608.6). The 7704 bp ORF encodes for a 2567-amino acid protein with a molecular mass of ~285 kDa that shares ~90% identity with the ~288 kDa homologue produced in mouse (Nishioka et al. 2002; Salamon et al. 2003). According to the NCBI database, an additional 7707-bp mRNA transcript has been curated for the human MYO18B gene, which lacks the 5′ non-coding region and includes three nucleotides that encode an additional glutamine residue at position 1020 within the protein sequence [NCBI, NM_001318245.1]. However, no data are currently available to validate or differentiate the occurrence or function of these two proteins.
Northern blot and PCR analyses have demonstrated that unlike Myo18A, which is ubiquitously expressed in most mammalian cells and tissues, Myo18B transcript expression is highly tissue specific. The highest level of expression is observed in cardiac and skeletal striated muscles, although Myo18B contributes as little as 0.005% of the total mRNA content in skeletal muscle (Nishioka et al. 2002; Salamon et al. 2003; Ajima et al. 2008). In addition, Myo18B mRNA is also found at low levels in other organs and tissues, including the testis, bone marrow, thymus, mammary glands, lungs (both fibroblastic and epithelial cell types), kidney, pancreas, placenta, prostate and brain. Whilst BLAST analyses have indicated that three potential splice isoforms exist in murine and human tissues, there is currently no experimental evidence demonstrating that MYO18B is susceptible to alternative mRNA splicing (Salamon et al. 2003).
Myo18B comprises a central region with a generic myosin configuration (Fig. 19.3). Within this core structure is a 767-amino acid motor domain (residues 565-1332), which encompasses both nucleotide- and actin-binding sites. Sequence analysis reveals that much like Myo18A, with which it shares ~40% protein identity, Myo18B also harbors amino acid substitutions in evolutionarily-conserved residues within the P-loop, switch-1, and switch-2; the regions essential for generating ATP-dependent force and movement along actin filaments (Fig. 19.2). Whilst these alignments indicate that Myo18B lacks motor ATPase activity, the precise function of the motor domain remains to be deciphered through biochemical analyses. Following the motor domain is a 69-amino acid neck region (residues 1333-1402), which contains two myosin light chain-binding IQ motifs stretching from amino acids 1343-1353 (FQAACKGFLSR) and 1369-1379 (IQKNVAVFLAV). At the present time, there is no information available regarding the interaction of Myo18B with either ELC or RLC. The generic region is completed with a coiled-coil tail domain between residues 1403-2085, which is essential in other myosins, including Myo18A, for heavy chain dimerization. Exhaustive domain analyses predict that this tail region also incorporates an ERM (ezrin-radixin-moesin) domain for plasma membrane interactions (residues 1458-1576), an AAA domain for association with chaperone-like ATPase proteins (residues 1434-1917 amino acids), and a putative A-motif for nucleotide binding at position 1818-1825 (Salamon et al. 2003). However, the functional capabilities of each of these proposed domains remains to be assessed. Flanking the core region are large N-terminal and C-terminal extensions, which encompass 564 and 480 amino acids, respectively. Unlike the Myo18Aα N-terminal extension, which contains a well-defined KE-rich region and a PDZ domain, the N-terminal extension of Myo18B does not resemble any known protein sequence. With the exception of a putative nuclear localization sequence (NLS) at residues 2377-2387, the C-terminal extension also bares no similarity with known protein domains (Salamon et al. 2003).
Schematic representation of the domain structure of human Myosin-18B illustrating the 564-amino acid N-terminal extension, generic motor domain, neck region containing two myosin light chain binding IQ motifs, coiled-coil tail domain predicted to enable dimerization, and C-terminal extension, which contains a putative nuclear localization sequence (NLS) at residues 2377–2387
Whilst the biochemical properties of Myo18B remain to be elucidated, cellular localization studies and experiments with truncated overexpression constructs provide some insight into the function of this unique protein. Using overexpression constructs as well as N-terminal- and C-terminal-specific antibodies, this protein has been localized to a number of specific sites within both nonmuscle and muscle cell types (c.f. 19.3.2). In nonmuscle cells specifically, Myo18B has been shown to localize in a punctate pattern throughout the cytoplasm, in membrane protrusions, and within stress fibers (Inoue et al. 2006; Ajima et al. 2007; Jiu et al. 2019). In this last locality, Myo18B is thought to assemble into a compact configuration at the end of NM2 bipolar bundles, where it aids in the fusion and coalescence of adjacent myosin stacks to facilitate actomyosin bundle maturation (Jiu et al. 2019). This is evident in CRISPR/Cas9-mediated MYO18B knockout osteosarcoma cells, where a subset of stress fibers known as transverse arcs are unable to assemble into thick mature ventral stress fibers. Defective stress fiber maturation in these cells leads to abnormal morphogenesis, contractility, and reduced migration. Utilizing a series of overexpression domain constructs, both the N-terminal extension and coiled-coil tail domains were identified as being essential for the integration of Myo18B within stress fibers. The N-terminal extension alone localizes to stress fibers and actin bundles (Ajima et al. 2008; Jiu et al. 2019), suggesting that this unique protein domain may harbor essential actin-interaction sites, as seen within the N-terminal extension of Myo18A (Isogawa et al. 2005; Taft et al. 2013). Punctate and cytoplasmic localization of an overexpression construct lacking the N-terminal extension further supports the pivotal role for the N-terminus in mediating actin interactions. Unlike the N-terminus, deletion of the C-terminal extension has no effect on stress fiber localization and overexpression of a C-terminal deletion construct is able to rescue the CRISPR/Cas9-mediated MYO18B-knockout phenotype in osteosarcoma cells. Interestingly, Myo18A overexpression is shown to partially rescue the Myo18B-knockout phenotype, although it is not intrinsically upregulated to compensate for reduced Myo18B protein levels.
Aside from actin, two other interaction partners have been identified and validated for Myo18B through yeast two-hybrid screening studies. In the first study, Sug1, a component of the 19S regulatory complex of the 26S proteasome was identified and the interaction was validated in COS-7 and C2C12 cells (Inoue et al. 2006). Colocalization analysis in HeLa cells revealed that cytoplasmic punctate Myo18B staining in nonmuscle cells partially overlaps with Sug1 staining. Accordingly, it was validated that Myo18B is susceptible to polyubiquitination and Sug1-mediated proteasomal degradation, although the precise mechanisms and physiological relevance of these findings remain undetermined. In a second report, the C-terminal tail of Myo18B was found to interact with the C-terminal half of Homer2, a post-synaptic adaptor protein, at membrane protrusions and stress fibers (Ajima et al. 2007). Expression of Homer2 was shown to enhance the ability of Myo18B to suppress anchorage-independent growth (cf. 19.3.3); however, the precise mechanisms mediating this effect are unresolved.
19.3.2 Myo18B in Striated Muscle Development and Assembly
In accordance with the strong expression profile of MYO18B in cardiac and skeletal muscles, a considerable portion of the literature on Myo18B is specifically focused on understanding its function in striated muscle tissues. At the level of the gene, MYO18B is distinct from many other human sarcomeric myosin genes, as it does not map to chromosomes 14 and 17, where cardiac and skeletal myosin heavy chain genes are clustered, but instead localizes to chromosome 22 (Salamon et al. 2003). Gene expression profiling studies in mice reveal that Myo18B expression is upregulated following the onset of myogenic differentiation, the process by which myotubes mature from myoblast precursors under the control of myocyte enhancer factor-2 (MEF2) transcription factors (Salamon et al. 2003; Ajima et al. 2008). A single study in Atlantic cod also indicates that the expression of this gene in skeletal muscle is linked to the transcription pattern of clock genes (Lazado et al. 2014). In the immortalized C2C12 mouse myoblast cell line, expression peaks at differentiation day 3 and remains constant throughout myotube maturation (Salamon et al. 2003). In mouse embryos, this translates to strong expression of MYO18B mRNA in heart tissue at embryonic days 9.5–11.5 and within myotomes and muscle mass at embryonic day 13.5 (Ajima et al. 2008).
Whilst a number of studies have reported that Myo18B adopts a regularly-spaced striated distribution pattern within sarcomeric structures in mature myocytes, its precise subcellular distribution throughout differentiation and within sarcomeres remains controversial. The first study examining Myo18B localization by Salamon et al. utilizing stable c-myc-tagged overexpression constructs and an N-terminal rabbit polyclonal antibody reported several distinct localization patterns. Specifically, Myo18B was observed in the cytoplasm in undifferentiated myoblasts, within the nucleus in undifferentiated myoblasts and select differentiated myotubes, and was prominently incorporated into A-bands within mature sarcomeric structures (Salamon et al. 2003). However, a follow-up study utilizing two antibodies raised against C-terminal residues (amino acids 2448-2518) found no indication of Myo18B within the nucleus and observed that it localized with α-actinin at Z-discs rather than with muscle myosin-2 in A-bands (Ajima et al. 2008). Aspects of both of these initial studies were reproduced in a third publication assessing GFP-tagged Myo18B localization in zebrafish. In this instance, Berger et al. reported that exogenous Myo18B is integrated into sarcomeric A-bands 3 days post fertilization (dpf), where it alternates with α-actinin (Berger et al. 2017). They also report that they do not observe a nuclear localization pattern for Myo18B; however, it is unclear if their analysis adequately assesses localization in undifferentiated myoblasts.
Regardless of the discrepancies in localization analyses, these studies firmly demonstrate that Myo18B is an essential component of muscle contractile machinery. In mice, homozygous knockout of Myo18B was found to be embryonically lethal with major defects evident at embryonic day 10.5 (Ajima et al. 2008). Knockout embryos had severe cardiac defects, including dilated pericardial cavities and internal hemorrhage or resorption, and ultrastructural analysis revealed severe abnormalities in the organization and alignment of myosin thick and actin thin filaments in muscle tissue. This gave the first indication that Myo18B is required for the development and maintenance of myofibril structure. Two recent studies linking mutations in the zebrafish myo18B gene with defective muscle integrity, cardiac deformities and reduced motile capacity have further demonstrated the essential role for Myo18B in sarcomere assembly (Berger et al. 2017; Gurung et al. 2017). In this model organism, myo18B is expressed exclusively in fast-twitch muscle fibres 1 dpf, is found within the fin and heart 2 dpf, and is present in the head and extraocular muscles 3 dpf. Accordingly, mechanical force measurements in zebrafish baring allelic deletions and mutations in myo18B reveal that this protein is essential for force generation in fast-twitch muscles and not slow-twitch musculature (Berger et al. 2017). Ultrastructural analysis of fast-twitch muscle tissue in these mutant fish (3 dpf) by transmission electron microscopy demonstrates that the registered alignment of thick and thin filaments into sarcomeric structures does not occur in the absence of functional Myo18B.
To date, two studies have implicated MYO18B mutations in human myopathies. In the first instance, Malfatti et al. reported that a homozygous missense mutation in MYO18B (c.6496G > T, p.Glu2166∗) was linked with nemaline myopathy, a rare and severe congenital muscle disorder characterized by the presence of nemaline bodies (rods) within patient skeletal muscle (Malfatti et al. 2015). In this instance, mRNA expression was not affected by the mutation; however, the resulting protein was truncated within the C-terminal extension and the full-length protein could not be detected with a C-terminal antibody. The patients presented with dysmorphic features, clinodactyly, severe hypotonia, muscle weakness, and cardiomyopathy. In the second published case study, two patients presenting with a novel form of Klippel-Feil anomaly that included myopathy and distinct facial features were found to have the same homozygous nonsense mutation in MYO18B (c.6905C > A), which induced a premature stop codon at residue 2302 (Alazami et al. 2015). This mutation induced nonsense-mediated mRNA decay with almost no MYO18B mRNA identified in patient lymphoblastic cells. Consistent with the mouse and zebrafish models of Myo18B knockdown and knockout, ultrastructural analysis revealed that loss of functional Myo18B in these patients culminated in an apparent loss of thick and thin filament banding in muscle biopsies, further demonstrating the essential role for Myo18B in sarcomere assembly.
19.3.3 MYO18B in Cancer Progression
Initial studies characterizing the MYO18B gene in cancer stemmed from the observation that allelic losses on chromosome 22q are frequently detected in human tumors. This suggested that an essential tumor suppressor gene is present within this chromosomal region. A series of studies by the work group of Jun Yokota demonstrated that MYO18B is effectively inactivated in lung, ovarian, and colorectal cell lines and primary tumor samples by homozygous deletions, mutations, and/or epigenetic silencing (Nishioka et al. 2002; Tani et al. 2004; Yanaihara et al. 2004; Nakano et al. 2005). Specifically, hypermethylation of CpG sites in the MYO18B promoter region and deacetylation of histone H3 and histone H4 all significantly inactivate MYO18B in these cancer subsets. In accordance with the initial hypothesis that MYO18B functions as a tumor suppressor gene, Myo18B overexpression or restoration was also found to impede tumor progression by suppressing anchorage-independent growth.
The tumor suppressive role for Myo18B is further supported by the handful of studies that have identified MYO18B mutations in various cancers. This includes patients with melanoma and pancreatic ductal adenocarcinoma (Bleeker et al. 2009), two siblings with familial lung cancer (Kukita et al. 2016), and a single patient with neuroendocrine cancer (Bhatla et al. 2016). Two additional reports further strengthen the link between MYO18B epigenetic modifications and cancer. Specifically, reduced MYO18B expression due to hypermethylation was linked to a reduced response to platinum-based chemotherapies in ovarian cancer cases (Tomar et al. 2017), and increased MYO18B expression due to hypomethylation in T-cell acute leukemia was associated with better patient prognosis (Haider et al. 2019). Both reports support a tumor suppressive role for MYO18B in cancer.
To date, a single study published in 2018 by Zhang et al. has reported a tumor promoting role for MYO18B in hepatocellular carcinoma progression (Zhang et al. 2018). Specifically, increased MYO18B expression in a large TCGA (The Cancer Genome Atlas) cohort and an independent validation cohort correlated with advanced disease and worse overall survival. In support of this, Myo18B knockdown by siRNA reduced cell proliferation, decreased tumor colony formation over time, slowed 2D-wound healing, and significantly decreased 3D-invasion in Transwell assays. This is in line with the effect of CRISPR/Cas9-mediated Myo18B knockout in osteosarcoma cells, which significantly impaired the ability for cells to undergo 3D-invasive migration, likely due to the defective maturation of NM2 stress fibers (Jiu et al. 2019). In the case of hepatocellular carcinoma cells, it was shown that Myo18B knockdown significantly affected signaling components essential for proliferation, apoptosis, and migrations. Specifically, knockdown suppressed the phosphorylation of PI3K, AKT, and mTOR, and resulted in an evident decrease in p70S6K expression.
19.3.4 MYO18B in Other Disease Contexts
In addition to the role of MYO18B in cancer and myopathies, this gene has also been linked to other diseases. One of the most significant findings was a report by Ludwig et al., which found that single nucleotide polymorphisms in MYO18B are indicative and causative of mathematical disability (Ludwig et al. 2013). The authors reported that people carrying a MYO18B risk-genotype displayed significantly lower depth in an area of the brain involved in numerical processing, the intraparietal sulcus (IPS). These results have however been questioned by a follow-up study, which reports that no link between MYO18B and mathematical disability could be found in four independent validation cohorts (Pettigrew et al. 2015). In addition, MYO18B single nucleotide variants have also been associated with schizophrenia in a genome-wide association study (Takata et al. 2013), and a heterozygous variant in MYO18B was identified in a single case of infant death, where the fetus had significant deformities, which included a muscle phenotype (Armes et al. 2018).
19.4 Perspectives
In the 20 years since class XVIII myosins were first discovered, there have been significant advances in resolving their functions at both the molecular and cellular levels. However, it is clear that we have only just begun to unravel the properties of members of this new branch within the myosin family, characterized by their unique N- and C-terminal extensions, as well as their apparent lack of active ATPase-driven motor function. A number of obvious questions remain to be addressed. This includes whether the two proteins in this family, Myo18A and Myo18B, have overlapping functions within common subcellular localities. This has been addressed in a single study by Jiu et al., where overexpression of Myo18A could slightly compensate for Myo18B knockout in stress fiber maturation; however, knockout did not induce an intrinsic compensatory increase in Myo18A (Jiu et al. 2019). With regard to surfactant protein A receptor, it is important to address how this cytoplasmic soluble protein is translocated and anchored to the plasma membrane. Further, does the PDZ domain, which is known to mediate membrane association, play a role in this function? With regard to Myo18B, it is clear that an in-depth biochemical analysis of the protein’s subdomains is required to completely understand the function of Myo18B in stress fiber maturation and sarcomere assembly. Furthermore, as a putative NLS has been identified within the C-terminal extension and a nuclear localization has been reported in myocytes, it would be of great interest to resolve whether Myo18B has a nuclear function. Whilst Salamon et al. proposed that Myo18B may function as a transcriptional regulator of myogenic differentiation (Salamon et al. 2003), this remains to be resolved in follow-up studies.
References
Ajima R, Kajiya K, Inoue T et al (2007) HOMER2 binds MYO18B and enhances its activity to suppress anchorage independent growth. Biochem Biophys Res Commun 356:851–856. https://doi.org/10.1016/j.bbrc.2007.03.060
Ajima R, Akazawa H, Kodama M et al (2008) Deficiency of Myo18B in mice results in embryonic lethality with cardiac myofibrillar aberrations. Genes Cells 13:987–999. https://doi.org/10.1111/j.1365-2443.2008.01226.x
Alazami AM, Kentab AY, Faqeih E et al (2015) A novel syndrome of Klippel-Feil anomaly, myopathy, and characteristic facies is linked to a null mutation in MYO18B. J Med Genet 52:400–404. https://doi.org/10.1136/jmedgenet-2014-102964
Armes JE, Williams M, Price G et al (2018) Application of whole genome sequencing technology in the investigation of genetic causes of Fetal, perinatal, and early infant death. Pediatr Dev Pathol 21:54–67. https://doi.org/10.1177/1093526617715528
Berg JS, Powell BC, Cheney RE (2001) A millennial myosin census. Mol Biol Cell 12:780–794
Berger J, Berger S, Li M, Currie PD (2017) Myo18b is essential for sarcomere assembly in fast skeletal muscle. Hum Mol Genet 26:ddx025. https://doi.org/10.1093/hmg/ddx025
Bhatla T, Dandekar S, Lu BY et al (2016) Genomic characterization of poorly differentiated neuroendocrine carcinoma in a Pediatric patient. J Pediatr Hematol Oncol 38:e21–e25. https://doi.org/10.1097/MPH.0000000000000463
Billington N, Beach JR, Heissler SM et al (2015) Myosin 18A coassembles with nonmuscle myosin 2 to form mixed bipolar filaments. Curr Biol 25:942–948. https://doi.org/10.1016/j.cub.2015.02.012
Bishé B, Syed GH, Field SJ, Siddiqui A (2012) Role of phosphatidylinositol 4-phosphate (PI4P) and its binding protein GOLPH3 in hepatitis C virus secretion. J Biol Chem 287:27637–27647. https://doi.org/10.1074/jbc.M112.346569
Bleeker FE, Lamba S, Rodolfo M et al (2009) Mutational profiling of cancer candidate genes in glioblastoma, melanoma and pancreatic carcinoma reveals a snapshot of their genomic landscapes. Hum Mutat 30:E451–E459. https://doi.org/10.1002/humu.20927
Bloemink MJ, Geeves MA (2011) Shaking the myosin family tree: biochemical kinetics defines four types of myosin motor. Semin Cell Dev Biol 22:961–967
Bonn BR, Rudolf A, Hornbruch-Freitag C et al (2013) Myosin heavy chain-like localizes at cell contact sites during Drosophila myoblast fusion and interacts in vitro with rolling pebbles 7. Exp Cell Res 319:402–416. https://doi.org/10.1016/j.yexcr.2012.12.005
Bruun K, Beach JR, Heissler SM et al (2017) Re-evaluating the roles of myosin 18Aα and F-actin in determining Golgi morphology. Cytoskeleton (Hoboken) 74:205–218. https://doi.org/10.1002/cm.21364
Buschman MD, Field SJ (2018) MYO18A: an unusual myosin. Adv Biol Regul 67:84–92. https://doi.org/10.1016/j.jbior.2017.09.005
Buschman MD, Xing M, Field SJ (2015) The GOLPH3 pathway regulates Golgi shape and function and is activated by DNA damage. Front Neurosci 9:362. https://doi.org/10.3389/fnins.2015.00362
Cao J, Li S, Shao M et al (2014) The PDZ-containing unconventional myosin XVIIIA regulates embryonic muscle integrity in zebrafish. J Genet Genomics 41:417–428. https://doi.org/10.1016/j.jgg.2014.06.008
Cao J-M, Cheng X-N, Li S-Q et al (2016) Identification of novel MYO18A interaction partners required for myoblast adhesion and muscle integrity. Sci Rep 6:36768. https://doi.org/10.1038/srep36768
Cross M, Csar XF, Wilson NJ et al (2004) A novel 110 kDa form of myosin XVIIIA (MysPDZ) is tyrosine-phosphorylated after colony-stimulating factor-1 receptor signalling. Biochem J 380:243–253. https://doi.org/10.1042/BJ20031978
De Masson A, Giustiniani J, Marie-Cardine A et al (2016) Identification of CD245 as myosin 18A, a receptor for surfactant a: a novel pathway for activating human NK lymphocytes. Oncoimmunology 5:e1127493. https://doi.org/10.1080/2162402X.2015.1127493
Dippold HC, Ng MM, Farber-Katz SE et al (2009) GOLPH3 bridges phosphatidylinositol-4- phosphate and actomyosin to stretch and shape the Golgi to promote budding. Cell 139:337–351. https://doi.org/10.1016/j.cell.2009.07.052
Dowler S, Currie RA, Campbell DG et al (2000) Identification of pleckstrin-homology-domain-containing proteins with novel phosphoinositide-binding specificities. Biochem J 351:19. https://doi.org/10.1042/0264-6021:3510019
Farber-Katz SE, Dippold HC, Buschman MD et al (2014) DNA Damage Triggers Golgi Dispersal via DNA-PK and GOLPH3. Cell 156:413–427. https://doi.org/10.1016/j.cell.2013.12.023
Furch M, Fujita-Becker S, Geeves MA et al (1999) Role of the salt-bridge between switch-1 and switch-2 of Dictyostelium myosin. J Mol Biol 290:797–809
Furusawa T, Ikawa S, Yanai N, Obinata M (2000) Isolation of a novel PDZ-containing myosin from hematopoietic supportive bone marrow stromal cell lines. Biochem Biophys Res Commun 270:67–75. https://doi.org/10.1006/bbrc.2000.2377
Geeves MA (2016) Review: the ATPase mechanism of myosin and actomyosin. Biopolymers 105:483–491. https://doi.org/10.1002/bip.22853
Geeves MA, Fedorov R, Manstein DJ (2005) Molecular mechanism of actomyosin-based motility. Cell Mol Life Sci 62:1462–1477. https://doi.org/10.1007/s00018-005-5015-5
Gurel PS, Hatch AL, Higgs HN (2014) Connecting the cytoskeleton to the endoplasmic reticulum and Golgi. Curr Biol 24:R660–R672. https://doi.org/10.1016/j.cub.2014.05.033
Gurung R, Ono Y, Baxendale S et al (2017) A zebrafish model for a human myopathy associated with mutation of the unconventional myosin MYO18B. Genetics 205:725–735. https://doi.org/10.1534/genetics.116.192864
Guzik-Lendrum S, Nagy A, Takagi Y et al (2011) Drosophila melanogaster myosin-18 represents a highly divergent motor with actin tethering properties. J Biol Chem 286:21755–21766
Guzik-Lendrum S, Heissler SM, Billington N et al (2013) Mammalian myosin-18A, a highly divergent myosin. J Biol Chem 288:9532–9548. https://doi.org/10.1074/jbc.M112.441238
Haider Z, Larsson P, Landfors M et al (2019) An integrated transcriptome analysis in T-cell acute lymphoblastic leukemia links DNA methylation subgroups to dysregulated TAL1 and ANTP homeobox gene expression. Cancer Med 8:311–324. https://doi.org/10.1002/cam4.1917
Horsthemke M, Nutter LMJ, Bachg AC et al (2019) A novel isoform of myosin 18A (Myo18Aγ) is an essential sarcomeric protein in mouse heart. J Biol Chem 294:7202–7218. https://doi.org/10.1074/jbc.RA118.004560
Hsu R-M, Tsai M-H, Hsieh Y-J et al (2010) Identification of MYO18A as a novel interacting partner of the PAK2/betaPIX/GIT1 complex and its potential function in modulating epithelial cell migration. Mol Biol Cell 21:287–301. https://doi.org/10.1091/mbc.E09-03-0232
Hsu R-M, Hsieh Y-J, Yang T-H et al (2014) Binding of the extreme carboxyl-terminus of PAK-interacting exchange factor β (βPIX) to myosin 18A (MYO18A) is required for epithelial cell migration. Biochim Biophys Acta, Mol Cell Res 1843:2513–2527. https://doi.org/10.1016/j.bbamcr.2014.06.023
Inoue T, Kon T, Ajima R et al (2006) MYO18B interacts with the proteasomal subunit Sug1 and is degraded by the ubiquitin-proteasome pathway. Biochem Biophys Res Commun 342:829–834. https://doi.org/10.1016/j.bbrc.2006.02.025
Isogawa Y, Kon T, Inoue T et al (2005) The N-terminal domain of MYO18A has an ATP-insensitive actin-binding site. Biochemistry 44:6190–6196
Jean Beltran PM, Mathias RA, Cristea IM (2016) A portrait of the human organelle proteome in space and time during cytomegalovirus infection. Cell Syst 3:361–373.e6. https://doi.org/10.1016/j.cels.2016.08.012
Ji H, Zhai Q, Zhu J et al (2000) A novel protein MAJN binds to Jak3 and inhibits apoptosis induced by IL-2 deprival. Biochem Biophys Res Commun 270:267–271. https://doi.org/10.1006/bbrc.2000.2413
Jiu Y, Kumari R, Fenix AM et al (2019) Myosin-18B promotes the assembly of myosin II stacks for maturation of contractile Actomyosin bundles. Curr Biol 29:81–92.e5. https://doi.org/10.1016/j.cub.2018.11.045
Kukita Y, Okami J, Yoneda-Kato N et al (2016) Homozygous inactivation of CHEK2 is linked to a familial case of multiple primary lung cancer with accompanying cancers in other organs. Mol Case Stud 2:a001032. https://doi.org/10.1101/mcs.a001032
Langer W, Sohler F, Leder G et al (2010) Exon array analysis using re-defined probe sets results in reliable identification of alternatively spliced genes in non-small cell lung cancer. BMC Genomics 11:676. https://doi.org/10.1186/1471-2164-11-676
Lazado CC, Nagasawa K, Babiak I et al (2014) Circadian rhythmicity and photic plasticity of myosin gene transcription in fast skeletal muscle of Atlantic cod (Gadus morhua). Mar Genomics 18:21–29. https://doi.org/10.1016/j.margen.2014.04.011
Levine TP, Munro S (2002) Targeting of Golgi-specific pleckstrin homology domains involves both PtdIns 4-kinase-dependent and -independent components. Curr Biol 12:695–704
Ludwig KU, Sämann P, Alexander M et al (2013) A common variant in Myosin-18B contributes to mathematical abilities in children with dyslexia and intraparietal sulcus variability in adults. Transl Psychiatry 3:e229. https://doi.org/10.1038/tp.2012.148
Makowska KA, Hughes RE, White KJ et al (2015) Specific Myosins control actin organization, cell morphology, and migration in prostate Cancer cells. Cell Rep 13:2118–2125. https://doi.org/10.1016/j.celrep.2015.11.012
Malfatti E, Böhm J, Lacène E et al (2015) A premature stop codon in MYO18B is associated with severe Nemaline myopathy with cardiomyopathy. J Neuromuscul Dis 2:219–227. https://doi.org/10.3233/JND-150085
McNally E, Sohn R, Frankel S, Leinwand L (1991) Expression of myosin and actin in Escherichia coli. Methods Enzymol 196:368–389
Mori K, Furusawa T, Okubo T et al (2003) Genome structure and differential expression of two isoforms of a novel PDZ-containing myosin (MysPDZ) (Myo18A). J Biochem 133:405–413
Mori K, Matsuda K, Furusawa T et al (2005) Subcellular localization and dynamics of MysPDZ (Myo18A) in live mammalian cells. Biochem Biophys Res Commun 326:491–498
Nakano T, Tani M, Nishioka M et al (2005) Genetic and epigenetic alterations of the candidate tumor-suppressor gene MYO18B, on chromosome arm 22q, in colorectal cancer. Genes Chromosomes Cancer 43:162–171. https://doi.org/10.1002/gcc.20180
Ng MM, Dippold HC, Buschman MD et al (2013) GOLPH3L antagonizes GOLPH3 to determine Golgi morphology. Mol Biol Cell 24:796–808. https://doi.org/10.1091/mbc.e12-07-0525
Nishioka M, Kohno T, Tani M et al (2002) MYO18B, a candidate tumor suppressor gene at chromosome 22q12.1, deleted, mutated, and methylated in human lung cancer. Proc Natl Acad Sci U S A 99:12269–12274. https://doi.org/10.1073/pnas.192445899
Odronitz F, Kollmar M (2007) Drawing the tree of eukaryotic life based on the analysis of 2,269 manually annotated myosins from 328 species. Genome Biol 8:R196
Onishi H, Kojima S, Katoh K et al (1998) Functional transitions in myosin: formation of a critical salt-bridge and transmission of effect to the sensitive tryptophan. Proc Natl Acad Sci U S A 95:6653–6658
Peckham M (2016) How myosin organization of the actin cytoskeleton contributes to the cancer phenotype. Biochem Soc Trans 44:1026–1034. https://doi.org/10.1042/BST20160034
Pettigrew KA, Fajutrao Valles SF, Moll K et al (2015) Lack of replication for the myosin-18B association with mathematical ability in independent cohorts. Genes Brain Behav 14:369–376. https://doi.org/10.1111/gbb.12213
Preller M, Manstein DJ (2013) Myosin structure, allostery, and mechano-chemistry. Structure 21:1911–1922. https://doi.org/10.1016/j.str.2013.09.015
Salamon M, Millino C, Raffaello A et al (2003) Human MYO18B, a novel unconventional myosin heavy chain expressed in striated muscles moves into the myonuclei upon differentiation. J Mol Biol 326:137–149
Samten B, Townsend JC, Sever-Chroneos Z et al (2008) An antibody against the surfactant protein a (SP-A)-binding domain of the SP-A receptor inhibits T cell-mediated immune responses to Mycobacterium tuberculosis. J Leukoc Biol 84:115–123. https://doi.org/10.1189/jlb.1207835
Scott KL, Kabbarah O, Liang M-C et al (2009) GOLPH3 modulates mTOR signalling and rapamycin sensitivity in cancer. Nature 459:1085–1090. https://doi.org/10.1038/nature08109
Szeliga J, Jordan J, Yang C-H et al (2005) Bacterial expression of recombinant MyoXVIIIA domains. Anal Biochem 346:179–181. https://doi.org/10.1016/j.ab.2005.07.021
Taft MH, Behrmann E, Munske-Weidemann L-C et al (2013) Functional characterization of human Myosin-18A and its interaction with F-actin and GOLPH3. J Biol Chem 288:30029–30041. https://doi.org/10.1074/jbc.M113.497180
Takata A, Iwayama Y, Fukuo Y et al (2013) A population-specific uncommon variant in GRIN3A associated with schizophrenia. Biol Psychiatry 73:532–539. https://doi.org/10.1016/j.biopsych.2012.10.024
Tan I, Yong J, Dong JM et al (2008) A tripartite complex containing MRCK modulates lamellar actomyosin retrograde flow. Cell 135:123–136. https://doi.org/10.1016/j.cell.2008.09.018
Tani M, Ito J, Nishioka M et al (2004) Correlation between histone acetylation and expression of the MYO18B gene in human lung cancer cells. Genes Chromosomes Cancer 40:146–151. https://doi.org/10.1002/gcc.20027
Tomar T, Alkema NG, Schreuder L et al (2017) Methylome analysis of extreme chemoresponsive patients identifies novel markers of platinum sensitivity in high-grade serous ovarian cancer. BMC Med 15:116. https://doi.org/10.1186/s12916-017-0870-0
Ussowicz M, Jaśkowiec A, Meyer C et al (2012) A three-way translocation of MLL, MLLT11, and the novel reciprocal partner gene MYO18A in a child with acute myeloid leukemia. Cancer Genet 205:261–265. https://doi.org/10.1016/j.cancergen.2012.02.006
Walz C, Chase A, Schoch C et al (2005) The t(8;17)(p11;q23) in the 8p11 myeloproliferative syndrome fuses MYO18A to FGFR1. Leuk Off J Leuk Soc Am Leuk Res Fund UK 19:1005–1009. https://doi.org/10.1038/sj.leu.2403712
Walz C, Haferlach C, Hänel A et al (2009) Identification of a MYO18A-PDGFRB fusion gene in an eosinophilia-associated atypical myeloproliferative neoplasm with a t(5;17)(q33-34;q11.2). Genes Chromosomes Cancer 48:179–183. https://doi.org/10.1002/gcc.20629
Xing M, Peterman MC, Davis RL et al (2016) GOLPH3 drives cell migration by promoting Golgi reorientation and directional trafficking to the leading edge. Mol Biol Cell 27:3828–3840. https://doi.org/10.1091/mbc.E16-01-0005
Yamashita RA, Sellers JR, Anderson JB (2000) Identification and analysis of the myosin superfamily in Drosophila: a database approach. J Muscle Res Cell Motil 21:491–505
Yanaihara N, Nishioka M, Kohno T et al (2004) Reduced expression ofMYO18B, a candidate tumor-suppressor gene on chromosome arm 22q, in ovarian cancer. Int J Cancer 112:150–154. https://doi.org/10.1002/ijc.20339
Yang C-H, Szeliga J, Jordan J et al (2005) Identification of the surfactant protein A receptor 210 as the unconventional myosin 18A. J Biol Chem 280:34447–34457. https://doi.org/10.1074/jbc.M505229200
Yang L, Carrillo M, Wu YM et al (2015) SP-R210 (Myo18A) isoforms as intrinsic modulators of macrophage priming and activation. PLoS One 10:e0126576. https://doi.org/10.1371/journal.pone.0126576
Zhang Z, Zhu J, Huang Y et al (2018) MYO18B promotes hepatocellular carcinoma progression by activating PI3K/AKT/mTOR signaling pathway. Diagn Pathol 13:85. https://doi.org/10.1186/s13000-018-0763-3
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Taft, M.H., Latham, S.L. (2020). Myosin XVIII. In: Coluccio, L. (eds) Myosins. Advances in Experimental Medicine and Biology, vol 1239. Springer, Cham. https://doi.org/10.1007/978-3-030-38062-5_19
Download citation
DOI: https://doi.org/10.1007/978-3-030-38062-5_19
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-38061-8
Online ISBN: 978-3-030-38062-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)