Abstract
Osteosarcoma is the most common bone sarcoma and is one of the cancer entities characterized by the highest level of heterogeneity in humans. This heterogeneity takes place not only at the macroscopic and microscopic levels, with heterogeneous micro-environmental components, but also at the genomic, transcriptomic and epigenetic levels. Recent investigations have revealed the existence in osteosarcoma of cancer cells with stemness properties. Cancer stem cells are characterized by their specific phenotype and low cycling capacity, and are linked to drug resistance, tumour growth and the metastatic process. In addition, cancer stem cells contribute to the enrichment of tumour heterogeneity. The present manuscript will describe the main characteristic features of cancer stem cells in osteosarcoma and will discuss their impact on maintaining tumour heterogeneity. Their clinical implications will also be briefly addressed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Tumour heterogeneity
- Gene mutation drivers
- Cancer stem-like cells
- Circulating tumour cells
- Monitoring
- Clinical trials
- Osteosarcoma
- Cancer stem cell
- Drug resistance
- Cell dormancy
11.1 Introduction
Osteosarcoma is part of the family of malignant bone sarcomas which originate from a common mesenchymal precursor located in the bone marrow, and known as mesenchymal stem cells (Brown et al. 2018a). Osteosarcoma is the main bone sarcoma in adolescents and young adults, with a peak of incidence at around 18 years old. Osteosarcomas are preferentially detected in the metaphysis of long bones and the tumour tissue is characterized by the presence of osteoid matrix produced by cancer cells (Fig. 11.1). Microscopic heterogeneity is the first marker for osteosarcoma with the presence of highly vascularized, necrotic, proliferating and osteoid foci. Depending on the morphological features of the cancer cells, osteosarcomas can be classified as osteoblastic, chondroblastic, fibroblastic or telangiectatic. Current treatment combines neo-adjuvant chemotherapy, surgery and adjuvant chemotherapy including at least three cytotoxic agents such as doxorubicin, methotrexate and ifosfamide. Unfortunately, prognosis remains poor and overall survival has stagnated in the last four decades (Heymann et al. 2016). Overall survival reaches 50–70% at 5 years depending on the series in the absence of detectable metastases, but drops to 30% when lung metastases are detected at the time of diagnosis.
Clonal evolution of osteosarcoma cells and their role in tumour heterogeneity. Initially formed by mono- or oligoclonal subclones, dominant clones appear progressively, resulting in marked heterogeneity in the tumour mass. Of these cancer cells, a subpopulation exhibits stemness markers and educates mesenchymal stem cells to release exosomes, which in turn increase the stem cell phenotype and upmodulate tumour growth and the development of metastases. These two-way communications enrich tumour heterogeneity and increase the risk of drug resistance. Immune cells, with their diversity and induction of local immune tolerance, complete the heterogeneity of the tumour mass
Tumour heterogeneity can be directly related to both the natural history of the cancer cells and to their dialogue with the protagonists in the local micro-environment (Mutsaers and Walkley 2014; Tang et al. 2008; Mohseny et al. 2009; Cortini et al. 2017; Alfranca et al. 2015). The local micro-environment is composed of numerous cell types, including immune (e.g. tumour-infiltrating lymphocytes and tumour-associated macrophages) (Heymann et al. 2017; Dumars et al. 2016; Théoleyre et al. 2005) and non-immune cells such as endothelial cells, fibroblasts and mesenchymal stem cells, which are spatially, temporally and functionally linked to cancer cells (Brown et al. 2018a). Cancer cells can control the behaviour of their neighbours, which in turn play a part in fuelling tumour growth and the metastatic process . Cancer cells are composed of numerous cell clones competing together to preserve the overall survival of their congeners through selective advantage. Some of these clones drive tumour initiation and are called cancer stem cells (Brown et al. 2017). Even if the term “stem cell” is not perfectly appropriate, it describes a subpopulation of cells capable of reconstituting the characteristics of all cancer cells detectable in the tumour mass. Consequently, cancer stem cells can generate a tumour mass after inoculation into an immunodeficient organism (Najafi et al. 2019).
The present review will discuss the main data available in the literature in favour of the existence of cancer stem-like cells in osteosarcoma, as well as their potential contribution to the enrichment of tumour heterogeneity . Their clinical impact in drug resistance will be also discussed.
11.2 Clonal Evolution of Cancer Cells in Osteosarcoma: A Combination of Oncogenic and Epigenetic Events
In parallel to histological heterogeneity, osteosarcoma is one of the most complex oncologic diseases in terms of genetic aberration. In 2014, Reimann et al. found wide genomic rearrangements in the tumour exome of a single case of osteosarcoma (Reimann et al. 2014). These authors detected 3,000 somatic single nucleotide variants, small indels and more than 2,000 copy number variants in diverse chromosomes. The osteosarcomas were thus characterized by a loss of heterozygosity. The complexity of the disease was confirmed by Bousquet et al. who studied a series of 44 osteosarcomas and observed recurrent somatic alterations to TP53 and RB1 and also detected 84 mutation points and 4 deletions related to 84 genes (Bousquet et al. 2016). Similarly, Smida et al. analysed 160 osteosarcoma samples by whole-genome sequencing in order to identify somatic copy number alterations. They found specific unstable genomic regions in which numerous tumour suppressor genes were included (e.g. TP53, RB1, WWOX and DLG2) (Smida et al. 2017). This very high number of alterations perfectly illustrates the genomic complexity of osteosarcomas. The development of cancer is sustained by two main theories: (1) the “linear model” theory , which is based on successive accumulations of oncogenic events in one cell leading to the development of a heterogeneous disease; (2) the branched evolution theory known as the “parallel model” , characterized by the parallel evolution of subclones which accumulate DNA alterations and also lead to a polyclonal tumour mass (Greaves and Maley 2012; Tellez-Gabriel et al. 2016). Of course, the nature and number of these oncogenic events drive tumour initiation as has been shown by Funes et al. who transformed mesenchymal stem cells using genetic alterations (Funes et al. 2007). They observed that four oncogenic hits made possible the formation of colonies in agar, although only five oncogenes were able to induce tumour development in immunodeficient mice. They also suggested that tumourigenesis of modified mesenchymal stem cells was dependent on the nature of the oncogene. For instance, disruption to the RB pathway was enough to induce anchorage-independent growth of mesenchymal stem cells (Funes et al. 2007). In addition, one oncogenic hit sensitized mesenchymal stem cells to carcinogenic agents such as pesticides and may have led to tumour development in immunocompromised mice (Hochane et al. 2017). In addition, the clonal evolution of cancers is tightly controlled by the selective pressure of the local micro-environment (e.g. immune infiltrate and hypoxia), and can be oriented under drug pressure toward resistant or tolerant cancer cells (Brown et al. 2018b; Vallette et al. 2018). Regardless of what the first oncogenic event is, a permissive local micro-environment is obligatory for protecting cancer-initiating cells against immune cells and fuelling these cells with adequate nutrients (de Groot et al. 2017).
Both types of clonal evolution have been described in osteosarcoma (Wang et al. 2019). Wang et al. analysed and compared 86 tumours in 10 osteosarcoma patients using whole exosome and genome sequencing. By analysing the architecture and relationships of the cancer subclones, they demonstrated a dynamic mutational process and, for the first time, two patterns of lung metastases—with a linear model in six patients and a branched model in four patients. Based on the low number of patients included, the co-existence of both models in a same patient can be excluded. The tumour evolution model has recently been enriched by a “plasticity” model identified in Ewing sarcoma (Franzetti et al. 2017). In Ewing sarcoma, the plasticity model is based on equilibrium between various cancer cell subclones differentially expressing the chimeric EWS1/FLI1 transcription factor, leading to major modifications in cell migration and invasion properties. The two populations create an ecosystem with dynamic fluctuation in cells differentially expressing the fusion protein depending on the stage of the disease.
Very recently, Gambera et al. established multicolour (RGB) p53−/− Rb−/− mouse mesenchymal stem cells (Gambera et al. 2018) that can form osteosarcomas when inoculated into bone micro-environment cells. They also deciphered the clonal evolution during tumour progression (Rubio et al. 2014). They identified two main steps in tumour progression. At an early stage of development (25 days), tumour growth is characterized by polyclonal expansion with no modification to the proportions of the coloured cells injected. At a late stage of tumour growth (50 days), Gambera et al. observed the emergence of dominant clones at the periphery of the tumour mass, corresponding to clonal evolution of the disease. Overall, these data provided evidence of marked clonal modifications in cancer cells from a polyclonal context to the formation of dominant clones which were oligoclonal and exhibited similar tumourigenesis properties (Gambera et al. 2018). In addition, the metastatic process to the lung was associated with an oligoclonal and monoclonal dynamic. Although this model cannot be transposed to humans, there is some evidence of dominant clones in osteosarcoma. In 2015, Kovac et al. investigated the evolutionary landscape using exosome sequencing in 31 osteosarcoma samples (Kovac et al. 2015). They identified 14 genes associated with a BRCAness signature as the main drivers for tumour development not exclusively expressed in all subclones. TP53 mutations were frequently observed in subclones. These authors hypothesized that osteosarcoma could be initiated by a mutation in TP53 or RB in one specific subclone (monoclonal disease), leading to chromosal instability and chromatide breakages, and to new oncogenic events in various subclones (polyclonal disease). PARP inhibitors may then be a therapeutic option in osteosarcoma (Engert et al. 2017). The existence of dominant subclones was confirmed by Chen et al. by studying a case report of a chemoresistant osteosarcoma sample in which they identified a clone associated with a new TP53-KPN3 translocation (Chen et al. 2016). The three models for cancer cell evolution are responsible for the considerable heterogeneity found in osteosarcoma and the emergence of dominant clones which evolve in a dynamic manner and in perfect symbiosis with their permissive ecosystem.
As shown in Ewing sarcoma, for which the heterogeneity of DNA methylation is a reflection of the spectrum of the disease (Surdez et al. 2017), epigenetic genetic alterations are observed in osteosarcomas and are associated with its pathogenesis (e.g. tumour growth and metastatic process ) (Feng et al. 2018; Sarver and Subramanian 2016; Georges et al. 2018). Epigenetic modulations can regulate osteosarcoma cell differentiation and can concomitantly interfere with their micro-environment (Itoh et al. 2018; Lamoureux et al. 2014; Li et al. 2018). For instance, Lamoureux et al. demonstrated that selective inhibition of bromodomain epigenetic signalling induced an inhibitory effect in primary tumour growth and simultaneously in osteoblasts and osteoclasts, two cell types found in the local micro-environment (Lamoureux et al. 2014). More recently, Li et al. gave evidence of epigenetic downregulation in osteosarcoma cells of CXCL12 (SDF-1) via DNA methyltransferase-1, related to their ability to form lung metastases and, interestingly, to their impairment of cytotoxic T-cells homing in on the tumour mass (Li et al. 2018). They found a correlation between CXCL12 expression and the overall survival of osteosarcoma patients. Tumour heterogeneity and clonal evolution of osteosarcomas are thus regulated by epigenetic events.
11.3 Presence of Cancer Stem Cells in Osteosarcoma: Their Functional Impact
The conventional theories for clonal evolution described above can be completed by the “cell origin” theory . In this theory, the first oncogenic event may occur in a cancer stem cell or in a cell in the non-side population (López-Lázaro 2018), with cancer as the end result of successive cell divisions in stem cells with cumulative DNA replication errors (e.g. mutations and epigenetic mistakes) making possible both the self-renewal of “differentiated” cancer cells and the maintenance of undifferentiated cells. The presence of a side population that excludes Hoechst 33342 dye has been demonstrated on osteosarcoma cell lines and in human primary osteosarcoma (Murase et al. 2009; Yang et al. 2011). These cells are able to regenerate both side- and non-side cells, show higher clonogenicity than non-side populations and sustained tumourigenicity. They have also shown increased multi-drug resistance and are phenotypically similar to stem cells thanks to the expression of Oct-4 and Nanog for instance. Through analogy with embryonic stem cells, this side population has been called cancer stem cells or stem-like cells. However, their immune-tolerant property, their low cycling characteristic and drug resistance have led to this population also being referred to as dormant, quiescent, tolerant and persister cells (Vallette et al. 2018). It has been suggested that cancer stem cells are unique subclones within a tumour, responsible for tumour progression, resistance to therapies and the initiation of metastases. This definition is supported by clinical cases showing metastases more than 20 years after complete remission (Halldorsson et al. 2009) or a local recurrent disease after inoculation of adipose tissue 13 years after complete remission (Perrot et al. 2010).
In the last few decades, numerous works have tried to identify specific markers and the properties of cancer stem cells in osteosarcoma (Table 11.1). Osteosarcoma cancer stem cells are supported by sox2, a stem cell transcription factor which inhibits the Hippo pathway (Basu-Roy et al. 2012, 2015). In addition to the expression of the stemness markers shared with embryonic stem cells, osteosarcoma cancer stem cells have been characterized by their ability to form cell spheroids in vitro, which are highly tumourigenic in vivo (Murase et al. 2009; Yang et al. 2011). Cancer stem-like cells expressed high levels of aldehyde deshydrogenase-1 (ALDH1) (Honoki et al. 2010; Greco et al. 2014). ALDH-1 expression was associated with resistance to chemotherapy (Honoki et al. 2010) and the metastatic potential of cancer cells (Greco et al. 2014). The receptors for stem cell growth factor (CD117) and stro-1 expressed by mesenchymal stem cells are expressed by osteosarcoma cancer stem cells and were associated with metastasis and drug resistance (Adhikari et al. 2010). CD133 was also linked to the stem cell phenotype in osteosarcoma (Tirino et al. 2008, 2011; He et al. 2012; Li et al. 2013; Fujiwara et al. 2014). CD133 or prominin-1 is a pentaspan transmembrane glycoprotein localized in cellular protrusions (Glumac and LeBeau 2018). Like ALDH1+ cells, CD133-expressing osteosarcoma cells displayed high tumourigenicity in vivo (Tirino et al. 2011). Its high expression in patients predicted lung metastases and consequently correlated with poor prognosis (He et al. 2012; Li et al. 2013; Fujiwara et al. 2014). Tian et al. demonstrated the expression of CD271, a low-affinity nerve growth factor receptor, by osteosarcoma cancer stem cells (Tian et al. 2014) and defective autophagy led to the suppression of the stem-like properties of CD271+ (Zhang et al. 2016). Numerous other factors (CBX3, KLF4, SATB2, etc.) summarized in Table 11.1 controlled the biological properties/maintenance of stem cells. The biology of osteosarcoma cancer stem cells is also under the control of epigenetic networks. Several recently identified microRNAs regulate stem cell phenotype and their invasion and migration properties by targeting specific molecular pathways, such as PTEN, POU5F1 Wnt or Jagged1 (Lu et al. 2017; Zhang et al. 2018a; Zou et al. 2017; Zhao et al. 2017; Guo et al. 2017; Di Fiore et al. 2016; La Noce et al. 2018).
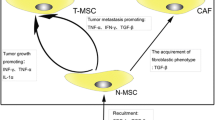
Osteosarcoma cancer stem cells are resistant to chemotherapy and radiotherapy, and can drive cancer recurrence (Gibbs et al. 2005; Fujii et al. 2009). Consequently, conventional chemotherapy impacts cancer stem cells and enriches the tumour mass in stem cells (Martins-Neves et al. 2012). They play a significant role in tumour heterogeneity through permanent enrichment of new mutated cancer cells and dominant subclones, and by regulating their local micro-environment. Cancer cells dialogue permanently with locally based partners. These communications include direct exchanges of small mediators using channels of the gap junction type (Tellez-Gabriel et al. 2017), with some selectivity. For instance, endothelial cells use gap junctions to communicate with osteosarcoma cells, and cancer cells do not communicate using this mode of communication with undifferentiated mesenchymal stem cells, unlike mesenchymal stem cells, which initiate their differentiation toward the osteoblast lineage. The dialogue between both cell types is controlled by acidosis. Acid-activated mesenchymal stem cells influence osteosarcoma cell behaviour such as their stemness properties (Avnet et al. 2017). Mesenchymal stem cells and cancer cells can also dialogue through the release of extracellular vesicles (Cortini et al. 2017; Baglio et al. 2017). Baglio et al. recently demonstrated the role of exosomes in osteosarcoma development and, more interestingly, that tumour cells educated mesenchymal stem cells by paracrine activity associating extracellular vesicles (Baglio et al. 2017). Tumour exosomes containing both IL-1 and TGFβ educated mesenchymal stem cells, which in turn promoted tumour growth and the development of lung metastases. In addition to identifying new therapeutic targets, these works show that osteosarcoma cells can regulate their micro-environment qualitatively and consequently enrich tumour heterogeneity. These data are reinforced by publications that underline the role of TGFβ in the stemness of osteosarcoma cells (Zhang et al. 2013; Lamora et al. 2016; Martins-Neves et al. 2016b). TGFβ1 is thus crucial for the differentiation of osteosarcoma cells for cancer toward cancer stem cells (Zhang et al. 2013). The second key molecular pathway for stemness in osteosarcoma is Wnt/beta catenin signalling, which supports stem cell formation (Lamora et al. 2016). Crosstalk between both pathways has been observed in both chronic inflammation and carcinogenesis (Martins-Neves et al. 2016b). Local immunity is also controlled by cell communications leading to an increase in tumour heterogeneity (Heymann et al. 2017). In parallel to the heterogeneity of cancer cells, immune heterogeneity with tumour-associated macrophages (Vallée and Lecarpentier 2018) and tumour-infiltrating lymphocytes (Sharma and Capobianco 2017) has been established and defines an immune-tolerant niche.
11.4 Tumour Heterogeneity, Cancer Stem Cells and New Therapeutic Options
Based on the data available in the literature, blocking agents have been developed as new therapeutic options for osteosarcoma patients to overcome drug resistance (Heymann et al. 2016; Dumars et al. 2016; Saraf et al. 2018; Makena et al. 2018). Thus, pimozide and resveratrol inhibit osteosarcoma cancer stem cells (Gonçalves et al. 2019; Peng and Jiang 2018). Pimozide blocks the epithelial-to-mesenchymal transition and both drugs interrupt the STAT-3 (IL-6 signalling pathways) and Wnt-β/catenin signalling modulated by TGBβ. Numerous therapeutic options target the TGFβ pathway. Miao et al. have recently developed single-walled carbon nanotubes to specifically inhibit TGFβ-induced osteosarcoma cell dedifferentiation and prevent the acquisition of the stem cell phenotype (Miao et al. 2017). Targeting the Wnt/beta catenin pathway may be a therapeutic alternative (Martins-Neves et al. 2018). Martins-Neves et al. used IWR-1, a tankyrase inhibitor to attenuate Wnt/beta catenin signalling in osteosarcoma cancer stem cells with promising results in pre-clinical mouse models (Martins-Neves et al. 2018). Blockading N-cadherin/NF-KB signalling also appears interesting with the administration of metformin to inhibiting the stem cell phenotype (Xu et al. 2017). Shang et al. revealed that metformin increased the sensitivity of stem cells to conventional chemotherapy and confirmed the advantages of this drug in the treatment of osteosarcoma (Shang et al. 2017). Metformin should be considered as a metabolic modulator of osteosarcoma cancer stem cells (Paiva-Oliveira et al. 2018). Targeting EGFR and CD133 (Chen et al. 2018), Sox9 (Qu et al. 2018), TSSSC3 and Src/AKt pathways (Yan et al. 2017) FGFR2 (Zhang et al. 2018b) has also been assessed recently in osteosarcoma and exhibited high efficacy by repressing the self-renewal of stem cells, tumour growth and the metastatic process. Immunotherapies are interesting potential future options (Heymann et al. 2016; Mesiano et al. 2018), as shown recently by Mesiano et al. (2018) and D’Angelo et al. (2018). Mesiano et al. used cytokine-induced killer cells which are effective against cancer stem cells in sarcoma (Mesiano et al. 2018). D’Angelo et al. developed autologous T cells expressing NY-ESO-1c259 expressed by synovial sarcomas (D’Angelo et al. 2018), and showed that patients with metastases treated with an affinity-enhanced T-cell receptor recognizing an HLA-A2-restricted NY-ESO-1/LAGE1a-derived peptide, increased the anti-tumour response by around 50%. In addition, circulating NY-ESO-1c259 T cells were detectable in blood for at least 6 months in all responders, and most administered NY-ESO-1c259 T cells exhibited an effector memory phenotype following ex vivo expansion (D’Angelo et al. 2018).
11.5 Conclusion
Cancer stem cells, which should be called cancer “stem-like” cells, are detectable in osteosarcoma. They contribute markedly to tumour heterogeneity and are responsible for drug resistance. Dialogue is established between cancer stem-like cells and their local micro-environment, and they are able to educate to facilitate their maintenance and development. This dialogue is a future potential target and there is drug development in combination with conventional chemotherapies (Fig. 11.1). Better characterization of cancer stem-like cells in osteosarcoma and their role in the clonal evolution of the disease is mandatory for improving the therapeutic response of poor responders, as well as for improving the overall survival of osteosarcoma patients which has changed little in the last four decades.
References
Adhikari AS, Agarwal N, Wood BM, Porretta C, Ruiz B, Pochampally RR, Iwakuma T (2010) CD117 and Stro-1 identify osteosarcoma tumor-initiating cells associated with metastasis and drug resistance. Cancer Res 70:4602–4612
Alfranca A, Martinez-Cruzado L, Tornin J, Abarrategi A, Amaral T, de Alava E, Menendez P, Garcia-Castro J, Rodriguez R (2015) Bone microenvironment signals in osteosarcoma development. Cell Mol Life Sci 72:3097–3113
Avnet S, Di Pompo G, Chano T, Errani C, Ibrahim-Hashim A, Gillies RJ, Donati DM, Baldini N (2017) Cancer-associated mesenchymal stroma fosters the stemness of osteosarcoma cells in response to intratumoral acidosis via NF-κB activation. Int J Cancer 140:1331–1345
Baglio SR, Lagerweij T, Pérez-Lanzón M, Ho XD, Léveillé N, Melo SA, Cleton-Jansen AM, Jordanova ES, Roncuzzi L, Greco M, van Eijndhoven MAJ, Grisendi G, Dominici M, Bonafede R, Lougheed SM, de Gruijl TD, Zini N, Cervo S, Steffan A, Canzonieri V, Martson A, Maasalu K, Köks S, Wurdinger T, Baldini N, Pegtel DM (2017) Blocking tumor-educated MSC paracrine activity halts osteosarcoma progression. Clin Cancer Res 23:3721–3733
Basu-Roy U, Seo E, Ramanathapuram L, Rapp TB, Perry JA, Orkin SH, Mansukhani A, Basilico C (2012) Sox2 maintains self-renewal of tumor-initiating cells in osteosarcomas. Oncogene 31:2270–2282
Basu-Roy U, Bayin NS, Rattanakorn K, Han E, Placantonakis DG, Mansukhani A, Basilico C (2015) Sox2 antagonizes the Hippo pathway to maintain stemness in cancer cells. Nat Commun 6:6411
Bousquet M, Noirot C, Accadbled F, Sales de Gauzy J, Castex MP, Brousset P, Gomez-Brouchet A (2016) Whole-exome sequencing in osteosarcoma reveals important heterogeneity of genetic alterations. Ann Oncol 27:738–744
Brown HK, Tellez-Gabriel M, Heymann D (2017) Cancer stem cells in osteosarcoma. Cancer Lett 386:189–195
Brown HK, Schiavone K, Gouin F, Heymann MF, Heymann D (2018a) Biology of bone sarcomas and new therapeutic developments. Calcif Tissue Int 102:174–195
Brown HK, Tellez-Gabriel M, Cartron PF, Vallette FM, Heymann MF, Heymann D (2018b) Characterization of circulating tumor cells as a reflection of the tumor heterogeneity: myth or reality? Drug Discov Today. https://doi.org/10.1016/j.drudis.2018.11.017
Chano T, Kita H, Avnet S, Lemma S, Baldini N (2018) Prominent role of RAB39A-RXRB axis in cancer development and stemness. Oncotarget 9:9852–9866
Chen KS, Kwon WS, Kim J, Heo SJ, Kim HS, Kim HK, Kim SH, Lee WS, Chung HC, Rha SY, Hwang TH (2016) A novel TP53-KPNA3 translocation defines a de novo treatment-resistant clone in osteosarcoma. Cold Spring Harb Mol Case Stud 2:a000992
Chen F, Zeng Y, Qi X, Chen Y, Ge Z, Jiang Z, Zhang X, Dong Y, Chen H, Yu Z (2018) Targeted salinomycin delivery with EGFR and CD133 aptamers based dual-ligand lipid-polymer nanoparticles to both osteosarcoma cells and cancer stem cells. Nanomedicine 14:2115–2127
Cortini M, Avnet S, Baldini N (2017) Mesenchymal stroma: role in osteosarcoma progression. Cancer Lett 405:90–99
D’Angelo SP, Melchiori L, Merchant MS, Bernstein D, Glod J, Kaplan R, Grupp S, Tap WD, Chagin K, Binder GK, Basu S, Lowther DE, Wang R, Bath N, Tipping A, Betts G, Ramachandran I, Navenot JM, Zhang H, Wells DK, Van Winkle E, Kari G, Trivedi T, Holdich T, Pandite L, Amado R, Mackall CL (2018) Antitumor activity associated with prolonged persistence of adoptively transferred NY-ESO-1 c259T cells in synovial sarcoma. Cancer Discov 8:944–957
de Groot AE, Roy S, Brown JS, Pienta KJ, Amend SR (2017) Revisiting seed and soil: examining the primary tumor and cancer cell foraging in metastasis. Mol Cancer Res 15:361–370
Di Fiore R, Drago-Ferrante R, Pentimalli F, Di Marzo D, Forte IM, Carlisi D, De Blasio A, Tesoriere G, Giordano A, Vento R (2016) Let-7d miRNA shows both antioncogenic and oncogenic functions in osteosarcoma-derived 3AB-OS cancer stem cells. J Cell Physiol 231:1832–1841
Dumars C, Ngyuen JM, Gaultier A, Lanel R, Corradini N, Gouin F, Heymann D, Heymann MF (2016) Dysregulation of macrophage polarization is associated with the metastatic process in osteosarcoma. Oncotarget 7:78343–78354
Engert F, Kovac M, Baumhoer D, Nathrath M, Fulda S (2017) Osteosarcoma cells with genetic signatures of BRCAness are susceptible to the PARP inhibitor talazoparib alone or in combination with chemotherapeutics. Oncotarget 8:48794–48806
Feng H, Tillman H, Wu G, Davidoff AM, Yang J (2018) Frequent epigenetic alterations in polycomb repressive complex 2 in osteosarcoma cell lines. Oncotarget 9:27087–27091
Franzetti GA, Laud-Duval K, van der Ent W, Brisac A, Irondelle M, Aubert S, Dirksen U, Bouvier C, de Pinieux G, Snaar-Jagalska E, Chavrier P, Delattre O (2017) Cell-to-cell heterogeneity of EWSR1-FLI1 activity determines proliferation/migration choices in Ewing sarcoma cells. Oncogene 36:3505–3514
Fujii H, Honoki K, Tsujiuchi T, Kido A, Yoshitani K, Takakura Y (2009) Sphere-forming stem-like cell populations with drug resistance in human sarcoma cell lines. Int J Oncol 34:1381–1386
Fujiwara T, Katsuda T, Hagiwara K, Kosaka N, Yoshioka Y, Takahashi RU, Takeshita F, Kubota D, Kondo T, Ichikawa H, Yoshida A, Kobayashi E, Kawai A, Ozaki T, Ochiya T (2014) Clinical relevance and therapeutic significance of microRNA-133a expression profiles and functions in malignant osteosarcoma-initiating cells. Stem Cells 32:959–973
Funes JM, Quintero M, Henderson S, Martinez D, Qureshi U, Westwood C, Clements MO, Bourboulia D, Pedley RB, Moncada S, Boshoff C (2007) Transformation of human mesenchymal stem cells increases their dependency on oxidative phosphorylation for energy production. Proc Natl Acad Sci U S A 104:6223–6228
Gambera S, Abarrategi A, González-Camacho F, Morales-Molina Á, Roma J, Alfranca A, García-Castro J (2018) Clonal dynamics in osteosarcoma defined by RGB marking. Nat Commun 9:3994
Georges S, Calleja LR, Jacques C, Lavaud M, Moukengue B, Lecanda F, Quillard T, Gabriel MT, Cartron PF, Baud’huin M, Lamoureux F, Heymann D, Ory B (2018) Loss of miR-198 and -206 during primary tumor progression enables metastatic dissemination in human osteosarcoma. Oncotarget 9:35726–35741
Gibbs CP, Kukekov VG, Reith JD, Tchigrinova O, Suslov ON, Scott EW, Ghivizzani SC, Ignatova TN, Steindler DA (2005) Stem-like cells in bone sarcomas: implications for tumorigenesis. Neoplasia 7:967–976
Glumac PM, LeBeau AM (2018) The role of CD133 in cancer: a concise review. Clin Transl Med 7:18
Gonçalves JM, Silva CAB, Rivero ERC, Cordeiro MMR (2019) Inhibition of cancer stem cells promoted by Pimozide. Clin Exp Pharmacol Physiol 46(2):116–125
Greaves M, Maley CC (2012) Clonal evolution in cancer. Nature 481:306–313
Greco N, Schott T, Mu X, Rothenberg A, Voigt C, McGough RL 3rd, Goodman M, Huard J, Weiss KR (2014) ALDH activity correlates with metastatic potential in primary sarcomas of bone. J Cancer Ther 5:331–338
Guo X, Yu L, Zhang Z, Dai G, Gao T, Guo W (2017) miR-335 negatively regulates osteosarcoma stem cell-like properties by targeting POU5F1. Cancer Cell Int 17:29
Guth AM, Deogracias M, Dow SW (2014) Comparison of cancer stem cell antigen expression by tumor cell lines and by tumor biopsies from dogs with melanoma and osteosarcoma. Vet Immunol Immunopathol 161:132–140
Halldorsson A, Brooks S, Montgomery S, Graham S (2009) Lung metastasis 21 years after initial diagnosis of osteosarcoma: a case report. J Med Case Rep 3:9298
He A, Qi W, Huang Y, Feng T, Chen J, Sun Y, Shen Z, Yao Y (2012) CD133 expression predicts lung metastasis and poor prognosis in osteosarcoma patients: a clinical and experimental study. Exp Ther Med 4:435–441
He A, Yang X, Huang Y, Feng T, Wang Y, Sun Y, Shen Z, Yao Y (2015) CD133(+) CD44(+) cells mediate in the lung metastasis of osteosarcoma. J Cell Biochem 116:1719–1729
Heymann MF, Brown HK, Heymann D (2016) Drugs in early clinical development for the treatment of osteosarcoma. Expert Opin Investig Drugs 25:1265–1280
Heymann MF, Lézot F, Heymann D (2017) The contribution of immune infiltrates and the local microenvironment in the pathogenesis of osteosarcoma. Cell Immunol. https://doi.org/10.1016/j.cellimm.2017.10.011
Hochane M, Trichet V, Pecqueur C, Avril P, Oliver L, Denis J, Brion R, Amiaud J, Pineau A, Naveilhan P, Heymann D, Vallette FM, Olivier C (2017) Low-dose pesticide mixture induces senescence in normal mesenchymal stem cells (MSC) and promotes tumorigenic phenotype in premalignant MSC. Stem Cells 35:800–811
Honoki K, Fujii H, Kubo A, Kido A, Mori T, Tanaka Y, Tsujiuchi T (2010) Possible involvement of stem-like populations with elevated ALDH1 in sarcomas for chemotherapeutic drug resistance. Oncol Rep 24:501–5015
Itoh H, Kadomatsu T, Tanoue H, Yugami M, Miyata K, Endo M, Morinaga J, Kobayashi E, Miyamoto T, Kurahashi R, Terada K, Mizuta H, Oike Y (2018) TET2-dependent IL-6 induction mediated by the tumor microenvironment promotes tumor metastasis in osteosarcoma. Oncogene 37:2903–2920
Kovac M, Blattmann C, Ribi S, Smida J, Mueller NS, Engert F, Castro-Giner F, Weischenfeldt J, Kovacova M, Krieg A, Andreou D, Tunn PU, Dürr HR, Rechl H, Schaser KD, Melcher I, Burdach S, Kulozik A, Specht K, Heinimann K, Fulda S, Bielack S, Jundt G, Tomlinson I, Korbel JO, Nathrath M, Baumhoer D (2015) Exome sequencing of osteosarcoma reveals mutation signatures reminiscent of BRCA deficiency. Nat Commun 6:8940
La Noce M, Paino F, Mele L, Papaccio G, Regad T, Lombardi A, Papaccio F, Desiderio V, Tirino V (2018) HDAC2 depletion promotes osteosarcoma’s stemness both in vitro and in vivo: a study on a putative new target for CSCs directed therapy. J Exp Clin Cancer Res 37:296
Lamora A, Talbot J, Mullard M, Brounais-Le Royer B, Redini F, Verrecchia F (2016) TGF-β Signaling in bone remodeling and osteosarcoma progression. J Clin Med 5(11):E96
Lamoureux F, Baud’huin M, Rodriguez Calleja L, Jacques C, Berreur M, Rédini F, Lecanda F, Bradner JE, Heymann D, Ory B (2014) Selective inhibition of BET bromodomain epigenetic signalling interferes with the bone-associated tumour vicious cycle. Nat Commun 5:3511
Li J, Zhong XY, Li ZY, Cai JF, Zou L, Li JM, Yang T, Liu W (2013) CD133 expression in osteosarcoma and derivation of CD133+ cells. Mol Med Rep 7:577–584
Li Y, Xian M, Yang B, Ying M, He Q (2017) Inhibition of KLF4 by statins reverses Adriamycin-induced metastasis and cancer stemness in osteosarcoma cells. Stem Cell Rep 8:1617–1629
Li B, Wang Z, Wu H, Xue M, Lin P, Wang S, Lin N, Huang X, Pan W, Liu M, Yan X, Qu H, Sun L, Li H, Wu Y, Teng W, Wang Z, Zhou X, Chen H, Poznansky MC, Ye Z (2018) Epigenetic regulation of CXCL12 plays a critical role in mediating tumor progression and the immune response in osteosarcoma. Cancer Res 78:3938–3953
Liu F, Li L, Li Y, Ma X, Bian X, Liu X, Wang G, Zhang D (2018) Overexpression of SENP1 reduces the stemness capacity of osteosarcoma stem cells and increases their sensitivity to HSVtk/GCV. Int J Oncol 53:2010–2020
López-Lázaro M (2018) The stem cell division theory of cancer. Crit Rev Oncol Hematol 123:95–113
Lu J, Song G, Tang Q, Yin J, Zou C, Zhao Z, Xie X, Xu H, Huang G, Wang J, Lee DF, Khokha R, Yang H, Shen J (2017) MiR-26a inhibits stem cell-like phenotype and tumor growth of osteosarcoma by targeting Jagged1. Oncogene 36:231–241
Makena MR, Ranjan A, Thirumala V, Reddy A (2018) Cancer stem cells: road to therapeutic resistance and strategies to overcome resistance. Biochim Biophys Acta Mol basis Dis. https://doi.org/10.1016/j.bbadis.2018.11.015
Martins-Neves SR, Lopes AO, do Carmo A, Paiva AA, Simoes PC, Abrunhosa AJ, Gomes CM (2012) Therapeutic implications of an enriched cancer stem-like cell population in a human osteosarcoma cell line. BMC Cancer 12:139
Martins-Neves SR, Corver WE, Paiva-Oliveira DI, van den Akker BE, Briaire-de-Bruijn IH, Bovee JV, Gomes CM, Cleton-Jansen AM (2016a) Osteosarcoma stem cells have active Wnt/beta-catenin and overexpress SOX2 and KLF4. J Cell Physiol 231:876–886
Martins-Neves SR, Paiva-Oliveira DI, Wijers-Koster PM, Abrunhosa AJ, Fontes-Ribeiro C, Bovee JV, Cleton-Jansen AM, Gomes CM (2016b) Chemotherapy induces stemness in osteosarcoma cells through activation of Wnt/beta-catenin signaling. Cancer Lett 370:286–295
Martins-Neves SR, Paiva-Oliveira DI, Fontes-Ribeiro C, Bovée JVMG, Cleton-Jansen AM, Gomes CMF (2018) IWR-1, a tankyrase inhibitor, attenuates Wnt/β-catenin signaling in cancer stem-like cells and inhibits in vivo the growth of a subcutaneous human osteosarcoma xenograft. Cancer Lett 414:1–15
Mesiano G, Grignani G, Fiorino E, Leuci V, Rotolo R, D’Ambrosio L, Salfi C, Gammaitoni L, Giraudo L, Pisacane A, Butera S, Pignochino Y, Basiricó M, Capozzi F, Sapino A, Aglietta M, Sangiolo D (2018) Cytokine induced killer cells are effective against sarcoma cancer stem cells spared by chemotherapy and target therapy. Oncoimmunology 7:e1465161
Miao Y, Zhang H, Pan Y, Ren J, Ye M, Xia F, Huang R, Lin Z, Jiang S, Zhang Y, Songyang Z, Zhang Y (2017) Single-walled carbon nanotube: one specific inhibitor of cancer stem cells in osteosarcoma upon downregulation of the TGFβ1 signaling. Biomaterials 149:29–40
Mohseny AB, Szuhai K, Romeo S, Buddingh EP, Briaire-de Bruijn I, de Jong D, van Pel M, Cleton-Jansen AM, Hogendoorn PC (2009) Osteosarcoma originates from mesenchymal stem cells in consequence of aneuploidization and genomic loss of Cdkn2. J Pathol 219:294–305
Murase M, Kano M, Tsukahara T, Takahashi A, Torigoe T, Kawaguchi S, Kimura S, Wada T, Uchihashi Y, Kondo T, Yamashita T, Sato N (2009) Side population cells have the characteristics of cancer stem-like cells/cancer-initiating cells in bone sarcomas. Br J Cancer 101:1425–1432
Mutsaers AJ, Walkley CR (2014) Cells of origin in osteosarcoma: mesenchymal stem cells or osteoblast committed cells? Bone 62:56–63
Najafi M, Farhood B, Mortezaee K (2019) Cancer stem cells (CSCs) in cancer progression and therapy. J Cell Physiol 234(6):8381–8395
Paiva-Oliveira DI, Martins-Neves SR, Abrunhosa AJ, Fontes-Ribeiro C, Gomes CMF (2018) Therapeutic potential of the metabolic modulator metformin on osteosarcoma cancer stem-like cells. Cancer Chemother Pharmacol 81:49–63
Peng L, Jiang D (2018) Resveratrol eliminates cancer stem cells of osteosarcoma by STAT3 pathway inhibition. PLoS One 13:e0205918
Perrot P, Rousseau J, Bouffaut AL, Rédini F, Cassagnau E, Deschaseaux F, Heymann MF, Heymann D, Duteille F, Trichet V, Gouin F (2010) Safety concern between autologous fat graft, mesenchymal stem cell and osteosarcoma recurrence. PLoS One 5:e10999
Qi XT, Li YL, Zhang YQ, Xu T, Lu B, Fang L, Gao JQ, Yu LS, Zhu DF, Yang B, He QJ, Ying MD (2018) KLF4 functions as an oncogene in promoting cancer stem cell-like characteristics in osteosarcoma cells. Acta Pharmacol Sin. https://doi.org/10.1038/s41401-018-0050-6
Qu H, Xue Y, Lian W, Wang C, He J, Fu Q, Zhong L, Lin N, Lai L, Ye Z, Wang Q (2018) Melatonin inhibits osteosarcoma stem cells by suppressing SOX9-mediated signaling. Life Sci 207:253–264
Reimann E, Kõks S, Ho XD, Maasalu K, Märtson A (2014) Whole exome sequencing of a single osteosarcoma case--integrative analysis with whole transcriptome RNA-seq data. Hum Genomics 8:20
Rubio R, Abarrategi A, Garcia-Castro J, Martinez-Cruzado L, Suarez C, Tornin J, Santos L, Astudillo A, Colmenero I, Mulero F, Rosu-Myles M, Menendez P, Rodriguez R (2014) Bone environment is essential for osteosarcoma development from transformed mesenchymal stem cells. Stem Cells 32:1136–1148
Saini V, Hose CD, Monks A, Nagashima K, Han B, Newton DL, Millione A, Shah J, Hollingshead MG, Hite KM, Burkett MW, Delosh RM, Silvers TE, Scudiero DA, Shoemaker RH (2012) Identification of CBX3 and ABCA5 as putative biomarkers for tumor stem cells in osteosarcoma. PLoS One 7:e41401
Saraf AJ, Fenger JM, Roberts RD (2018) Osteosarcoma: accelerating progress makes for a hopeful future. Front Oncol 8:4
Sarver AE, Subramanian S (2016) Imprinting defects in osteosarcoma: DNA- and chromatin-modifying drugs hold promise for osteosarcoma therapy. Epigenomics 8:885–888
Shang D, Wu J, Guo L, Xu Y, Liu L, Lu J (2017) Metformin increases sensitivity of osteosarcoma stem cells to cisplatin by inhibiting expression of PKM2. Int J Oncol 50:1848–1856
Sharma A, Capobianco E (2017) Immuno-oncology integrative networks: elucidating the influences of osteosarcoma phenotypes. Cancer Inform 16:1176935117721691
Smida J, Xu H, Zhang Y, Baumhoer D, Ribi S, Kovac M, von Luettichau I, Bielack S, O’Leary VB, Leib-Mösch C, Frishman D, Nathrath M (2017) Genome-wide analysis of somatic copy number alterations and chromosomal breakages in osteosarcoma. Int J Cancer 141:816–828
Surdez D, Guillemot D, Lapouble E, Freneaux P, Champigneulle J, Bouvier R, Walder D, Ambros IM, Hutter C, Sorz E, Amaral AT, de Álava E, Schallmoser K, Strunk D, Rinner B, Liegl-Atzwanger B, Huppertz B, Leithner A, de Pinieux G, Terrier P, Laurence V, Michon J, Ladenstein R, Holter W, Windhager R, Dirksen U, Ambros PF, Delattre O, Kovar H, Bock C, Tomazou EM (2017) DNA methylation heterogeneity defines a disease spectrum in Ewing sarcoma. Nat Med 23:386–395
Tang N, Song WX, Luo J, Haydon RC, He TC (2008) Osteosarcoma development and stem cell differentiation. Clin Orthop Relat Res 466:2114–2130
Tellez-Gabriel M, Ory B, Lamoureux F, Heymann MF, Heymann D (2016) Tumour heterogeneity: the key advantages of single-cell analysis. Int J Mol Sci 17(12):E2142
Tellez-Gabriel M, Charrier C, Brounais-Le Royer B, Mullard M, Brown HK, Verrecchia F, Heymann D (2017) Analysis of gap junctional intercellular communications using a dielectrophoresis-based microchip. Eur J Cell Biol 96:110–118
Théoleyre S, Mori K, Cherrier B, Passuti N, Gouin F, Rédini F, Heymann D (2005) Phenotypic and functional analysis of lymphocytes infiltrating osteolytic tumors: use as a possible therapeutic approach of osteosarcoma. BMC Cancer 5:123
Tian J, Li X, Si M, Liu T, Li J (2014) CD271+ osteosarcoma cells display stem-like properties. PLoS One 9:e98549
Tirino V, Desiderio V, d’Aquino R, De Francesco F, Pirozzi G, Graziano A, Galderisi U, Cavaliere C, De Rosa A, Papaccio G, Giordano A (2008) Detection and characterization of CD133+ cancer stem cells in human solid tumours. PLoS One 3:e3469
Tirino V, Desiderio V, Paino F, De Rosa A, Papaccio F, Fazioli F, Papaccio G (2011) Human primary bone sarcomas contain CD133+ cancer stem cells displaying high tumorigenicity in vivo. FASEB J 25:2022–2030
Vallée A, Lecarpentier Y (2018) Crosstalk between peroxisome proliferator-activated receptor gamma and the canonical WNT/β-catenin pathway in chronic inflammation and oxidative stress during carcinogenesis. Front Immunol 9:745
Vallette FM, Olivier C, Lézot F, Oliver L, Cochonneau D, Lalier L, Cartron PF, Heymann D (2018) Dormant, quiescent, tolerant and persister cells: four synonyms for the same target in cancer. Biochem Pharmacol. https://doi.org/10.1016/j.bcp.2018.11.004
Wang L, Park P, Zhang H, La Marca F, Lin CY (2011) Prospective identification of tumorigenic osteosarcoma cancer stem cells in OS99-1 cells based on high aldehyde dehydrogenase activity. Int J Cancer 128:294–303
Wang J, Wang D, Niu X, Wang Z, Song CL, Huang Z, Chen KN, Duan J, Bai H, Xu J, Zhao J, Wang Y, Zhuo M, Xie XS, Kang X, Tian Y, Cai L, Han JF, An T, Sun Y, Gao S, Zhao J, Ying J, Wang L, He J (2019) Multiregion sequencing reveals the genetic heterogeneity and evolutionary history of osteosarcoma and matched pulmonary metastases. Cancer Res 79(1):7–20
Xi X, Wu Q, Bao Y, Zhong X, Dai X, Lin H (2019) Overexpression of TBL1XR1 confers tumorigenic capability and promotes recurrence of osteosarcoma. Eur J Pharmacol 844:259–267
Xu HY, Fang W, Huang ZW, Lu JC, Wang YQ, Tang QL, Song GH, Kang Y, Zhu XJ, Zou CY, Yang HL, Shen JN, Wang J (2017) Metformin reduces SATB2-mediated osteosarcoma stem cell-like phenotype and tumor growth via inhibition of N-cadherin/NF-kB signaling. Eur Rev Med Pharmacol Sci 21(20):4516–4528
Yan GN, Tang XF, Zhang XC, He T, Huang YS, Zhang X, Meng G, Guo DY, Lv YF, Guo QN (2017) TSSC3 represses self-renewal of osteosarcoma stem cells and Nanog expression by inhibiting the Src/Akt pathway. Oncotarget 8:85628–85641
Yang M, Yan M, Zhang R, Li J, Luo Z (2011) Side population cells isolated from human osteosarcoma are enriched with tumor-initiating cells. Cancer Sci 102:1774–1781
Yu L, Liu S, Zhang C, Zhang B, Simoes BM, Eyre R, Liang Y, Yan H, Wu Z, Guo W, Clarke RB (2013) Enrichment of human osteosarcoma stem cells based on hTERT transcriptional activity. Oncotarget 4:2326–2338
Zhang H, Wu H, Zheng J, Yu P, Xu L, Jiang P, Gao J, Wang H, Zhang Y (2013) Transforming growth factor beta1 signal is crucial for dedifferentiation of cancer cells to cancer stem cells in osteosarcoma. Stem Cells 31:433–446
Zhang D, Zhao Q, Sun H, Yin L, Wu J, Xu J, He T, Yang C, Liang C (2016) Defective autophagy leads to the suppression of stem-like features of CD271+ osteosarcoma cells. J Biomed Sci 23:82
Zhang RM, Tang T, Yu HM, Yao XD (2018a) LncRNA DLX6-AS1/miR-129-5p/DLK1 axis aggravates stemness of osteosarcoma through Wnt signaling. Biochem Biophys Res Commun 507:260–266
Zhang W, Zhao JM, Lin J, Hu CZ, Zhang WB, Yang WL, Zhang J, Zhang JW, Zhu J (2018b) Adaptive fibrogenic reprogramming of osteosarcoma stem cells promotes metastatic growth. Cell Rep 24:1266–1277
Zhao D, Chen Y, Chen S, Zheng C, Hu J, Luo S (2017) MiR-19a regulates the cell growth and apoptosis of osteosarcoma stem cells by targeting PTEN. Tumour Biol 39:1010428317705341
Zou Y, Huang Y, Yang J, Wu J, Luo C (2017) miR-34a is downregulated in human osteosarcoma stem-like cells and promotes invasion, tumorigenic ability and self-renewal capacity. Mol Med Rep 15:1631–1637
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Schiavone, K., Garnier, D., Heymann, MF., Heymann, D. (2019). The Heterogeneity of Osteosarcoma: The Role Played by Cancer Stem Cells. In: Birbrair, A. (eds) Stem Cells Heterogeneity in Cancer. Advances in Experimental Medicine and Biology, vol 1139. Springer, Cham. https://doi.org/10.1007/978-3-030-14366-4_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-14366-4_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-14365-7
Online ISBN: 978-3-030-14366-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)