Abstract
Considerable progress has been made in both germline and somatic characterization of genetic alterations in neuroblastoma patients. Indeed, predisposition genes that account for a proportion of familial and syndromic cases have been identified, and genome-wide association studies have retrieved a number of susceptibility loci. Moreover, genome-wide sequencing, copy-number, and expression studies conducted on tumors have detected recurrent gene modifications, profiles, and signatures that have strong implications for the therapeutic stratification of patients. The identification of major players in neuroblastoma (NB) oncogenesis, including MYCN and ALK as well as other less frequently observed mutations, provides insights into the mechanisms of NB development with potential impact on the clinical management of patients.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
To date, the precise etiology of neuroblastoma is unknown, and unlike many adult malignancies, environmental factors are not thought to play a major role, although predisposing effects of prenatal exposures to potentially toxic substances warrant further investigation. However, genetic factors, both at constitutional and somatic levels, are thought to play a major role in neuroblastoma development [1].
2 Hereditary Genetic Factors
Several observations corroborate the hypothesis of a role of underlying hereditary genetic factors in the etiology of neuroblastoma.
First, although rare and representing less than 1% of all cases [2], familial neuroblastomas have been described. Mutations of gain of function in the tyrosine kinase domain of the ALK anaplastic lymphoma kinase gene have been detected in the majority of familial cases [3, 4]. This is thought to be associated with an autosomal-dominant pattern of inheritance with incomplete penetrance.
Second, neuroblastoma can appear in association with different clinical syndromes. Neural crest-related developmental disorders associated with an increased risk of developing neuroblastoma have been linked to inactivating mutations in the PHOX2B gene, a major regulator of neural crest development, identified as the first neuroblastoma predisposition mutation [5, 6]. While expansions of the second polyalanine sequence of PHOX2B are mainly observed in patients with a curse of Ondine (also called CCHS, Congenital Central Hypoventilation Syndrome) associated with a low risk of peripheral neuroblastic tumors, non-expansive mutations with Hirschsprung’s disease are associated with a higher risk of developing neuroblastoma. Other associations between neuroblastic tumors and cancer susceptibility syndromes include neurofibromatosis type 1 (NF1) [7], characterized by constitutive activation of the RAS–MAPK pathway, as well as Noonan syndrome with PTPN11 gene involvement.
Third, genome-wide association studies have shown that peripheral neuroblastic tumors may occur in the context of underlying genetic factors, as it has been demonstrated that different polymorphic alleles, localized at different genome loci, influence oncogenesis [8,9,10,11,12,13]. Of a weak individual impact on the initiation of the disease (with a relative risk of 1.5–2.0 compared to the global population), these polymorphic alleles can cooperate in an individual patient to promote malignant transformation during neurological development, and some genes targeted by these polymorphic alleles play a role in the pathogenesis of neuroblastoma, including BARD1, LMO1, DUSP12, DDX4, HACE1, and LIN28B. The polymorphic alleles described show a correlation with high-risk or low-risk disease, indicating that favorable and unfavorable forms of neuroblastoma may represent distinct entities in terms of genetic events that initiate tumorigenesis.
In addition, rare cases with various aberrant constitutional karyotypes have been described in patients with neuroblastoma, including constitutional copy-number abnormalities, balanced and unbalanced translocations, and specific chromosomal deletions, including deletions of chromosome 1p [14]. In total, there are probably as yet undiscovered additional genes that predispose to neuroblastoma when they are altered in the germline. It is important to note that, to date, there are no clinically validated guidelines for determining who should be screened for germline mutations, nor how to monitor patients or families with known susceptibility alleles.
2.1 Somatic Genetic Alterations
2.1.1 Copy-Number Alterations
With only 1–2% of neuroblastomas occurring in a familial context or predisposition, over 98% of all cases occur sporadically. A large number of recurrent somatic genetic alterations have been found in neuroblastoma, the most common being quantitative genomic alterations with gains or losses in genetic material. These genetic abnormalities are related to distinct biological and clinical subgroups of the disease.
The amplification of the oncogene MYCN, located at chromosome 2p24.1, is observed in about 25% of neuroblastomas and 40% of high-risk tumors [15]. It remains one of the most important genetic alterations associated with advanced stages of disease, with an aggressive phenotype and poor survival [1]. It is the first genetic marker used in clinical practice for risk stratification and treatment adaptation [16]. Closely associated with poor survival in patients with localized disease and in infants, its prognostic impact in metastatic disease of older children with an overall poor outcome is less clear [16, 17]. At a cytogenetic level, amplification of the MYCN oncogene occurs either as double-minute chromosomes (DM) or homogeneously staining regions (HSR), which contain between 10 and over 100 ectopic copies of the MYCN oncogene. The oncogenic role of MYCN has been clearly demonstrated as its ectopic expression in the neural crest is sufficient to drive neuroblastoma tumorigenesis in zebrafish and mice models [18]. Its oncogenic role is based on an enhancement of the expression of genes involved in cell proliferation, and on the repression of genes involved in differentiation and apoptosis [19].
Other recurrent amplifications concern the ALK gene on chromosome 2p23, as well as amplicons of chromosome 12q13–14 encompassing, among others, the MDM2 and CDK4 genes [20]. Recent data indicate that NB with focal amplifications other than MYCN might present with atypical clinical features and a poorer outcome [21].
Other recurrent structural alterations recurrently observed in neuroblastoma concern segmental chromosome alterations (SCA) corresponding to unbalanced chromosome translocations, including deletions of chromosome arms 1p, 3p, 4p, and 11q, and gains of 1q, 2p, or 17q. Although several recurrently altered chromosome regions have been identified, chromosome breakpoints are not recurrent but lie scattered over wide genomic regions. Deletion of 1p36 is observed in 20–35% of cases and is associated with poor survival in multivariate analyses as well as with aggressive disease markers [22, 23]. Deletions of 11q in a consensus region at 11q23 occur in approximately 40% of cases and are inversely correlated with MYCN amplification, identifying a molecularly distinct high-risk patient subset, characterized by advanced stage, older age, as well as a higher genomic instability with a higher number of chromosome breakpoints [24, 25]. Gains of chromosome 17q21-qter represent the most frequent genetic alteration in neuroblastoma, occurring in 70% of tumors. Numerous studies have reported that 17q gain is significantly associated with advanced stage of disease, increased patient age, MYCN amplification, as well as other unfavorable genetic parameters [26].
Although intense decade-long research has focused on the identification of hypothetical tumor-suppressor genes or oncogenes in recurrently altered regions of chromosome loss or gain, the smallest regions of overlap remain quite extensive and to date do not point to single-gene candidates as tumor suppressors or oncogenes, suggesting that an overall imbalance of copy-number regions is of importance in neuroblastoma oncogenesis.
Although the individual segmental chromosome alterations have been shown to correlate with outcome, importantly, the overall genomic profile has been shown to be of prognostic impact in neuroblastoma [27,28,29,30]. Whereas an overall genomic copy-number profile characterized by numerical chromosome alterations, consisting of gains or losses of whole chromosomes, is associated with a favorable outcome, segmental chromosome alterations of any chromosome region, without or with numerical chromosome alterations, are associated with advanced stage of disease, with an increased age at diagnosis and, importantly, a higher risk of relapse in multivariate analyses [27, 28, 31]. In addition to the determination of MYCN amplifications status, the overall genomic copy-number profile determined by array comparative genomic hybridization (CGH) or single nucleotide polymorphism (SNP) array is now considered part of routine work up in particular in low- and intermediate-risk neuroblastoma and might be used for treatment stratification within prospective clinical protocols [1].
Higher-resolution copy-number analyses have also revealed smaller recurrent interstitial events. Indeed, SNP arrays have identified alterations on chromosome 9p, with homo- or hemizygous deletions encompassing the CDKN2A gene [32]. Other sporadic copy-number alterations include focal TERT gains [33] and microdeletions encompassing the PTPRD gene [34].
More complex rearrangements resulting from chromothripsis, corresponding to massive genomic rearrangements acquired in a single catastrophic event, have been described in neuroblastoma, but their association with other genomic and clinical subtypes remains to be determined [35, 36].
2.1.2 Single-Gene Mutations
Recent next-generation sequencing approaches have indicated that most neuroblastomas harbor only few mutations, with an average of 10–20 predicted non-synonymous variations in coding regions per genome, indicating an exonic mutation frequency of 0.2–0.4 per Mb. [35, 37,38,39] The frequency of somatic events strongly correlates with tumor stage, higher-stage tumors harboring a higher number of mutations.
The most frequent recurrent somatic mutation in neuroblastoma concerns the gene ALK (anaplastic lymphoma kinase), with mutations activating the tyrosine kinase domain in approximately 10% of all cases at diagnosis. [3, 4] The somatic ALK-1174 mutation appears to contribute to a more aggressive phenotype, but unlike ALK-1275 mutations, these specific mutations are not found in familial neuroblastoma, suggesting that they are not tolerated in the germline [40, 41]. The oncogenic role of activating ALK mutations in neuroblastoma has been demonstrated in vitro and in vivo in both zebrafish and mouse models, with coexpression of ALK-F1174L and MYCN producing a synergistic effect for neuroblastoma tumorigenesis in mice. New-generation small-molecule inhibitors targeting the activated kinase domain of ALK are now available, making this a promising target for molecular therapy, possibly in combination therapies, but still requiring more specific development [40, 42, 43].
Other recurrent mutations in neuroblastoma target distinct cellular pathways and include PTPN11 mutations (in 3% of cases), as well as genes involved in cytoskeleton maintenance, neuritogenesis, and other regulators of the RAC/Rho pathway [35].
Interestingly, genes involved in chromatin remodeling have been found to be targeted in a significant number of cases, either by mutations or by structural variations, including mutations in the ARID1A/ARID1B genes [38]. Somatic alterations of ATRX, consisting either of mutations or small interstitial deletions, are associated with an increase in telomere length and with an absence of the ATRX protein in the nucleus. ATRX alterations appear to be more frequent in older children and occur in mutually exclusive fashion with MYCN amplifications [39]. ATRX mutations are associated with activation of a telomere maintenance mechanism termed alternate lengthening of telomeres (ALT), which may be associated with primary chemotherapy resistance.
Recurrent genomic rearrangements of the promoter region of the telomerase reverse transcriptase (TERT) gene on chromosome 5p15.33 have been described in >10% of neuroblastoma cases, with structural rearrangements of TERT resulting from chromothripsis in some cases [44, 45]. These rearrangements, which are associated with increased TERT expression, target regions immediately up- and downstream of TERT, and position the TERT coding sequence to strong enhancer elements, resulting in massive chromatin remodeling and DNA methylation of the affected region. [44] Occurring in mutually exclusive fashion with MYCN amplification and ATRX mutations, these rearrangements define a further subgroup of high-risk disease, with TERT rearrangements (23%), ATRX deletions (11%), and MYCN amplifications (37%) identifying three almost non-overlapping groups of high-stage neuroblastoma, each associated with very poor prognosis.
Thus, a large number of high-risk neuroblastomas are affected by genetic alterations of either MYCN, TERT, or ATRX, all of which converge to an activation of telomere-lengthening mechanisms either by direct activation or by ALT, leading to a capacity of near-infinite cell proliferation. [46] Advances in the development of inhibitors of these pathways and their evaluation in clinical trials will lead to new treatment opportunities.
Future studies will determine if the association between these major players in neuroblastoma oncogenesis with distinct genetic profiles and mutational patterns might serve for the definition of different risk groups in particular in high-risk neuroblastoma.
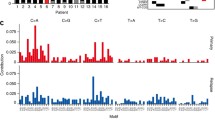
Overall, large-scale sequencing efforts have highlighted distinct mutational signatures which are thought to reflect distinct biochemical cellular processes [47]. In neuroblastoma, in some cases a predominance of C > T transitions, termed mutational signature 1, are observed, with an association with age. Other mutational signatures observed in neuroblastoma, although rarer, concern the canonical double-stranded break signature linked to mutations in BRCA1 or BRCA2 or to a “BRCAness” phenotype [48]. Altogether, these studies underline the heterogeneity of somatic genetic alterations in neuroblastoma and highlight the importance to pursue efforts of molecular characterization.
2.1.3 Expression Profiles
In addition to genetic changes, neuroblastoma can also be characterized by specific expression profiles. Indeed, to date, a large number of studies have focused on the analysis of differential expression patterns in NB, seeking to identify expression patterns that might be enable to distinguish patients with different clinical courses and thus define different prognostic groups in high-risk disease, and to potentially identify new therapeutic targets.
Thus, different expression signatures, based on 144-gene or 59-gene signatures, reliably distinguished patients with distinct clinical courses, with the strongest difference observed in non-high-risk disease [49, 50]. Using real-time PCR expression data, an expression signature based on only three genes (CHD5, PAFAH1B1, and NME1) has been able to discriminate patients with different outcomes [51]. Among high-risk patients, an expression profile based on 55 genes defined patient populations with divergent outcome [52]. Differential expression signature depending on MYCN has led to a definition of a 157-gene signature which identified NB patients with poor prognosis independent of the genomic MYCN status, those without MYCN amplification presenting stabilization of MYCN at the protein level [53].
More recently, based on the hypothesis that tumor-associated inflammatory cells might contribute to the differences in age-dependent outcome of patients with metastatic NB, expression of genes representing tumor-associated macrophages, such as CD33/CD16/IL6R/IL10/FCGR3, contributed to a novel 14-gene tumor classification score. Progression-free survival was 47% versus 12% for patients with a low- versus a high-risk score, indicating that interactions between tumor and inflammatory cells may contribute to an aggressive metastatic NB phenotype [54].
In addition to messenger RNA, expression levels of non-coding RNAs are also highly variable. Micro-RNAs function as regulators of gene expression at the posttranscriptional level in diverse cellular processes and constitute the most widely studied non-coding RNA molecules in NB. MYCN modulates the expression of several classes of non-coding RNAs, especially some micro-RNAs, and it can also regulate the expression of long non-coding RNAs such as T-UCRs (Transcribed UltraConserved Regions) and non-coding RNA, whereas other long non-coding RNAs remain to be characterized in NB. The landscape of T-UCRs in NB has been studied recently and has revealed a correlation with the MYCN status, and preliminary studies have suggested that T-UCR-based expression signatures might distinguish short-from long-term survivors in high risk NB [55].
The miRNA expression pattern can also be used to classify NB patients according to survival [56, 57]. An advantage of the study of miRNA rather than mRNA expression signatures is linked to the greatest stability of miRNAs, and thus to an analytical feasibility even with formalin-fixed, paraffin-embedded (FFPE) samples as opposed to frozen samples [57]. The miRNA expression pattern can also be used to classify NB patients according to survival [56, 57].
The paucity of recurrent genetic mutations as compared to adult tumors indicates that additional mechanisms such as epigenetic alterations may play an important role in the molecular pathogenesis of these developmental tumors. Alterations in DNA methylation represent one of the most common molecular events in neoplasia, and CpG-island hypermethylation of gene promoters is a frequent mechanism for functional inactivation of relevant tumor-associated genes in neuroblastoma. Promoter methylation patterns which are associated with patient subgroups and distinct clinical features have been identified [58]. Further studies are now necessary to determine whether these genome-wide methylation patterns correlate with outcome and other prognostic molecular markers in NB patients.
Taken together, to date, many studies have demonstrated the feasibility of expression profiling of mRNA, miRNA, other non-coding RNAs, or epigenetic modifiers in order to determine different prognostic subgroups among NB patients. However, there is little, if any, overlap between the genes of the different signatures rendering cross-study comparisons unfeasible. Furthermore, although most expression signatures clearly distinguish prognostic groups in the overall population, differences in survival among high-risk patients are frequently not very marked. The routine setup of real-time determination of expression profiles in a prospective clinical trial setting, and their interpretation, remains a clear challenge.
2.1.4 Spatial and Temporal Heterogeneity
Neuroblastoma presents important spatial and temporal heterogeneity. Spatial heterogeneity has been recorded for several recurrent somatic genetic alterations in neuroblastoma. Indeed, MYCN alterations might occur only in a subset of neuroblastic cells of a given tumor [59]. Segmental chromosome alterations might also vary between different neuroblastic cell populations [60, 61]. Furthermore, mutations can also be observed at a heterogeneous level. In some tumors, low-level mutated allele fractions for ALK driver mutations have been observed [62].
In neuroblastoma, temporal heterogeneity can also occur. Indeed, genetic alterations may evolve over time and clonal evolution is common, leading to the acquisition of somatic alterations in known oncogenic pathways, some of which may be targeted. ALK-activating mutations, in some instance present in a minor subclone at diagnosis, might emerge at relapse [63]. Furthermore, activation of the MAPK pathway and other signaling pathways for epithelial–mesenchymal transition (EMT) processes may appear during a relapse and represent promising targets for targeted molecular treatment approaches [64, 65]. In total, spatial and temporal genetic heterogeneity plays an important role in neuroblastoma.
However, multi-site tumor biopsies, or sequential biopsies from the same tumor, can only rarely be realized, and recently liquid biopsies have emerged as a very promising tool to explore somatic genetic alterations with regards to both spatial and temporal heterogeneity. Indeed, circulating tumor DNA, a fraction of cell-free DNA, can readily be extracted from plasma of neuroblastoma patients. This can serve for the detection of MYCN amplification [66]. Copy-number alterations or mutations such as ALK can also be detected in ctDNA. [67,68,69] More recently application of whole-exome sequencing techniques to sequential ctDNA samples from NB patients has provided further evidence of the importance of clonal evolution in the progression of neuroblastoma, enabling the description of resistant clones emerging at the time of relapse [70].
Altogether, as neuroblastoma is in general associated with a low mutational burden and only few recurrently occurring mutations, it might be considered a copy-number disease, with large chromosome segments contributing to oncogenesis by gene dosage effects. Further ongoing efforts will enable to determine whether epigenetic changes also play a role in neuroblastoma oncogenesis.
References
Matthay KK, Maris JM, Schleiermacher G, Nakagawara A, Mackall CL, Diller L, Weiss WA. Neuroblastoma. Nat Rev Dis Primers. 2016;2:16078.
Shojaei-Brosseau T, Chompret A, Abel A, de Vathaire F, Raquin M-A, Brugières L, Feunteun J, Hartmann O, Bonaïti-Pellié C. Genetic epidemiology of neuroblastoma: a study of 426 cases at the Institut Gustave-Roussy in France. Pediatr Blood Cancer. 2004;42:99–105.
Mossé YP, Laudenslager M, Longo L, Cole KA, Wood A, Attiyeh EF, Laquaglia MJ, Sennett R, Lynch JE, Perri P, Laureys G, Speleman F, et al. Identification of ALK as a major familial neuroblastoma predisposition gene. Nature. 2008;455:930–5.
Janoueix-Lerosey I, Lequin D, Brugières L, Ribeiro A, de Pontual L, Combaret V, Raynal V, Puisieux A, Schleiermacher G, Pierron G, Valteau-Couanet D, Frebourg T, et al. Somatic and germline activating mutations of the ALK kinase receptor in neuroblastoma. Nature. 2008;455:967–70.
Trochet D, Bourdeaut F, Janoueix-Lerosey I, Deville A, de Pontual L, Schleiermacher G, Coze C, Philip N, Frébourg T, Munnich A, Lyonnet S, Delattre O, et al. Germline mutations of the paired-like homeobox 2B (PHOX2B) gene in neuroblastoma. Am J Hum Genet. 2004;74:761–4.
Trochet D, O’Brien LM, Gozal D, Trang H, Nordenskjöld A, Laudier B, Svensson P-J, Uhrig S, Cole T, Niemann S, Munnich A, Gaultier C, et al. PHOX2B genotype allows for prediction of tumor risk in congenital central hypoventilation syndrome. Am J Hum Genet. 2005;76:421–6.
Brems H, Beert E, de Ravel T, Legius E. Mechanisms in the pathogenesis of malignant tumours in neurofibromatosis type 1. Lancet Oncol. 2009;10:508–15.
Bosse KR, Diskin SJ, Cole KA, Wood AC, Schnepp RW, Norris G, Nguyen LB, Jagannathan J, Laquaglia M, Winter C, Diamond M, Hou C, et al. Common variation at BARD1 results in the expression of an oncogenic isoform that influences neuroblastoma susceptibility and oncogenicity. Cancer Res. 2012;72:2068–78.
Diskin SJ, Hou C, Glessner JT, Attiyeh EF, Laudenslager M, Bosse K, Cole K, Mossé YP, Wood A, Lynch JE, Pecor K, Diamond M, et al. Copy number variation at 1q21.1 associated with neuroblastoma. Nature. 2009;459:987–91.
Maris JM, Mosse YP, Bradfield JP, Hou C, Monni S, Scott RH, Asgharzadeh S, Attiyeh EF, Diskin SJ, Laudenslager M, Winter C, Cole KA, et al. Chromosome 6p22 locus associated with clinically aggressive neuroblastoma. N Engl J Med. 2008;358:2585–93.
Capasso M, Devoto M, Hou C, Asgharzadeh S, Glessner JT, Attiyeh EF, Mosse YP, Kim C, Diskin SJ, Cole KA, Bosse K, Diamond M, et al. Common variations in BARD1 influence susceptibility to high-risk neuroblastoma. Nat Genet. 2009;41:718–23.
Nguyen LB, Diskin SJ, Capasso M, Wang K, Diamond MA, Glessner J, Kim C, Attiyeh EF, Mosse YP, Cole K, Iolascon A, Devoto M, et al. Phenotype restricted genome-wide association study using a gene-centric approach identifies three low-risk neuroblastoma susceptibility Loci. PLoS Genet. 2011;7:e1002026.
Wang K, Diskin SJ, Zhang H, Attiyeh EF, Winter C, Hou C, Schnepp RW, Diamond M, Bosse K, Mayes PA, Glessner J, Kim C, et al. Integrative genomics identifies LMO1 as a neuroblastoma oncogene. Nature. 2011;469:216–20.
Vandepoele K, Andries V, Van Roy N, Staes K, Vandesompele J, Laureys G, De Smet E, Berx G, Speleman F, van Roy F. A constitutional translocation t(1;17)(p36.2;q11.2) in a neuroblastoma patient disrupts the human NBPF1 and ACCN1 genes. PLoS One. 2008;3:e2207.
Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM. Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science. 1984;224:1121–4.
Campbell K, Gastier-Foster JM, Mann M, Naranjo AH, Van Ryn C, Bagatell R, Matthay KK, London WB, Irwin MS, Shimada H, Granger MM, Hogarty MD, et al. Association of MYCN copy number with clinical features, tumor biology, and outcomes in neuroblastoma: a report from the Children’s Oncology Group. Cancer. 2017;123:4224–35.
Bagatell R, Beck-Popovic M, London WB, Zhang Y, Pearson ADJ, Matthay KK, Monclair T, Ambros PF, Cohn SL, International Neuroblastoma Risk Group. Significance of MYCN amplification in international neuroblastoma staging system stage 1 and 2 neuroblastoma: a report from the International Neuroblastoma Risk Group database. J Clin Oncol Off J Am Soc Clin Oncol. 2009;27:365–70.
Weiss WA, Aldape K, Mohapatra G, Feuerstein BG, Bishop JM. Targeted expression of MYCN causes neuroblastoma in transgenic mice. EMBO J. 1997;16:2985–95.
Hansford LM, Thomas WD, Keating JM, Burkhart CA, Peaston AE, Norris MD, Haber M, Armati PJ, Weiss WA, Marshall GM. Mechanisms of embryonal tumor initiation: distinct roles for MycN expression and MYCN amplification. Proc Natl Acad Sci U S A. 2004;101:12664–9.
Depuydt P, Boeva V, Hocking TD, Cannoodt R, Ambros IM, Ambros PF, Asgharzadeh S, Attiyeh EF, Combaret V, Defferrari R, Fischer M, Hero B, et al. Genomic amplifications and distal 6q loss: novel markers for poor survival in high-risk neuroblastoma patients. J Natl Cancer Inst. 2018;110:1084.
Guimier A, Ferrand S, Pierron G, Couturier J, Janoueix-Lerosey I, Combaret V, Mosseri V, Thebaud E, Gambart M, Plantaz D, Marabelle A, Coze C, et al. Clinical characteristics and outcome of patients with neuroblastoma presenting genomic amplification of loci other than MYCN. PLoS One. 2014;9:e101990.
Caron H, Spieker N, Godfried M, Veenstra M, van Sluis P, de Kraker J, Voûte P, Versteeg R. Chromosome bands 1p35-36 contain two distinct neuroblastoma tumor suppressor loci, one of which is imprinted. Genes Chromosomes Cancer. 2001;30:168–74.
Schleiermacher G, Delattre O, Peter M, Mosseri V, Delonlay P, Vielh P, Thomas G, Zucker J, Magdelénat H, Michon J. Clinical relevance of loss heterozygosity of the short arm of chromosome 1 in neuroblastoma: a single-institution study. Int J Cancer. 1996;69:73–8.
Attiyeh EF, London WB, Mossé YP, Wang Q, Winter C, Khazi D, McGrady PW, Seeger RC, Look AT, Shimada H, Brodeur GM, Cohn SL, et al. Chromosome 1p and 11q deletions and outcome in neuroblastoma. N Engl J Med. 2005;353:2243–53.
Carén H, Kryh H, Nethander M, Sjöberg R-M, Träger C, Nilsson S, Abrahamsson J, Kogner P, Martinsson T. High-risk neuroblastoma tumors with 11q-deletion display a poor prognostic, chromosome instability phenotype with later onset. Proc Natl Acad Sci U S A. 2010;107:4323–8.
Bown N, Cotterill S, Lastowska M, O’Neill S, Pearson AD, Plantaz D, Meddeb M, Danglot G, Brinkschmidt C, Christiansen H, Laureys G, Speleman F, et al. Gain of chromosome arm 17q and adverse outcome in patients with neuroblastoma. N Engl J Med. 1999;340:1954–61.
Janoueix-Lerosey I, Schleiermacher G, Michels E, Mosseri V, Ribeiro A, Lequin D, Vermeulen J, Couturier J, Peuchmaur M, Valent A, Plantaz D, Rubie H, et al. Overall genomic pattern is a predictor of outcome in neuroblastoma. J Clin Oncol. 2009;27:1026–33.
Schleiermacher G, Michon J, Huon I, d’Enghien C, Klijanienko J, Brisse H, Ribeiro A, Mosseri V, Rubie H, Munzer C, Thomas C, Valteau-Couanet D, et al. Chromosomal CGH identifies patients with a higher risk of relapse in neuroblastoma without MYCN amplification. Br J Cancer. 2007;97:238–46.
Vandesompele J, Baudis M, De Preter K, Van Roy N, Ambros P, Bown N, Brinkschmidt C, Christiansen H, Combaret V, Lastowska M, Nicholson J, O’Meara A, et al. Unequivocal delineation of clinicogenetic subgroups and development of a new model for improved outcome prediction in neuroblastoma. J Clin Oncol Off J Am Soc Clin Oncol. 2005;23:2280–99.
Tomioka N, Oba S, Ohira M, Misra A, Fridlyand J, Ishii S, Nakamura Y, Isogai E, Hirata T, Yoshida Y, Todo S, Kaneko Y, et al. Novel risk stratification of patients with neuroblastoma by genomic signature, which is independent of molecular signature. Oncogene. 2008;27:441–9.
Coco S, Theissen J, Scaruffi P, Stigliani S, Moretti S, Oberthuer A, Valdora F, Fischer M, Gallo F, Hero B, Bonassi S, Berthold F, et al. Age-dependent accumulation of genomic aberrations and deregulation of cell cycle and telomerase genes in metastatic neuroblastoma. Int J Cancer. 2012;131:1591–600.
Caren H, Erichsen J, Olsson L, Enerback C, Sjoberg RM, Abrahamsson J, Kogner P, Martinsson T. High-resolution array copy number analyses for detection of deletion, gain, amplification and copy-neutral LOH in primary neuroblastoma tumors: four cases of homozygous deletions of the CDKN2A gene. BMC Genomics. 2008;9:353.
Cobrinik D, Ostrovnaya I, Hassimi M, Tickoo SK, Cheung IY, Cheung N-KV. Recurrent pre-existing and acquired DNA copy number alterations, including focal TERT gains, in neuroblastoma central nervous system metastases. Genes Chromosomes Cancer. 2013;52:1150–66.
Stallings RL, Nair P, Maris JM, Catchpoole D, McDermott M, O’Meara A, Breatnach F. High-resolution analysis of chromosomal breakpoints and genomic instability identifies PTPRD as a candidate tumor suppressor gene in neuroblastoma. Cancer Res. 2006;66:3673–80.
Molenaar JJ, Koster J, Zwijnenburg DA, van Sluis P, Valentijn LJ, van der Ploeg I, Hamdi M, van Nes J, Westerman BA, van Arkel J, Ebus ME, Haneveld F, et al. Sequencing of neuroblastoma identifies chromothripsis and defects in neuritogenesis genes. Nature. 2012;483:589–93.
Boeva V, Jouannet S, Daveau R, Combaret V, Pierre-Eugène C, Cazes A, Louis-Brennetot C, Schleiermacher G, Ferrand S, Pierron G, Lermine A, Rio Frio T, et al. Breakpoint features of genomic rearrangements in neuroblastoma with unbalanced translocations and chromothripsis. PLoS One. 2013;8:e72182.
Pugh TJ, Morozova O, Attiyeh EF, Asgharzadeh S, Wei JS, Auclair D, Carter SL, Cibulskis K, Hanna M, Kiezun A, Kim J, Lawrence MS, et al. The genetic landscape of high-risk neuroblastoma. Nat Genet. 2013;45:279–84.
Sausen M, Leary RJ, Jones S, Wu J, Reynolds CP, Liu X, Blackford A, Parmigiani G, Diaz LA, Papadopoulos N, Vogelstein B, Kinzler KW, et al. Integrated genomic analyses identify ARID1A and ARID1B alterations in the childhood cancer neuroblastoma. Nat Genet. 2013;45:12–7.
Cheung N-KV, Zhang J, Lu C, Parker M, Bahrami A, Tickoo SK, Heguy A, Pappo AS, Federico S, Dalton J, Cheung IY, Ding L, et al. Association of age at diagnosis and genetic mutations in patients with neuroblastoma. JAMA. 2012;307:1062–71.
Bresler SC, Weiser DA, Huwe PJ, Park JH, Krytska K, Ryles H, Laudenslager M, Rappaport EF, Wood AC, McGrady PW, Hogarty MD, London WB, et al. ALK mutations confer differential oncogenic activation and sensitivity to ALK inhibition therapy in neuroblastoma. Cancer Cell. 2014;26:682–94.
De Brouwer S, De Preter K, Kumps C, Zabrocki P, Porcu M, Westerhout EM, Lakeman A, Vandesompele J, Hoebeeck J, Van Maerken T, De Paepe A, Laureys G, et al. Meta-analysis of neuroblastomas reveals a skewed ALK mutation spectrum in tumors with MYCN amplification. Clin Cancer Res. 2010;16:4353–62.
Wood AC, Krytska K, Ryles HT, Infarinato NR, Sano R, Hansel TD, Hart LS, King FJ, Smith TR, Ainscow E, Grandinetti KB, Tuntland T, et al. Dual ALK and CDK4/6 Inhibition Demonstrates Synergy against Neuroblastoma. Clin Cancer Res. 2017;23:2856–68.
Infarinato NR, Park JH, Krytska K, Ryles HT, Sano R, Szigety KM, Li Y, Zou HY, Lee NV, Smeal T, Lemmon MA, Mossé YP. The ALK/ROS1 inhibitor PF-06463922 overcomes primary resistance to crizotinib in ALK-driven neuroblastoma. Cancer Discov. 2016;6:96–107.
Valentijn LJ, Koster J, Zwijnenburg DA, Hasselt NE, van Sluis P, Volckmann R, van Noesel MM, George RE, Tytgat GAM, Molenaar JJ, Versteeg R. TERT rearrangements are frequent in neuroblastoma and identify aggressive tumors. Nat Genet. 2015;47:1411–4.
Peifer M, Hertwig F, Roels F, Dreidax D, Gartlgruber M, Menon R, Krämer A, Roncaioli JL, Sand F, Heuckmann JM, Ikram F, Schmidt R, et al. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature. 2015;526:700–4.
Hertwig F, Peifer M, Fischer M. Telomere maintenance is pivotal for high-risk neuroblastoma. Cell Cycle. 2016;15:311–2.
Alexandrov LB, Nik-Zainal S, Wedge DC, Aparicio SAJR, Behjati S, Biankin AV, Bignell GR, Bolli N, Borg A, Børresen-Dale A-L, Boyault S, Burkhardt B, et al. Signatures of mutational processes in human cancer. Nature. 2013;500:415–21.
Gröbner SN, Worst BC, Weischenfeldt J, Buchhalter I, Kleinheinz K, Rudneva VA, Johann PD, Balasubramanian GP, Segura-Wang M, Brabetz S, Bender S, Hutter B, et al. The landscape of genomic alterations across childhood cancers. Nature. 2018;555:321–7.
Oberthuer A, Hero B, Berthold F, Juraeva D, Faldum A, Kahlert Y, Asgharzadeh S, Seeger R, Scaruffi P, Tonini GP, Janoueix-Lerosey I, Delattre O, et al. Prognostic impact of gene expression-based classification for neuroblastoma. J Clin Oncol. 2010;28:3506–15.
Vermeulen J, De Preter K, Naranjo A, Vercruysse L, Van Roy N, Hellemans J, Swerts K, Bravo S, Scaruffi P, Tonini GP, De Bernardi B, Noguera R, et al. Predicting outcomes for children with neuroblastoma using a multigene-expression signature: a retrospective SIOPEN/COG/GPOH study. Lancet Oncol. 2009;10:663–71.
Garcia I, Mayol G, Ríos J, Domenech G, Cheung N-KV, Oberthuer A, Fischer M, Maris JM, Brodeur GM, Hero B, Rodríguez E, Suñol M, et al. A three-gene expression signature model for risk stratification of patients with neuroblastoma. Clin Cancer Res. 2012;18:2012–23.
Asgharzadeh S, Pique-Regi R, Sposto R, Wang H, Yang Y, Shimada H, Matthay K, Buckley J, Ortega A, Seeger RC. Prognostic significance of gene expression profiles of metastatic neuroblastomas lacking MYCN gene amplification. J Natl Cancer Inst. 2006;98:1193–203.
Valentijn LJ, Koster J, Haneveld F, Aissa RA, van Sluis P, Broekmans MEC, Molenaar JJ, van Nes J, Versteeg R. Functional MYCN signature predicts outcome of neuroblastoma irrespective of MYCN amplification. Proc Natl Acad Sci U S A. 2012;109:19190–5.
Asgharzadeh S, Salo JA, Ji L, Oberthuer A, Fischer M, Berthold F, Hadjidaniel M, Liu CW-Y, Metelitsa LS, Pique-Regi R, Wakamatsu P, Villablanca JG, et al. Clinical significance of tumor-associated inflammatory cells in metastatic neuroblastoma. J Clin Oncol. 2012;30:3525–32.
Mestdagh P, Fredlund E, Pattyn F, Rihani A, Van Maerken T, Vermeulen J, Kumps C, Menten B, De Preter K, Schramm A, Schulte J, Noguera R, et al. An integrative genomics screen uncovers ncRNA T-UCR functions in neuroblastoma tumours. Oncogene. 2010;29:3583–92.
Schulte JH, Schowe B, Mestdagh P, Kaderali L, Kalaghatgi P, Schlierf S, Vermeulen J, Brockmeyer B, Pajtler K, Thor T, de Preter K, Speleman F, et al. Accurate prediction of neuroblastoma outcome based on miRNA expression profiles. Int J Cancer. 2010;127:2374–85.
De Preter K, Mestdagh P, Vermeulen J, Zeka F, Naranjo A, Bray I, Castel V, Chen C, Drozynska E, Eggert A, Hogarty MD, Izycka-Swieszewska E, et al. miRNA expression profiling enables risk stratification in archived and fresh neuroblastoma tumor samples. Clin Cancer Res. 2011;17:7684–92.
Decock A, Ongenaert M, Hoebeeck J, De Preter K, Van Peer G, Van Criekinge W, Ladenstein R, Schulte JH, Noguera R, Stallings RL, Van Damme A, Laureys G, et al. Genome-wide promoter methylation analysis in neuroblastoma identifies prognostic methylation biomarkers. Genome Biol. 2012;13:R95.
Berbegall AP, Bogen D, Pötschger U, Beiske K, Bown N, Combaret V, Defferrari R, Jeison M, Mazzocco K, Varesio L, Vicha A, Ash S, et al. Heterogeneous MYCN amplification in neuroblastoma: a SIOP Europe Neuroblastoma Study. Br J Cancer. 2018;118:1502.
Abbasi MR, Rifatbegovic F, Brunner C, Mann G, Ziegler A, Pötschger U, Crazzolara R, Ussowicz M, Benesch M, Ebetsberger-Dachs G, Chan GCF, Jones N, et al. Impact of disseminated neuroblastoma cells on the identification of the relapse-seeding clone. Clin Cancer Res. 2017;23:4224–32.
Karlsson J, Valind A, Holmquist Mengelbier L, Bredin S, Cornmark L, Jansson C, Wali A, Staaf J, Viklund B, Øra I, Börjesson A, Backman T, et al. Four evolutionary trajectories underlie genetic intratumoral variation in childhood cancer. Nat Genet. 2018;50:944–50.
Bellini A, Bernard V, Leroy Q, Rio Frio T, Pierron G, Combaret V, Lapouble E, Clement N, Rubie H, Thebaud E, Chastagner P, Defachelles A, et al. Deep sequencing reveals occurrence of subclonal ALK mutations in neuroblastoma at diagnosis. Clin Cancer Res. 2015;21:4913–21.
Schleiermacher G, Javanmardi N, Bernard V, Leroy Q, Cappo J, Rio Frio T, Pierron G, Lapouble E, Combaret V, Speleman F, de Wilde B, Djos A, et al. Emergence of new ALK mutations at relapse of neuroblastoma. J Clin Oncol. 2014;32:2727–34.
Schramm A, Köster J, Assenov Y, Althoff K, Peifer M, Mahlow E, Odersky A, Beisser D, Ernst C, Henssen AG, Stephan H, Schröder C, et al. Mutational dynamics between primary and relapse neuroblastomas. Nat Genet. 2015;47:872–7.
Eleveld TF, Oldridge DA, Bernard V, Koster J, Daage LC, Diskin SJ, Schild L, Bentahar NB, Bellini A, Chicard M, Lapouble E, Combaret V, et al. Relapsed neuroblastomas show frequent RAS-MAPK pathway mutations. Nat Genet. 2015;47:864–71.
Combaret V, Audoynaud C, Iacono I, Favrot M-C, Schell M, Bergeron C, Puisieux A. Circulating MYCN DNA as a tumor-specific marker in neuroblastoma patients. Cancer Res. 2002;62:3646–8.
Chicard M, Boyault S, Colmet Daage L, Richer W, Gentien D, Pierron G, Lapouble E, Bellini A, Clement N, Iacono I, Bréjon S, Carrere M, et al. Genomic copy number profiling using circulating free tumor DNA highlights heterogeneity in neuroblastoma. Clin Cancer Res. 2016;22:5564–73.
Combaret V, Bréjon S, Iacono I, Schleiermacher G, Pierron G, Ribeiro A, Bergeron C, Marabelle A, Puisieux A. Determination of 17q gain in patients with neuroblastoma by analysis of circulating DNA. Pediatr Blood Cancer. 2011;56:757–61.
Van Roy N, Van Der Linden M, Menten B, Dheedene A, Vandeputte C, Van Dorpe J, Laureys G, Renard M, Sante T, Lammens T, De Wilde B, Speleman F, et al. Shallow whole genome sequencing on circulating cell-free DNA allows reliable noninvasive copy-number profiling in neuroblastoma patients. Clin Cancer Res. 2017;23:6305–14.
Chicard M, Colmet Daage L, Clement N, Danzon A, Bohec M, Bernard V, Baulande S, Bellini A, Deveau P, Pierron G, Lapouble E, Janoueix-Lerosey I, et al. Whole exome sequencing of cell-free DNA reveals temporo-spatial heterogeneity and identifies treatment-resistant clones in neuroblastoma. Clin Cancer Res. 2018;24:939.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Schleiermacher, G. (2020). Biology of Neuroblastoma. In: Sarnacki, S., Pio, L. (eds) Neuroblastoma. Springer, Cham. https://doi.org/10.1007/978-3-030-18396-7_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-18396-7_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-18395-0
Online ISBN: 978-3-030-18396-7
eBook Packages: MedicineMedicine (R0)