Abstract
Lupins provide an insightful model for plant domestication with five species domesticated over a wide range of time and geography. The most intensively studied species is narrow-leafed lupin, a twentieth-century domesticate where the addition of each successive domestication trait was documented in the scientific literature. Foundational to the advances made in our understanding of lupin domestication was the availability of excellent genetic resources: Well-annotated wild seed collections, published pedigrees of Australian narrow-leafed lupin cultivars and a suite of wild × domesticated cross populations. Rapid developments in genomic technologies culminating in the reference genome for narrow-leafed lupin have greatly increased our understanding of the origins of domesticated lupins, how diversity has been profoundly affected and the molecular control of domestication genes. This chapter provides an overview of our current understanding of lupin domestication and how this knowledge can equip lupin breeders to create more diverse and productive cultivars.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others

8.1 Background
The legume genus Lupinus is special in many respects, not least its extraordinary history of domestication of several species across a wide range of time and geography. Lupinus encompasses around 275 species distributed across the Mediterranean region and North Africa (‘Old World’ lupins) and the Americas (‘New World’ lupins) (Hughes and Eastwood 2006). While Old World lupins represent just 13 of those 275 species, four Old World species can be considered fully domesticated, while several others show signs of historic cultivation and selection (Swiecicki and Swiecicki 1995). The oldest fully domesticated species is white lupin (L. albus L.). There is clear evidence that white lupin was cultivated in Egypt by 300 BC and possibly as early as 2000–1000 BC (Wolko et al. 2011). Three further Old World species are more modern domesticates: Narrow-leafed lupin (L. angustifolius L.), yellow lupin (L. luteus L.) and West Australian blue lupin or sandplain lupin (L. cosentinii Guss.; often mistaken for L. digitatus Forsk.). Despite the majority of Lupinus species being from the New World, just one—Andean lupin (L. mutabilis Sweet)—has been domesticated, likely 3000–4000 years ago.
Today, the most widely cultivated species are L. angustifolius and L. albus, while L. luteus and L. mutabilis are niche crops, and L. cosentinii is not currently cultivated to our knowledge. The production of lupin seeds as an agricultural product still occurs mainly in Australia but also in parts of Europe, Africa and South America. Although production has fluctuated over the last 20 years, over a million tonnes are produced every year. In 2017, the largest producers were Australia (1,031,425 t), Poland (168,678 t) and the Russian Federation (161,680 tonnes) (FAO 2017).
It is easy to understand why lupins attracted the attention of early farmer and hunter-gatherers: their seeds are large and highly nutritious with protein contents of around 30–45%, comparable to soybean (Lucas et al. 2015). Wild types and landraces are high in bitter quinolizidine alkaloids but humans quickly learned to remove most of the alkaloids by soaking and rinsing in water. As a snack food some residual bitterness in the lupin seeds provides a pleasant, distinctive flavour, which remains popular in Spain (‘altramuces’), Italy (‘lupini’), Ethiopia (‘gibto’), Egypt and Sudan (‘termes’) and South America (‘tarwi’ or ‘chocho’). Naturally low alkaloid, ‘sweet’ cultivars have been developed, which now represent most of the lupins cultivated worldwide. Sweet cultivars are grown primarily for grain for animal feed but increasingly as a healthy adjunct to the human diet in breads and pastries, or to provide a gluten-free alternative to wheat flour (Gresta et al. 2017).
The Lupinus genus has been the subject of genomic studies since the late 1990s (Wolko et al. 2011). Most extensive genomic research has focused on the most widely cultivated species L. angustifolius, which culminated in the publication of a high-quality reference genome (Hane et al. 2017). This chapter explores how the genomic resources in L. angustifolius are enabling a greater understanding of lupin domestication, which is becoming an increasingly insightful model for crop domestication and species evolution more generally.
8.2 Multiple Lupin Domestications Spanning Time and Space
Domestication can be defined as the taming of wild plants and animals to become more productive for humans, enabling the development of trade specializations and burgeoning human populations (Diamond 2002). It involved progressively accumulating domestication traits that made the plants increasingly more useful and productive to people. The founding father of the discipline, Nikolai Vavilov, described this as the ‘homologous series in inherited variation’ (Vavilov 1951) and which is now known as the ‘domestication syndrome’ (Hammer 1984). These traits included reduced fruit dehiscence, increased apical dominance, removal of seed dormancy, altered time of flowering and maturity, and reduced bitter compounds in seeds (Doebley et al. 2006). The domestication of lupin occurred several times throughout human history and across wide geographical regions (Gladstones 1998).
8.2.1 Ancient Lupin Domestication
The first records that suggest lupin had been adopted and adapted for use within human culture are from Greek and Roman texts. However, it is believed that lupins had been cultivated around the Mediterranean much earlier, having spread from the place of initial cultivation, Egypt (Gladstones 1974). Archaeological remains of L. albus have been found in Greece and Cyprus dating from around the Bronze Age (Zohary et al. 2012). It is believed that it was white lupin that the ancient Greek writers Hippocrates (400–356 B.C) and Theophrastus (372–288 B.C) record, discussing soil type and harvesting requirements for the crop. The Roman writer, the elder Cato (243–149 B.C) referred to its use as a cattle feed and as a green manure, and the poet Virgil writes of its use in crop rotation with cereals (Hondelmann 1984). It was at some point during this early cultivation that the initial domestication of white lupin would have occurred, selecting the permeable seed coats to aid even germination and non-shattering pods to reduce wastage during harvest (Gladstones 1970). The history of lupin use and domestication in the ‘New World’ is harder to follow as there are fewer records. The early cultivation of L. mutabilis in the Andes of South America has been dated to around 700 B.C. (Hondelmann 1984). Later, it was the Incas who used lupin extensively in crop rotations until the Spanish conquest in the early sixteenth century (Wolko et al. 2011). Similar traits as those in L. albus would have been selected for in the early domestication and adoption of L. mutabilis into South American agriculture, and there are no true wild lines remaining without these domestication traits (Eastwood and Hughes 2008).
Lupin cultivation and domestication underwent a renaissance in the eighteenth century. This was by royal decree in Prussia as a means of soil improvement using L. albus. This species did not thrive in the Northern European climate and was replaced successfully with L. luteus, which was used for seed production for animal feed as well as soil improvement in crop rotations (Wolko et al. 2011; Hondelmann 1984). L. angustifolius was also introduced to Europe over the following 100 years and along with L. luteus was taken up by farmers in Northern Europe as both species had good frost tolerance and suitable maturity timing compared to L. albus (Wolko et al. 2011).
8.2.2 Modern Lupin Domestication
The driver for the modern era domestication of lupins was to find sweet varieties which were low in alkaloids (Hondelmann 1984). Up to this point, all lupins were bitter and the consumption of seed was only possible after soaking the seeds for a period of time in water. If sweet varieties could be developed, then it could open up a greater use of the crop for animal feed as well as for humans without the risk of toxicity. The first recorded discovery of sweet plants for both L. luteus and L. angustifolius was in the late 1920s by German scientist and plant breeder Dr. Reinhold von Sengbusch. This was only possible after the development of a simple, high-throughput assay to detect the presence of alkaloids (Hondelmann 1984; Gladstones 1970). The identification of sweet types of L. albus was subsequently achieved and this, along with breeding for early maturity, was carried out in 1930–1940s in northern Europe leading to varieties such as Nahrquell being released post-war in West Germany (Gladstones 1970).
At the same time, other key seed traits—indehiscence, water permeability (soft seededness) and white colouring—were included in the selection and proved successful in L. luteus. Lupin breeding also began in Poland in the 1930s, focused mainly on L. luteus and L. angustifolius but it was not until after the Second World War that interest for lupins grew, particularly in the Mediterranean, Australasia and South Africa. In the 1950s, a breeding programme was established in Western Australia for L. angustifolius where the full domestication of this species was achieved by incorporating domestication genes from several sources (Gladstones 1977) (Table 8.1). A L. angustifolius breeding programme was also set-up in the USA in the 1940s and continued to the 1960s with advances made in disease resistance, particularly to anthracnose and grey leaf mould. These varieties and knowledge were then combined into the Australian breeding programme (Gladstones 1977). Yield improvements then became the focus for the breeding efforts as the domestication process had been completed.
L. mutabilis was another lupin species for which von Sengbusch produced sweet types in the 1930s, as other domestication traits such as non-shattering were already in place. Mutation breeding for sweetness using ethyl methanesulfonate (EMS) was also attempted later on, lowering alkaloid levels to around 0.2–0.3% (Clements et al. 2008; Williams et al. 1984). However, it was breeding work based on a natural mutant by von Baer and Gross in Chile that led to the production of an extremely low-alkaloid cultivar, Inti (Gross et al. 1988). Breeding was then continued in Western Australia from 1999, focusing on flowering time and male sterility. The Australian Lupin Collection containing a number of L. mutabilis accessions with differing characteristics provided additional traits, which could be combined into breeding programmes for the continual improvement of L. mutabilis cultivars (Clements et al. 2008).
L. cosentinii domestication and breeding was undertaken by Gladstones in the 1950s, around a century after it had been initially introduced to the country for flour production. It was a good choice for domestication as it had naturalized well into the Western Australian environment and thrived on infertile, sandy soils as well as having some drought tolerance (Gladstones 1970). By this time, it was used mainly for soil improvement and sheep feed. Domestication traits that were targeted for L. cosentinii improvement were low-alkaloid seed (from artificial mutagenesis), non-shattering pods, early flowering and soft seededness, from natural mutations (Gladstones and Francis 1965; Gladstones and Hill 1969; Gladstones 1958, 1967). All of these were incorporated into the cultivar ‘Erregulla’. However, it was not widely taken up due to problems with deformed seeds and reduced seed filling (Cowling et al. 1998). While this species is not currently grown to any appreciable extent, it may provide a useful legume rotation crop in a drying climate, as there is anecdotal evidence of drought tolerance in this species (Gladstones 1970).
8.3 Genetic and Genomic Resources Supporting Domestication Research in Lupin
Lupins provide an excellent model for understanding domestication genes due to the multiple independent domestications across wide spatial (Europe, South America and Australia) and temporal (from 4000 to 50 years ago) ranges. Two crucial features supporting domestication studies are the availability of extensive and well-annotated seed collections made for Lupinus species, and the relatively small diploid genomes (2C = 1.16–2.44 pg, equivalent to around 600–1200 Mbp per haploid genome; (Naganowska et al. 2003)), which makes them tractable to genomic analyses.
8.3.1 Genetic Resources
Lupin breeders and researchers are blessed with excellent germplasm resources, especially for the domesticated species, L. angustifolius, L. albus, L. luteus and L. mutabilis. The value of seed collections is not only related to the number of accessions (estimated to be over 36,000 in the largest 40 collections (Wolko et al. 2011)), but also to the geographic spread and annotated passport data (which is very good for a large proportion of accessions). The largest and best characterized collection is located in Perth, Australia, the majority of which is currently being transferred to the Australian Grain Genebank in Horsham, Australia for long-term conservation (Sally Norton, pers. comm.). These international collections cover both Old World and New World species, wild and landrace types as well as many breeding lines. These seed collections are an invaluable resource for understanding plant domestication as well as a source of genetic variation for important agronomic traits such as abiotic and biotic stress tolerance (Berger et al. 2017).
Recombinant inbred line (RIL) populations have been created for L. angustifolius, L. albus and L. luteus (Berger et al. 2013), which are valuable for investigating the genetic basis of domestication traits. RIL populations are produced by crossing contrasting parental lines to produce an F1 hybrid, which is self-pollinated to produce a large F2 population. Each F2 individual is then subjected to inbreeding by a process of single seed descent to generate an inbred population (typically F8 generation) in which traits of interest have segregated. The value of RIL populations is the capacity to generate unlimited seed for replicated phenotyping and sharing with research collaborators. The main reference RIL populations for L. angustifolius, L. albus and L. luteus were generated from wide crosses at the Department of Primary Industries and Regional Development (DPIRD, Perth, Australia) (Wolko et al. 2011; Berger et al. 2013). All three populations segregate for domestication traits as they were generated by crossing a domesticated parent with a wild (L. angustifolius and L. luteus) or partially domesticated landrace (L. albus) parent.
The L. angustifolius RIL population developed from a cross between 83A:476 (an Australian breeding line) and P27255 (a Moroccan wild accession) has been particularly instrumental for understanding lupin domestication, through the provision of genetic maps to locate domestication genes (Boersma et al. 2005; Nelson et al. 2006, 2010; Kroc et al. 2014; Kamphuis et al. 2015; Zhou et al. 2018) and ultimately as the genetic backbone for the first lupin reference genome (Hane et al. 2017), a key resource for domestication gene discovery. The L. albus and L. luteus RIL populations are now being used to further our understanding of domestication in those species (Matthew N. Nelson et al. unpublished data).
8.3.2 Genomic Resources
As the most widely grown lupin species, genomic resources are most advanced for L. angustifolius. Starting from humble beginnings with protein isozyme markers (Wolko and Weeden 1989), genomic resources for L. angustifolius have grown in scale and complexity as technology has evolved. Transcriptomic (that is, sets of expressed gene sequences) resource development began with cloning and sequencing genes expressed in seed tissues (Nelson et al. 2006), then exploded in scale with the advent of next generation sequencing (NGS) platforms, resulting in comprehensive transcriptomes for seed, leaf, flower, pod, stem and root organs (Kamphuis et al. 2015; Foley et al. 2011, 2015; Cannon et al. 2015; Kroc et al. 2019; Yang et al. 2017). For a detailed review of lupin transcriptome studies, see Chap. 5. Two genomic bacterial artificial chromosomes (BAC) libraries based on cultivars Sonet (from Poland; (Kasprzak et al. 2006)) and Tanjil (from Australia; (Gao et al. 2011)) proved to be useful tools for gene discovery before the availability of whole genome surveys (based on Tanjil; (Kamphuis et al. 2015, Yang et al. 2013)) and then comprehensive Tanjil genome assemblies (Zhou et al. 2018; Hane et al. 2017). The Tanjil genome assembly is currently being improved with long sequence-read technology, and a pan-genome is also being developed that will represent species-wide genome diversity through incorporating portions of the L. angustifolius genome that are absent in Tanjil but present in domesticated and wild accessions (Karam B. Singh, pers. comm.; Sect. 3.6.1).
While not yet as comprehensive as for L. angustifolius, genomic resources have also rapidly developed for other lupin species (see Chaps. 3 and 5). Indeed, the first lupin transcriptomic resources were generated to explore L. albus cluster roots (Uhde-Stone et al. 2003), which was followed up later with richer next generation datasets (O'Rourke et al. 2013; Wang et al. 2014; Secco et al. 2014). Other L. albus transcriptomes were produced to explore seed storage proteins and for genetic marker development (Foley et al. 2015; Książkiewicz et al. 2017). Transcriptome sequences were used for marker discovery and exploring organ abscission in yellow lupin (Parra-González et al. 2012; Glazinska et al. 2017), and its chloroplast genome was sequenced (Martin et al. 2014). In the broadest sampling reported yet, Nevado et al. (2016) sequenced transcriptomes from 55 New World lupin species in order to understand adaptive evolution of rapidly speciating lupins in the New World. One of those species—L. polyphyllus—had been sequenced earlier by Cannon et al. (2015) as part of the 1000 Plants (Leebens-Mack et al. 2019).
High-quality reference genomes are being prepared using long sequence-read technologies and optical mapping for L. albus (Hufnagel et al. 2019) and L. luteus (Joshua Udall, pers. comm.). These are expected to be as useful as the L. angustifolius genome has already proven to be. Taken together, these genomic resources provide powerful tools for lupin domestication gene discovery.
8.4 Lupin Domestication Gene Discovery
8.4.1 Lupinus angustifolius
Identifying the genes controlling domestication traits is important for basic understanding of plant evolution but also for improving crops through plant breeding. One of the key constraints in accessing trait diversity in wild relatives for breeding purposes is the poor agronomic performance of early generations of progeny from crosses between breeding lines and wild relatives (Cowling et al. 2009). The availability of diagnostic molecular markers based on domestication genes transforms the speed and efficiency of the conversion of wild material into a suitable domesticated background in which the agronomic value of wild alleles can be measured. Understanding how domestication genes operate may also help to refine and improve the domestication genes themselves, either through prospecting for natural allelic diversity in those genes or by biotechnological intervention through transgenics or targeted mutation. For example, a modified set of phenology genes could be used to expand the adaptation of crops to new or changing climatic regions (Mousavi‐Derazmahalleh et al. 2019; Taylor et al. 2019).
Our most advanced understanding of lupin domestication genes comes from studies of L. angustifolius. This twentieth-century domestication is special in that each event was recorded at the time in the scientific literature (Table 8.1). There are five domestication traits controlled by six major genes: soft seededness (mollis), seed indehiscence (lentus and tardus), low alkaloid (iucundus), early flowering through removal of vernalization requirement (Ku and Julius) and white flower/green vegetative organ pigmentation (leucospermus) as a marker for domestication (wild types having blue flowers and a purple or red tinge throughout the vegetative organs) (Nelson et al. 2006; Taylor et al. 2019). Several studies have mapped each of the six domestication genes to the 83A:476 x P27255 reference genetic map with increasing resolution as marker technology improved and the size of the RIL population expanded (Boersma et al. 2005, 2009; Hane et al. 2017; Kamphuis et al. 2015; Kroc et al. 2014; Nelson et al. 2006, 2010; Zhou et al. 2018). These studies shed light on the chromosomal location of domestication genes but fell short of identifying the causal genes underlying the domestication traits.
The first clue about the identity of a lupin domestication gene was found by Kroc et al. (2014). Alignment of the L. angustifolius genetic map to the genome of the model legume Medicago truncatula revealed a cluster of three homologues of the flowering time gene, FT, on Chromosome 7 of M. truncatula at the equivalent map position as the Ku locus in L. angustifolius. Kroc et al. (2014) developed FT gene-based markers and mapped them back into L. angustifolius. One of the FT markers (FTc) mapped precisely to the Ku locus. This lead was followed up by Nelson et al. (2017) who were able to confirm that the FT homologue LanFTc1 not only mapped perfectly to the Ku locus but a 1.4 kb deletion in its promoter region was perfectly correlated with vernalization responsiveness in a panel of 216 wild and domesticated accessions of L. angustifolius. Of four FT homologues found in the L. angustifolius, only LanFTc1 showed elevated gene expression across a range of organ types in response to vernalization treatment in the vernalization responsive accession P27255. Taylor et al. (2019) were then able to demonstrate conclusively that the 1.4 kb deletion was the causal variant responsible for the loss of vernalization responsiveness, presumably due to a loss of regulatory sequence(s) that represses LanFTc1 expression. This explanation was supported by the discovery of a smaller, partly overlapping 1.2 kb deletion in the LanFTc1 promoter region of a wild accession from Israel that showed an intermediate flowering time phenotype. This discovery offers the exciting prospect of an allelic series of LanFTc1 that can be used as a simple breeding tool for targeting lupin varieties to specific climatic regions and flowering time in lupins (see Chap. 9 for more detailed discussion).
The low-alkaloid trait is another important target for gene identification for breeding in all lupin crop species. Quinolizidine alkaloids (QA) are responsible for the bitterness found in the Lupinus genus but the specific QAs differ between each of the species. In L. angustifolius, the major QA is lupanine (Frick et al. 2017; Wink et al. 1995). While several low-alkaloid genes have been discovered in L. angustifolius, only one is used in cultivars: iucundus (Table 8.1). It was first discovered by Von Sengbusch in 1928 and found to be a single recessive gene (Hackbarth 1957; Von Sengbusch 1942). However, it was not until the development of genetic tools that the iucundus region could be explored and the alkaloid biosynthesis pathway further understood. Li et al. (2011) identified markers linked to iucundus, which could be used for marker-assisted selection in wild × domesticated introgressive crossing programmes for broadening genetic diversity in breeding pools. Even more useful would be a perfectly predictive marker based on the causal gene mutation underlying iucundus. The iucundus gene was mapped to a 746 Kb region on chromosome NLL-07 (Hane et al. 2017). Kroc et al. (2019) used a transcriptomic approach to identify a strong candidate gene for iucundus in this interval: RAP2-7, an ethylene responsive transcription factor. A less promising candidate gene in the same region could not be fully ruled out: DHDPS, a 4-hydroxytetrahydrodipicolinate synthase gene. Further validation work will be required to confirm the causal mutation underlying iucundus. Three other alkaloid biosynthesis genes genetically unlinked to iucundus have been identified in L. angustifolius. The first step in the alkaloid biosynthesis pathway was found to be a lysine decarboxylase (LDC; Bunsupa et al. (2012)) and more recently the second step was identified as a copper amine oxidase (CAO; Yang et al. (2017)). A third gene, which role is not yet fully understood, is an acyl transferase (LaAT; Bunsupa et al. (2011)). The quinolizidine biosynthetic pathway has yet to be elucidated in any species, but most progress achieved to date has been in L. angustifolius, which functions as a model for other species. In this regard there has been recent progress made on the genetic factors affecting the biosynthetic pathway and how it responds to some biotic (Frick et al. 2019) and abiotic stresses (Frick et al. 2018) in L. angustifolius.
8.4.2 Lupinus albus
The reference mapping population for L. albus (Kiev Mutant x P27174 RIL population) segregates for just two domestication traits: low alkaloid (controlled by the pauper locus) and early flowering. The first genetic map for L. albus mapped at low resolution the pauper locus and two quantitative trait loci (QTL) for flowering time (Phan et al. 2007). This map was modestly improved by Vipin et al. (2013) although this did not provide more insight into the domestication traits. More recently, Książkiewicz et al. (2017) used genotyping by sequencing (GBS) in the same RIL population to generate a much improved, high-resolution map. They located pauper in a well-defined interval on linkage group ALB18 and identified a candidate gene residing in that region—LaAT, a gene previously identified in L. angustifolius by Bunsupa et al. (2011). The two flowering time QTL previously identified by Phan et al. (2007) were confirmed and furthermore were demonstrated to be involved in the vernalization responsive (Książkiewicz et al. 2017). One of these QTL may respond to the previously reported brevis locus (Gladstones 1970) but this has yet to be confirmed. An additional weak QTL was identified, which was not vernalization related. The improved L. albus genetic map was aligned to the L. angustifolius reference genome but interestingly none of the mapped loci corresponded to the positions of the equivalent iucundus and Ku loci in L. angustifolius. However, the L. angustifolius genome provided candidate genes for the flowering time QTLs, as subsequently described by Rychel et al. (2019). This example illustrates both the value and limitations of the L. angustifolius reference genome sequence for domestication gene research in other legume species. The reference genome for L. albus based on the cultivar Amiga (Hufnagel et al. 2019) is helping identify the genes underlying pauper and other low-alkaloid mutant loci in an international collaboration between France, Denmark, Poland, UK and Australia (Nelson et al. unpublished data).
8.4.3 Lupinus luteus
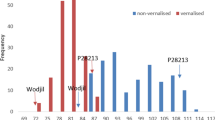
The reference RIL population for L. luteus was developed from a cross between the Australian cultivar Wodjil (a selection from the Polish cultivar Teo) (French et al. 2001) and P28213 (wild accession from the Azores) (Iqbal et al. 2019). This population segregates for the complete suite of domestication traits: soft seededness, seed indehiscence, vernalization responsiveness in flowering, alkaloid content and flower colour (yellow versus orange). The first map for L. luteus was recently released (Iqbal et al. 2019) and analysis of domestication traits is underway (see Chap. 11). Domestication gene discovery will be greatly facilitated by the availability of a reference genome, which is currently under development (Joshua Udall, pers. comm.).
8.4.4 Other Lupin Species
To our knowledge, little progress has been made in other lupin species to identify domestication genes. Foundational resources such as RIL populations should be developed between wild and domesticated accessions of both L. mutabilis and L. cosentinii. Mining of available transcriptomic datasets (see above) may provide some initial leads to follow-up in more comprehensive experiments.
8.5 Genetic Consequences of Domestication on Genome Diversity
The domestication of grain crops involves a series of population bottlenecks as new domestication alleles undergo extreme selection pressure (Doebley et al. 2006). This leads to a reduction in genetic diversity, which takes time to recover through gene flow from wild populations and spontaneous mutations. It is therefore to be expected that the genetic diversity of a young, twentieth-century domesticate such as L. angustifolius will have very depleted diversity compared to its wild ancestors. This was indeed found to be the case in a diversity analysis of 1,248 wild and 95 domesticated accessions using low-resolution Diversity Arrays Technology (DArT) genotyping (Berger et al. 2012). Figure 8.1 graphically illustrates the small portion of diversity captured in Australian and European cultivars compared to wild accessions collected across the Mediterranean Basin. This highlighted the need to understand where useful genetic diversity can be found among wild accessions (Berger et al. 2013).

Redrawn from data presented by Berger et al. (2012)
Domesticated cultivars of L. angustifolius contain a small proportion of species diversity. This multidimensional scaling plot was based on diversity measured at 137 DArT marker loci in 1,248 wild (black crosses) and 95 domesticated (Australian varieties represented by red circles and European varieties represented by blue triangles) accessions.
A detailed analysis of 142 wild accessions using high-resolution single nucleotide polymorphism (SNP) genotyping revealed that accessions from the western Mediterranean region were more diverse and that there had been an historic eastward migration during which there was a shift in phenological adaptation to warmer, lower rainfall environments (Mousavi-Derazmahalleh et al. 2018a). This provides valuable guidance for lupin breeders to identify untapped sources of genetic and adaptive diversity for lupin improvement. Mousavi-Derazmahalleh et al. (2018b) went further to demonstrate that the western Mediterranean region provided the founder populations for the domestication of lupin, which had been suspected previously based on morphological observations (Gladstones 1998). Another important finding was the much higher linkage disequilibrium evidence in domesticated compared to wild accessions, meaning that plant breeder efforts to accumulate beneficial alleles will be hampered by unwanted linkage to unfavourable alleles (Mousavi-Derazmahalleh et al. 2018b). Only by introducing wild diversity into breeding programmes will such unwanted linkages be broken up over time. Interestingly, a search for footprints of selection around domestication trait loci proved inconclusive, which may have been due to the recentness of L. angustifolius domestication.
Less is known about the impact of domestication on the genome diversity of other lupin crop species. Gilbert et al. (1999) investigated the genetic diversity present in 40 L. albus accessions using ISSR-PCR. The small sample size, the repeatability of the marker technology limitations and lack of useful passport information accompanying accessions severely limited the conclusions that could be drawn from this study. In a more comprehensive study of 94 landrace and cultivar accessions, Raman et al. (2008, 2014) found that L. albus landraces clustered separately from modern cultivars and that within landraces, Ethiopian landraces were the most distinct. Annicchiarico et al. (2010) investigated agronomic and phenological diversity in a more globally representative collection of L. albus landraces. Current work is underway to extend this work using high-resolution genotyping (Paolo Annicchiarico, pers. comm.). Iqbal et al. (2012) used AFLPs to investigate diversity, population structure and linkage disequilibrium. They found that there was some clustering among the accessions, but this could not be related to geographic origin due to lack of information and the probable high rate of transfer of germplasm across the world. Their findings also showed a weak population structure and a low level of linkage disequilibrium, which can be helpful for follow on experiments such as association mapping. In more focused analyses, Atnaf et al. (2015) and Atnaf et al. (2017), explored agronomic, phenological and low-resolution marker diversity in Ethiopian landraces. So far, no study has included wild accessions, which can now only be found in Greece (known as graecus types; (Gladstones 1998)). Currently, we are investigating molecular and phenological diversity in a large global collection of wild, landrace and cultivar accessions from 15 countries, which we believe will provide insights into the origin and genetic consequences of L. albus domestication (M. Nelson, unpublished data).
8.6 Closing Remarks
The genomics revolution has provided powerful new tools to answer basic questions about crop domestication, an insightful model for species evolution. The reference genome sequence of L. angustifolius provides a valuable resource for identifying domestication genes and understanding the effects of domestication on genome-wide diversity in lupin crop species. These discoveries provide the knowledge and the genetic tools needed by lupin breeders and pre-breeders to introduce much-needed genetic and adaptive diversity into lupin crops.
References
Annicchiarico P, Harzic N, Carroni AM (2010) Adaptation, diversity, and exploitation of global white lupin (Lupinus albus L.) landrace genetic resources. Field Crops Res 119:114–124
Atnaf M, Tesfaye K, Dagne K, Wegari D (2015) Extent and pattern of genetic diversity in Ethiopian white lupin landraces for agronomical and phenological traits. Afr Crop Sci J 23:327–341
Atnaf M, Yao N, Martina K, Dagne K, Wegary D, Tesfaye K (2017) Molecular genetic diversity and population structure of Ethiopian white lupin landraces: implications for breeding and conservation. PLoS ONE 12:e0188696
Berger JD, Buirchell BJ, Luckett DJ, Nelson MN (2012) Domestication bottlenecks limit genetic diversity and constrain adaptation in narrow-leafed lupin (Lupinus angustifolius L.). Theor Appl Genet 124:637–652
Berger JD, Clements JC, Nelson MN, Kamphuis LG, Singh KB, Buirchell B (2013) The essential role of genetic resources in narrow-leafed lupin improvement. Crop Pasture Sci 64:361–373
Berger JD, Shrestha D, Ludwig C (2017) Reproductive strategies in Mediterranean legumes: trade-offs between phenology, seed size and vigor within and between wild and domesticated Lupinus species collected along aridity gradients. Front Plant Sci 8:548
Boersma JG, Pallotta M, Li C, Buirchell BJ, Sivasithamparam K, Yang H (2005) Construction of a genetic linkage map using MFLP and identification of molecular markers linked to domestication genes in narrow-leafed lupin (Lupinus angustifolius L.). Cell Mol Biol Lett 10:331–344
Boersma J, Nelson M, Sivasithamparam K, Yang H (2009) Development of sequence-specific PCR markers linked to the Tardus gene that reduces pod shattering in narrow-leafed lupin (Lupinus angustifolius L.). Mol Breed 23:259–267
Bunsupa S, Okada T, Saito K, Yamazaki M (2011) An acyltransferase-like gene obtained by differential gene expression profiles of quinolizidine alkaloid-producing and nonproducing cultivars of Lupinus angustifolius. Plant Biotechnol 28:89–94
Bunsupa S, Katayama K, Ikeura E, Oikawa A, Toyooka K, Saito K, Yamazaki M (2012) Lysine decarboxylase catalyzes the first step of quinolizidine alkaloid biosynthesis and coevolved with alkaloid production in Leguminosae. Plant Cell 24:1202–1216
Cannon SB, McKain MR, Harkess A, Nelson MN, Dash S, Deyholos MK, Peng Y, Joyce B, Stewart CN, Rolf M, Kutchan T, Tan X, Chen C, Zhang Y, Carpenter E, Wong GK-S, Doyle JJ, Leebens-Mack J (2015) Multiple polyploidy events in the early radiation of nodulating and nonnodulating legumes. Mol Biol Evol 32:193–210
Clements J, Sweetingham M, Smith L, Francis G, Thomas G, Sipsas S (2008) Crop improvement in Lupinus mutabilis for Australian agriculture-progress and prospects. Lupins for health and wealth. In: Proceedings of the 12th international lupin conference, Fremantle, Western Australia. International Lupin Association, pp 244–250, 14–18 Sept 2008
Cowling WA, Huyghe C, Święcicki W (1998) Lupin breeding. In: Gladstones JS, Atkins CA, Hamblin J (eds) Lupins as crop plants: biology, production, and utilization. CAB International, Wallingford, UK
Cowling WA, Buirchell BJ, Falk DE (2009) A model for introducing novel genetic diversity from wild relatives into elite crop populations. Crop Pasture Sci 60:1009–1015
Diamond J (2002) Evolution, consequences and future of plant and animal domestication. Nature 418:700–707
Doebley JF, Gaut BS, Smith BD (2006) The molecular genetics of crop domestication. Cell 127:1309–1321
Eastwood RJ, Hughes CE (2008) Origins of domestication of Lupinus mutabilis in the Andes. Lupins for health and wealth. In: Proceedings of the 12th international lupin conference, pp 14–18
FAO (2017) FAOSTAT: crop production data. Food and Agriculture Organisation of the United Nations. https://www.fao.org/faostat/en/#data/QC. Accessed Dec 2018
Foley RC, Gao LL, Spriggs A, Soo LY, Goggin DE, Smith PM, Atkins CA, Singh KB (2011) Identification and characterisation of seed storage protein transcripts from Lupinus angustifolius. BMC Plant Biol 11:59
Foley RC, Jimenez-Lopez JC, Kamphuis LG, Hane JK, Melser S, Singh KB (2015) Analysis of conglutin seed storage proteins across lupin species using transcriptomic, protein and comparative genomic approaches. BMC Plant Biol 15:106
Forbes I, Wells HD (1968) Hard and soft seededness in Blue Lupine, Lupinus angustifolius L.: inheritance and phenotype classification 1. Crop Sci 8:195–197
French R, Sweetingham M, Shea G (2001) A comparison of the adaptation of yellow lupin (Lupinus luteus L.) and narrow-leafed lupin (L. angustifolius L.) to acid sandplain soils in low rainfall agricultural areas of Western Australia. Aust J Agric Res 52:945–954
Frick KM, Kamphuis LG, Siddique KH, Singh KB, Foley RC (2017) Quinolizidine alkaloid biosynthesis in lupins and prospects for grain quality improvement. Front Plant Sci 8:87
Frick KM, Foley RC, Kamphuis LG, Siddique KHM, Garg G, Singh KB (2018) Characterisation of the genetic factors affecting quinolizidine alkaloid biosynthesis and its response to abiotic stress in narrow-leafed lupin (Lupinus angustifolius L.). Plant Cell Environ 41:2155–2168
Frick KM, Foley R, Siddique KHM, Singh KB, Kamphuis LG (2019) The role of jasmonate signalling in quinolizidine alkaloid biosynthesis, wounding and aphid predation response in narrow-leafed lupin. Funct Plant Biol 46:443–454
Gao LL, Hane JK, Kamphuis LG, Foley R, Shi BJ, Atkins CA, Singh KB (2011) Development of genomic resources for the narrow-leafed lupin (Lupinus angustifolius): construction of a bacterial artificial chromosome (BAC) library and BAC-end sequencing. BMC Genomics 12:521
Gilbert J, Lewis R, Wilkinson M, Caligari P (1999) Developing an appropriate strategy to assess genetic variability in plant germplasm collections. Theor Appl Genet 98:1125–1131
Gladstones J (1958) Induction of mutation in the west Australian blue lupin (Lupinus digitatus Forsk.) by X-irradiation. Aust J Agric Res 9:473–482
Gladstones J (1967) Selection for economic characters in Lupinus angustifolius and L. digitatus. Aust J Exp Agric 7:360–366
Gladstones JS (1970) Lupins as crop plants. Field Crop Abstr 23:123–148
Gladstones JS (1974) Lupins of the Mediterranean region and Africa. Western Australia Department of Agriculture. Technical Bulletin
Gladstones JS (1977) The narrow-leafed lupin in Western Australia (Lupinus angustifolius L.). Western Australian Department of Agriculture
Gladstones J (1998) Distribution, origin, taxonomy, history and importance. In: Gladstones JS, Atkins CA, Hamblin J (eds) Lupins as crop plants: biology, production and utilization. CAB International, Wallingford, UK
Gladstones J, Francis C (1965) Low-alkaloid mutants of Lupinus digitatus Forsk. Nature 207:553
Gladstones J, Hill G (1969) Selection for economic characters in Lupinus angustifolius and L. digitatus. 2. Time of flowering. Aust J Exp Agric 9:213–220
Glazinska P, Wojciechowski W, Kulasek M, Glinkowski W, Marciniak K, Klajn N, Kesy J, Kopcewicz J (2017) De novo Transcriptome profiling of flowers, flower pedicels and pods of Lupinus luteus (yellow lupine) reveals complex expression changes during organ abscission. Front Plant Sci 8:641
Gresta F, Wink M, Prins U, Abberton M, Capraro J, Scarafoni A, Hill G (2017) Lupins in European cropping systems. In: Murphy-Bokern D, Stoddard FL, Watson CA (eds) Legumes in cropping systems. The Centre for Agriculture and Bioscience International
Gross R, von Baer E, Koch F, Marquard R, Trugo L, Wink M (1988) Chemical composition of a new variety of the Andean lupin (Lupinus mutabilis cv. Inti) with low-alkaloid content. J Food Compos Anal 1:353–361
Hackbarth J (1957) Die Gene der Lupinenarten. II. Schmalblättrige Lupine (Lupinus angustifolius L.). Z Pflanzenzüchl 37:81–95
Hackbarth J, Troll H-J (1959) Lupinen als Körnerleguminosen und Futterpflanzen. Handbuch der Pflanzenzüchtung 2:1–51
Hallqvist C (1921) The inheritance of the flower coloour and the seed colour in Lupinus angustifolius. Hereditas 2:299–363
Hammer K (1984) Das Domestikationssyndrom. Die Kulturpflanze 32:11–34
Hane JK, Ming Y, Kamphuis LG, Nelson MN, Garg G, Atkins CA, Bayer PE, Bravo A, Bringans S, Cannon S, Edwards D, Foley R, Gao L-L, Harrison MJ, Huang W, Hurgobin B, Li S, Liu C-W, McGrath A, Morahan G, Murray J, Weller J, Jian J, Singh KB (2017) A comprehensive draft genome sequence for lupin (Lupinus angustifolius), an emerging health food: insights into plant–microbe interactions and legume evolution. Plant Biotechnol J 15:318–330
Hondelmann W (1984) The lupin—ancient and modern crop plant. Theor Appl Genet 68:1–9
Hufnagel B, Marques A, Soriano A, Marquès L, Divol F, Doumas P, Sallet E, Mancinotti D, Carrere S, Marande W, Arribat S, Keller J, Huneau C, Blein T, Aime D, Laguerre M, Taylor J, Schubert V, Nelson M, Geu-Flores F, Crespi M, Gallardo-Guerrero K, Delaux P-M, Salse J, Bergès H, Guyot R, Gouzy J, Péret B (2019) Genome sequence of the cluster root forming white lupin. bioRxiv:708917
Hughes C, Eastwood R (2006) Island radiation on a continental scale: exceptional rates of plant diversification after uplift of the Andes. Proc Natl Acad Sci 103:10334–10339
Iqbal MJ, Mamidi S, Ahsan R, Kianian SF, Coyne CJ, Hamama AA, Narina SS, Bhardwaj HL (2012) Population structure and linkage disequilibrium in Lupinus albus L. germplasm and its implication for association mapping. Theor Appl Genet 125:517–530
Iqbal M, Huynh M, Udall J, Kilian A, Adhikari K, Berger J, Erskine W, Nelson MN (2019) The first genetic map for yellow lupin enables genetic dissection of adaptation traits in an orphan grain legume crop. BMC Genetics 20:68
Kamphuis LG, Hane JK, Nelson MN, Gao L, Atkins CA, Singh KB (2015) Transcriptome sequencing of different narrow-leafed lupin tissue types provides a comprehensive uni-gene assembly and extensive gene-based molecular markers. Plant Biotechnol J 13:14–25
Kasprzak A, Safár J, Janda J, Dolezel J, Wolko B, Naganowska B (2006) The bacterial artificial chromosome (BAC) library of the narrow-leafed lupin (Lupinus angustifolius L.). Cell Mol Biol Lett 11:396–407
Kroc M, Koczyk G, Święcicki W, Kilian A, Nelson MN (2014) New evidence of ancestral polyploidy in the Genistoid legume Lupinus angustifolius L. (narrow-leafed lupin). Theor Appl Genet 127:1237–1249
Kroc M, Koczyk G, Kamel KA, Czepiel K, Fedorowicz-Strońska O, Krajewski P, Kosińska J, Podkowiński J, Wilczura P, Święcicki W (2019) Transcriptome-derived investigation of biosynthesis of quinolizidine alkaloids in narrow-leafed lupin (Lupinus angustifolius L.) highlights candidate genes linked to iucundus locus. Sci Rep 9:2231
Książkiewicz M, Nazzicari N, Yang HA, Nelson MN, Renshaw D, Rychel S, Ferrari B, Carelli M, Tomaszewska M, Stawiński S, Naganowska B, Wolko B, Annicchiarico P (2017) A high-density consensus linkage map of white lupin highlights synteny with narrow-leafed lupin and provides markers tagging key agronomic traits. Sci Rep 7:15335
Leebens-Mack JH, Barker MS, Carpenter EJ, Deyholos MK, Gitzendanner MA, Graham SW, Grosse I, Li Z, Melkonian M, Mirarab S, Porsch M, Quint M, Rensing SA, Soltis DE, Soltis PS, Stevenson DW, Ullrich KK, Wickett NJ, DeGironimo L, Edger PP, Jordon-Thaden IE, Joya S, Liu T, Melkonian B, Miles NW, Pokorny L, Quigley C, Thomas P, Villarreal JC, Augustin MM, Barrett MD, Baucom RS, Beerling DJ, Benstein RM, Biffin E, Brockington SF, Burge DO, Burris JN, Burris KP, Burtet-Sarramegna V, Caicedo AL, Cannon SB, Çebi Z, Chang Y, Chater C, Cheeseman JM, Chen T, Clarke ND, Clayton H, Covshoff S, Crandall-Stotler BJ, Cross H, dePamphilis CW, Der JP, Determann R, Dickson RC, Di Stilio VS, Ellis S, Fast E, Feja N, Field KJ, Filatov DA, Finnegan PM, Floyd SK, Fogliani B, García N, Gâteblé G, Godden GT, Goh F, Greiner S, Harkess A, Heaney JM, Helliwell KE, Heyduk K, Hibberd JM, Hodel RGJ, Hollingsworth PM, Johnson MTJ, Jost R, Joyce B, Kapralov MV, Kazamia E, Kellogg EA, Koch MA, Von Konrat M, Könyves K, Kutchan TM, Lam V, Larsson A, Leitch AR, Lentz R, Li F-W, Lowe AJ, Ludwig M, Manos PS, Mavrodiev E, McCormick MK, McKain M, McLellan T, McNeal JR, Miller RE, Nelson MN, Peng Y, Ralph P, Real D, Riggins CW, Ruhsam M, Sage RF, Sakai AK, Scascitella M, Schilling EE, Schlösser E-M, Sederoff H, Servick S, Sessa EB, Shaw AJ, Shaw SW, Sigel EM, Skema C, Smith AG, Smithson A, Stewart CN, Stinchcombe JR, Szövényi P, Tate JA, Tiebel H, Trapnell D, Villegente M, Wang C-N, Weller SG, Wenzel M, Weststrand S, Westwood JH, Whigham DF, Wu S, Wulff AS, Yang Y, Zhu D, Zhuang C, Zuidof J, Chase MW, Pires JC, Rothfels CJ, Yu J, Chen C, Chen L, Cheng S, Li J, Li R, Li X, Lu H, Ou Y, Sun X, Tan X, Tang J, Tian Z, Wang F, Wang J, Wei X, Xu X, Yan Z, Yang F, Zhong X, Zhou F, Zhu Y, Zhang Y, Ayyampalayam S, Barkman TJ, Nguyen N-p, Matasci N, Nelson DR, Sayyari E, Wafula EK, Walls RL, Warnow T, An H, Arrigo N, Baniaga AE, Galuska S, Jorgensen SA, Kidder TI, Kong H, Lu-Irving P, Marx HE, Qi X, Reardon CR, Sutherland BL, Tiley GP, Welles SR, Yu R, Zhan S, Gramzow L, Theißen G, Wong GK-S (2019) One thousand plant transcriptomes and the phylogenomics of green plants. One thousand plant transcriptomes initiative. Nature 574:679–685
Li X, Yang H, Buirchell B, Yan G (2011) Development of a DNA marker tightly linked to low-alkaloid gene iucundus in narrow-leafed lupin (Lupinus angustifolius L.) for marker-assisted selection. Crop Pasture Sci 62:218–224
Lucas MM, Stoddard FL, Annicchiarico P, Frías J, Martínez-Villaluenga C, Sussmann D, Duranti M, Seger A, Zander PM, Pueyo JJ (2015) The future of lupin as a protein crop in Europe. Front Plant Sci 6:705
Martin GE, Rousseau-Gueutin M, Cordonnier S, Lima O, Michon-Coudouel S, Naquin D, de Carvalho JF, Aïnouche M, Salmon A, Aïnouche A (2014) The first complete chloroplast genome of the Genistoid legume Lupinus luteus: evidence for a novel major lineage-specific rearrangement and new insights regarding plastome evolution in the legume family. Ann Bot 113:1197–1210
Mikolajczyk J (1966) Genetic studies in Lupinus angustifolius. Part III. Inheritance of the alkaloid content, seed hardness and length of the growing season in blue lupin. Genet Pol 7:181–196
Mousavi-Derazmahalleh M, Bayer PE, Nevado B, Hurgobin B, Filatov D, Kilian A, Kamphuis LG, Singh KB, Berger JD, Hane JK, Edwards D, Erskine W, Nelson MN (2018a) Exploring the genetic and adaptive diversity of a pan-mediterranean crop wild relative: narrow-leafed lupin. Theor Appl Genet 131:887–901
Mousavi-Derazmahalleh M, Nevado B, Bayer PE, Filatov DA, Hane JK, Edwards D, Erskine W, Nelson MN (2018b) The western Mediterranean region provided the founder population of domesticated narrow-leafed lupin. Theor Appl Genet 131:2543–2554
Mousavi-Derazmahalleh M, Bayer PE, Hane JK, Valliyodan B, Nguyen HT, Nelson MN, Erskine W, Varshney RK, Papa R, Edwards D (2019) Adapting legume crops to climate change using genomic approaches. Plant Cell Environ 42:6–19
Naganowska B, Wolko B, Sliwińska E, Kaczmarek Z (2003) Nuclear DNA content variation and species relationships in the genus Lupinus (Fabaceae). Ann Bot 92:349–355
Nelson M, Phan H, Ellwood S, Moolhuijzen P, Hane J, Williams A, O'Lone C, Fosu-Nyarko J, Scobie M, Cakir M, Jones M, Bellgard M, Ksiazkiewicz M, Wolko B, Barker S, Oliver R, Cowling W (2006) The first gene-based map of Lupinus angustifolius L.- location of domestication genes and conserved synteny with Medicago truncatula. Theor Appl Genet 113:225–238
Nelson MN, Moolhuijzen PM, Boersma JG, Chudy M, Lesniewska K, Bellgard M, Oliver RP, Swiecicki W, Wolko B, Cowling WA, Ellwood SR (2010) Aligning a new reference genetic map of Lupinus angustifolius with the genome sequence of the model legume, Lotus japonicus. DNA Res 17:73–83
Nelson MN, Książkiewicz M, Rychel S, Besharat N, Taylor CM, Wyrwa K, Jost R, Erskine W, Cowling WA, Berger JD, Batley J, Weller JL, Naganowska B, Wolko B (2017) The loss of vernalization requirement in narrow-leafed lupin is associated with a deletion in the promoter and de-repressed expression of a Flowering Locus T (FT) homologue. New Phytol 213:220–232
Nevado B, Atchison GW, Hughes CE, Filatov DA (2016) Widespread adaptive evolution during repeated evolutionary radiations in New World lupins. Nat Commun 7:12384
O’Rourke JA, Yang SS, Miller SS, Bucciarelli B, Liu J, Rydeen A, Bozsoki Z, Uhde-Stone C, Tu ZJ, Allan D, Gronwald JW, Vance CP (2013) An RNA-Seq transcriptome analysis of orthophosphate-deficient white lupin reveals novel insights into phosphorus acclimation in plants. Plant Physiol 161:705–724
Parra-González LB, Aravena-Abarzúa GA, Navarro-Navarro CS, Udall J, Maughan J, Peterson LM, Salvo-Garrido HE, Maureira-Butler IJ (2012) Yellow lupin (Lupinus luteus L.) transcriptome sequencing: molecular marker development and comparative studies. BMC Genomics 13:425
Phan HTT, Ellwood SR, Adhikari K, Nelson MN, Oliver RP (2007) The first genetic and comparative map of white lupin (Lupinus albus L.): identification of QTLs for anthracnose resistance and flowering time, and a locus for alkaloid content. DNA Res 14:59–70
Raman R, Luckett DJ, Raman H (2008) Estimation of genetic diversity in albus lupin (Lupinus albus L.) using DArT and genic markers. International Lupin Association, Canterbury
Raman R, Cowley RB, Raman H, Luckett DJ (2014) Analyses using SSR and DArT molecular markers reveal that Ethiopian accessions of white lupin (Lupinus albus L.) represent a unique genepool. Open J Genet 4:87
Rychel S, Książkiewicz M, Tomaszewska M, Bielski W, Wolko B (2019) FLOWERING LOCUS T, GIGANTEA, SEPALLATA, and FRIGIDA homologs are candidate genes involved in white lupin (Lupinus albus L.) early flowering. Mol Breed 39:43
Secco D, Shou H, Whelan J, Berkowitz O (2014) RNA-seq analysis identifies an intricate regulatory network controlling cluster root development in white lupin. BMC Genomics 15:230
Swiecicki W, Swiecicki W (1995) Domestication and breeding improvement of narrow-leafed lupin (L. angustifolius L.). J Appl Genet 2:155–167
Taylor CM, Kamphuis LG, Zhang W, Garg G, Berger JD, Mousavi-Derazmahalleh M, Bayer PE, Edwards D, Singh KB, Cowling WA, Nelson MN (2019) INDEL variation in the regulatory region of the major flowering time gene LanFTc1 is associated with vernalization response and flowering time in narrow-leafed lupin (Lupinus angustifolius L.). Plant Cell Environ 42:174–187
Uhde-Stone C, Gilbert G, Johnson JM, Litjens R, Zinn KE, Temple SJ, Vance CP, Allan DL (2003) Acclimation of white lupin to phosphorus deficiency involves enhanced expression of genes related to organic acid metabolism. Plant Soil 248:99–116
Vavilov NI (1951) The origin, variation, immunity and breeding of cultivated plants. Chron Bot 13:1–366
Vipin CA, Luckett DJ, Harper JD, Ash GJ, Kilian A, Ellwood SR, Phan HT, Raman H (2013) Construction of integrated linkage map of a recombinant inbred line population of white lupin (Lupinus albus L.). Breed Sci 63:292–300
von Sengbusch R (1942) Sweet lupins and oil lupins. The history of the origin of some new crop plants. Landwirtschaftlic Jahrb 91:719–880
Wang Z, Straub D, Yang H, Kania A, Shen J, Ludewig U, Neumann G (2014) The regulatory network of cluster-root function and development in phosphate-deficient white lupin (Lupinus albus) identified by transcriptome sequencing. Physiol Plant 151:323–338
Williams W, Harrison JEM, Jayasekera S (1984) Genetical control of alkaloid production in Lupinus mutabilis and the effect of a mutant allele Mutal isolated following chemical mutagenesis. Euphytica 33:811–817
Wink M, Meißner C, Witte L (1995) Patterns of quinolizidine alkaloids in 56 species of the genus Lupinus. Phytochemistry 38:139–153
Wolko B, Weeden N (1989) Estimation of Lupinus genome polyploidy on the basis of isozymic loci number. Genet Pol 30
Wolko B, Clements JC, Naganowska B, Nelson MN, Yang H (2011) Lupinus. In: Kole C (ed) Wild crop relatives: genomic and breeding resources. Springer, Heidelberg
Yang H, Tao Y, Zheng Z, Zhang Q, Zhou G, Sweetingham MW, Howieson JG, Li C (2013) Draft genome sequence, and a sequence-defined genetic linkage map of the legume crop species Lupinus angustifolius L. PLoS ONE 8:e64799
Yang T, Nagy I, Mancinotti D, Otterbach SL, Andersen TB, Motawia MS, Asp T, Geu-Flores F (2017) Transcript profiling of a bitter variety of narrow-leafed lupin to discover alkaloid biosynthetic genes. J Exp Bot 68:5527–5537
Zhou G, Jian J, Wang P, Li C, Tao Y, Li X, Renshaw D, Clements J, Sweetingham M, Yang H (2018) Construction of an ultra-high density consensus genetic map, and enhancement of the physical map from genome sequencing in Lupinus angustifolius. Theor Appl Genet 131:209–223
Zohary D, Hopf M, Weiss E (2012) Domestication of plants in the old world: the origin and spread of domesticated plants in Southwest Asia, Europe, and the Mediterranean basin. Oxford University Press on Demand
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Taylor, J.L., De Angelis, G., Nelson, M.N. (2020). How Have Narrow-Leafed Lupin Genomic Resources Enhanced Our Understanding of Lupin Domestication?. In: Singh, K., Kamphuis, L., Nelson, M. (eds) The Lupin Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-030-21270-4_8
Download citation
DOI: https://doi.org/10.1007/978-3-030-21270-4_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-21269-8
Online ISBN: 978-3-030-21270-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)