Abstract
Compared to other cells, sperm undergo dramatic remodeling of their chromatin during late spermiogenesis in which approximately 95% of histones are removed and replaced with protamines. Despite this large-scale remodeling, key developmental genes, some miRNA genes, and imprinted genes retain their association with histone. The developmental genes have a unique epigenetic signature, termed bivalency, that poises the genes for embryonic activation. Anomalies in that epigenetic poising signature, either in the form of DNA methylation aberrations, improper protamination, or altered histone modifications, are associated with infertility and reduced embryogenesis capability. Additionally, some small noncoding RNAs are retained, while others are actively added to the sperm and appear to affect embryogenesis. Therefore, initial studies have begun to formulate pathways by which the sperm epigenome can be used as a diagnostic tool in the clinic. While in their infancy, these assays likely portend improved diagnostics and added information for patients and clinicians. Recent studies also highlight the possibility that the sperm epigenome can be used to evaluate lifestyle and environmental risks to the patient and potentially to the offspring.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Epigenetics, a term first coined by Conrad Waddington in the 1940s, is generally defined as having three major components. First, epigenetic “marks” are stable, non-DNA coding (polymorphisms or mutations) changes that, second, affect gene expression. Third, epigenetic alterations are heritable, generally defined as being passed at least to the F3 generation (Waddington 1942; Holliday 2006). Epigenetics is both practically and historically linked to the field of reproductive biology, since such gene expression changes are the key to cellular differentiation during embryonic development and since the term “epigenetics” is derived from “epigenesis,” the embryological concept of “stepwise” development of the embryo that has been discussed since the times of Aristotle, but particularly debated in the seventeenth and eighteenth centuries during the debates of the “preformation theory” of development versus the epigenesis theory of development (Harvey 1651, 1653). Since the term epigenetics was first coined, epigenetics has moved into a much broader realm, including the study of environmental influences on gene expression and disease etiologies (Jirtle and Skinner 2007; Deans and Maggert 2015).
Numerous studies have now demonstrated the powerful effects of alterations of the epigenome due to environmental or lifestyle factors on subsequent health. Among the best examples are studies evaluating famine or diet alterations at specific developmental time periods, such as the prenatal, perinatal, and peri-pubertal periods. For example, children born during the 1944–1945 Dutch famine period have been shown to have an increased risk of heart disease and obesity if their mother was exposed to the famine, apparently due to a change in DNA methylation to the insulin-like growth factor 2 (ILGF2) gene (Painter et al. 2005; Heijmans et al. 2008). In another example, paternal grandsons of prepubertal boys exposed to famine in the Overkalix region of Sweden have been shown to have increased mortality, although no effects were seen for the paternal grandmother or maternal grandparents (Pembrey et al. 2006). These studies highlight the ability of environmental changes, diet in these cases, to alter the epigenome through sperm, as well as highlighting the potential of abnormal epigenetics to affect the health of offspring and progeny.
As is often the case, the first studies of the sperm epigenome were not aimed at defining the normal sperm epigenome in normal development, rather the result of a possible associated disease risk. The earliest studies of the sperm epigenome were the result of a reported increase in the incidence of imprinting diseases, a form of epigenetic abnormality, in the offspring of individuals conceived using intracytoplasmic sperm injection (ICSI) during assisted reproductive therapy (ART) (Cox et al. 2002; Kobayashi et al. 2007; Le Bouc et al. 2010). Imprinting diseases are the result of improper sex-determined allelic methylation, and some studies demonstrated that the sperm of men undergoing ICSI had increased levels of abnormal DNA methylation, associated with decreased sperm counts. These studies opened the door to a deeper evaluation of the sperm epigenome and the role it may play in embryogenesis and the health of the offspring. While still in its infancy, the role of the sperm epigenome in embryogenesis is becoming better understood, as well as the role of certain lifestyle and environmental factors in altering it. This chapter explores these advances in terms of the potential to offer improved diagnosis and treatment of infertility through ART and in terms of ultimately better understanding the transmission of health risk to offspring through the sperm epigenome.
The Sperm Epigenome: Protamination and Histone Modifications
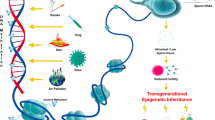
During late spermiogenesis, sperm undergo a dramatic remodeling of the sperm chromatin in which approximately 95% of the histones are sequentially replaced, first by transition proteins and then by protamines (P1 and P2) (see Fig. 3.1). The replacement of histones facilitates a higher order of DNA packaging, up to 20 times more than DNA in somatic cells, and is useful in providing a compact sperm head consistent with sperm motility requirements. Lastly, it is believed that the compaction of the DNA into toroidal structures also protects the DNA from oxidative stress while the sperm traverse the female reproductive tract. In fertile men, P1 and P2 replace histones with an approximate ratio of 1:1 (Carrell and Liu 2001), and alterations in the P1/P2 ratio reflect abnormal protamination and are associated with reduced semen quality, increased sperm DNA fragmentation, and reduced fertilization capabilities and embryo implantation in couples undergoing IVF (Aoki et al. 2005, 2006a; Carrell et al. 2008; Hammoud et al. 2009a; Carrell 2012).
An overview of the sperm epigenome. This figure highlights the repackaging of the sperm chromatin with protamines, with interspersed histones at key loci, including developmental gene promoters. The packaging of the genome with protamines facilitates higher-order chromatin compaction, including the formation of toroids. The figure shows the three key components of the sperm epigenome: histone modifications, DNA methylation, and the pool of RNAs, some of which are miRNAs and tRFs
Using a mouse model, intracytoplasmic sperm injection (ICSI) of sperm containing altered histone/protamine ratios and high DNA fragmentation resulted in embryos with a lower competency (Cho et al. 2003). Generally, there is a consensus that aberrant protamination is associated with increased DNA fragmentation and reduced embryo quality (Carrell and Liu 2001; Aoki et al. 2006a, b, c; Cho et al. 2003).
In addition to the role of protamination on protecting sperm DNA, the replacement of 95% of histones begs the question of if there is a role for the remaining histones—perhaps an epigenetic role? Stated differently, for such an evolutionary important aspect of reproduction, why would protamination be so inefficient as to leave 5% of the genome non-protaminated? Lastly, one would also hypothesize that if there were a role for retained histones, their loci of retention would be consistent and may suggest a biological role. Studies by Hammoud et al. were initially undertaken to help answer those questions and included genome-wide analysis of the loci of histone retention, specific histone modification analysis, and evaluation of DNA methylation status genome-wide and found that retained histones are found at consistent and deliberate locations throughout the sperm genome, including key developmental genes, poising these genes for activation during early embryogenesis (Hammoud et al. 2009b). These findings imply that proper protamination, and likewise normal retention of specific histones, is not only important in regard to a reflection of male-factor (MF) infertility status, as a reflection of abnormal spermatogenesis, and in regard to protecting the genome from DNA damage but also suggested a role in paternal contributions to normal embryogenesis.
The hypothesis that retention of sperm histones at key developmental loci is of biological significance is also dependent on proper histone modifications, since specific modifications can either facilitate or preclude transcription by making gene promoters accessible or inaccessible to transcription factors. Histone tail modifications are a major class of epigenetic regulators in somatic cells. Briefly, acetylation of H3 and H4 as well as methylation of H3K4 results in an “open” state of genes that facilitates transcription. Conversely, methylation of H3K9 and H3K27 and deacetylation of H3 and H4 drive a chromatin state which silences genes at those loci (Jenkins and Carrell 2011; Jenuwein and Allis 2001). Hammoud et al., and subsequently others, demonstrated that in human sperm, the modifications of histones, associated with developmental genes, are unique in that there is bivalency as both marks containing both H3K4me3 activation marks and H3K27 silencing marks are present, similar to what is found in some embryonic stem cell gene loci (Hammoud et al. 2009b). This arrangement suggests a “gene poising” of key genes involved in embryonic development. Interestingly, many IVF patients with altered embryogenesis capability have been shown to exhibit defects in this poising pattern (Hammoud et al. 2011). Furthermore, this unique poising pattern of embryonic developmental genes has been confirmed in zebrafish, an evolutionarily distant species (Murphy et al. 2018a).
The Sperm Epigenome: DNA Methylation
DNA methylation is the major regulator of gene transcription and technically easier to evaluate than histone modifications; therefore, more studies have reported DNA methylation status of the human sperm than those evaluating histone modifications. While targeted sequencing studies can be employed following bisulfate conversion, many studies have used arrays to screen many loci and evaluate possible associations between DNA methylation alterations and various phenotypes of male infertility. The arrays offer the advantages of ease of use, as well as screening many possible CpG loci. Hammoud et al. and others have demonstrated that the normal male sperm epigenome has variable methylation at the key developmental loci described above that contain bivalent histone modifications, thus strengthening the poising hypothesis (Hammoud et al. 2009b). Furthermore, several early studies have observed DNA methylation aberrations in sperm with abnormal chromatin packaging, sperm from men who generate embryos of poor quality while undergoing in vitro fertilization, as well as infertile men (Hammoud et al. 2010, 2011; Aston et al. 2012, 2015; Nanassy and Carrell 2011).
Sperm DNA methylation is also important in terms of genomic imprinting, a system in which certain genes are methylated or demethylated based on whether the locus is inherited from the father or mother. Prior to zygotic genome activation in the early embryo, DNA methylation patterns acquired from the sperm and oocyte are actively and passively demethylated and then reset later in the primordial germ cells (Messerschmidt et al. 2014). This process has led some to minimize the potential importance of DNA methylation in epigenetic inheritance; however, it is known that not only imprinted genes but other regions of the genome escape this reprogramming event in early embryos, and at least in the case of imprinted loci, the methylation signature provided to the embryo by sperm is maintained (Messerschmidt et al. 2014; Reik and Walter 2001). Some methylation signatures beyond imprinted regions are retained in the embryo and are involved in modulating development and affecting phenotype transgenerationally. Such signatures have been identified, including methylation patterns inherited via sperm (Guibert et al. 2012; Illum et al. 2018; Seisenberger et al. 2012). One study found that the female offspring of male rats consuming a high-fat diet displayed multiple characteristics consistent with metabolic phenotypes, including reduced birthweight, decreased pancreatic beta-cell mass, and glucose intolerance (Barbosa et al. 2015). DNA methylation analysis was conducted on the sperm from the F0 high-fat-fed rats and their F1 male offspring, and multiple methylation alterations were observed when compared to control rats, and in fact many differentially methylated regions were concordantly observed in both the F0 and F1 male sperm, suggesting a possible mechanism for transgenerational inheritance of metabolic disease .
Given the early data described above in which methylation errors at imprinted loci were found to be more common in men with abnormal spermatogenesis, and particularly men with low sperm counts, it was imperative to evaluate the methylation status of DNA from sperm of men with broader types of male-associated infertility. In one such early study, Aston et al. evaluated several thousand loci, using an early array system, in sperm from men with either aberrant protamination or men with unexplained poor embryogenesis while undergoing IVF27. Interestingly, more than 7% of all loci evaluated were abnormally methylated in these patients, and more than 60% of imprinted loci were aberrantly methylated. This study was supported by numerous other studies and focused attention on the potential use of sperm DNA methylation analysis as a potential screen for IVF embryogenesis outcome, as will be discussed below (Jenkins et al. 2016a; Karaca et al. 2017; Laqqan and Hammadeh 2018; Santi et al. 2017).
The Sperm Epigenome: Small, Noncoding RNAs
Although gene transcription does not occur in a mature sperm, sperm contain many RNA species that are stable in the embryo following fertilization (Ostermeier et al. 2004; Pessot et al. 1989). Sperm RNAs include remnant mRNAs from spermatogenesis (Ostermeier et al. 2002, 2004), mRNAs that may be functionally important to the developing embryo (Ostermeier et al. 2002; Jodar et al. 2015; Sendler et al. 2013), and a variety of noncoding RNAs (Krawetz et al. 2011). Recently, much of the research on sperm epigenetic factors and epigenetic-mediated inheritance has focused on the small, noncoding RNAs (Conine et al. 2018; Liu et al. 2012; Chen et al. 2016; Sharma et al. 2016; Zhang et al. 2018).
Studies in the mouse have shown that there are two major sources of noncoding RNAs in sperm. First, sperm contain a large number of piRNAs that are remnants of spermatogenesis. Second, during epididymal transit, there appears to be a significant remodeling of the RNAs, with some RNAs apparently removed in the caput epididymis and then subsequently replaced, along with other species of RNAs. This replacement and remodeling appears to occur largely through epididymosomes, exosomes that are secreted in the epididymis and attach to sperm and transfer their contents. These epididymosomes transfer a large contingent of miRNAs and tRNA fragments (tRFs), as well as proteins necessary in the acquisition of sperm motility and fertilization ability. Interestingly, the RNA payload of a sperm isolated from the cauda epididymis is similar to a sperm isolated from the testis, but RNAs are removed in the caput epididymis and then are subsequently replaced via the epidiymosomes (Sharma et al. 2018).
In an elegant study by Conine et al., it was shown that some of the RNA species that are lost and subsequently regained by sperm cells transiting the epididymis are associated with improper embryonic implantation as well as gross defects in embryonic development47. Using caput and cauda sperm to generate mouse embryos, Conine et al. reported that caput-derived embryos showed significantly reduced rates of successful implantation, gross embryo morphology defects, and a reduced number of viable offspring (Conine et al. 2018). They then showed that the embryos could be “rescued” by injection of the miRNA fraction of epididymosomes. These results suggest that the miRNAs delivered to sperm during epididymal transit are required for proper preimplantation gene expression in the mouse (Conine et al. 2018). The miRNAs injected included miR-34c, which has previously been shown by another group to be essential for the first cleavage division in mouse embryos (Liu et al. 2012). These two studies strongly suggest a role for spermatozoal RNAs, specifically miRNAs, in embryogenesis.
Studies evaluating the effects of diet changes have implicated spermatozoal RNAs in epigenetic inheritance of metabolic disease from fathers. Altered metabolic phenotypes have been observed in the offspring of male mice consuming high-fat or low-protein diets. These offspring display a phenotype characterized by glucose intolerance and impaired insulin secretion (Barbosa et al. 2015; Chen et al. 2016; Sharma et al. 2016; Carone et al. 2010; Ng et al. 2010). Spermatozoal RNAs, especially tRFs whose effects appear to be mediated by DNA methyltransferase 2 (DNMT2), have been implicated in this form of epigenetic inheritance (Chen et al. 2016; Sharma et al. 2016; Zhang et al. 2018). Jodar et al. rescued diet-affected zygotes by microinjection with RNAs isolated from control-diet sperm (Sharma et al. 2016). These studies highlight the ability of spermatozoal RNAs to affect gene expression patterns in the embryo and the possible role of sperm RNAs in epigenetic inheritance.
Using the Sperm Epigenome to Predict Fertility or ART Outcome
The bivalent poising of the human sperm epigenome strongly suggests a role in normal embryogenesis, and similar poising motifs are observed in diverse species, again suggesting an evolutionary role of importance (Hammoud et al. 2009b; Wu et al. 2011). Additionally, numerous studies have reported associations among abnormal sperm DNA methylation, histone replacement, histone modifications, or RNA complements with reduced male fertility, altered spermatogenesis, and altered embryogenesis capability during IVF (Jenkins et al. 2017; Gannon et al. 2014). Therefore, the possible use of the sperm epigenome as a diagnostic tool for couples undergoing infertility evaluation is apparent and has begun to be the focus of some researchers (Carrell 2012).
Aston et al. initially set out to determine the predictive role of sperm methylation patterns in patients who had undergone IVF treatment (Aston et al. 2015). Previous IVF patients were classified on whether their sperm generally generated good-quality embryos (normal blastocyst morphology) and positive pregnancies or generated an unusually high rate of poor morphological quality embryos. Sperm from these two groups were compared to sperm from men of known fertility and analyzed using machine-learning techniques to develop predictive algorithms. Surprisingly, this study found that predictive models based on methylation array data from these groups were highly predictive of male fertility status. In other words, IVF patients, who were not exclusively classified as having male-factor infertility, could with good accuracy be identified from men of known fertility. Interestingly, the most accurate algorithm was able to predict fertility status using a relatively low number of CpG loci, which were enriched for imprinted loci. This finding was surprising but strengthened by a concurrent study in which time to pregnancy was evaluated in young couples not presumed to be experiencing infertility and which also identified DNA methylation markers apparently predictive of fecundity (Jenkins et al. 2016b).
Additionally, hierarchical clustering was capable of identifying clusters containing IVF patients and poor embryo quality samples based on methylation array data, and a predictive algorithm was developed (Aston et al. 2015). While the methylation changes observed between these groups were not biased toward genomic regions of any particular annotation category, such as imprinted regions, these data show that global alterations in sperm methylation can be predictive of male fertility status and potentially embryo quality during IVF treatment (Aston et al. 2015). The data also imply that, as is well understood, embryogenesis includes a broad complement of gene pathways, and it is likely that poor embryogenesis is not the result of a dominant defective pathway, but rather may include a diverse set of defects in a myriad of pathways in a cohort of patients. The initial loci reported by Aston et al. were used by Abbasi et al. to develop a simplified and accurate testing platform for possible patient testing (Abbasi et al. 2018). The utility of such platforms will become apparent with further usage.
Recently, Denomme et al. studied on the epigenetic evaluation of embryos derived from male-factor (MF) infertility patients compared to controls and reported a dysregulation of DNA methylation in the embryos of the MF-derived embryos, including in genes involved in regulation of cellular metabolic processes (Denomme et al. 2018). While the overall pregnancy rates were similar for the two groups, the MF-derived embryos with altered methylation were associated with an increased miscarriage rate. In a separate study that eliminated female-factor confounders by using only couples employing donor oocytes, this group also found differences in sperm DNA methylation and miRNAs, as well as embryonic gene expression, in “good embryo quality” patients versus “poor embryo quality” patients (Denomme et al. 2017). These studies strengthen the potential use of methylation data to predict IVF outcome and fertility.
At present, several studies are underway with the intent of validating the above studies. Similarly, other aspects of reproduction and infertility are also being evaluated. For example, the role of inherited DNA methylation defects in unexplained, recurrent miscarriage is one area of intense interest (Ankolkar et al. 2012; Rotondo et al. 2012; Spinelli et al. 2019). Additionally, studies are beginning to identify abnormalities in sperm DNA methylation associated with environmental exposures, including bisphenol A (Dere et al. 2018), mercury (Lu et al. 2018), pesticides (Pallotta et al. 2019; Skinner et al. 2018), phthalates (Tian et al. 2018), vinclozolin (Beck et al. 2017), tobacco (Murphy et al. 2018b), and other chemicals (Siddeek et al. 2018). Interestingly, studies have also reported changes in the sperm methylome associated with paternal aging, including a stepwise increase in the number of abnormally methylated loci as during aging, beginning in approximately the mid-30s (Jenkins et al. 2013, 2014). This issue is of clinical relevance due to the increasing societal trend of delaying pregnancy until later paternal ages.
The concept that the sperm epigenome is altered by environmental exposures, age, and life events and decisions, coupled with the emerging understanding that epigenetic defects can be transmitted transgenerationally, suggests that in the future it may be able to screen potential fathers for sperm epigenetic abnormalities with the objective of identifying potential risks to offspring and progeny, in addition to infertility risk, and motivating preconception lifestyle changes to mitigate that risk (Jenkins et al. 2018). Such uses of the sperm epigenome are distant but may provide additional benefits to patients and offspring.
Conclusions
This chapter has provided a brief overview of the sperm epigenome and its potential use as a diagnostic tool to predict male infertility, to assess sperm competency in developing normal embryos, and possibly as a means to assess environmental and lifestyle risks. With the growth of the field in the past 10 years, it is likely that the future is bright in regard to meaningful advances that will directly benefit patients and clinicians, as well as aid society at large.
The existing data clearly show an implied mechanism for the sperm epigenome in regulating or facilitating early embryogenesis. Additionally, the studies described above clearly show associations with aberrant sperm epigenomes and diminished embryogenesis capacity. Lastly, early studies have demonstrated an ability to predict embryogenesis ability. At this time, the field awaits independent, large-scale validation studies before these technologies can be implemented in the clinic. Similarly, the sperm epigenome also provides a historical record of spermatogenesis, and numerous studies have shown that altered spermatogenesis is associated with altered sperm DNA methylation, particularly of imprinted genes. However, validation studies are also needed in this regard before sperm epigenetic testing can be used as a screen of fertility status. The use of the sperm epigenome as a toxicology tool and as a means to assess risk to progeny is a more distant goal, but very intriguing. While associations are strong, studies are needed to better understand the biology involved in DNA reprogramming in the embryo and fetus and the means of transmission to progeny. Most importantly, it will be imperative that such assays quantify risk in a clinically useful manner.
To this point, sperm DNA methylation has been the major focus of studies evaluating the possible development of diagnostic tools. This is due to cost and technical feasibility issues. However, it is important that evaluation of histone modifications and RNAs continue to be a focus, since it is likely that epigenetic pathways using these markers are of high relevance to reproduction.
References
Abbasi M, Smith AD, Swaminathan H, Sangngern P, Douglas A, Horsager A, Carrell DT, Uren PJ (2018) Establishing a stable, repeatable platform for measuring changes in sperm DNA methylation. Clin Epigenetics 10(1):119. https://doi.org/10.1186/s13148-018-0551-7
Ankolkar M et al (2012) Methylation analysis of idiopathic recurrent spontaneous miscarriage cases reveals aberrant imprinting at H19 ICR in normozoospermic individuals. Fertil Steril 98:1186–1192. https://doi.org/10.1016/j.fertnstert.2012.07.1143
Aoki VW, Liu L, Carrell DT (2005) Identification and evaluation of a novel sperm protamine abnormality in a population of infertile males. Hum Reprod 20:1298–1306. https://doi.org/10.1093/humrep/deh798
Aoki VW et al (2006a) Sperm protamine 1/protamine 2 ratios are related to in vitro fertilization pregnancy rates and predictive of fertilization ability. Fertil Steril 86:1408–1415. https://doi.org/10.1016/j.fertnstert.2006.04.024
Aoki VW, Emery BR, Liu L, Carrell DT (2006b) Protamine levels vary between individual sperm cells of infertile human males and correlate with viability and DNA integrity. J Androl 27:890–898. https://doi.org/10.2164/jandrol.106.000703
Aoki VW, Liu L, Carrell DT (2006c) A novel mechanism of protamine expression deregulation highlighted by abnormal protamine transcript retention in infertile human males with sperm protamine deficiency. Mol Hum Reprod 12:41–50. https://doi.org/10.1093/molehr/gah258
Aston KI, Punj V, Liu L, Carrell DT (2012) Genome-wide sperm deoxyribonucleic acid methylation is altered in some men with abnormal chromatin packaging or poor in vitro fertilization embryogenesis. Fertil Steril 97:285–292. https://doi.org/10.1016/j.fertnstert.2011.11.008
Aston KI et al (2015) Aberrant sperm DNA methylation predicts male fertility status and embryo quality. Fertil Steril 104:1388–1397 e1381–1385. https://doi.org/10.1016/j.fertnstert.2015.08.019
Barbosa TD et al (2015) Paternal chronic high-fat diet consumption reprogrammes the gametic epigenome and induces transgenerational inheritance of metabolic disorder. Diabetologia 58:S162–S163
Beck D, Sadler-Riggleman I, Skinner MK (2017) Generational comparisons (F1 versus F3) of vinclozolin induced epigenetic transgenerational inheritance of sperm differential DNA methylation regions (epimutations) using MeDIP-Seq. Environ Epigenet 3. https://doi.org/10.1093/eep/dvx016
Carone BR et al (2010) Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell 143:1084–1096. https://doi.org/10.1016/j.cell.2010.12.008
Carrell DT (2012) Epigenetics of the male gamete. Fertil Steril 97:267–274. https://doi.org/10.1016/j.fertnstert.2011.12.036
Carrell DT, Liu L (2001) Altered protamine 2 expression is uncommon in donors of known fertility, but common among men with poor fertilizing capacity, and may reflect other abnormalities of spermiogenesis. J Androl 22:604–610
Carrell DT, Emery BR, Hammoud S (2008) The aetiology of sperm protamine abnormalities and their potential impact on the sperm epigenome. Int J Androl 31:537–545. https://doi.org/10.1111/j.1365-2605.2008.00872.x
Chen Q et al (2016) Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 351:397–400. https://doi.org/10.1126/science.aad7977
Cho C et al (2003) Protamine 2 deficiency leads to sperm DNA damage and embryo death in mice. Biol Reprod 69:211–217. https://doi.org/10.1095/biolreprod.102.015115
Conine CC, Sun F, Song L, Rivera-Pérez JA, Rando OJ (2018) Small RNAs gained during epididymal transit of sperm are essential for embryonic development in mice. Dev Cell 46:470–480.e473. https://doi.org/10.1016/j.devcel.2018.06.024
Cox GF et al (2002) Intracytoplasmic sperm injection may increase the risk of imprinting defects. Am J Hum Genet 71:162–164. https://doi.org/10.1086/341096
Deans C, Maggert KA (2015) What do you mean, “epigenetic”? Genetics 199:887–896. https://doi.org/10.1534/genetics.114.173492
Denomme MM, McCallie BR, Parks JC, Schoolcraft WB, Katz-Jaffe MG (2017) Alterations in the sperm histone-retained epigenome are associated with unexplained male factor infertility and poor blastocyst development in donor oocyte IVF cycles. Hum Reprod 32:2443–2455. https://doi.org/10.1093/humrep/dex317
Denomme MM et al (2018) Inheritance of epigenetic dysregulation from male factor infertility has a direct impact on reproductive potential. Fertil Steril 110:419–428 e411. https://doi.org/10.1016/j.fertnstert.2018.04.004
Dere E et al (2018) Effects of continuous bisphenol A exposure from early gestation on 90day old rat testes function and sperm molecular profiles: a CLARITY-BPA consortium study. Toxicol Appl Pharmacol 347:1–9. https://doi.org/10.1016/j.taap.2018.03.021
Gannon JR, Emery BR, Jenkins TG, Carrell DT (2014) The sperm epigenome: implications for the embryo. Adv Exp Med Biol 791:53–66. https://doi.org/10.1007/978-1-4614-7783-9_4
Guibert S, Forne T, Weber M (2012) Global profiling of DNA methylation erasure in mouse primordial germ cells. Genome Res 22:633–641. https://doi.org/10.1101/gr.130997.111
Hammoud S, Liu L, Carrell DT (2009a) Protamine ratio and the level of histone retention in sperm selected from a density gradient preparation. Andrologia 41:88–94. https://doi.org/10.1111/j.1439-0272.2008.00890.x
Hammoud SS et al (2009b) Distinctive chromatin in human sperm packages genes for embryo development. Nature 460:473–478. https://doi.org/10.1038/nature08162
Hammoud SS, Purwar J, Pflueger C, Cairns BR, Carrell DT (2010) Alterations in sperm DNA methylation patterns at imprinted loci in two classes of infertility. Fertil Steril 94:1728–1733. https://doi.org/10.1016/j.fertnstert.2009.09.010
Hammoud SS et al (2011) Genome-wide analysis identifies changes in histone retention and epigenetic modifications at developmental and imprinted gene loci in the sperm of infertile men. Hum Reprod 26:2558–2569. https://doi.org/10.1093/humrep/der192
Harvey W (1651) Exercitationes de generatione animalium. Typis, London
Harvey W (1653) Anatomical exercitations concerning the generation of living creatures to which are added particular discourses of births and of conceptions, &c. (James Young, for Octavian Pulleyn)
Heijmans BT et al (2008) Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci U S A 105:17046–17049. https://doi.org/10.1073/pnas.0806560105
Holliday R (2006) Epigenetics: a historical overview. Epigenetics 1:76–80
Illum LRH, Bak ST, Lund S, Nielsen AL (2018) DNA methylation in epigenetic inheritance of metabolic diseases through the male germ line. J Mol Endocrinol 60:R39–R56. https://doi.org/10.1530/JME-17-0189
Jenkins TG, Carrell DT (2011) The paternal epigenome and embryogenesis: poising mechanisms for development. Asian J Androl 13:76–80. https://doi.org/10.1038/aja.2010.61
Jenkins TG, Aston KI, Cairns BR, Carrell DT (2013) Paternal aging and associated intraindividual alterations of global sperm 5-methylcytosine and 5-hydroxymethylcytosine levels. Fertil Steril 100:945–951. https://doi.org/10.1016/j.fertnstert.2013.05.039
Jenkins TG, Aston KI, Pflueger C, Cairns BR, Carrell DT (2014) Age-associated sperm DNA methylation alterations: possible implications in offspring disease susceptibility. PLoS Genet 10:e1004458. https://doi.org/10.1371/journal.pgen.1004458
Jenkins TG et al (2016a) Teratozoospermia and asthenozoospermia are associated with specific epigenetic signatures. Andrology 4:843–849. https://doi.org/10.1111/andr.12231
Jenkins TG et al (2016b) Decreased fecundity and sperm DNA methylation patterns. Fertil Steril 105:51–57 e51–53. https://doi.org/10.1016/j.fertnstert.2015.09.013
Jenkins TG, Aston KI, James ER, Carrell DT (2017) Sperm epigenetics in the study of male fertility, offspring health, and potential clinical applications. Syst Biol Reprod Med 63:69–76. https://doi.org/10.1080/19396368.2016.1274791
Jenkins TG, Aston KI, Carrell DT (2018) Sperm epigenetics and aging. Transl Androl Urol 7:S328–S335. https://doi.org/10.21037/tau.2018.06.10
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293:1074–1080. https://doi.org/10.1126/science.1063127
Jirtle RL, Skinner MK (2007) Environmental epigenomics and disease susceptibility. Nat Rev Genet 8:253–262. https://doi.org/10.1038/nrg2045
Jodar M et al (2015) Absence of sperm RNA elements correlates with idiopathic male infertility. Sci Transl Med 7:295re296. https://doi.org/10.1126/scitranslmed.aab1287
Karaca MZ et al (2017) Association between methylenetetrahydrofolate reductase (MTHFR) gene promoter hypermethylation and the risk of idiopathic male infertility. Andrologia 49(7). https://doi.org/10.1111/and.12698
Kobayashi H et al (2007) Aberrant DNA methylation of imprinted loci in sperm from oligospermic patients. Hum Mol Genet 16:2542–2551. https://doi.org/10.1093/hmg/ddm187
Krawetz SA et al (2011) A survey of small RNAs in human sperm. Hum Reprod 26:3401–3412. https://doi.org/10.1093/humrep/der329
Laqqan M, Hammadeh ME (2018) Aberrations in sperm DNA methylation patterns of males suffering from reduced fecundity. Andrologia 50(3). https://doi.org/10.1111/and.12913
Le Bouc Y et al (2010) Epigenetics, genomic imprinting and assisted reproductive technology. Ann Endocrinol (Paris) 71:237–238. https://doi.org/10.1016/j.ando.2010.02.004
Liu WM et al (2012) Sperm-borne microRNA-34c is required for the first cleavage division in mouse. Proc Natl Acad Sci U S A 109:490–494. https://doi.org/10.1073/pnas.1110368109
Lu Z et al (2018) Urine mercury levels correlate with DNA methylation of imprinting gene H19 in the sperm of reproductive-aged men. PLoS One 13:e0196314. https://doi.org/10.1371/journal.pone.0196314
Messerschmidt DM, Knowles BB, Solter D (2014) DNA methylation dynamics during epigenetic reprogramming in the germline and preimplantation embryos. Genes Dev 28:812–828. https://doi.org/10.1101/gad.234294.113
Murphy PJ, Wu SF, James CR, Wike CL, Cairns BR (2018a) Placeholder nucleosomes underlie germline-to-embryo DNA methylation reprogramming. Cell 172:993–1006 e1013. https://doi.org/10.1016/j.cell.2018.01.022
Murphy SK et al (2018b) Cannabinoid exposure and altered DNA methylation in rat and human sperm. Epigenetics 13:1208. https://doi.org/10.1080/15592294.2018.1554521
Nanassy L, Carrell DT (2011) Abnormal methylation of the promoter of CREM is broadly associated with male factor infertility and poor sperm quality but is improved in sperm selected by density gradient centrifugation. Fertil Steril 95:2310–2314. https://doi.org/10.1016/j.fertnstert.2011.03.096
Ng SF et al (2010) Chronic high-fat diet in fathers programs beta-cell dysfunction in female rat offspring. Nature 467:963–966. https://doi.org/10.1038/nature09491
Ostermeier GC, Dix DJ, Miller D, Khatri P, Krawetz SA (2002) Spermatozoal RNA profiles of normal fertile men. Lancet 360:772–777. https://doi.org/10.1016/S0140-6736(02)09899-9
Ostermeier GC, Miller D, Huntriss JD, Diamond MP, Krawetz SA (2004) Reproductive biology: delivering spermatozoan RNA to the oocyte. Nature 429:154. https://doi.org/10.1038/429154a
Painter RC, Roseboom TJ, Bleker OP (2005) Prenatal exposure to the Dutch famine and disease in later life: an overview. Reprod Toxicol 20:345–352. https://doi.org/10.1016/j.reprotox.2005.04.005
Pallotta MM et al (2019) In vitro exposure to CPF affects bovine sperm epigenetic gene methylation pattern and the ability of sperm to support fertilization and embryo development. Environ Mol Mutagen 60:85–95. https://doi.org/10.1002/em.22242
Pembrey ME et al (2006) Sex-specific, male-line transgenerational responses in humans. Eur J Hum Genet 14:159–166. https://doi.org/10.1038/sj.ejhg.5201538
Pessot CA et al (1989) Presence of RNA in the sperm nucleus. Biochem Biophys Res Commun 158:272–278
Reik W, Walter J (2001) Genomic imprinting: parental influence on the genome. Nat Rev Genet 2:21–32. https://doi.org/10.1038/35047554
Rotondo JC et al (2012) Methylenetetrahydrofolate reductase gene promoter hypermethylation in semen samples of infertile couples correlates with recurrent spontaneous abortion. Hum Reprod 27:3632–3638. https://doi.org/10.1093/humrep/des319
Santi D, De Vincentis S, Magnani E, Spaggiari G (2017) Impairment of sperm DNA methylation in male infertility: a meta-analytic study. Andrology 5:695–703. https://doi.org/10.1111/andr.12379
Seisenberger S et al (2012) The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol Cell 48:849–862. https://doi.org/10.1016/j.molcel.2012.11.001
Sendler E et al (2013) Stability, delivery and functions of human sperm RNAs at fertilization. Nucleic Acids Res 41:4104–4117. https://doi.org/10.1093/nar/gkt132
Sharma U et al (2016) Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science 351:391–396. https://doi.org/10.1126/science.aad6780
Sharma U et al (2018) Small RNAs are trafficked from the epididymis to developing mammalian sperm. Dev Cell 46:481–494.e486. https://doi.org/10.1016/j.devcel.2018.06.023
Siddeek B, Mauduit C, Simeoni U, Benahmed M (2018) Sperm epigenome as a marker of environmental exposure and lifestyle, at the origin of diseases inheritance. Mutat Res 778:38–44. https://doi.org/10.1016/j.mrrev.2018.09.001
Skinner MK et al (2018) Alterations in sperm DNA methylation, non-coding RNA and histone retention associate with DDT-induced epigenetic transgenerational inheritance of disease. Epigenetics Chromatin 11:8. https://doi.org/10.1186/s13072-018-0178-0
Spinelli P et al (2019) Identification of the novel Ido1 imprinted locus and its potential epigenetic role in pregnancy loss. Hum Mol Genet 28:662. https://doi.org/10.1093/hmg/ddy383
Tian M, Liu L, Zhang J, Huang Q, Shen H (2018) Positive association of low-level environmental phthalate exposure with sperm motility was mediated by DNA methylation: a pilot study. Chemosphere 220:459–467. https://doi.org/10.1016/j.chemosphere.2018.12.155
Waddington CH (1942) The epigenotype. Endeavour 1:18–20
Wu SF, Zhang H, Cairns BR (2011) Genes for embryo development are packaged in blocks of multivalent chromatin in zebrafish sperm. Genome Res 21:578–589. https://doi.org/10.1101/gr.113167.110
Zhang Y et al (2018) Dnmt2 mediates intergenerational transmission of paternally acquired metabolic disorders through sperm small non-coding RNAs. Nat Cell Biol 20:535–540. https://doi.org/10.1038/s41556-018-0087-2
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Carrell, D.T. (2019). The Sperm Epigenome: Implications for Assisted Reproductive Technologies. In: Baldi, E., Muratori, M. (eds) Genetic Damage in Human Spermatozoa. Advances in Experimental Medicine and Biology, vol 1166. Springer, Cham. https://doi.org/10.1007/978-3-030-21664-1_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-21664-1_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-21663-4
Online ISBN: 978-3-030-21664-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)