Abstract
“Lipid raft” is the term applied to transient lipid domains organized laterally within the plane of biological membranes. These structures are thought to range in size from tens to hundreds of nanometers and to be compositionally enriched in high-melting temperature lipids. These characteristics create different physical environments within the “rafts” and in the continuous lipid phase that surrounds them. Based on the principle of hydrophobic mismatch and other properties, membrane proteins preferentially partition into, or out of, the raft, facilitating protein interactions, which in turn regulate downstream cellular processes. Though better understood in the context of the eukaryotic cell membrane, microbial membranes also appear to utilize similar principles to organize their membranes – though their lipid compositions differ considerably from those of eukaryotic membranes. Raft-like structures have been directly observed in microorganisms such as Bacillus subtilis, and a number of functional roles for rafts are being investigated. Among these roles is the organization of proteins associated with antibiotic resistance in Staphylococcus aureus, further highlighting the importance of lipids and lipid rafts, in biology and medicine.
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
1 Introduction
Microorganisms have an emerging and fascinating role to play in the story of lipid rafts. That story begins with the notion that the cell membrane is not the archetypical homogeneous fluid mosaic that was envisioned by Singer and Nicholson (1972). Through the 1970s (Yu et al. 1973; Shimshick and McConnell 1973) and 1980s (Simons and Van Meer 1988), evidence of lipid sorting and detergent-resistant regions within the cell membrane gained wider acceptance which resulted in the proposal of the lipid raft hypothesis (Simons and Ikonen 1997) in the late 1990s. Evidence for lateral organization of microbial membranes emerged in the literature shortly thereafter (Matsumoto et al. 2006), though studies had indicated of the possibility of lateral phase separation earlier (Linden et al. 1973). For example, a heterogeneous distribution of lipids was observed in the septa of mycobacteria (Christensen et al. 1999). In E. coli, membrane domains were visualized in dividing cells (Fishov and Woldringh 1999) with the poles and septa enriched in cardiolipin (Mileykovskaya and Dowhan 2000). Cardiolipin domains were also observed in B. subtilis (Kawai et al. 2004), and more recently isotopic labeling and neutron scattering methods directly probed the lateral heterogeneity of the acyl chains of lipids in B. subtilis (Nickels et al. 2017b) (Fig. 1).
The bacterial cell membrane for a gram-positive organism such as B. subtilis. Gram-positive organisms possess a single-cell membrane, surrounded by a cell wall, and do not typically contain membrane-bound organelles. Bacterial membranes also contain lateral heterogeneities consistent with lipid rafts, which recent experiments indicated are around 40 nm in diameter. (Adapted from Nickels et al. 2017b)
The lipid raft hypothesis asserts that regions within the cell membrane differ in lipid composition and physical properties. Membrane proteins are thought to partition preferentially between lipid domains and the continuous phase that surrounds them, enhancing certain associations of proteins while suppressing others (Sezgin et al. 2017b; Lingwood and Simons 2010). These domains are thought to facilitate a range of biochemical processes (Brown and London 1998), including cell adhesion and migration (del Pozo et al. 2004), cell recognition (Pierce 2002), protein sorting (Lingwood and Simons 2010; Jacobson et al. 2007), synaptic transmission (Hering et al. 2003), cytoskeletal organization (Villalba et al. 2001), signal transduction (Simons and Toomre 2000), and apoptosis (Gajate and Mollinedo 2001). Recently, it has been suggested that lipid rafts function as reservoirs of high-melting temperature membrane components, causing rafts to act as buffers of membrane physical properties (Nickels et al. 2019), a concept which may be applicable to other environmental perturbation such as solvents or small amphiphilic molecules. Because of these biological roles, rafts have been studied extensively in both animal cells (Verkleij et al. 1973; Allen et al. 2007) and microbes (López and Kolter 2010; García-Fernández et al. 2017; Nickels et al. 2017b). But the canonical view of lipid rafts is a mammalian/eukaryotic perspective. Microbial membranes differ from those of eukaryotes in terms of composition, implying that the lipids domains between eukaryotes and prokaryotes differ in composition, even though they may perform similar roles.
As a result, a new frontier of understanding in lateral membrane organization involving bacterial membranes is rapidly approaching. Recent studies suggest lipid rafts are of interest to medical researchers for a variety of phenomena, specifically bacterial and viral pathogenesis (van der Goot and Harder 2001; Dick et al. 2012; García-Fernández et al. 2017; Epand and Epand 2009). For example, work by Lopez (García-Fernández et al. 2017) shows that targeting raft organization using statin class drugs diminishes antibiotic resistance in methicillin-resistant Staphylococcus aureus – a pathogen that is often hospital-associated and notoriously difficult to treat, due to its resistance to common antibiotics.
In this chapter, we will discuss the evolving story of lipid rafts in microorganisms and describe some of the current concepts of lipid raft structure and function in microbial systems.
2 Structure of Lipid Rafts
We can consider lipid raft structure from a number of perspectives. The physical size of domains is an obvious metric to describe rafts in any system, but one can also consider the lipid composition and protein content. Each perspective is a valid way to view the structure embedded within the cellular membrane and to frame how rafts contribute to biological function.
From a eukaryotic perspective, lipid rafts are commonly understood as being heterogeneous (Pike 2004) and dynamic (Samsonov et al. 2001) structures ranging in size between 10 and 100 nanometers (nm) (Pralle et al. 2000) and composed primarily of “high-melting temperature” lipids, but also rich in sterols, carbohydrates, and proteins (Brown and London 2000). However, microbial systems deviate from this picture in several important respects. Compositionally, many microbial systems lack the sphingolipids and sterols found in eukaryotic cells. Bacterial cells are also substantially smaller and possess cell walls and other complex membrane-associated structures, such as flagella.
The size of lipid rafts has been a contentious question since the articulation of the lipid raft hypothesis. Much of the debate stems from an inability to detect nanoscale structures due to limitations in the methods available to study them. More specifically, with the optical resolution limits and the lack of clear rafts in systems which were thought to contain rafts, it was speculated that lipid rafts were nanoscopic in size (Varma and Mayor 1998) quite early on. Besides their small size, rafts present little contrast with the surrounding lipid membrane, presenting an additional analytical challenge in determining a definitive size. However, it is highly unlikely that there is any definitive size for lipid rafts at all. Evidence from model lipid mixtures shows that domain size varies strongly with lipid composition (de Almeida et al. 2005). It is important to keep in mind, therefore, that rafts are expected to be heterogeneous both within a single-cell based due to local composition and curvature fluctuation and across many types of biological membrane which vary widely in composition (Van Meer et al. 2008). This is important to provide context to the reported ~40 nm size of rafts observed for Bacillus subtilis cell membranes (Nickels et al. 2017b) which certainly does not reflect a universal lipid raft size, but rather an average of raft sizes within this specific organism at this specific temperature. The sensitivity of raft size to local composition is illustrated in a simple comparison of model binary and ternary lipid mixtures which produce ordered domain ~20–100 nm and > ~75–100 nm in size, respectively (de Almeida et al. 2005). And while additional factors such as mechanical properties and the inclusion of other molecules also influence the size of these domains, it is widely thought that lipid-lipid interactions are the dominant force.
2.1 Methods
Determining the structural details of lateral lipid organization and lipid rafts is an analytical challenge. There is little inherent contrast between lipid rafts and the surrounding lipids which can be used for observation. The experimental and computational approaches to understanding rafts in microbial systems are broadly similar to those that have been used to study eukaryotic rafts and model membranes. There are a range of direct physical methods performed ex vivo or on model systems such as mass spectrometry (Kraft et al. 2006), nuclear magnetic resonance (NMR) (Veatch et al. 2004), various fluorescence spectroscopy (Bacia et al. 2004; Schwille et al. 1999) and microscopy (Zipfel et al. 2003; Baumgart et al. 2007b; Stöckl and Herrmann 2010; Zacharias et al. 2002; Korlach et al. 1999; Baumgart et al. 2003) methods, atomic force microscopy (AFM) (Yuan et al. 2002; Johnston 2007; Tokumasu et al. 2003; Goksu et al. 2009), and x-ray (Mills et al. 2008) and neutron scattering (Pencer et al. 2005; Nickels et al. 2015a, 2017b; Heberle et al. 2013).
Among these methods for observing lateral organization of lipid bilayer components, fluorescence-based approaches are possibly the most widely used. Although limited by the diffraction limit, fluorescence microscopy (Stöckl and Herrmann 2010) is able to interrogate domain size, shape, and physical properties (Baumgart et al. 2003) based on the partitioning of dye molecules into the different phases (Klymchenko and Kreder 2014) and the diffusion and ligand binding of raft lipid analogs in model and cellular plasma membranes (Sezgin et al. 2012). Fluorescence recovery after photobleaching (FRAP) (Wu et al. 1977) and fluorescence correlation spectroscopy (Bacia et al. 2004) are particularly useful approaches for studying lateral diffusion in bilayers (Derzko and Jacobson 1980) and distinguishing the liquid-ordered phase in raft-forming models (Mouritsen and Jørgensen 1994; Almeida et al. 1992). FRAP works by photobleaching a defined area and measuring the rate at which fluorophores diffuse back into the bleached spot. Because of the small size of lipid rafts, many tools have been employed to probe length scales smaller than the diffraction limit. Confocal and two-photon fluorescence (Gaus et al. 2003) microscopy have been used to directly image large domains, and Förster resonance energy transfer (FRET) (Jares-Erijman and Jovin 2003) has proven extremely useful for studying domain formation and phase behavior (Zhao et al. 2007; de Almeida et al. 2005; Feigenson 2009), as well as identifying raft-associating molecules (Kenworthy et al. 2000). Similarly, other multiphoton methods (Bagatolli and Gratton 2000), a range of super-resolution microscopy techniques (Simons and Gerl 2010), and single molecule (Schütz et al. 2000) approaches are also being developed to overcome the diffraction limit of optical techniques.
Vibrational spectroscopy is another workhorse method that has been applied to questions of lipid organization (Mendelsohn and Moore 1998). The vibrational frequencies of specific functional groups are sensitive to the local environment of lipids and proteins. Because of this, shifts in vibrational frequency and relative amplitudes can be descriptive of lipid order, given sufficient signal to noise ratios. The most widespread vibrational spectroscopy method, infrared spectroscopy (IR), has been used in several studies to detect raft-like domains in model membrane systems (Schultz and Levin 2008; Lewis and McElhaney 2013). Raman scattering has been used on lipid systems for many years (Czamara et al. 2015) with recent developments being applied to asymmetric and laterally organized lipid bilayers. Tip-enhanced Raman spectroscopy is an attractive way to enhance the observable signal, especially when combined with isotopic substitutions (Opilik et al. 2011). Multiphoton Raman methods, such as coherent anti-Stokes Raman spectroscopy (Potma and Xie 2005; Li et al. 2005), have also been used to observe lateral demixing in the plane of model lipid membranes. Multiphoton methods may provide some advantages over standard Raman in terms of lowering excitation power, resulting in less potential damage to the sample. Nonlinear methods, such as sum frequency vibrational spectroscopy, have been used to study bilayer asymmetry (Brown and Conboy 2013) and lipid phase behavior (Liu and Conboy 2004). Lipid asymmetry is another key organizational feature of the cell membrane (Nickels et al. 2015b). This method clearly observes a spectral indication of bilayer asymmetry, but the resulting flip-flop rates appear to be quite a bit faster than other methods (Heberle et al. 2016). This may be due to the presence of defects in the supported lipid bilayers needed for this method. It has been known for several decades that pore formations dramatically increase the rate of lipid flip-flop (Fattal et al. 1994). Lipid diffusion through such defects would be a competing mechanism with true transmembrane flip-flop in such systems.
Here we focus on experimental approaches used to study microbial membranes, but molecular dynamics (MD) simulations are another important approach that is beginning to be used. We mention this because the next two methods we will discuss, nuclear magnetic resonance (NMR) and scattering (x-ray and neutron), are the most conducive to comparison with MD simulation. The time and length scales of the resulting observations are nearly the same as what current simulations can probe. NMR, specifically solid-state NMR for lipid systems, provides detailed information about molecular identity, local structure and structural uncertainty, and molecular motions (Davis 1983; Seelig and Seelig 1974) based on the response of atoms to an applied magnetic field. Several different elements (Henderson et al. 1974; Petrache et al. 2000; Davis 1983) are used for NMR in the direct study of lipids, commonly including 1H, 2H, 13C, and 31P, and there are many methods and protocols within the overall umbrella of NMR. The addition of membrane insoluble paramagnetic compounds is useful for determination of bilayer asymmetry (Solomon 1955; Bloembergen 1957). If a paramagnetic shift reagent is added to the bulk solution outside of the vesicle, such as a lanthanide ion, there will be a detectible shift in the NMR spectra for one leaflet (Su et al. 2008; Solomon 1955; Bloembergen 1957). Shift reagents, also used to track asymmetric membrane protein insertion (Su et al. 2008), were recently been used to track lipid flip-flop in asymmetric vesicles (Heberle et al. 2016), addressing some potential issues described in the preceding paragraph.
For the case of lateral organization, 2H NMR has played an important role in quantifying the compositional changes within phase separating model lipid mixtures (Veatch et al. 2004). It should be noted that these authors were not comfortable ascribing this result to strict phase coexistence, citing the possibility of critical fluctuations within a single phase. Subsequent observations of nanoscopic compositional heterogeneities in similar model lipid compositions using scattering methods indicate that there is a population with ensemble properties of a nanoscopic size and local compositions resembling the distinct lipid phases. Carbon-deuterium order parameters of lipid acyl chains are a tremendously useful reporter on the structural order of the fatty acid tails in the hydrophobic region of a lipid bilayer. This parameter is then an obvious target to connect NMR experimental results and MD simulations describing lipid acyl chain order because the carbon-deuterium order parameter reflects the average angle between the C-D bond and the bilayer normal. The results of NMR and MD simulations seem to be in good agreement at present, and while comparisons of order parameters for lipid head groups are not yet well modeled, current efforts can be followed at http://nmrlipids.blogspot.com. Parameters describing lipid acyl chain order can also be obtained by fluorescence depolarization (Heyn 1979), yet the requirement of a fluorescent probe does introduce an outside element, perturbing the system and potentially affecting the result. NMR has the benefit of using isotopic labels which are far less perturbing. Using such labeling, one may access the T1 relaxation time revealing similar information about lipid ordering (Brown et al. 1983). Heteronuclear, or cross polarization, methods where polarization is transferred between two atoms are another strategy for obtaining information about the ordering of the lipid tails (Hong et al. 1995). The use of magic angle spinning (MAS) conditions should also be mentioned as a way of dramatically enhancing the signal of the system and avoiding the necessity of oriented samples (Gross et al. 1997). MD simulation and NMR are also well matched for the description of local motions of lipids in bilayers. MD simulations can compute diffusion confidents directly from the atomic displacements in their trajectories. The pulsed field gradient method has been used to measure the lateral diffusion rate of lipids and cholesterol in the plane of the bilayer (Orädd and Lindblom 2004; Scheidt et al. 2005). This has been applied to the dynamical properties of the lipids within model raft-forming mixtures as well (Orädd et al. 2005; Lindblom et al. 2006).
Neutron and x-ray scattering are powerful structural biology techniques that have been used extensively in the determination of the lipid membrane structure (Marquardt et al. 2015) (Fig. 2). Scattering methods probe important molecular length scales (Å to 100s of nm), and inelastic scattering techniques are able to probe molecular motions occurring from the order of 100s of nanoseconds to fractions of a picosecond. The coherent elastic scattering of both x-rays and neutrons reflects the pair distribution functions of the atoms making up the sample. Such pair distribution functions reflect the molecular structure and can be directly computed from MD simulations. Indeed, the lamellar structure of certain lipid phases (Luzzati and Husson 1962) and liquid-like conformations of the lipid tails in the hydrophobic region (Luzzati and Husson 1962; Tardieu et al. 1973; Engelman 1971) were observed by x-ray diffraction. X-ray diffraction played a critical role in quantifying the linear relationship between acyl chain length and bilayer thickness in lipid bilayers (Lewis and Engelman 1983) and remains a critical structural technique for biomembranes (Nagle and Tristram-Nagle 2000). X-ray diffraction is sensitive to the existence of lateral organization for some lipid systems (Koenig et al. 1997), including complex lipid mixtures (Majewski et al. 2001). The way that lateral organization can be implied from x-ray scattering observations (Heftberger et al. 2015) is usually the presence of multiple d-spacings at lower scattering wave vectors.
Neutron scattering is an emerging method for the study of lipid rafts in bacteria. (a) Neutrons scatter differently from the isotopes of hydrogen, 1H and 2H (deuterium). By careful isotopic substitutions, it is possible to manipulate the scattering contrast of an entire living bacterial cell, such that contrast between rafts and the surrounding membrane is observed. (b) The lamellar form factor of the cell membrane can be observed by labeling a partially deuterated cell’s membrane with 1H fatty acids, revealing the bilayer thickness in the living cell. (c) Raft structures can be detected based on differences in partitioning of neutron scattering length density matched fatty acid mixtures in the cell membrane. Here we see the excess scattering induced by unequal partitioning of the isotopes of hydrogen. (Adapted from Nickels et al. 2017b)
Similar to x-ray scattering, neutron scattering is a diffraction technique that can probe length scales from 100s of nm to Å; however, there are some important distinctions. Firstly, neutrons scatter from the atomic nucleus with a strength denoted in the nuclear scattering length, b, whereas x-rays scatter from the electron density. Secondly, thermal and cold neutrons are more often used to study dynamics via inelastic scattering. The observed changes in momentum (differences in the velocity of the incident and scattered neutron) can be directly related to the collective and self-motions of the atoms in the sample (Bee 1988). Note that inelastic scattering of x-rays is also a useful method to probe atomic motions, but typically at very high frequencies and longer wave vectors than neutrons, making the method less useful for lipids. The neutron scattering length, b, varies widely between elements and even between the isotopes of a single element (Sears 1992), in contrast to x-ray scattering where electron density varies systematically with atomic number. This has two important implications in regard to the study of lipids and complex lipid structures. Firstly, this means that there is a substantial contribution to the observed scattering from low atomic number atoms such as hydrogen. This matters because lipids are so hydrogen-rich. Indeed, neutrons have been a vital tool in studying the detail structure of biomembranes (Büldt et al. 1978, 1979; Zaccai et al. 1979), including the location of another hydrogen-rich molecule in biological systems, water (Zaccai et al. 1975; Nickels and Katsaras 2015), and the motions of water around biointerfaces (Toppozini et al. 2015; Nickels et al. 2012; Perticaroli et al. 2017; König et al. 1994; Settles and Doster 1996; Swenson et al. 2008). The inelastic scattering of neutrons provides additional information in the time domain about the motions of lipid molecules over a wide dynamic range from the collective undulations (Watson and Brown 2010; Zilman and Granek 1996) and thickness fluctuations (Woodka et al. 2012) of the bilayer, to lateral diffusion of lipids (Pfeiffer et al. 1989) and local motions of the individual lipids (Pfeiffer et al. 1989; König et al. 1992; Swenson et al. 2008), to the vibrational regime (Rheinstädter et al. 2004). These motions are correlated to important physical properties of lipid membranes, such as the bending modulus (Zilman and Granek 1996).
The second important point is isotopic substitution, specifically the isotopes of hydrogen (1H) and deuterium (2H). These molecules can be deployed to achieve “contrast-matching ” conditions (Jacrot 1976). In the case of complex lipid compositions, contrast matching typically means introducing deuterium substituted lipids in carefully calculated amounts into specific lipid phases. Calculations of atomic/isotopic content are used to match the scattering length density of a water/heavy water solvent mixture to a single given phase (Nickels et al. 2015a) or the theoretical match point of bilayer when perfectly mixed (Heberle et al. 2013; Nickels et al. 2017b; Pencer et al. 2005). In the latter case, a null scattering condition should be observed, and deviations from this imply lateral demixing of the bilayer. This approach has been used in other guises for many applications however, for example, in the study of proteins (Jacrot 1976), lipids (Knoll et al. 1981), nucleic acids (Pardon et al. 1975), polymers (Nickels et al. 2016), and other materials. There is recent work using this kind of neutron scattering data to carefully study the sizes of lipid domains using these contrast-matching approaches (Heberle et al. 2013; Nickels et al. 2015a). Indeed, contrast-matched vesicles are useful to study how the modulus of nanoscopic lipid domains is distinct from the surround phase of phase-separated bilayer (Nickels et al. 2015a). This work suggested that it is possible that phase separation can be influenced by the local distribution of elastic moduli. Again, the observed dynamical features are directly connected to the time dependence of the self and pair correlation functions, properties accessible in MD simulations. By defining those correlation functions for specific modes, one can extract dynamical terms analogous to these experimental observations.
Not surprisingly, technical limitations frustrate many of these powerful physical techniques in the biologically relevant living cellular environment. Ex vivo approaches to get around these limitations, such as the study of giant plasma membrane vesicles derived from living cells via chemical treatment, show clear indications of large-scale liquid-liquid phase separation (Baumgart et al. 2007a). Also, natural lipid extracts (Nickels et al. 2017a) and synthetic model lipid mixtures which exhibit microscopic liquid-liquid phase coexistence (Veatch and Keller 2002, 2003; Dietrich et al. 2001; Edidin 2003) have made careful study of the physical processes behind lipid rafts a tractable problem (Jacobson et al. 2007).
It is clearly desirable to study lipid rafts in living cell membranes, resulting in a number of experimental approaches being employed. Early in the study of lipid rafts, detergent resistance was found to be a useful way to isolate raft lipids and raft cargo in both model lipid mixtures and cells (Ahmed et al. 1997). Detergent resistance (Lingwood and Simons 2007; London and Brown 2000) uses various detergents, such as Triton-X100, to solubilize regions of the cell membrane. Interestingly, not all portions of the membrane are disrupted resulting in detergent-soluble and detergent-resistant membrane fractions. These fractions can subsequently be interrogated via mass spectrometry and other biochemical assays (Lingwood and Simons 2007). A recent example of this approach might be the isolation of staphyloxanthin in the detergent-resistant Staphylococcus aureus membrane (García-Fernández et al. 2017). Another indirect method which was important in the early study of lipid rafts is cholesterol depletion. Cholesterol depletion studies demonstrated changes in cellular behavior or activity (Kabouridis et al. 2000) based on the removal of cholesterol, presumably from lipid rafts. However, the vast majority of microorganisms do not contain sterols in their cell membranes, instead using functional analogs such as hopanoids (Sáenz et al. 2012) and isoprenoid lipids (Boucher et al. 2004; García-Fernández et al. 2017). High-resolution, two-photon fluorescence spectroscopy is one such approach at tracking lipid raft cargo in living cells (Gaus et al. 2003). The method is based on the differential partitioning of the small hydrophobic dye laurodan (Owen et al. 2012). Other fluorescence-based methods using protein-linked molecules showed the cycling of raft-associated membrane components in living cells (Nichols et al. 2001). These methods provide powerful, if indirect, evidence of the existence of lipid rafts and optical length scale structural information. Recently, neutron scattering was used to directly observe lateral demixing in the cell membrane of a living bacterium, confirming the presence of nanoscopic heterogeneities consistent with the concept of lipid rafts (Nickels et al. 2017b).
2.2 Composition
The composition of lipid rafts is often understood in the context of high cholesterol and sphingomyelin/high-melting lipids. This is a eukaryote centric view, however, and does not apply to the majority of microbial membranes. From a bacterial membrane perspective, one frequently thinks lipids such as phosphatidylglycerol (PG), phosphatidylethanolamine (PE), and cardiolipin (CL) are common in the membrane of E. coli (Gidden et al. 2009; Parsons and Rock 2013), the gram-negative bacterial model, and in that of B subtilis (Gidden et al. 2009; Nickels et al. 2017a; Den Kamp et al. 1969), a typical gram-positive model organism. Yet, there are many other molecules to consider (Fig. 3). Microbial membranes are overwhelmingly diverse in terms of their lipid compositions (Sohlenkamp and Geiger 2016), even in the model organisms such as E. coli and B. subtilis. In the context of lipid rafts, these other lipids, such as hopanoids and prenyl lipids, are important as analogs to sterols (Sáenz et al. 2012). Other varieties of lipid include ornithine lipids, sulfonolipids, or the many varieties of glycolipids (GL). Free fatty acids and other small molecules can also be found in some bacterial membranes. Ether lipids are also found in a range of bacterial species. Common in archaea, bacterial ether lipids contain fatty acyl and acyl modifications in the sn-1 and sn-2 positions. The distribution of these and other less frequent bacterial membrane components across the range of organisms is summarized well by Sohlenkamp and Geiger (Sohlenkamp and Geiger 2016) in Table 1 of their excellent review.
Bacteria contain a wide range of lipids. This includes typical glycerophospholipids such as phosphatidylglycerol (PG), phosphatidylethanolamine (PE), and cardiolipin (CL), as well as the metabolic intermediate diacylglycerol (DAG). Terpenoids and prenyl lipids (like staphyloxanthin) are found in many bacterial membranes, some of which are thought to be important analogs to sterols. Lipopolysaccharides are typically found in the outer membrane of gram-negative species. A range of fatty acids are also employed across bacterial species, including branched chain and cyclopropane fatty acids
Glycerophospholipids are the most common lipid species in most cell membranes. This group includes those mentioned above, PE, PG, and CL, as well as other molecules such as phosphatidylserine (PS), phosphatidylinositol (PI), phosphatidylcholine (PC), and the lysyl versions of these lipids, such as lysyl-PG (LPG). Diacylglycerol (DAG) is another component of bacterial membranes and an important metabolic intermediate in lipid synthesis (Carrasco and Mérida 2007). DAG can be enzymatically modified to obtain phosphatidic acid (PA) and subsequently cytidine diphosphate-diacylglycerol (CDP-DAG) which is the central metabolite in the production of glycerophospholipids. Because of this, DAG is the backbone of GLs and the anchor for lipoteichoic acid, an important structural component of the cell wall (Percy and Gründling 2014). Lipopolysaccharides (LPS), such as Lipid A, are typically found in the outer membrane of gram-negative bacteria. LPS is often a toxin that triggers innate immunity (Alexander and Rietschel 2001) in humans and is reported to cluster in mammalian lipid rafts (Pfeiffer et al. 2001).
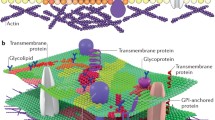
Local differences between the global lipid composition and the local composition are of course the central feature of lateral organization (Fig. 4). The physical properties of lipids drive such differences, with acyl thickness and spontaneous curvature mismatches as potential driving forces for raft formation. DAG, for instance, may have a role in raft formation due to its negative curvature, while lysyl lipids have a large positive spontaneous curvature. Other lipids, such as CL (Mileykovskaya and Dowhan 2000) and PE (Nishibori et al. 2005), have been demonstrated to localize in the cell membrane of bacterial cells. But the correlation from raft-forming lipid mixtures is still being investigated in living bacteria, and most anticipate that there will be a far greater diversity of raft-forming compositions than is seen in mammalian membranes.
Schematic of a lipid raft with associated transmembrane protein. This simplified composition belies the reality that much remains to be discovered about the varied compositions of lipid rafts in bacteria. Some bacterial membranes are thought to contain rafts enriched in phosphatidylethanolamine (PE) (Nishibori et al. 2005), cardiolipin (CL) (Mileykovskaya and Dowhan 2000), or some terpenoids, like staphyloxanthin (Sáenz et al. 2012) (protein shown is a LPS transporter from Pseudomonas aeruginosa (Botos et al. 2016))
The dependence of rafts upon sterols in eukaryotes, typically cholesterol in humans, is a distinguishing feature of lateral organization. Ergosterol is thought to regulate membrane properties to much the same degree in fungi, protozoans, and yeasts (Xu et al. 2001), with observations that a membrane with little to no cholesterol will exhibit little to no raft formation (Rothberg et al. 1990). Sterols are major component of the membrane in eukaryotes, and their presence and absence throughout the lipid bilayer plays an important role in the functions and behaviors of not only lipid rafts but with the membrane as a whole. Their amphipathic structure allows them to interact with both the nonpolar acyl chain regions and polar head group regions of the cellular membrane. Sterols also have a substantial influence over the biophysical properties of the membrane, such as bilayer fluidity, thickness (Kusumi et al. 1983; Vance and Vance 1985) permeability, bending modulus, and protein complex conglomeration (Haines 2001; Nickels et al. 2015a; Simons and Toomre 2000).
Unsurprisingly, prokaryotes are thought to utilize a range of compounds to perform similar functions. Sterols are produced in a limited number of microorganisms; while some organisms have been shown to scavenge sterols from the extracellular environment. We will discuss one such case for B. burgdorferi later in this chapter. Isoprenoid lipids, also called terpenoids, are a broad class of compounds that are thought to play some role in raft formation in bacterial membranes. Their functional convergence has been long speculated, and experimental evidence is now emerging (Sáenz et al. 2012). In fact, sterols are synthesized from isoprenoid precursors such as squalene.
In prokaryotic membranes isoprenoid lipids are increasingly being associated with rafts and raft-associated cellular functions, such as membrane trafficking, protein localization, and signal transduction (Lopez and Koch 2017). Hopanoids are a specific class of isoprenoid lipid – triterpenoids – which has been strongly implied to function analogously to sterols. Hopanoids are a diverse set of molecules found widely among bacteria (Rohmer et al. 1984). Diplopterol is one example used by Saenz and coworkers (Sáenz et al. 2012) to illustrate experimentally the biophysical similarity. They compared the membrane ordering effect in model sphingomyelin/cholesterol and sphingomyelin/diplopterol bilayers. This is not a universal effect across all hopanoids, however, as was pointed out subsequently in simulations of PC lipid bilayers containing diplotene or bacteriohopanetetrol. There, the authors showed how the latter molecule accumulated in the bilayer midplane, having no ordering effect at all (Poger and Mark 2013). However, such effects are also seen for the location of vitamin E (Marquardt et al. 2013) as a function of chain length, so it is possible that bacteriohopanetetrol would partition differently in other model bilayers.
Carotenoids are another subclass of isoprenoid lipid – also called tetraterpenoids – based on eight isoprene units. Carotenoids are important molecules in photosynthetic organisms due to their light-absorbing properties, especially in the 400–550 nm wavelength range. Carotenoids typically span the lipid bilayer and do not have as many ring structures as hopanoids. Nonetheless, carotenoids have also been implicated in the maintenance of lipid rafts. Garcia-Fernandez et al. showed that carotenoids, or absence thereof, are a key factor in formation of rafts using a carotenoid-deficient Staphylococcus aureus mutant. They demonstrate the pivotal role rafts play in the mechanism of antibiotic resistance (García-Fernández et al. 2017).
Proteins are another major component of the cell membrane, with integral membrane proteins accounting for 20–30% of genes (Krogh et al. 2001). This is true for bacteria as much as for eukaryotes (Daley et al. 2005). In the context of this chapter, we are interested in the organization of membrane proteins and their recruitment to rafts. Membrane proteins broadly come in two categories: integral membrane proteins and peripheral membrane proteins. Integral membrane proteins can only be removed by membrane disruption. This includes proteins which cross the full bilayer like ion channels or receptors involved in cell adhesion or active transport of metabolites. Transmembrane domains typically contain hydrophobic residues and have conserved sequences, making their identification possible from sequence analysis tools. Peripheral membrane proteins are those which can be extracted from the membrane without fully disrupting the bilayer and do not typically cross the full bilayer but are instead associated with one leaflet through acylation, ionic interactions, or specific affinity to membrane components.
Membrane proteins are sensitive to bilayer thickness (Andersen and Koeppe 2007), and this is thought to be the energetic driver of protein partitioning between raft and “sea” phases of the cell membrane. These phases present different hydrophobic thickness which will provide better solvation matching to the hydrophobic portions of membrane proteins. Because of this, there are structural motifs which have become associated and are potentially predictive for proteins which are found in lipid rafts. The set of proteins described by the acronym, SPFH, is one such domain, describing the group of proteins stomatin, prohibitin, flotillin, and HflK/C (Browman et al. 2007). Evidence has shown that proteins containing these SPFH segments may interact with other proteins to localize binding partners to the raft and impact their organization once inside it (Langhorst et al. 2005, 2007; Browman et al. 2007). Another important category of raft-associated proteins is the glycosylphosphatidylinositol (GPI)-anchored proteins. These are integral membrane proteins, typically containing long acyl chain and doubly acyl chain modifications; and though it varies greatly based on activity, GPI-anchored proteins have been reported to exist within rafts at concentrations up to 15 times greater than the surrounding membrane (Varma and Mayor 1998).
3 Functions of Lipid Rafts
Lipid rafts are the subject of ongoing research, and their role in biological systems is varied and evolving. Some aspects of lipid rafts remain under debate, such as the mechanism of raft formation, phase behavior (Feigenson 2009), or active biological process (Yethiraj and Weisshaar 2007; Turner et al. 2005). Yet the core concept connects lipid organization to protein co-localization, and protein co-localization to cellular function is now widely accepted. The physical properties of coexisting lipid phases and their incorporated proteins create distinct characteristics of both raft and non-raft regions. It is hypothesized (Nickels et al. 2019) that rafts may function to stabilize physical properties against rapid changes in temperature or solvent. Membrane thickness, curvature, tension, and other properties are products of these interactions. Further, the partitioning of proteins in the bilayer based on their structural features along with active sorting methods allows for the regulation of a range of cellular processes. This mediation influences signaling pathways, transport mechanisms, cellular division, and other membrane functions.
3.1 Defining Membrane Physical Properties
The understanding of lipid rafts as coexisting lipid phases in the cell membrane plays a key role in defining the physical properties of rafts and maintaining the properties of the cell membrane as a whole. As noted above, bacterial membranes are compositionally diverse (Sohlenkamp and Geiger 2016), with a given organism using potentially hundreds of lipids. In order to simplify the system for biophysical study, model lipid bilayers have emerged as an excellent resource for the study of lipid bilayer phase separation, with a rich literature emerging since the year 2000. Commonly using ternary lipid mixtures (Feigenson 2009; Veatch and Keller 2003; Marsh 2009) containing one high-melting lipid, one low-melting lipid, and cholesterol, these studies identified the liquid-ordered phase, Lo, and the liquid-disordered phase, Ld, as analogs to the biologically relevant raft phase. LO and LD phases have numerous physical differences beyond their composition, such as the local lipid order (acyl chain, tilt, splay), lateral compressibility, hydrophobic thickness, and bending modulus (Nickels et al. 2015a). Moreover, the precise combination of these properties contribute to the interfacial energy between the phases, quantified in the line tension and partition coefficients of the various molecules within the bilayer (Honerkamp-Smith et al. 2008; Kuzmin et al. 2005; García-Sáez et al. 2007).
As might be guessed from the nomenclature, lipids in the Lo phase are more ordered as defined by a number of order parameters. In general, this means that lipid acyl tails in the Lo phase have on average a more trans configuration and are more likely to be oriented parallel to the bilayer normal. In the case of typical diacyl lipid molecule, the acyl tails have parallel orientations with respect to each other when compared to lipids in the Ld phase. This ties quite directly into differences in bilayer thickness, compressibility, and bending modulus. The bending modulus describes the resistance of the bilayer to deformation out of the plane of the bilayer and is directly related to the lateral compressibility and bilayer thickness. The bending modulus and the related Gaussian modulus have an influence on the global bilayer free energy (Helfrich 1973) and have been indicated as one potential driver of lateral organization (Seul and Andelman 1995; Nickels et al. 2015a).
Membrane protein hydrophobic mismatch is a common concept for discussion of lipid rafts discussed in the prior and subsequent sections of this chapter; but lateral pressure profiles exerted upon membrane proteins are another potentially important influence of lipid rafts. For example, it has been suggested that the insertion of ketamine in lipid bilayers changes the opening probability for model ion channels (Jerabek et al. 2010), supporting the lateral pressure mechanism of anesthesia (Cantor 1997). Changes in protein activity based upon conformational changes induced by the lateral pressure profile of the lipid environment are another way that lipid rafts can influence cellular behavior. Finally, bending modulus and the diffusivity of proteins within the membrane can be influenced by the existence of distinct lipid phases. Indeed, partitioning of lipids between phases continually changes as a function of temperature leading to a thermal buffering of bilayer rigidity and bilayer fluidity (Nickels et al. 2019) .
3.2 Protein Partitioning
The functionality of lipid rafts is premised upon their role as a mechanism of preferential protein sorting. The origins of this mechanism are rooted in fundamental thermodynamic interactions. When two membrane domains with differing hydrophobic thickness form in the presence of a protein, the protein will preferentially accumulate within the bilayer region which more closely matches their hydrophobic thickness (Fig. 5). This will tend to co-localize proteins with similar characteristics, leading to a clear mechanism by which multiple proteins can be segregated together in a certain membrane region.
The physical characteristics that help define the hydrophobic thickness of a lipid raft are governed by a combination of factors which depend upon composition of lipids found in the specified domain, namely, the lengths of the acyl tails of its component lipids, the chemistry of the lipid head groups and their interaction with water, and the lateral packing of the constituent lipids. The less that membrane proteins and lipids must adapt to reduce any exposure of hydrophobic regions to the aqueous environment outside of the bilayer, the more energetically advantageous the protein partitioning will be. Various mechanisms are employed in order to minimize these mismatches and resultant strain energies. For example, small conformational changes in trans- and intramembrane proteins can alter their overall length and reduce the impact of these mismatches. Additionally, local deformation of the lipid bilayer allows for reduction of line tension at the phase boundary.
Protein partitioning into a lipid raft is just one step in sequence of events leading to various cellular functions. A raft associated protein frequently collocates with other proteins to form protein complexes. Detergent resistance has identified a number of protein complexes prokaryotic cells. In B. subtilis the resistant protein complexes include signaling proteins, biosynthesis machinery, and transporter complexes (López and Kolter 2010; Niño and Marc 2013). Further study revealed that there was direct interaction between these proteins and flotillins, like FloT and FloA, indicating co-localization of specific proteins. Functional dependences were observed as protein function was inhibited when flotillins (FloT and FloA) were not present (López and Kolter 2010; Niño and Marc 2013; Schneider et al. 2015). Flotillins belong to the SPFH (stomatin/prohibitin/flotillin/HflK/C) superfamily of proteins, which share a region of undetermined function but similar sequence (Tavernarakis et al. 1999). Flotillins are distinct in their coiled structures at the C-terminal (Schroeder et al. 1994).
In many single-cell organisms, flotillins are thought to mark lipid raft locations (Fig. 6). Through the use of electron microscopy and staining-based methods, these proteins can be easily observed to localized domains across the cellular membrane (Lang and Philp 1998; Stuermer et al. 2001). These domains tend to be uniform in diameter and very similar in size to lipid rafts, approx. 100 nm across. While they relatively evenly spaced across cellular membranes, they tend to conglomerate more at cellular contact sites (Stuermer et al. 2001). Flotillins have been observed to co-localize with other GPI-anchored proteins like F3/contactin and others, suggesting that they are involved in the ferrying of GPI-anchored proteins across the cell membrane. Additionally, they appear to closely interact with kinases, transport, and several types of receptor signaling mechanisms (Stuermer et al. 2001; Wakasugi et al. 2004).
Localization of flotillin-GFP fusion protein to membrane rafts in B. subtilis cells expressing Pspac-cfp reporter. Fluorescent microscopy image from Bramkamp and Lopez (2015)
3.3 Selected Examples
We now discuss several specific examples of rafts, or raft-like domains, either in the membranes of, or interacting with, microorganisms. Lipid rafts in both eukaryotes and bacteria alike play important roles in the mechanism of pathology. Their ability to co-localize specific complexes of proteins is especially useful as a vector for cellular interaction and invasion of a host cell, or responding to an environmental challenge, such as an antibiotic. Moreover, microbial pathogens have been able to evolve many complex strategies to avoid immunological detection and responses within their hosts.
3.3.1 Vibrio cholera and Cholera
Vibrio cholera is a gram-negative bacterium that causes the disease chorea. This bacterium produces a protein toxin called cholera toxin and is associated with a membrane binding B-subunit. CTxB (cholera toxin B-subunit) has become an important and frequently studied example of raft-associated proteins. CTxB can bind up to five GM1 gangliosides, by targeting the toxin to host cells. The GM1 gangliosides tend to be associated with lipid raft regions of the plasma membrane, and subsequent endocytosis can occur through GM1-rich caveola structures (Parton 1994) or clathrin-coated pits (Nichols 2003). Once internalized, a reverse translocation from the plasma membrane to the Golgi complex occurs (Nichols et al. 2001). The A1 and A2 subunits of CTxB are cleaved, triggering the activation of the A1 enzymatic fragment. This fragment activates and locks the G protein of Gsα in a GTP-bound form, resulting in a continuous activation and stimulation of the enzyme adenylate cyclase to produce cAMP. This triggers a large excretion of water and ions out of infected cells through the opening of cAMP sensitive channels.
The targeting mechanism of CTxB to GM1-enriched areas of the plasma membrane allows them to be used as lipid raft markers (Nichols et al. 2001) in a productive use of the molecule. Due to the multivalent binding, the toxin has the capability of recognizing the underlying membrane structure by cross-linking small and ephemeral lipid rafts (Day and Kenworthy 2015). The enrichment of their receptors in rafts suggests that bacterial products such as toxins bind preferentially to detergent-resistant highly ordered plasma membrane regions to access the cell (Sezgin et al. 2017a). This suggests that CTxB actively changes the properties of the membrane upon binding. Fluorescence resonance energy transfer (FRET) microscopy can then be used to detect the presence of lipid rafts. FRET was detected between molecules of the glycosphingolipids GM1 labeled with cholera toxin B-subunit and between antibody-labeled GPI-anchored proteins, exhibiting these raft markers are in submicrometer proximity in the plasma membrane. In the plasma membrane, lipid rafts either exist only as transiently stabilized structures or, if stable, comprise at most a minor fraction of the cell surface (Kenworthy et al. 2000). CTxB thus serves as a useful model for understanding the properties and functions of protein-stabilized domains.
3.3.2 Borrelia burgdorferi
While each bacterial and eukaryotic membrane has its own unique composition of lipids in order to facilitate lipid raft and protein complex formation, some bacteria have been found to go to extreme lengths to create proper functioning conditions for lipid rafts. One example is that of Borrelia burgdorferi. Known as the bacteria that cause Lyme disease, the Borrelia genus was found to contain types of glycolipids rarely found in bacterial cells and thought only to be used in eukaryotic cellular bilayers. Of three specific glycolipids identified in B. burgdorferi, two were found to contain cholesterol. These glycolipids were identified as cholesteryl 6-O-acyl-β-D-galactopyranoside (ACGal), cholesteryl-β-D-galactopyranoside (CGal), and mono-α-galactosyl-diacylglycerol (MGalD) (Ben-Menachem et al. 2003). Upon investigation these glycolipids were also found to exist in other Borrelia species as well, showing a similarity in functional regulation of the lipid bilayer across the entire genus. This presence of cholesterols in the Borrelia genus suggests that the characteristic microdomains of sterol-, or sterol analog, rich areas found in eukaryotic bilayers may be found inside bacterial bilayers as well. Further, the complexes of proteins found in the liquid-ordered phase of B. burgdorferi suggest a sensory or signaling function due to the environmental factors (LaRocca et al. 2010).
Fascinatingly, B. burgdorferi lack the cellular machinery to produce these molecules. This means that the acquisition of sterols for the outer membranes of these cells is a process which must be undergone via lipid transfer from a host eukaryotic cell. This is supported by its inability to synthesize long-chain-saturated, unsaturated fatty acids, or cholesterol analogs, paired with the cell’s lipid membrane concentration mirroring those of its host fluid or tissue (Johnson 1977). The similarity in lipid composition of B. burgdorferi and its eukaryotic host allow for lipid-lipid interactions between the two membranes, especially in lipid raft domains, where the similar interactions would be highest. These spirochetes use this interaction as an important mechanism for nutrition acquisition, which includes the sterol and other fatty acid stripping from the host cell (Crowley et al. 2013). This interaction was observed by Crowley through the use of fluorescently labeled fatty acids before and after Borrelia interaction with host cells. The presence of labeled sterols on both host and B. burgdorferi after incubation confirmed this hypothesis (Crowley et al. 2013). This mechanism is thought to be responsible for the cellular damage that results from Lyme disease.
3.3.3 Staphylococcus aureus
The human pathogen S. aureus, commonly known as MRSA, is a notable example of complex microdomain interactions. This bacterium uniquely expresses a single flotillin, FloA (Marshall and Wilmoth 1981; Pelz et al. 2005; Wieland et al. 1994), and has been able to develop an ability to successfully resist antibiotic treatments, causing its iconic hard to treat infections in hospitals. Antibiotics such as methicillin (a β-lactam antibiotic) have been shown to be ineffective versus strains of S. aureus via the production of low-affinity penicillin-binding protein (PBP2a). This protein, by binding with the antibiotic, is able to prevent it from inhibiting PBP active sites, which are essential to cellular division. In this way PBP2a allows MRSA strains to replicate and grow in the presence of antibiotics (Kreiswirth et al. 1993).
A key component of this mechanism is FloA, as it identified to be the singular flotillin driving protein recruitment in these microdomains (Marshall and Wilmoth 1981; Pelz et al. 2005). Preferential binding of FloA to staphyloxanthin (a lipid produced in S. aureus) leads to its oligomerization in staphyloxanthin-rich microdomains. As these areas grow and assemble FMMs, PBP2a is taken as protein cargo of FloA, concentrated in these regions, and is ultimately used to inhibit antibiotics (García-Fernández et al. 2017). As FloA conglomeration increases scaffold-forming activities in the bilayer, an increase in PBP2a organization is also seen. This relationship is important because the flotillin dependence of PBP2a localization means that the lack of FloA causes inhibition of protein recruitment. By disrupting the FMM formation in S. aureus, the antibiotic resistance of MRSA bacteria was decreased significantly, and conventional antibiotic therapy again became effective (García-Fernández et al. 2017).
4 Research Needs and Conclusions
The term “microbial membrane” refers to an almost impossible diverse array of biomembranes. There are a huge variety of lipid compositions (Sohlenkamp and Geiger 2016) that need to be investigated, and efforts to describe the properties of the molecules involved in microbial lipid rafts should be supported. Isoprenylated species, cyclic and branched chain fatty acids, methylated lipid head group chemistries, and glycosylated lipids anchoring a cell wall all differ substantially from the eukaryotic picture. Understanding properties like the hydrophobic thickness and spontaneous curvature of these molecules will enable a more direct comparison to the better studied, mammalian centric, semi-canonical view of lipid rafts.
Beyond compositional considerations, another vital and broad-reaching research need is for robust and reliable methods for detecting membrane domains and identifying protein content. Fluorescent strategies relying on co-localization of proteins have convincingly proven the existence and functional roles of lipid domains (Kenworthy et al. 2000). Unfortunately, the resolution limits of optical microscopy impact the utility of these methods in determining the sizes of these structures. Advances in super-resolution microscopy are overcoming these challenges (Huang et al. 2008; Hess et al. 2006), but the need for exogenous fluorescent probes remains potentially problematic.
Neutron scattering has been a valuable, and fluorophore free, tool for studying lipid raft structure and physical properties both in vitro (Nickels et al. 2015a) and in vivo (Nickels et al. 2017b). These studies rely upon exploiting the differences in scattering length density of deuterated and hydrogenated molecules. Because microbial membranes have differing compositions from eukaryotes, the commercially available deuterated lipids and fatty acids are insufficient, making the availability of deuterated microbial lipids and lipid precursors as another pressing need. Lipid extracts contain the rich compositional diversity of the biological membrane, and many microbes can tolerate the highly deuterated culture conditions needed to produce deuterated biomass. Because of this, lipid extracts are a great source of some deuterated materials, as well as being excellent ways to study the physical properties of native microbial cell membranes (Nickels et al. 2017a).
Lipid rafts are currently understood as an organizational motif in the cell membrane of living cells. Rafts are thought to preferentially sort protein cargo based on the physical properties of the membrane phases. The biochemistry that this organization facilitates is deeply involved in the regulation of cellular activity, making these structures critical for proper cellular function and potential avenues for pathogenic interactions.
References
Ahmed SN, Brown DA, London E (1997) On the origin of sphingolipid/cholesterol-rich detergent-insoluble cell membranes: physiological concentrations of cholesterol and sphingolipid induce formation of a detergent-insoluble, liquid-ordered lipid phase in model membranes. Biochemistry 36:10944–10953
Alexander C, Rietschel ET (2001) Invited review: bacterial lipopolysaccharides and innate immunity. J Endotoxin Res 7:167–202
Allen JA, Halverson-Tamboli RA, Rasenick MM (2007) Lipid raft microdomains and neurotransmitter signalling. Nat Rev Neurosci 8:128–140
Almeida PF, Vaz WL, Thompson T (1992) Lateral diffusion and percolation in two-phase, two-component lipid bilayers. Topology of the solid-phase domains in-plane and across the lipid bilayer. Biochemistry 31:7198–7210
Andersen OS, Koeppe RE (2007) Bilayer thickness and membrane protein function: an energetic perspective. Annu Rev Biophys Biomol Struct 36:107–130
Bacia K, Scherfeld D, Kahya N, Schwille P (2004) Fluorescence correlation spectroscopy relates rafts in model and native membranes. Biophys J 87:1034–1043
Bagatolli L, Gratton E (2000) Two photon fluorescence microscopy of coexisting lipid domains in giant unilamellar vesicles of binary phospholipid mixtures. Biophys J 78:290–305
Baumgart T, Hess ST, Webb WW (2003) Imaging coexisting fluid domains in biomembrane models coupling curvature and line tension. Nature 425:821–824
Baumgart T, Hammond AT, Sengupta P, Hess ST, Holowka DA, Baird BA, Webb WW (2007a) Large-scale fluid/fluid phase separation of proteins and lipids in giant plasma membrane vesicles. Proc Natl Acad Sci 104:3165–3170
Baumgart T, Hunt G, Farkas ER, Webb WW, Feigenson GW (2007b) Fluorescence probe partitioning between L o/L d phases in lipid membranes. Biochim Biophys Acta Biomembr 1768:2182–2194
Bee M (1988) Quasielastic neutron scattering: principles and applications in solid state chemistry, biology, and materials science. Adam Hilger, Bristol
Ben-Menachem G, Kubler-Kielb J, Coxon B, Yergey A, Schneerson R (2003) A newly discovered cholesteryl galactoside from Borrelia burgdorferi. Proc Natl Acad Sci U S A 100:7913–7918
Bloembergen N (1957) Proton relaxation times in paramagnetic solutions. J Chem Phys 27:572–573
Botos I, Majdalani N, Mayclin SJ, Mccarthy JG, Lundquist K, Wojtowicz D, Barnard TJ, Gumbart JC, Buchanan SK (2016) Structural and functional characterization of the LPS transporter LptDE from gram-negative pathogens. Structure 24:965–976
Boucher Y, Kamekura M, Doolittle WF (2004) Origins and evolution of isoprenoid lipid biosynthesis in archaea. Mol Microbiol 52:515–527
Bramkamp M, Lopez D (2015) Exploring the existence of lipid rafts in Bacteria. Microbiol Mol Biol Rev 79:81–100
Browman DT, Hoegg MB, Robbins SM (2007) The SPFH domain-containing proteins: more than lipid raft markers. Trends Cell Biol 17:394–402
Brown KL, Conboy JC (2013) Lipid flip-flop in binary membranes composed of phosphatidylserine and phosphatidylcholine. J Phys Chem B 117:15041–15050
Brown D, London E (1998) Functions of lipid rafts in biological membranes. Annu Rev Cell Dev Biol 14:111–136
Brown DA, London E (2000) Structure and function of sphingolipid-and cholesterol-rich membrane rafts. J Biol Chem 275:17221–17224
Brown MF, Ribeiro AA, Williams GD (1983) New view of lipid bilayer dynamics from 2H and 13C NMR relaxation time measurements. Proc Natl Acad Sci 80:4325–4329
Büldt G, Gally H, Seelig A, Seelig J, Zaccai G (1978) Neutron diffraction studies on selectively deuterated phospholipid bilayers. Nature 271:182
Büldt G, Gally H, Seelig J, Zaccai G (1979) Neutron diffraction studies on phosphatidylcholine model membranes: I. Head group conformation. J Mol Biol 134:673–691
Cantor RS (1997) The lateral pressure profile in membranes: a physical mechanism of general anesthesia. Biochemistry 36:2339–2344
Carrasco S, Mérida I (2007) Diacylglycerol, when simplicity becomes complex. Trends Biochem Sci 32:27–36
Christensen H, Garton NJ, Horobin RW, Minnikin DE, Barer MR (1999) Lipid domains of mycobacteria studied with fluorescent molecular probes. Mol Microbiol 31:1561–1572
Crowley JT, Toledo AM, Larocca TJ, Coleman JL, London E, Benach JL (2013) Lipid exchange between Borrelia burgdorferi and host cells. PLoS Pathog 9:e1003109
Czamara K, Majzner K, Pacia MZ, Kochan K, Kaczor A, Baranska M (2015) Raman spectroscopy of lipids: a review. J Raman Spectrosc 46:4–20
Daley DO, Rapp M, Granseth E, Melén K, Drew D, Von Heijne G (2005) Global topology analysis of the Escherichia coli inner membrane proteome. Science 308:1321–1323
Davis JH (1983) The description of membrane lipid conformation, order and dynamics by 2H-NMR. Biochim Biophys Acta Rev Biomembr 737:117–171
Day CA, Kenworthy AK (2015) Functions of cholera toxin B-subunit as a raft cross-linker. Essays Biochem 57:135–145
De Almeida RF, Loura LM, Fedorov A, Prieto M (2005) Lipid rafts have different sizes depending on membrane composition: a time-resolved fluorescence resonance energy transfer study. J Mol Biol 346:1109–1120
Del Pozo MA, Alderson NB, Kiosses WB, Chiang H-H, Anderson RG, Schwartz MA (2004) Integrins regulate Rac targeting by internalization of membrane domains. Science 303:839–842
Den Kamp JO, Redai I, Van Deenen L (1969) Phospholipid composition of Bacillus subtilis. J Bacteriol 99:298–303
Derzko Z, Jacobson K (1980) Comparative lateral diffusion of fluorescent lipid analogs in phospholipid multibilayers. Biochemistry 19:6050–6057
Dick RA, Goh SL, Feigenson GW, Vogt VM (2012) HIV-1 Gag protein can sense the cholesterol and acyl chain environment in model membranes. Proc Natl Acad Sci 109:18761–18766
Dietrich C, Bagatolli L, Volovyk Z, Thompson N, Levi M, Jacobson K, Gratton E (2001) Lipid rafts reconstituted in model membranes. Biophys J 80:1417–1428
Edidin M (2003) The state of lipid rafts: from model membranes to cells. Annu Rev Biophys Biomol Struct 32:257–283
Engelman DM (1971) Lipid bilayer structure in the membrane of Mycoplasma laidlawii. J Mol Biol 58:153–165
Epand RM, Epand RF (2009) Domains in bacterial membranes and the action of antimicrobial agents. Mol BioSyst 5:580–587
Fattal E, Nir S, Parente RA, Szoka FC Jr (1994) Pore-forming peptides induce rapid phospholipid flip-flop in membranes. Biochemistry 33:6721–6731
Feigenson GW (2009) Phase diagrams and lipid domains in multicomponent lipid bilayer mixtures. Biochim Biophys Acta Biomembr 1788:47–52
Fishov I, Woldringh CL (1999) Visualization of membrane domains in Escherichia coli. Mol Microbiol 32:1166–1172
Gajate C, Mollinedo F (2001) The antitumor ether lipid ET-18-OCH3 induces apoptosis through translocation and capping of Fas/CD95 into membrane rafts in human leukemic cells. Blood 98:3860–3863
García-Fernández E, Koch G, Wagner RM, Fekete A, Stengel ST, Schneider J, Mielich-Süss B, Geibel S, Markert SM, Stigloher C, Lopez D (2017) Membrane microdomain disassembly inhibits MRSA antibiotic resistance. Cell 171:1354–1367.e20
García-Sáez AJ, Chiantia S, Schwille P (2007) Effect of line tension on the lateral organization of lipid membranes. J Biol Chem 282:33537–33544
Gaus K, Gratton E, Kable EP, Jones AS, Gelissen I, Kritharides L, Jessup W (2003) Visualizing lipid structure and raft domains in living cells with two-photon microscopy. Proc Natl Acad Sci 100:15554–15559
Gidden J, Denson J, Liyanage R, Ivey DM, Lay JO (2009) Lipid compositions in Escherichia coli and Bacillus subtilis during growth as determined by MALDI-TOF and TOF/TOF mass spectrometry. Int J Mass Spectrom 283:178–184
Goksu EI, Vanegas JM, Blanchette CD, Lin WC, Longo M (2009) AFM for structure and dynamics of biomembranes. Biochim Biophys Acta 1788:254–266
Gross JD, Warschawski DE, Griffin RG (1997) Dipolar recoupling in MAS NMR: a probe for segmental order in lipid bilayers. J Am Chem Soc 119:796–802
Haines TH (2001) Do sterols reduce proton and sodium leaks through lipid bilayers? Prog Lipid Res 40:299–324
Heberle FA, Petruzielo RS, Pan J, Drazba P, KučErka N, Standaert RF, Feigenson GW, Katsaras J (2013) Bilayer thickness mismatch controls domain size in model membranes. J Am Chem Soc 135:6853–6859
Heberle FA, Marquardt D, Doktorova M, Geier B, Standaert RF, Heftberger P, Kollmitzer B, Nickels JD, Dick RA, Feigenson GW (2016) Sub-nanometer structure of an asymmetric model membrane: interleaflet coupling influences domain properties. Langmuir 32:5195–5200
Heftberger P, Kollmitzer B, Rieder AA, Amenitsch H, Pabst G (2015) In situ determination of structure and fluctuations of coexisting fluid membrane domains. Biophys J 108:854–862
Helfrich W (1973) Elastic properties of lipid bilayers: theory and possible experiments. Zeitschrift für Naturforschung C 28:693–703
Henderson TO, Glonek T, Myers TC (1974) Phosphorus-31 nuclear magnetic resonance spectroscopy of phospholipids. Biochemistry 13:623–628
Hering H, Lin C-C, Sheng M (2003) Lipid rafts in the maintenance of synapses, dendritic spines, and surface AMPA receptor stability. J Neurosci 23:3262–3271
Hess ST, Girirajan TP, Mason MD (2006) Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys J 91:4258–4272
Heyn MP (1979) Determination of lipid order parameters and rotational correlation times from fluorescence depolarization experiments. FEBS Lett 108:359–364
Honerkamp-Smith AR, Cicuta P, Collins MD, Veatch SL, Den Nijs M, Schick M, Keller SL (2008) Line tensions, correlation lengths, and critical exponents in lipid membranes near critical points. Biophys J 95:236–246
Hong M, Schmidt-Rohr K, Pines A (1995) NMR measurement of signs and magnitudes of CH dipolar couplings in lecithin. J Am Chem Soc 117:3310–3311
Huang B, Wang W, Bates M, Zhuang X (2008) Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319:810–813
Jacobson K, Mouritsen OG, Anderson RG (2007) Lipid rafts: at a crossroad between cell biology and physics. Nat Cell Biol 9:7–14
Jacrot B (1976) The study of biological structures by neutron scattering from solution. Rep Prog Phys 39:911
Jares-Erijman EA, Jovin TM (2003) FRET imaging. Nat Biotechnol 21:1387–1395
Jerabek H, Pabst G, Rappolt M, Stockner T (2010) Membrane-mediated effect on ion channels induced by the anesthetic drug ketamine. J Am Chem Soc 132:7990–7997
Johnson RC (1977) The Spirochetes. Annu Rev Microbiol 31:89–106
Johnston LJ (2007) Nanoscale imaging of domains in supported lipid membranes. Langmuir 23:5886–5895
Kabouridis PS, Janzen J, Magee AL, Ley SC (2000) Cholesterol depletion disrupts lipid rafts and modulates the activity of multiple signaling pathways in T lymphocytes. Eur J Immunol 30:954–963
Kawai F, Shoda M, Harashima R, Sadaie Y, Hara H, Matsumoto K (2004) Cardiolipin domains in Bacillus subtilis Marburg membranes. J Bacteriol 186:1475–1483
Kenworthy AK, Petranova N, Edidin M (2000) High-resolution FRET microscopy of cholera toxin B-subunit and GPI-anchored proteins in cell plasma membranes. Mol Biol Cell 11:1645–1655
Klymchenko AS, Kreder R (2014) Fluorescent probes for lipid rafts: from model membranes to living cells. Chem Biol 21:97–113
Knoll W, Ibel K, Sackmann E (1981) Small-angle neutron scattering study of lipid phase diagrams by the contrast variation method. Biochemistry 20:6379–6383
Koenig BW, Strey HH, Gawrisch K (1997) Membrane lateral compressibility determined by NMR and x-ray diffraction: effect of acyl chain polyunsaturation. Biophys J 73:1954–1966
König S, Pfeiffer W, Bayerl T, Richter D, Sackmann E (1992) Molecular dynamics of lipid bilayers studied by incoherent quasi-elastic neutron scattering. J Phys II 2:1589–1615
König S, Sackmann E, Richter D, Zorn R, Carlile C, Bayerl T (1994) Molecular dynamics of water in oriented DPPC multilayers studied by quasielastic neutron scattering and deuterium-nuclear magnetic resonance relaxation. J Chem Phys 100:3307
Korlach J, Schwille P, Webb WW, Feigenson GW (1999) Characterization of lipid bilayer phases by confocal microscopy and fluorescence correlation spectroscopy. Proc Natl Acad Sci 96:8461–8466
Kraft ML, Weber PK, Longo ML, Hutcheon ID, Boxer SG (2006) Phase separation of lipid membranes analyzed with high-resolution secondary ion mass spectrometry. Science 313:1948–1951
Kreiswirth B, Kornblum J, Arbeit R, Eisner W, Maslow J, Mcgeer A, Low D, Novick R (1993) Evidence for a clonal origin of methicillin resistance in Staphylococcus aureus. Science 259:227–230
Krogh A, Larsson B, Von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Kusumi A, Tsuda M, Akino T, Ohnishi S, Terayama Y (1983) Protein-phospholipid-cholesterol interaction in the photolysis of invertebrate rhodopsin. Biochemistry 22:1165–1170
Kuzmin PI, Akimov SA, Chizmadzhev YA, Zimmerberg J, Cohen FS (2005) Line tension and interaction energies of membrane rafts calculated from lipid splay and tilt. Biophys J 88:1120–1133
Lang S, Philp JC (1998) Surface-active lipids in rhodococci. Antonie Van Leeuwenhoek 74:59–70
Langhorst MF, Reuter A, Stuermer CAO (2005) Scaffolding microdomains and beyond: the function of reggie/flotillin proteins. Cell Mol Life Sci 62:2228–2240
Langhorst MF, Solis GP, Hannbeck S, Plattner H, Stuermer CAO (2007) Linking membrane microdomains to the cytoskeleton: regulation of the lateral mobility of reggie-1/flotillin-2 by interaction with actin. FEBS Lett 581:4697–4703
Larocca TJ, Crowley JT, Cusack BJ, Pathak P, Benach J, London E, Garcia-Monco JC, Benach JL (2010) Cholesterol lipids of Borrelia burgdorferi form lipid rafts and are required for the bactericidal activity of a complement-independent antibody. Cell Host Microbe 8:331–342
Lewis BA, Engelman DM (1983) Lipid bilayer thickness varies linearly with acyl chain length in fluid phosphatidylcholine vesicles. J Mol Biol 166:211–217
Lewis RN, Mcelhaney RN (2013) Membrane lipid phase transitions and phase organization studied by Fourier transform infrared spectroscopy. Biochim Biophys Acta Biomembr 1828:2347–2358
Li L, Wang H, Cheng J-X (2005) Quantitative coherent anti-stokes Raman scattering imaging of lipid distribution in coexisting domains. Biophys J 89:3480–3490
Lindblom G, Orädd G, Filippov A (2006) Lipid lateral diffusion in bilayers with phosphatidylcholine, sphingomyelin and cholesterol: an NMR study of dynamics and lateral phase separation. Chem Phys Lipids 141:179–184
Linden CD, Wright KL, Mcconnell HM, Fox CF (1973) Lateral phase separations in membrane lipids and the mechanism of sugar transport in Escherichia coli. Proc Natl Acad Sci 70:2271–2275
Lingwood D, Simons K (2007) Detergent resistance as a tool in membrane research. Nat Protoc 2:2159–2165
Lingwood D, Simons K (2010) Lipid rafts as a membrane-organizing principle. Science 327:46–50
Liu J, Conboy JC (2004) Phase transition of a single lipid bilayer measured by sum-frequency vibrational spectroscopy. J Am Chem Soc 126:8894–8895
London E, Brown DA (2000) Insolubility of lipids in triton X-100: physical origin and relationship to sphingolipid/cholesterol membrane domains (rafts). Biochim Biophys Acta Biomembr 1508:182–195
Lopez D, Koch G (2017) Exploring functional membrane microdomains in bacteria: an overview. Curr Opin Microbiol 36:76–84
López D, Kolter R (2010) Functional microdomains in bacterial membranes. Genes Dev 24:1893–1902
Luzzati V, Husson F (1962) The structure of the liquid-crystalline phases of lipid-water systems. J Cell Biol 12:207–219
Majewski J, Kuhl T, Kjaer K, Smith G (2001) Packing of ganglioside-phospholipid monolayers: an x-ray diffraction and reflectivity study. Biophys J 81:2707–2715
Marquardt D, Williams JA, KučErka N, Atkinson J, Wassall SR, Katsaras J, Harroun TA (2013) Tocopherol activity correlates with its location in a membrane: a new perspective on the antioxidant vitamin E. J Am Chem Soc 135:7523–7533
Marquardt D, Heberle FA, Nickels JD, Pabst G, Katsaras J (2015) On scattered waves and lipid domains: detecting membrane rafts with X-rays and neutrons. Soft Matter 11:9055–9072
Marsh D (2009) Cholesterol-induced fluid membrane domains: a compendium of lipid-raft ternary phase diagrams. Biochim Biophys Acta Biomembr 1788:2114–2123
Marshall JH, Wilmoth GJ (1981) Pigments of Staphylococcus aureus, a series of triterpenoid carotenoids. J Bacteriol 147:900–913
Matsumoto K, Kusaka J, Nishibori A, Hara H (2006) Lipid domains in bacterial membranes. Mol Microbiol 61:1110–1117
Mendelsohn R, Moore DJ (1998) Vibrational spectroscopic studies of lipid domains in biomembranes and model systems. Chem Phys Lipids 96:141–157
Mileykovskaya E, Dowhan W (2000) Visualization of phospholipid domains in Escherichia coli by using the cardiolipin-specific fluorescent dye 10-N-nonyl acridine orange. J Bacteriol 182:1172–1175
Mills TT, Toombes GE, Tristram-Nagle S, Smilgies D-M, Feigenson GW, Nagle JF (2008) Order parameters and areas in fluid-phase oriented lipid membranes using wide angle X-ray scattering. Biophys J 95:669–681
Mouritsen OG, Jørgensen K (1994) Dynamical order and disorder in lipid bilayers. Chem Phys Lipids 73:3–25
Nagle JF, Tristram-Nagle S (2000) Structure of lipid bilayers. Biochim Biophys Acta Rev Biomembr 1469:159–195
Nichols BJ (2003) GM1-containing lipid rafts are depleted within Clathrin-coated pits. Curr Biol 13:686–690
Nichols BJ, Kenworthy AK, Polishchuk RS, Lodge R, Roberts TH, Hirschberg K, Phair RD, Lippincott-Schwartz J (2001) Rapid cycling of lipid raft markers between the cell surface and Golgi complex. J Cell Biol 153:529–542
Nickels JD, Katsaras J (2015) Water and lipid bilayers. In: Membrane hydration. Springer International Publishing, Switzerland
Nickels JD, O’neill H, Hong L, Tyagi M, Ehlers G, Weiss KL, Zhang Q, Yi Z, Mamontov E, Smith JC (2012) Dynamics of protein and its hydration water: neutron scattering studies on fully deuterated GFP. Biophys J 103:1566–1575
Nickels JD, Cheng X, Mostofian B, Stanley C, Lindner B, Heberle FA, Perticaroli S, Feygenson M, Egami T, Standaert RF, Smith JC, Myles DAA, Ohl M, Katsaras J (2015a) Mechanical properties of Nanoscopic lipid domains. J Am Chem Soc 137:15772–15780
Nickels JD, Smith JC, Cheng X (2015b) Lateral organization, bilayer asymmetry, and inter-leaflet coupling of biological membranes. Chem Phys Lipids 192:87–99
Nickels JD, Atkinson J, Papp-Szabo E, Stanley C, Diallo SO, Perticaroli S, Baylis B, Mahon P, Ehlers G, Katsaras J, Dutcher JR (2016) Structure and hydration of highly-branched, monodisperse phytoglycogen nanoparticles. Biomacromolecules 17:735–743
Nickels JD, Chatterjee S, Mostofian B, Stanley CB, Ohl M, Zolnierczuk P, Schulz R, Myles DA, Standaert RF, Elkins JG (2017a) Bacillus subtilis lipid extract, A branched-chain fatty acid model membrane. J Phys Chem Lett 8:4214–4217
Nickels JD, Chatterjee S, Stanley CB, Qian S, Cheng X, Myles DAA, Standaert RF, Elkins JG, Katsaras J (2017b) The in vivo structure of biological membranes and evidence for lipid domains. PLoS Biol 15:e2002214
Nickels JD, Smith MD, Alsop RJ, Himbert S, Yahya A, Cordner D, Zolnierczuk P, Stanley CB, Katsaras J, Cheng X, Rheinstadter MC (2019) Lipid rafts: buffers of cell membrane physical properties. J Phys Chem B 123:2050–2056
Niño BJ, Marc B (2013) Flotillins functionally organize the bacterial membrane. Mol Microbiol 88:1205–1217
Nishibori A, Kusaka J, Hara H, Umeda M, Matsumoto K (2005) Phosphatidylethanolamine domains and localization of phospholipid synthases in Bacillus subtilis membranes. J Bacteriol 187:2163–2174
Opilik L, Bauer T, Schmid T, Stadler J, Zenobi R (2011) Nanoscale chemical imaging of segregated lipid domains using tip-enhanced Raman spectroscopy. Phys Chem Chem Phys 13:9978–9981
Orädd G, Lindblom G (2004) Lateral diffusion studied by pulsed field gradient NMR on oriented lipid membranes. Magn Reson Chem 42:123–131
Orädd G, Westerman PW, Lindblom G (2005) Lateral diffusion coefficients of separate lipid species in a ternary raft-forming bilayer: a Pfg-NMR multinuclear study. Biophys J 89:315–320
Owen DM, Rentero C, Magenau A, Abu-Siniyeh A, Gaus K (2012) Quantitative imaging of membrane lipid order in cells and organisms. Nat Protoc 7:24
Pardon J, Worcester D, Wooley J, Tatchell K, Van Holde K, Richards B (1975) Low-angle neutron scattering from chromatin subunit particles. Nucleic Acids Res 2:2163–2176
Parsons JB, Rock CO (2013) Bacterial lipids: metabolism and membrane homeostasis. Prog Lipid Res 52:249–276
Parton RG (1994) Ultrastructural localization of gangliosides; GM1 is concentrated in caveolae. J Histochem Cytochem 42:155–166
Pelz A, Wieland K-P, Putzbach K, Hentschel P, Albert K, Götz F (2005) Structure and biosynthesis of Staphyloxanthin from Staphylococcus aureus. J Biol Chem 280:32493–32498
Pencer J, Mills T, Anghel V, Krueger S, Epand RM, Katsaras J (2005) Detection of submicron-sized raft-like domains in membranes by small-angle neutron scattering. Eur Phys J E 18:447–458
Percy MG, Gründling A (2014) Lipoteichoic acid synthesis and function in gram-positive Bacteria. Annu Rev Microbiol 68:81–100
Perticaroli S, Ehlers G, Stanley CB, Mamontov E, O’neill H, Zhang Q, Cheng X, Myles DA, Katsaras J, Nickels JD (2017) Description of hydration water in protein (green fluorescent protein) solution. J Am Chem Soc 139:1098–1105
Petrache HI, Dodd SW, Brown MF (2000) Area per lipid and acyl length distributions in fluid phosphatidylcholines determined by 2H NMR spectroscopy. Biophys J 79:3172–3192
Pfeiffer W, Henkel T, Sackmann E, Knoll W, Richter D (1989) Local dynamics of lipid bilayers studied by incoherent quasi-elastic neutron scattering. EPL (Europhys Lett) 8:201
Pfeiffer A, Böttcher A, Orsó E, Kapinsky M, Nagy P, Bodnár A, Spreitzer I, Liebisch G, Drobnik W, Gempel K (2001) Lipopolysaccharide and ceramide docking to CD14 provokes ligand-specific receptor clustering in rafts. Eur J Immunol 31:3153–3164
Pierce SK (2002) Lipid rafts and B-cell activation. Nat Rev Immunol 2:96–105
Pike L (2004) Lipid rafts: heterogeneity on the high seas. Biochem J 378:281–292
Poger D, Mark AE (2013) The relative effect of sterols and hopanoids on lipid bilayers: when comparable is not identical. J Phys Chem B 117:16129–16140
Potma EO, Xie XS (2005) Direct visualization of lipid phase segregation in single lipid bilayers with coherent anti-stokes Raman scattering microscopy. ChemPhysChem 6:77–79
Pralle A, Keller P, Florin E-L, Simons K, Hörber J (2000) Sphingolipid–cholesterol rafts diffuse as small entities in the plasma membrane of mammalian cells. J Cell Biol 148:997–1008
Rheinstädter M, Ollinger C, Fragneto G, Demmel F, Salditt T (2004) Collective dynamics of lipid membranes studied by inelastic neutron scattering. Phys Rev Lett 93:108107
Rohmer M, Bouvier-Nave P, Ourisson G (1984) Distribution of hopanoid triterpenes in prokaryotes. Microbiology 130:1137–1150
Rothberg KG, Ying Y-S, Kamen BA, Anderson R (1990) Cholesterol controls the clustering of the glycophospholipid-anchored membrane receptor for 5-methyltetrahydrofolate. J Cell Biol 111:2931–2938
Sáenz JP, Sezgin E, Schwille P, Simons K (2012) Functional convergence of hopanoids and sterols in membrane ordering. Proc Natl Acad Sci 109:14236–14240
Samsonov AV, Mihalyov I, Cohen FS (2001) Characterization of cholesterol-sphingomyelin domains and their dynamics in bilayer membranes. Biophys J 81:1486–1500
Scheidt HA, Huster D, Gawrisch K (2005) Diffusion of cholesterol and its precursors in lipid membranes studied by 1H pulsed field gradient magic angle spinning NMR. Biophys J 89:2504–2512
Schneider J, Mielich-Süss B, Böhme R, Lopez D (2015) In vivo characterization of the scaffold activity of flotillin on the membrane kinase KinC of Bacillus subtilis. Microbiology 161:1871–1887
Schroeder R, London E, Brown D (1994) Interactions between saturated acyl chains confer detergent resistance on lipids and glycosylphosphatidylinositol (GPI)-anchored proteins: GPI-anchored proteins in liposomes and cells show similar behavior. Proc Natl Acad Sci 91:12130–12134
Schultz ZD, Levin IW (2008) Lipid microdomain formation: characterization by infrared spectroscopy and ultrasonic velocimetry. Biophys J 94:3104–3114
Schütz GJ, Kada G, Pastushenko VP, Schindler H (2000) Properties of lipid microdomains in a muscle cell membrane visualized by single molecule microscopy. EMBO J 19:892–901
Schwille P, Korlach J, Webb WW (1999) Fluorescence correlation spectroscopy with single-molecule sensitivity on cell and model membranes. Cytometry A 36:176–182
Sears VF (1992) Neutron scattering lengths and cross sections. Neutron News 3:26–37
Seelig A, Seelig J (1974) Dynamic structure of fatty acyl chains in a phospholipid bilayer measured by deuterium magnetic resonance. Biochemistry 13:4839–4845
Settles M, Doster W (1996) Anomalous diffusion of adsorbed water: a neutron scattering study of hydrated myoglobin. Faraday Discuss 103:269–279
Seul M, Andelman D (1995) Domain shapes and patterns: the phenomenology of modulated phases. Science 267:476–483
Sezgin E, Levental I, Grzybek M, Schwarzmann G, Mueller V, Honigmann A, Vn B (2012) Partitioning, diffusion, and ligand binding of raft lipid analogs in model and cellular plasma membranes. Biochim Biophys Acta 1818:1777–1784
Sezgin E, Levental I, Mayor S, Eggeling C (2017a) The mystery of membrane organization: composition, regulation and physiological relevance of lipid rafts. Nat Rev Mol Cell Biol 18:361–374
Sezgin E, Levental I, Mayor S, Eggeling C (2017b) The mystery of membrane organization: composition, regulation and roles of lipid rafts. Nat Rev Mol Cell Biol 18:361
Shimshick EJ, Mcconnell HM (1973) Lateral phase separation in phospholipid membranes. Biochemistry 12:2351–2360
Simons K, Gerl MJ (2010) Revitalizing membrane rafts: new tools and insights. Nat Rev Mol Cell Biol 11:688–699
Simons K, Ikonen E (1997) Functional rafts in cell membranes. Nature 387:569–572
Simons K, Toomre D (2000) Lipid rafts and signal transduction. Nat Rev Mol Cell Biol 1:31–39
Simons K, Van Meer G (1988) Lipid sorting in epithelial cells. Biochemistry 27:6197–6202
Singer S, Nicolson GL (1972) The fluid mosaic model of the structure of cell membranes. Science 18:720–31
Sohlenkamp C, Geiger O (2016) Bacterial membrane lipids: diversity in structures and pathways. FEMS Microbiol Rev 40:133–159
Solomon I (1955) Relaxation processes in a system of two spins. Phys Rev 99:559
Stöckl M, Herrmann A (2010) Detection of lipid domains in model and cell membranes by fluorescence lifetime imaging microscopy. Biochim Biophys Acta 1798:1444–1456
Stuermer CA, Lang DM, Kirsch F, Wiechers M, Deininger S-O, Plattner H (2001) Glycosylphosphatidyl inositol-anchored proteins and fyn kinase assemble in noncaveolar plasma membrane microdomains defined by reggie-1 and-2. Mol Biol Cell 12:3031–3045
Su Y, Mani R, Hong M (2008) Asymmetric insertion of membrane proteins in lipid bilayers by solid-state NMR paramagnetic relaxation enhancement: a cell-penetrating peptide example. J Am Chem Soc 130:8856–8864
Swenson J, Kargl F, Berntsen P, Svanberg C (2008) Solvent and lipid dynamics of hydrated lipid bilayers by incoherent quasielastic neutron scattering. J Chem Phys 129:045101
Tardieu A, Luzzati V, Reman F (1973) Structure and polymorphism of the hydrocarbon chains of lipids: a study of lecithin-water phases. J Mol Biol 75:711–733
Tavernarakis N, Driscoll M, Kyrpides NC (1999) The SPFH domain: implicated in regulating targeted protein turnover in stomatins and other membrane-associated proteins. Trends Biochem Sci 24:425–427
Tokumasu F, Jin AJ, Feigenson GW, Dvorak JA (2003) Nanoscopic lipid domain dynamics revealed by atomic force microscopy. Biophys J 84:2609–2618
Toppozini L, Roosen-Runge F, Bewley RI, Dalgliesh RM, Perring T, Seydel T, Glyde HR, Garcia Sakai V, Rheinstadter MC (2015) Anomalous and anisotropic nanoscale diffusion of hydration water molecules in fluid lipid membranes. Soft Matter 11:8354–8371
Turner MS, Sens P, Socci ND (2005) Nonequilibrium raftlike membrane domains under continuous recycling. Phys Rev Lett 95:168301
Van Der Goot FG, Harder T (2001) Raft membrane domains: from a liquid-ordered membrane phase to a site of pathogen attack. Semin Immunol 13:89–97
Van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9:112
Vance JE, Vance DE (1985) The role of phosphatidylcholine biosynthesis in the secretion of lipoproteins from hepatocytes. Can J Biochem Cell Biol 63:870–881
Varma R, Mayor S (1998) GPI-anchored proteins are organized in submicron domains at the cell surface. Nature 394:798
Veatch SL, Keller SL (2002) Organization in lipid membranes containing cholesterol. Phys Rev Lett 89:268101
Veatch SL, Keller SL (2003) Separation of liquid phases in giant vesicles of ternary mixtures of phospholipids and cholesterol. Biophys J 85:3074–3083
Veatch S, Polozov I, Gawrisch K, Keller S (2004) Liquid domains in vesicles investigated by NMR and fluorescence microscopy. Biophys J 86:2910–2922
Verkleij A, Zwaal R, Roelofsen B, Comfurius P, Kastelijn D, Van Deenen L (1973) The asymmetric distribution of phospholipids in the human red cell membrane. A combined study using phospholipases and freeze-etch electron microscopy. Biochim Biophys Acta Biomembr 323:178–193
Villalba M, Bi K, Rodriguez F, Tanaka Y, Schoenberger S, Altman A (2001) Vav1/Rac-dependent actin cytoskeleton reorganization is required for lipid raft clustering in T cells. J Cell Biol 155:331–338
Wakasugi K, Nakano T, Kitatsuji C, Morishima I (2004) Human neuroglobin interacts with flotillin-1, a lipid raft microdomain-associated protein. Biochem Biophys Res Commun 318:453–460
Watson MC, Brown FL (2010) Interpreting membrane scattering experiments at the mesoscale: the contribution of dissipation within the bilayer. Biophys J 98:L9–L11
Wieland B, Feil C, Gloria-Maercker E, Thumm G, Lechner M, Bravo JM, Poralla K, Götz F (1994) Genetic and biochemical analyses of the biosynthesis of the yellow carotenoid 4,4′-diaponeurosporene of Staphylococcus aureus. J Bacteriol 176:7719–7726
Woodka AC, Butler PD, Porcar L, Farago B, Nagao M (2012) Lipid bilayers and membrane dynamics: insight into thickness fluctuations. Phys Rev Lett 109:058102
Wu E, Jacobson K, Papahadjopoulos D (1977) Lateral diffusion in phospholipid multibilayers measured by fluorescence recovery after photobleaching. Biochemistry 16:3936–3941
Xu X, Bittman R, Duportail G, Heissler D, Vilcheze C, London E (2001) Effect of the structure of natural sterols and Sphingolipids on the formation of ordered Sphingolipid/sterol domains (rafts): comparison of cholesterol to plant, fungal, and disease-associated sterols and comparison of sphingomyelin, cerebrosides, and ceramide. J Biol Chem 276:33540–33546
Yethiraj A, Weisshaar JC (2007) Why are lipid rafts not observed in vivo? Biophys J 93:3113–3119
Yu J, Fischman DA, Steck TL (1973) Selective solubilization of proteins and phospholipids from red blood cell membranes by nonionic detergents. J Cell Biochem 1:233–248
Yuan C, Furlong J, Burgos P, Johnston LJ (2002) The size of lipid rafts: an atomic force microscopy study of ganglioside GM1 domains in sphingomyelin/DOPC/cholesterol membranes. Biophys J 82:2526–2535
Zaccai G, Blasie J, Schoenborn B (1975) Neutron diffraction studies on the location of water in lecithin bilayer model membranes. Proc Natl Acad Sci 72:376–380
Zaccai G, Büldt G, Seelig A, Seelig J (1979) Neutron diffraction studies on phosphatidylcholine model membranes: II Chain conformation and segmental disorder. J Mol Biol 134:693–706
Zacharias DA, Violin JD, Newton AC, Tsien RY (2002) Partitioning of lipid-modified monomeric GFPs into membrane microdomains of live cells. Science 296:913–916
Zhao J, Wu J, Heberle FA, Mills TT, Klawitter P, Huang G, Costanza G, Feigenson GW (2007) Phase studies of model biomembranes: complex behavior of DSPC/DOPC/cholesterol. Biochim Biophys Acta Biomembr 1768:2764–2776
Zilman A, Granek R (1996) Undulations and dynamic structure factor of membranes. Phys Rev Lett 77:4788
Zipfel WR, Williams RM, Christie R, Nikitin AY, Hyman BT, Webb WW (2003) Live tissue intrinsic emission microscopy using multiphoton-excited native fluorescence and second harmonic generation. Proc Natl Acad Sci 100:7075–7080
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this entry
Cite this entry
Nickels, J.D., Hogg, J., Cordner, D., Katsaras, J. (2020). Lipid Rafts in Bacteria: Structure and Function. In: Goldfine, H. (eds) Health Consequences of Microbial Interactions with Hydrocarbons, Oils, and Lipids. Handbook of Hydrocarbon and Lipid Microbiology . Springer, Cham. https://doi.org/10.1007/978-3-030-15147-8_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-15147-8_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-15146-1
Online ISBN: 978-3-030-15147-8
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences