Abstract
Understanding genotype–phenotype dependency is a universal aim for all life sciences. While the complete genotype–phenotype relations remain challenging to resolve, metabolic phenotypes are moving within the reach through genome-scale metabolic model simulations. Genome-scale metabolic models are available for commonly investigated yeasts, such as model eukaryote and domesticated fermentation species Saccharomyces cerevisiae, and automatic reconstruction methods facilitate obtaining models for any sequenced species. The models allow for investigating genotype–phenotype relations through simulations simultaneously considering the effects of nutrient availability, and redox and energy homeostasis in cells. Genome-scale models also offer frameworks for omics data integration to help to uncover how the translation of genotypes to the apparent phenotypes is regulated at different levels. In this chapter, we provide an overview of the yeast genome-scale metabolic models and the simulation approaches for using these models to interrogate genotype–phenotype relations. We review the methodological approaches according to the underlying biological reasoning in order to inspire formulating novel questions and applications that the genome-scale metabolic models could contribute to. Finally, we discuss current challenges and opportunities in the genome-scale metabolic model simulations.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Genome-scale metabolic model
- Genotype–phenotype dependency
- Yeast metabolism
- Metabolic flux
- Strain design
5.1 Introduction to Genome-Scale Metabolic Models
Since the early distinction of genotypes from phenotypes (Johannsen 1911) life science research has sought for understanding their dependency. The dependency is inherently complex and dynamic. Single genotype may manifest several phenotypes (i.e., clonal heterogeneity) and different genotypes may translate to indistinguishable observable phenotypes. While the complete genotype–phenotype dependencies are challenging to resolve, metabolic phenotypes are moving within the reach through genome-scale metabolic model simulations. A genome-scale metabolic model is a description of the complete biochemical conversion potential encoded in an organism’s genome as a network of reactions (Fig. 5.1). The stoichiometries of these reactions form mass conservation constraints of cellular metabolism. When a biological optimality principle (e.g., fast cell growth) is additionally introduced, a steady-state metabolic phenotype can be simulated using powerful linear programming solvers. Such simulations holistically consider cellular resource, energy, and redox requirements for biochemical synthesis. A myriad of applications has been derived from the original undecorated phenotype simulation. The applications vary from simulating metabolic genotype–phenotype dependencies for finding cancer drug targets to designing genotype manipulations for achieving desired phenotypes in microbial hosts for industrial biotechnology needs.
Metabolic capacity of cells represented as a network of reactions or further as stoichiometric matrix allows simulations of metabolic phenotypes using linear programming. Metabolic steady-state assumption renders the system of metabolite mass balances linear. Reaction capacity and thermodynamic constraints can be included and limit the space of feasible metabolic phenotypes (i.e., metabolic fluxes)
Yeasts, unicellular eukaryotes, are suitable hosts for industrial biotechnology owing to their robustness against harsh growth environments, established genetic engineering tools for several species, and eukaryotic protein modification. They have scientific relevance also as simpler model system for higher cells and some yeasts are pathogenic causing difficult infections. Furthermore, yeasts, Saccharomyces cerevisiae, in particular, have been domesticated for food and beverage fermentations and baking already since ancient times. While S. cerevisiae is by far the most well studied and broadly used yeast in applications, several other species attract considerable interest as well. For instance, Pichia pastoris is a widely used protein production host, Kluyveromyces lactis is known for beta-galactosidase synthesis, Yarrowia lipolytica is an oleaginous yeast attractive for lipid production, Scheffersomyces stipitis is a naturally xylose-utilizing yeast, and pathogenic yeasts Candida tropicalis and Candida glabrata cause difficult infections urging for more efficient treatments to be developed. The variety of yeast species of scientific and application interest can be expected to broaden following the rise of CRISPR/Cas9 and other generally applicable genetic engineering tools such as synthetic expression system universal for fungi (Rantasalo et al. 2018). Genome sequences are already available for a large variety of yeasts. Reference genomes for 98 yeast species are available from NCBI (www.ncbi.nlm.nih.gov/genome).
5.1.1 Genome-Scale Metabolic Model Reconstruction
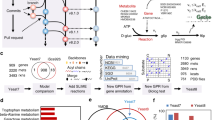
Genome sequence is the starting point for reconstructing a genome-scale metabolic model. Semi-automatic reconstruction methods are available for building the first drafts of genome-scale metabolic models from the genome sequences (Swainston et al. 2011; Agren et al. 2013; Pitkänen et al. 2014; Castillo et al. 2016; Dias et al. 2015). The quality of draft reconstructions after the semi-automatic processes is strongly dependent on the comprehensiveness and quality of the source reaction database used. The reaction database has to contain links from the reactions to corresponding gene/protein sequences either within the database or by proving adequate identifiers such as EC numbers for external mapping. Reactions need to essentially be atom balanced for mass conservation in the reconstructed model. Popular reaction databases for genome-scale metabolic model reconstruction include Kegg (Kanehisa et al. 2017), Rhea (Morgat et al. 2017), MetaCyc (Caspi et al. 2014), BiGG (Schellenberger et al. 2010), and Reactome (Fabregat et al. 2018). A confidence score for the presence of a reaction from the reaction database in the metabolic repertoire of the species is derived by most of the semi-automatic reconstruction methods. Then, the high scoring reactions are pulled to the model after which gap filling algorithms are used for introducing lower scoring reactions that are essential for the in silico synthesis of biomass. Gap filling benefits greatly from experimental data on the growth of the species under different nutrient environments (Tramontano et al. 2018). Alternatively, to the two-phase process of introducing high scoring reactions followed by gap filling for a functional model, a single step process of carving out the organism-specific metabolic network from a universal gapless model (CarveMe) has recently been proposed (Machado et al. 2018). When the universal model is well curated, simulatable species-specific models are fast to reconstruct using CarveMe (Machado et al. 2018). Further, using a universal model standardizes the quality of input reaction data for reconstructing different species models. However, there are also other sources of uncertainty in the model reconstruction such as the quality of the genome and the annotations, and the availability of similar annotated sequences in databases. Given the data, several models of a species could score equally well in the automatic reconstruction. Therefore, an approach has been suggested for simulating an ensemble of equally likely models simultaneously instead of a single reconstruction (Biggs and Papin 2017). Yet, evaluating the quality of models reconstructed for less well-studied non-model species is challenging. The reconstruction algorithms themselves can be evaluated against manually curated models and experimental data on model organisms such as metabolic gene knockout phenotypes. Metabolic gene knockout phenotypes can be simulated using the gene annotations of the models. The genes are annotated to the reactions whose catalyzing enzymes they encode. Preferably, the gene annotations include also Boolean rules describing whether the genes annotated to the reaction encode isoenzymes (i.e., OR rule) or whether they form a complex whose all components are required for activity (i.e., AND rule). Thereby, the Boolean rules allow propagating the genetic state into reaction activity state for performing mutant phenotype simulations. Simulated mutant phenotypes can be compared against experimental deletion mutant phenotypes for validating models. Though many metrics have been proposed for assessing the quality of reconstructed models (Sanchez and Nielsen 2015; Lopes and Rocha 2017), experimental growth and phenotype data are necessary for true evaluation (Tramontano et al. 2018).
5.2 Yeast Genome-Scale Models
Several genome-scale metabolic models have been reconstructed for S. cerevisiae during the last 15 years. The first S. cerevisiae model was created in 2003 by Föster et al. 2003 and was named iFF708 after the main developers and the number of genes supporting the reactions in the model. Slightly different and variable numbers of genes were annotated to metabolic reactions in the three following S. cerevisiae models (iND750, iLL672, and iIN800) derived directly from iFF708. Creating the first consensus model for S. cerevisiae was a collaborative effort. It was built on the iLL672 and iMM904 models (derived from iND750 model) and published in 2008 (Herrgård et al. 2008). After several updates of, in particular, lipid metabolism and transport reactions, the consensus model version 7 was published in 2013 by Aung et al. (2018). Since then the consensus yeast model has gone through several smaller updates (https://github.com/SysBioChalmers/yeast-GEM). Heavner and Price (2015) compared the 12 (S. cerevisiae) metabolic models created from 2003 until 2015. Though the coverage (i.e., number of genes annotated) and predictive power (i.e., in terms of gene essentiality predictions) had increased over time, the coverage of the models does not always correlate with the predictive ability. Extensive models annotating higher number of genes do not necessarily have better essentiality prediction capabilities than simpler ones. Introducing additional minor activity encoding genes may decrease the predictive capacity if the encoded enzymes cannot alone sustain the corresponding reactions (Pereira et al. 2016). However, in addition to using the models for predictive simulations of genotype–phenotype translation, the genome-scale metabolic models can also be seen as knowledge bases containing all known biochemical conversion potential of the organism. Including the minor activity encoding genes and the corresponding reactions in a model are valuable for a knowledge base or a biochemical interaction network use. In conclusion, the several genome-scale metabolic models of S. cerevisiae have been developed and evolved independently for different purposes and none of them is generally the best.
Genome-scale metabolic models have been reconstructed, and manually curated, also for other yeasts than S. cerevisiae (Fig. 5.2). The models have commonly been reconstructed in a comparative manner using an S. cerevisiae model as a template. The reconstruction tool RAVEN especially supports the comparative reconstruction using an S. cerevisiae and CoReCo exploits species relatedness in scoring the reactions (Pitkänen et al. 2014). The models for industrially relevant species K. lactis, P. pastoris, S. stipitis, and Y. lipolytica, and for pathogenic C. glabrata have been derived using S. cerevisiae models as templates. For pathogenic C. tropicalis and for scientifically relevant S. pombe model reconstructions no S. cerevisiae framework has been reported. In addition, a large set of draft fungal models, including yeast models, reconstructed using CoReCo (Pitkänen et al. 2014; Castillo et al. 2016) are available in the BioModels database (Chelliah et al. 2015). In addition to the BioModels database and the developer’s specific sites, genome-scale metabolic models for various species can be downloaded from other public databases such as BiGG database (http://bigg.ucsd.edu/) (King et al. 2016).
5.3 Methods for Metabolic Phenotype Simulations Derived from Flux Balance Analysis (FBA)
A myriad of methods for performing phenotype simulations using genome-scale metabolic models derived from Flux Balance Analysis (FBA) (Varma and Palsson 1994). FBA solves a linear programming problem of optimizing biologically relevant objective function (typically growth) under metabolic steady-state mass conservation, enzyme capacity, and thermodynamic constraints. Steady-state assumption implies that the intracellular metabolite concentrations are constant (i.e., their time derivatives are zero). Thus, the steady-state assumption renders the problem linear (Fig. 5.2) and eliminates the need to describe the reaction kinetics that are functions of reactant abundances often with several unknown parameters. The steady-state assumption linearizing the problem is well justified for many metabolic states. Particularly well the steady-state assumption holds when microbial cells divide unlimited by the external conditions or grow in continuous cultivations under constant conditions. Under these conditions, FBA-optimized growth yields have been found to closely match experimental observation in microbial species (Edwards et al. 2001). Yet, other optimality principles than growth such as maximization of energy generation in terms of ATP have been suggested and evaluated (Schuetz et al. 2007). Model simulations of optimizing defined objective functions take globally into account cellular energy and redox balancing requirements when fulfilling mass balance, enzyme capacity, and thermodynamic constraints in the whole metabolic network. Enzyme capacity and thermodynamic constraints are introduced into the FBA problem as flux upper and lower bounds. Commonly, the sign of flux value describes the net flux direction of the reaction but alternatively forward and backward reactions can be separately represented in the model. When thermodynamics do not allow for a particular reaction direction under cellular conditions (Flamholz et al. 2012), the flux bounds can be assigned accordingly for simulations.
Phenotype simulations with FBA and derived tools and genome-scale metabolic model manipulations are facilitated with frameworks supporting method development and/or tools with higher level interfaces for analysis (Table 5.1). While Python-based frameworks, relying on COBRApy (Ebrahim et al. 2013), are currently the primary choice of developers, there are R (R Development Core Team 2018) (Sybil (Gelius-Dietrich 2013)) and MATLAB (www.mathworks.com) (COBRA toolbox, (Schellenberger et al. 2011; Heirendt et al. 2017)) based frameworks available as well. The frameworks and tools commonly offer interfaces to external LP (and commonly also Mixed-Integer Linear Programming (MILP) and Quadratic Programming (QP)) solvers (e.g., glpk (www.gnu.org/software/glpk/), cplex (www.ibm.com/analytics/cplex-optimizer), gurobi (www.gurobi.com)) to be recruited for different applications. External libraries may also be engaged by the tools, in particular, for manipulating models in common Systems Biology Markup Language (SBML) format (Hucka et al. 2003) (SBML toolbox (Keating et al. 2006), libSBML (Bornstein et al. 2008)). Tools with higher level interfaces allow also experimental scientists analyzing metabolism with genome-scale models and designing genotype manipulations, as will be reviewed below.
Genome-scale metabolic model simulations using FBA with alternative, other than biological design principle mimicking, objectives can be used to explore an organism’s metabolic potential, possible metabolic states it may have. For instance, under the given mass balance, enzyme capacity, and thermodynamic constraints, the optimal theoretical yields of biotechnologically relevant molecules can be solved with simulations. The simulations can be done by assigning alternative nutritional conditions mimicking different growth media or bioconversion substrates. In case substrate utilization rates are available, they can be introduced to the models as exchange fluxes between cells and the environment, and FBA can be used to predict optimal steady-state growth (1/h) and specific production rates (mmol/(g cell dry weight * h)) instead of yields. While the optimal value solved for the chosen objective by FBA (i.e., yield or rate) is global and unique, the other fluxes (i.e., variables of the optimization problem) may adopt different values under optimality. Thus, there may be several, alternative, yet equally optimal metabolic phenotypes in terms of the defined objective function.
5.3.1 Parsimonious Flux Balance Analysis (pFBA)
Parsimonious Flux Balance Analysis (pFBA) aims at reducing the set of alternative equally optimal flux states in a biologically relevant way (Lewis et al. 2010). pFBA derives from FBA and includes a bi-level optimization where first the biological design objective (e.g., growth) is optimized after which, under the optimality condition, another linear programming problem is solved to minimize the sum of the fluxes. The flux-sum minimization in pFBA can be seen biologically relevant in optimizing the enzyme usage, and thereby the cellular resource utilization. Flux-sum minimization efficiently omits futile flux cycle artifacts from the returned flux vector. Yet, fluxes may adopt alternative values also under pFBA optimality.
5.3.2 Flux Variability Analysis (FVA)
The ranges of possible values fluxes may adopt under particular optimality can be assessed with Flux Variability Analysis (FVA) (Burgard and Maranas 2001; Mahadevan and Schilling 2003). FVA can be performed under the optimality of the assigned objective (i.e., commonly growth) or different levels of it. The computation involves solving two subsequent linear programming problems, minimization and maximization, for each of the fluxes. The fluxes whose ranges do not pass zero are coupled to the objective and can thus be considered essential for the particular objective. General analysis of flux coupling in a metabolic network is derived from FVA (Burgard et al. 2004).
5.3.3 Simulating Mutant Cell Phenotypes
The above FBA-derived simulation approaches assume optimal distribution of flux in the metabolic network. In case of FBA simulation with an objective function mimicking biological optimality principle, the premise is justified by evolutionary optimization of organism’s metabolism (Ibarra et al. 2002). However, mutant strains engineered in laboratory cannot be assumed to function optimally. Minimization of Metabolic Adjustment (MoMA) approach was developed to simulate the metabolic state of such engineered mutant strains (Segrè et al. 2002). MoMA solves a quadratic optimization problem of minimizing the flux differences to a reference flux state (i.e., wild-type flux state) given the constraints arising from the engineered modifications to the strain (e.g., gene deletions). There is also a linearized version, linear Minimization of Metabolic Adjustment (lMoMA) of the algorithm (Burgard et al. 2003; Becker et al. 2007). In biological sense MoMA and lMoMA assume that the wild-type regulation is still driving the distribution of metabolic fluxes in engineered but not evolutionarily streamlined strains. Wild-type regulation-driven flux distribution in engineered cells is also simulated with Minimization of Metabolites Balance (MiMBl) algorithm (Brochado et al. 2012). In contrast to MoMA and lMoMA, MiMBl is independent of the stoichiometric representation of the reactions. While multiplicating the stoichiometric coefficients of particular reaction(s) (which does not affect the reaction stoichiometry or elemental balance) would alter the output of MoMA computation, MiMBl solution would be unaffected. MiMBl computation minimizes the flux distribution difference to the wild-type state in terms of metabolite turnovers instead of fluxes. Yet another approach for simulating the metabolic state of engineered, but not evolved organisms is Regulatory On/Off Minimization (ROOM) algorithm (Shlomi et al. 2005). ROOM minimizes the number of fluxes that are changed in mutant cells compared to wild-type cells. The underlying premise in ROOM is the same as in MoMA, lMoMA, and MiMBl in assuming that the wild-type regulation drives the distribution of fluxes in a non-evolved mutant strain. In ROOM simulations, it is further assumed that the mutant metabolic state is reached through only the necessary transient metabolic changes mediated by the regulatory network. The necessary changes are simulated with ROOM by solving a Mixed-Integer Linear Programming (MILP) problem.
5.4 Examples of Genotype–Phenotype Simulations: Single and Double Gene KOs
The above-introduced simulation tools using genome-scale metabolic models allow predicting phenotype effects following from gene deletions (Förster et al. 2003). In silico metabolic gene deletions are propagated through the Boolean gene-reaction rules into reaction activities. If a regulatory model is integrated as in rFBA approach (Covert et al. 2001; Herrgård et al. 2006), the regulatory gene deletions can be first propagated to the status of metabolic genes through the regulatory Boolean rules, and then through the metabolic model’s gene-reaction rules into reaction activity states. The phenotype simulation is then performed with updated reaction activity states. FBA or another simulation algorithm, not assuming the metabolism in mutant could necessarily become optimized, can be used. In case the simulated growth is negligible, the deleted gene is predicted essential. Double gene deletion simulations predict in silico synthetic lethal gene pairs (Suthers et al. 2009). Since experimental screens of gene deletion mutants in model organisms are available in genome-scale, comparison to in silico model predicted essentialities and synthetic lethalities can be used for validating metabolic model reconstruction algorithms.
5.5 In Silico Metabolic Engineering—Strain Design
Since the genome-scale metabolic models allow predicting translation of genotype to phenotype, they can be used to design genotype manipulations leading to desired phenotypes. Overproducer phenotypes are especially sought for industrial biotechnology applications. While native strains are evolved to distribute the available resources for growth and survival, feasible industrial production using a microbial fermentation process requires cells to divert substantial resources to product synthesis. Diverting cellular resources toward production is the aim of metabolic engineering of the industrial biotechnology host organisms, like yeasts, in addition to introducing the production pathways in case of heterologous products. Strategies to achieve the desired metabolic flux re-regulation diverting resources efficiently to the production pathway can be computationally designed using genome-scale metabolic models. An elegant solution for the inherent competition of growth and product synthesis for resources is to align those objectives through metabolic network modifications. Aligning the growth and production objectives in cells can be achieved with specific metabolic gene deletions resulting in growth-coupled production. The specific metabolic gene deletions reduce the metabolic network in such a way that the cells cannot grow (optimally or at all) unless they simultaneously synthesize the product. In other words, some growth essential pathway produces the desired product as an unavoidable side stream. OptKnock was the pioneering method for finding growth–product coupling creating deletion targets using metabolic models (Burgard et al. 2003). It was implemented as a bi-level MILP. An alternative implementation of in silico growth–product coupling design is OptGene in which the phenotype simulation is embedded in a genetic algorithm allowing for nonlinear design objectives and searching larger target gene sets (Patil et al. 2005; Asadollahi et al. 2009). OptGene has been used successfully to design, for example, succinate and terpenoid overproducing S. cerevisiae strains (Otero et al. 2013; Asadollahi et al. 2009). For vanillin production in S. cerevisiae (in form of vanillin glycoside to reduce toxicity), OptGene was used to identify deletion targets out of which GDH1 (glutamate dehydrogenase encoding) and PDC1 (pyruvate decarboxylase encoding) deletions were experimentally implemented and evaluated (Brochado et al. 2010). Single deletion mutants, a double deletion mutant, and a double deletion mutant with GDH2 overexpression to improve nitrogen assimilation defect in gdh1\(\Delta \) were constructed. The mutant strains except single gdh1\(\Delta \) mutant showed 1.5 fold increase in vanillin glucoside yield in batch cultures compared to the non-host metabolism optimized strain. Furthermore, optimizing the synthetic, four-step, production pathway of vanillin glucoside in S. cerevisiae did not improve the production, before the OptGene identified targets to optimize the host metabolism were implemented (Brochado et al. 2010; Brochado and Patil 2013). Later, Tepper and Shlomi (2010) released their RobustKnock version for extracting such growth–product coupling creating deletions that force product synthesis with an additional optimization step (Tepper and Shlomi 2010). Growth–product coupling creating manipulations to genome fix the relative yields of biomass and target product. However, the rates are amenable for improvement through Adaptive Laboratory Evolution (ALE) of the mutant strains. While faster growing cells are selected for, the coupled production rate is improved on the side (Otero et al. 2013). If the growth–product coupling relies on a carbon–carbon bond cleaving reaction splitting a precursor for growth and production, the coupling is likely to be very robust in ALE. An Anchor reaction producing an essential precursor for growth and another product convertible to the target product is biochemically essential for a growth–product coupled reduced metabolic network (Jouhten et al. 2017). Carbon–carbon bond cleaving Anchor reactions are a subset of all possible Anchors. Growth-coupled succinate production in S. cerevisiae relies on carbon–carbon bond cleaving isocitrate lyase as an Anchor reaction (Otero et al. 2013). The initial production rate after the metabolic network reduction for growth–product coupling was substantially improved with ALE along with relieving glycine auxotrophy (Table 5.2).
Metabolic network manipulations for achieving growth–product coupling are identifiable also with elementary-mode analysis methods (Schuster and Hilgetag 1994; Schuster et al. 2000; Trinh and Srienc 2009; Unrean et al. 2010; Hädicke and Klamt 2011). Elementary modes are minimal sets of reactions allowing a steady-state operation (Heinrich and Schuster 1998). Engineering strategies are designed for disabling undesired elementary modes while retaining the desired ones (Hädicke and Klamt 2011). Introducing flux capacity constraints to the elementary-mode framework, as in FBA-derived methods, is enabled using Elementary Flux Vectors (EFVs) allowing also designing growth–product coupling strategies (Urbanczik 2007; Klamt and Mahadevan 2015). The scalability of searching metabolic engineering strategies in silico using elementary-modes-based approaches has been limited but is improving through algorithmic developments (von Kamp and Klamt 2014). Currently, minimum sets of genetic engineering targets can be exhaustively identified enabling evaluations also in yeast hosts. Beyond identifying growth–product coupling strategies, genome-scale metabolic models allow designing also other kinds of engineering strategies for improving production. While the methods for designing strategies to optimize the cellular fluxes for production are broadly reviewed elsewhere (e.g., Maia et al. (2016)) many of them are yet to be evaluated for yeasts. Among the variety of approaches, there are methods for identifying not only knockouts but also up- and downregulation targets for improving production. OptReg identifies combined strategies of deletions, overexpressions, and downregulations for host optimization as bi-level MILP solutions (Pharkya and Maranas 2006). Similarly, OptForce identifies combined strategies in a comparative manner against the wild-type flux status by classifying reactions based on the type of manipulation they require for optimizing production (Ranganathan et al. 2010). Flux Scanning based on Enforced Objective Flux (FSEOF) considers the wild-type flux status by identifying upregulation engineering targets as genes annotated to reactions whose flux is increased in silico when the production objective is enforced while biological objective (i.e., growth) prevails (Choi et al. 2010). FSEOF-identified targets have successfully been implemented in P. pastoris yeast for improving protein production (Nocon et al. 2014). The strain improvement strategies may also benefit from augmenting metabolic models with additional information on metabolic enzymes or wild-type phenotype. For instance, k-OptForce integrates available enzyme kinetic information to improve predictions by considering metabolite concentration effects on the distribution of fluxes (Chowdhury et al. 2014). OptFlux allows using gene expression data for using a comparative approach against the wild type for identifying overexpression and downregulation targets in a metaheuristic optimization framework (Gonçalves et al. 2012). Importantly, considering the wild-type gene expression data allows relieving the optimality assumption from the native operation of cells allowing a comparative strain design also in secondary metabolic pathways (Kim et al. 2016). Accordingly, transcriptomics-based Strain Optimization Tool (tSOT) identifies the metabolic engineering targets by considering the wild-type flux regulatory status inferred from gene expression data (Kim et al. 2016). However, a word of caution though, the gene expression status of central metabolic enzymes may not very well reflect the actual flux status in yeast cells as (Machado and Herrgård 2014) observed when integrating gene expression data to genome-scale metabolic models.
5.6 Integrating Omics Data into Models
Genome-scale metabolic models offer frameworks for integrating omics data since they connect metabolic genes/proteins to reaction fluxes through which biochemical conversion of metabolites occurs. Fluxes together with metabolite abundances are the metabolic phenotype determined by and reciprocally regulating the underlying transcriptional and translational states in a cell. Evolutionarily shaped cellular regulation can vary the metabolic phenotypes within the ultimate limits of the laws of mass conservation and chemical thermodynamics. Therefore, transcriptomics, proteomics, or metabolomics data have been integrated to the models for shrinking the space of feasible metabolic states to improve flux estimation outcomes. Indeed, flux predictions would often benefit from specific constraints representing the regulation of the metabolic network utilization under particular conditions (e.g., repression of respiration in S. cerevisiae on high glucose). Several methods have been developed for inferring the flux states from gene expression data, the most abundantly available omics data type. iMAT (Shlomi et al. 2008), GiMME (Becker and Palsson 2008), GIM3E (Schmidt et al. 2013), RELATCH (Kim and Reed 2012), and INIT (Agren et al. 2012) methods derive expected or allowable flux states from the gene expression data. However, flux estimation could also be misled by gene expression data (Machado and Herrgård 2014) as post-transcriptional regulation of metabolic phenotypes is prevalent. Consequently, additional constraints derived from proteomics data integrated with enzyme-specific turnover numbers (kcat) (Sanchez et al. 2017; Vazquez and Oltvai 2016) have allowed reproducing, using model simulations, metabolic phenotypes (e.g., overflow metabolism) that are not well captured with plain FBA or apparent in gene expression data. Further, time derivatives of extracellular metabolites in a cell culture (i.e., rates of consumption and production) can readily be integrated into the models as bounds on exchange fluxes between cells and environment, allowing simulations of consistent intracellular flux states (Mo et al. 2009). However, while the exchange flux, gene expression, and proteomics data derived constraints can directly be assigned to the fluxes in models, integration of intracellular metabolite abundance data to steady-state simulations is less straightforward. Metabolite concentrations can be used to refine reaction thermodynamics for resolving feasible reaction directions (Henry et al. 2007; Kümmel et al. 2006). Further, constraints for flux changes have been derived from relative metabolomics data through the connectivity of metabolites with several reactions in the metabolic network (Sajitz-Hermstein et al. 2016). Vice versa, metabolite concentration changes can be predicted using gene expression data and the network neighborhood (Zelezniak et al. 2014). When the metabolite concentration change prediction from gene expression data and network connectivity fails, the particular metabolite is likely to be connected to a post-transcriptionally regulated enzyme (Zelezniak et al. 2014). Likely post-transcriptionally regulated enzymes can similarly be identified in disagreements of gene expression data and flux estimates (Shlomi et al. 2008). Thus, omics data integration with model simulations allows also uncovering how the cells have achieved the observed metabolic phenotypes. Recently, (Strucko et al. 2018) uncovered in molecular detail how S. cerevisiae achieved an efficiently glycerol-utilizing phenotype through Adaptive Laboratory Evolution (ALE). Classical genetic crossing, genome-scale metabolic model simulations, whole genome sequencing, and omics analyses revealed involvement of all levels of cellular regulation, in a pathway-dependent manner, in achieving the glycerol utilization trait. The ALE for glycerol utilization was performed for a laboratory strain of S. cerevisiae, commonly lacking the ability to grow on glycerol in absence of amino acid supplementation. Interestingly, some wild S. cerevisiae strains can grow on glycerol as the sole carbon source, and the metabolic network structure of S. cerevisiae does not object the conversion of glycerol to biomass even without amino acids being provided. By gradually decreasing the amino acid supplementation, evolved lineages growing on glycerol as the sole carbon source were obtained (Strucko et al. 2018). Whole genome sequencing of evolved lineages revealed mutations that arose during the ALE. Few metabolic genes and genes involving osmoregulation controlling glycerol accumulation in cells had been repeatedly hit by mutations. A lineage not having loss-of-function mutations in osmoregulation involved genes was characterized in controlled bioreactors and analyzed on different omics levels (i.e., RNA sequencing, proteomics, and metabolomics). Further, genome-scale metabolic model simulations were run for identifying the necessary but minimum re-regulation of wild-type metabolic fluxes for achieving an optimally glycerol-utilizing phenotype. The identified necessary flux changes were overlaid with the mutated genes and the omics data on the metabolic network. The model simulations had revealed a necessary downregulation of TCA cycle activity while maintaining respiratory function for glycerol utilization which was in perfect concordance with the otherwise obscure KGD1 (encoding alpha-ketoglutarate dehydrogenase in the TCA cycle) loss-of-function mutation gained repeatedly in ALE. Further, the model simulations predicted also an activation of GABA shunt bypass of the TCA cycle for optimizing glycerol utilization. Indeed, reactant ratios from metabolomics data were in agreement with the GABA shunt activation. In addition, gene/protein expression changes were in agreement with the model simulated prediction of decreased TCA cycle flux. In conclusion, the flux change predictions with model simulations effectively reconciliated the separate observations in omics data and the genes repeatedly mutated in ALE.
5.7 Regulation of Yeast Metabolism: Key Nodes and Their Impact on Flux Distribution—Future Directions of Reincorporating These into Models
While metabolic models have greatly improved our ability to systematically map genotype–phenotype relations, they have also brought forward key gaps in the understanding of the complex interactions between different metabolic pathways and between metabolic and regulatory processes. This becomes evident when considering the dramatically reduced performance of genome-scale metabolic models from well predicting the essentiality of single genes to the low accuracy in predicting genetic interactions (Brochado et al. 2012). A major limitation of the models, especially when tackling higher order complex interactions, is the large degrees of freedom, i.e., multiple ways that the resource (carbon and other elemental) fluxes can be distributed in the cell. Without considering additional constraints imposed by protein abundance and activity status (e.g., phosphorylation), metabolite concentrations, and allosteric regulations, the models will not be able to narrow down the predictions on the actual routes operating in cells. Different approaches have been proposed toward constraining the solution space of metabolic models for improving the accuracy of predictions in a biologically sound manner. These include knowledge-based heuristics imposing constraints on flux distribution at key branch points (Pereira et al. 2016), constraining the fraction of protein resources allocated to metabolic processes (Sanchez et al. 2017), imposing a constraint on maximum Gibbs energy dissipation from cells (Niebel et al. 2019), and large-scale kinetic models that include metabolite concentrations and enzyme kinetic parameters (Chakrabarti et al. 2013; Stanford et al. 2013; Smallbone et al. 2010). The last mentioned would be an ideal approach encompassing various complexities in their mechanistic detail. Yet, the lack of reliable in vivo data on enzyme kinetics, metabolite concentrations, and enzyme/metabolite distributions within a cell limit the use of kinetic modeling to well-studied conditions and relatively small perturbations. Further, introducing a constraint on Gibbs energy dissipation to the metabolic models is computationally demanding as it results into nonlinear and non-convex model. Thus, the first two approaches are likely to be the most fruitful in the near future. Indeed, the distribution of major metabolic fluxes in yeast cells are tied to the redox and energy cofactor balance, which, in turn, are closely coupled with the flux distribution in pentose phosphate pathway and pyruvate nodes. The former largely determines the NADPH production and the latter affects NADH and ATP turnover. Indeed, a recent study (Yu et al. 2018) elegantly demonstrates this by replacing ethanol production by fatty acid production. Given that ethanol accumulation is a hallmark of yeast metabolism, this is a remarkable feat and yet can be understood in terms of redox balance rewiring. Along similar lines, an approach considering protein allocation constraint has suggested that lower protein requirement of ATP generation through fermentation is the trade-off factor underlying the switch from respirative to fermentative metabolism at higher glucose utilization rates in yeast (Nilsson and Nielsen 2016). The ongoing efforts in expanding the models to incorporate transcriptional and translational processes (Yang et al. 2018) are likely to complement the abovementioned approaches in expanding the scope of metabolic models as well as in improving their accuracy which is capturing complex metabolic traits.
References
Acevedo A, Conejeros R, Aroca G (2017) Ethanol production improvement driven by genome-scale metabolic modeling and sensitivity analysis in Scheffersomyces stipitis. Plos One 12(6):e0180074. https://doi.org/10.1371/journal.pone.0180074
Agren R, Bordel S, Mardinoglu A, Pornputtapong N, Nookaew I, Nielsen J (2012) Reconstruction of genome-scale active metabolic networks for 69 human cell types and 16 cancer types using INIT. PLoS Comput Biol 8(5):e1002518. https://doi.org/10.1371/journal.pcbi.1002518
Agren R, Liu L, Shoaie S, Vongsangnak W, Nookaew I, Nielsen J (2013) The RAVEN toolbox and its use for generating a genome-scale metabolic model for Penicillium chrysogenum. PLoS Comput Biol 9(3):e1002980. https://doi.org/10.1371/journal.pcbi.1002980
Asadollahi MA, Maury J, Patil KR, Schalk M, Clark A, Nielsen J (2009) Enhancing sesquiterpene production in Saccharomyces cerevisiae through in silico driven metabolic engineering. Metab Eng 11(6):328–334. https://doi.org/10.1016/j.ymben.2009.07.001
Aung HW, Henry SA, Walker LP (2018) SysBioChalmers/yeast-GEM: the consensus gem for Saccharomyces cerevisiae. https://github.com/SysBioChalmers/yeast-GEM
Becker SA, Feist AM, Mo ML, Hannum G, Palsson BØ, Herrgard MJ (2007) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox. Nat Protoc 2(3):727–738
Becker SA, Palsson BO (2008) Context-specific metabolic networks are consistent with experiments. PLoS Comput Biol 16 4(5):e1000082. https://doi.org/10.1371/journal.pcbi.1000082
Biggs MB, Papin JA (2017) Managing uncertainty in metabolic network structure and improving predictions using EnsembleFBA. PLoS Comput Biol 13(3):e1005413. https://doi.org/10.1371/journal.pcbi.1005413
Bornstein BJ, Keating SM, Jouraku A, Hucka M (2008) LibSBML: an api library for SBML. Bioinformatics 24(6):880–881. https://doi.org/10.1093/bioinformatics/btn051
Borodina I, Kildegaard KR, Jensen NB, Blicher TH, Maury J, Sherstyk S, et al (2015) Establishing a synthetic pathway for high-level production of 3-hydroxypropionic acid in Saccharomyces cerevisiae via \(\beta \)-alanine. Metab Eng 27:57–64. https://www.sciencedirect.com/science/article/pii/S1096717614001256
Brochado AR, Andrejev S, Maranas CD, Patil KR (2012) Impact of stoichiometry representation on simulation of genotype-phenotype relationships in metabolic networks. PLoS Comput Biol 8(11):e1002758. https://doi.org/10.1371/journal.pcbi.1002758
Brochado AR, Matos C, Møller BL, Hansen J, Mortensen UH, Patil KR (2010) Improved vanillin production in baker’s yeast through in silico design. Microb Cell Factories 9:84. https://doi.org/10.1186/1475-2859-9-84
Brochado AR, Patil KR (2013) Overexpression of O-methyltransferase leads to improved vanillin production in baker’s yeast only when complemented with model-guided network engineering. Biotechnol Bioeng 110(2):656–659. https://doi.org/10.1002/bit.24731
Bro C, Regenberg B, Förster J, Nielsen J (2006) In silico aided metabolic engineering of Saccharomyces cerevisiae for improved bioethanol production. Metab Eng 8(2):102–111. https://www.sciencedirect.com/science/article/pii/S1096717605000789
Burgard AP, Maranas CD (2001) Probing the performance limits of the Escherichia coli metabolic network subject to gene additions or deletions. Biotech Bioeng 74(5):364–37. https://doi.org/10.1002/bit.1127
Burgard AP, Nikolaev EV, Schilling CH, Maranas CD (2004) Flux coupling analysis of genome-scale metabolic network reconstructions. Genome Res 14(2):301–312
Burgard AP, Pharkya P, Maranas CD (2003) OptKnock: a bilevel programming framework for identifying gene knockout strategies for microbial strain optimization. Biotech Bioeng 84(6):647–657
Cardenas J, Da Silva NA (2014) Metabolic engineering of Saccharomyces cerevisiae for the production of triacetic acid lactone. Metab Eng 25:194–203. https://www.sciencedirect.com/science/article/pii/S1096717614000998
Cardoso JGR, Jensen K, Lieven C, Hansen ASL, Galkina S, Beber M et al (2018) Cameo: a python library for computer aided metabolic engineering and optimization of cell factories. ACS Synth Biol 7(4):1163–1166. https://doi.org/10.1021/acssynbio.7b00423
Caspi R, Altman T, Billington R, Dreher K, Foerster H, Fulcher CA, et al (2014) The MetaCyc database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Research. 2014 1;42(D1):D459–D471. https://doi.org/10.1093/nar/gkt1103
Castañeda MT, Nuñez S, Garelli F, Voget C, De Battista H (2018) Comprehensive analysis of a metabolic model for lipid production in Rhodosporidium toruloides. J Biotechnol 280:11–18. https://www.sciencedirect.com/science/article/pii/S0168165618301536
Castillo S, Barth D, Arvas M, Pakula TM, Pitkänen E, Blomberg P, et al (2016) Whole-genome metabolic model of Trichoderma reesei built by comparative reconstruction. Biotechnology for biofuels 9:252. http://www.ncbi.nlm.nih.gov/pubmed/27895706http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC5117618
Cautha SC, Gowen CM, Lussier FX, Gold ND, Martin VJJ, Mahadevan R (2013) Model-driven design of a Saccharomyces cerevisiae platform strain with improved tyrosine production capabilities. In: IFAC Proceedings, vol 46 no 31, pp 221–226. https://www.sciencedirect.com/science/article/pii/S1474667016313982
Chakrabarti A, Miskovic L, Soh KC, Hatzimanikatis V (2013) Towards kinetic modeling of genome-scale metabolic networks without sacrificing stoichiometric, thermodynamic and physiological constraints. Biotechnol J 8(9):1043–1057. https://doi.org/10.1002/biot.201300091
Chelliah V, Juty N, Ajmera I, Ali R, Dumousseau M, Glont M, et al (2015) BioModels: ten-year anniversary. Nucleic Acids Res 43(D1):D542–D548. http://academic.oup.com/nar/article/43/D1/D542/2439069/BioModels-tenyear-anniversary
Chen X, Xu G, Xu N, Zou W, Zhu P, Liu L, et al (2013) Metabolic engineering of Torulopsis glabrata for malate production. Metab Eng 19:10–16. https://www.sciencedirect.com/science/article/pii/S1096717613000505
Choi HS, Lee SY, Kim TY, Woo HM (2010) In silico identification of gene amplification targets for improvement of lycopene production. Appl Environ Microbiol 76(10):3097–3105. https://doi.org/10.1128/AEM.00115-10
Chowdhury A, Zomorrodi AR, Maranas CD (2014) k-OptForce: integrating kinetics with flux balance analysis for strain design. PLoS Comput Biol 10(2):e1003487. https://doi.org/10.1371/journal.pcbi.1003487
Covert MW, Schilling CH, Palsson B (2001) Regulation of gene expression in flux balance models of metabolism. J Theor Biol 213(1):73–88. https://www.sciencedirect.com/science/article/pii/S0022519301924051
Curran KA, Leavitt JM, Karim AS, Alper HS (2013) Metabolic engineering of muconic acid production in Saccharomyces cerevisiae. Metab Eng 15:55–66. https://www.sciencedirect.com/science/article/pii/S1096717612001139
Cvijovic M, Olivares-Hernandez R, Agren R, Dahr N, Vongsangnak W, Nookaew I et al (2010) BioMet toolbox: genome-wide analysis of metabolism. Nucleic Acids Res 38(Web Server issue):W144–W149. https://doi.org/10.1093/nar/gkq404
Dias O, Rocha M, Ferreira EC, Rocha I (2015) Reconstructing genome-scale metabolic models with merlin. Nucleic Acids Res 43(8):3899–910. http://www.ncbi.nlm.nih.gov/pubmed/25845595. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4417185
Ebrahim A, Lerman JA, Palsson BO, Hyduke DR (2013) COBRApy: constraints-based reconstruction and analysis for python. BMC Syst Biol 7:74. https://doi.org/10.1186/1752-0509-7-74
Edwards JS, Ibarra RU, Palsson BO (2001) In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat Biotech 19(2):125–130
Fabregat A, Jupe S, Matthews L, Sidiropoulos K, Gillespie M, Garapati P, et al (2018) The reactome pathway knowledgebase. Nucleic Acids Res 46(Database issue):D649. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5753187/
Feng J, Yang J, Li X, Guo M, Wang B, Yang St, et al (2017) Reconstruction of a genome-scale metabolic model and in silico analysis of the polymalic acid producer Aureobasidium pullulans CCTCC M2012223. Gene 607:1–8. https://www.sciencedirect.com/science/article/pii/S0378111916310459
Flamholz A, Noor E, Bar-Even A, Milo R (2012) EQuilibrator—The biochemical thermodynamics calculator. Nucleic Acids Res 40(Database issue):D770–D775. https://doi.org/10.1093/nar/gkr874
Förster J, Famili I, Fu P, Palsson BØ, Nielsen J (2003) Genome-scale reconstruction of the Saccharomyces cerevisiae metabolic network. Genome Res 3(2):244–53. http://www.ncbi.nlm.nih.gov/pubmed/12566402. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC420374
Förster J, Famili I, Palsson BO, Nielsen J (2003) Large-scale evaluation of in silico gene deletions in Saccharomyces cerevisiae. Omics J Integr Biol 7(2):193–202
Garcia-Albornoz M, Thankaswamy-Kosalai S, Nilsson A, Väremo L, Nookaew I, Nielsen J (2014) BioMet Toolbox 2.0: Genome-wide analysis of metabolism and omics data. Nucleic Acids Res 42(Web Server issue):W175–W181. https://doi.org/10.1093/nar/gku371
Gelius-Dietrich G (2013) sybil—efficient constrained based modelling in r. bmc systems biology
Gold ND, Gowen CM, Lussier FX, Cautha SC, Mahadevan R, Martin VJJ (2015) Metabolic engineering of a tyrosine-overproducing yeast platform using targeted metabolomics. Microb Cell fact 14:73. http://www.ncbi.nlm.nih.gov/pubmed/26016674. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4458059
Gonçalves E, Pereira R, Rocha I, Rocha M (2012) Optimization approaches for the in silico discovery of optimal targets for gene over/underexpression. J Comput Biol 19(2):102–114. https://doi.org/10.1089/cmb.2011.0265
Gruchattka E, Kayser O (2015) In vivo validation of in silico predicted metabolic engineering strategies in yeast: disruption of alpha-ketoglutarate dehydrogenase and expression of atp-citrate lyase for terpenoid production. PLOS One. 10(12):e0144981. https://doi.org/10.1371/journal.pone.0144981
Hädicke O, Klamt S (2011) Computing complex metabolic intervention strategies using constrained minimal cut sets. Metab Eng 13(2):204–213. https://doi.org/10.1016/j.ymben.2010.12.004
Heavner BD, Price ND (2015) Comparative analysis of yeast metabolic network models highlights progress, opportunities for metabolic reconstruction. PLOS Comput Biol 11(11):e1004530. https://doi.org/10.1371/journal.pcbi.1004530
Heinrich R, Schuster S (1998) The modelling of metabolic systems. Structure, control and optimality. BioSystems 47(1–2):61–77
Heirendt L, Arreckx S, Pfau T, Mendoza SN, Richelle A, Heinken A, et al (2017) Creation and analysis of biochemical constraint-based models: the COBRA Toolbox v3.0. arXiv:1710.04038
Henry CS, Broadbelt LJ, Hatzimanikatis V (2007) Thermodynamics-based metabolic flux analysis. Biophys J 92(5):1792–1805
Herrgård MJ, Lee BS, Portnoy V, Palsson BØ (2006) Integrated analysis of regulatory and metabolic networks reveals novel regulatory mechanisms in Saccharomyces cerevisiae. Genome Res 16(5):627–635
Herrgård MJ, Swainston N, Dobson P, Dunn WB, Arga KY, Arvas M, et al. (2008) A consensus yeast metabolic network reconstruction obtained from a community approach to systems biology. Nat Biotechnol 26(10):1155–1160
Hucka M, Finney A, Sauro HM, Bolouri H, Doyle JC, Kitano H et al (2003) The systems biology markup language (SBML): a medium for representation and exchange of biochemical network models. Bioinformatics 19(4):524–531
Ibarra RU, Edwards JS, Palsson BO (2002) Escherichia coli K-12 undergoes adaptive evolution to achieve in silico predicted optimal growth. Nature 420(6912):186–189
Johannsen W (1911) The genotype conception of heredity. Am Nat. 45(531):129–159. http://www.jstor.org/stable/2455747
Jouhten P, Huerta-Cepas J, Bork P, Patil KR (2017) Metabolic anchor reactions for robust biorefining. Metab Eng 40:1–4. https://doi.org/10.1016/j.ymben.2017.02.010
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45(D1):D353–D361. https://doi.org/10.1093/nar/gkw1092
Kavscek M, Bhutada G, Madl T, Natter K (2015) Optimization of lipid production with a genome-scale model of Yarrowia lipolytica. BMC Syst Biol 9:72. http://www.ncbi.nlm.nih.gov/pubmed/26503450. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4623914
Keating SM, Bornstein BJ, Finney A, Hucka M (2006) SBMLToolbox: an SBML toolbox for MATLAB users. Bioinformatics 22(10):1275–1277
Kildegaard KR, Jensen NB, Schneider K, Czarnotta E, Özdemir E, Klein T, et al (2016) Engineering and systems-level analysis of Saccharomyces cerevisiae for production of 3-hydroxypropionic acid via malonyl-CoA reductase-dependent pathway. Microb Cell Fact 15(1):53. http://www.microbialcellfactories.com/content/15/1/53
Kim J, Reed JL (2012) RELATCH: relative optimality in metabolic networks explains robust metabolic and regulatory responses to perturbations. Genome Biol 13(9):R78. https://doi.org/10.1186/gb-2012-13-9-r78
Kim M, Yi JS, Lakshmanan M, Lee DY, Kim BG (2016) Transcriptomics-based strain optimization tool for designing secondary metabolite overproducing strains of Streptomyces coelicolor. Biotech Bioeng 113(3):651–660. https://doi.org/10.1002/bit.25830
King ZA, Lu J, Dräger A, Miller P, Federowicz S, Lerman JA et al (2016) BiGG models: a platform for integrating, standardizing and sharing genome-scale models. Nucleic Acids Res 44(D1):D515–D522. https://doi.org/10.1093/nar/gkv1049
Klamt S, Mahadevan R (2015) On the feasibility of growth-coupled product synthesis in microbial strains. Metab Eng 30:166–178. https://doi.org/10.1016/j.ymben.2015.05.006
Klamt S, Saez-Rodriguez J, Gilles ED (2007) Structural and functional analysis of cellular networks with CellNetAnalyzer. BMC Syst Biol 1:2
Klamt S, von Kamp A (2011) An application programming interface for CellNetAnalyzer. BioSystems 105(2):162–168. https://doi.org/10.1016/j.biosystems.2011.02.002
Koivuranta K, Castillo S, Jouhten P, Ruohonen L, Penttila M, Wiebe MG (2018) Enhanced triacylglycerol production with genetically modified trichosporon oleaginosus. Front Microbiol 9:1337. https://doi.org/10.3389/fmicb.2018.01337/full
Kümmel A, Panke S, Heinemann M (2006) Putative regulatory sites unraveled by network-embedded thermodynamic analysis of metabolome data. Mol Syst Biol 2:2006.0034
Latendresse M, Krummenacker M, Trupp M, Karp PD (2012) Construction and completion of flux balance models from pathway databases. Bioinformatics 28(3):388–396. https://doi.org/10.1093/bioinformatics/btr681
Lewis NE, Hixson KK, Conrad TM, Lerman JA, Charusanti P, Polpitiya AD, et al (2007) Omic data from evolved E. coli are consistent with computed optimal growth from genome-scale models. Mol Syst Biol 6(1):390. http://www.ncbi.nlm.nih.gov/pubmed/20664636. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC2925526
Li S, Gao X, Xu N, Liu L, Chen J (2014) Enhancement of acetoin production in Candida glabrata by in silico-aided metabolic engineering. Microb Cell Fact 13(1):55. Available from: https://doi.org/10.1186/1475-2859-13-55
Lopes H, Rocha I (2017) Genome-scale modeling of yeast: chronology, applications and critical perspectives. FEMS Yeast Res 17(5). https://doi.org/10.1093/femsyr/fox050/3950252
Machado D, Andrejev S, Tramontano M, Patil KR (2018) Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res 46(15):7542–7553. https://doi.org/10.1093/nar/gky537
Machado D, Herrgård M (2014) Systematic evaluation of methods for integration of transcriptomic data into constraint-based models of metabolism. PLoS Comput Biol 10(4):e1003580. https://doi.org/10.1371/journal.pcbi.1003580
Mahadevan R, Schilling CH (2003) The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metab Eng 5(4):264–276. https://www.sciencedirect.com/science/article/pii/S1096717603000582
Maia P, Rocha M, Rocha I (2016) In silico constraint-based strain optimization methods: the quest for optimal cell factories. Microbiol Mol Biol Rev MMBR 80(1):45–67. http://www.ncbi.nlm.nih.gov/pubmed/26609052. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC4711187
Meadows AL, Hawkins KM, Tsegaye Y, Antipov E, Kim Y, Raetz L, et al (2016) Rewriting yeast central carbon metabolism for industrial isoprenoid production. Nature 537(7622):694–697. http://www.nature.com/articles/nature19769
Misra A, Conway MF, Johnnie J, Qureshi TM, Lige B, Derrick AM, et al (2013) Metabolic analyses elucidate non-trivial gene targets for amplifying dihydroartemisinic acid production in yeast. Front Microbiol 200. http://www.ncbi.nlm.nih.gov/pubmed/23898325. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC3724057
Mo ML, Palsson B, Herrgård MJ (2009) Connecting extracellular metabolomic measurements to intracellular flux states in yeast. BMC Syst Biol 3:37. https://doi.org/10.1186/1752-0509-3-37
Morgat A, Lombardot T, Axelsen KB, Aimo L, Niknejad A, Hyka-Nouspikel N, et al (2017) Updates in Rhea—an expert curated resource of biochemical reactions. Nucleic Acids Res 45(D1):D415–D418. https://doi.org/10.1093/nar/gkw990
Ng CY, Jung My, Lee J, Oh MK (2012) Production of 2,3-butanediol in Saccharomyces cerevisiae by in silico aided metabolic engineering. Microb Cell Fact 11(1):68. https://doi.org/10.1186/1475-2859-11-68. http://www.ncbi.nlm.nih.gov/pubmed/22640729. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC3442981
Niebel B, Leupold S, Heinemann M (2019) An upper limit on Gibbs energy dissipation governs cellular metabolism. Nat Metab 1:125–132
Nilsson A, Nielsen J (2016) Metabolic trade-offs in yeast are caused by F1F0-ATP synthase. Sci Rep 6:22264. https://doi.org/10.1038/srep22264.
Nocon J, Steiger MG, Pfeffer M, Sohn SB, Kim TY, Maurer M, et al (2014) Model based engineering of Pichia pastoris central metabolism enhances recombinant protein production. Metab Eng 24:129–138. https://www.sciencedirect.com/science/article/pii/S1096717614000706
Otero JM, Cimini D, Patil KR, Poulsen SG, Olsson L, Nielsen J (2013) Industrial systems biology of Saccharomyces cerevisiae enables novel succinic acid cell factory. PLoS One 8(1):e54144. https://doi.org/10.1371/journal.pone.0054144
Patil KR, Rocha I, Förster J, Nielsen J (2005) Evolutionary programming as a platform for in silico metabolic engineering. BMC Bioinf 6:308
Pereira R, Nielsen J, Rocha I (2016) Improving the flux distributions simulated with genome-scale metabolic models of Saccharomyces cerevisiae. Metab Eng Commun 3:153–163. https://doi.org/10.1016/j.meteno.2016.05.002
Pharkya P, Maranas CD (2006) An optimization framework for identifying reaction activation/inhibition or elimination candidates for overproduction in microbial systems. Metab Eng 8(1):1–13
Pitkänen E, Jouhten P, Hou J, Syed MF, Blomberg P, Kludas J, et al (2014) Comparative Genome-Scale Reconstruction of Gapless Metabolic Networks for Present and Ancestral Species. PLoS Comput Biol 10(2):e1003465. https://doi.org/10.1371/journal.pcbi.1003465
R Development Core Team (2018). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Ranganathan S, Suthers PF, Maranas CD (2010) OptForce: An optimization procedure for identifying all genetic manipulations leading to targeted overproductions. PLoS Comput Biol 6(4):e1000744. https://doi.org/10.1371/journal.pcbi
Rantasalo A, Landowski CP, Kuivanen J, Korppoo A, Reuter L, Koivistoinen O et al (2018) A universal gene expression system for fungi. Nucleic Acids Res 46(18):e111. https://doi.org/10.1093/nar/gky558
Rocha I, Maia P, Evangelista P, Vilaça P, Soares S, Pinto JP et al (2010) OptFlux: an open-source software platform for in silico metabolic engineering. BMC Syst Biol 4:45. https://doi.org/10.1186/1752-0509-4-45
Rosdi N, Abdullah A (2014) Limiting and excreting metabolites of succinate production in S. cerevisiae using flux balance analysis. In: 2014 8th Malaysian software engineering conference (MySEC). IEEE, pp 279–283. http://ieeexplore.ieee.org/document/6986029/
Sajitz-Hermstein M, Töpfer N, Kleessen S, Fernie AR, Nikoloski Z (2016) IReMet-flux: Constraint-based approach for integrating relative metabolite levels into a stoichiometric metabolic models. In: Bioinformatics 32(17):i755–i762. https://doi.org/10.1093/bioinformatics/btw465
Sanchez BJ, Nielsen J (2015) Genome scale models of yeast: towards standardized evaluation and consistent omic integration. Integr Biol 7(8):846–858. http://xlink.rsc.org/?DOI=C5IB00083A
Sanchez BJ, Zhang C, Nilsson A, Lahtvee PJ, Kerkhoven EJ, Nielsen J (2017) Improving the phenotype predictions of a yeast genome-scale metabolic model by incorporating enzymatic constraints. Mol Syst Biol 13(8):935. http://www.ncbi.nlm.nih.gov/pubmed/28779005. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC5572397
Schellenberger J, Park JO, Conrad TC, Palsson BØ (2010) BiGG: a Biochemical Genetic and Genomic knowledgebase of large scale metabolic reconstructions. BMC Bioinformatics 11:213
Schellenberger J, Que R, Fleming RMT, Thiele I, Orth JD, Feist AM, et al (2011) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA toolbox v2.0. Nat Protoc 6(9):1290–1307. https://doi.org/10.1038/nprot.2011.308
Schmidt BJ, Ebrahim A, Metz TO, Adkins JN, Palsson B, Hyduke DR (2013) GIM3E: condition-specific models of cellular metabolism developed from metabolomics and expression data. Bioinf 29(22):2900–2908. https://doi.org/10.1093/bioinformatics/btt493
Schuetz R, Kuepfer L, Sauer U (2007) Systematic evaluation of objective functions for predicting intracellular fluxes in Escherichia coli. Mol Syst Biol 3:119
Schuster S, Fell DA, Dandekar T (2000) A general definition of metabolic pathways useful for systematic organization and analysis of complex metabolic networks. Nat Biotechnol 18(3):326–332
Schuster S, Hilgetag C (1994) On elementary flux modes in biochemical reaction systems at steady state. J Biol Syst 2(2):165–182
Segrè D, Vitkup D, Church GM (2002) Analysis of optimality in natural and perturbed metabolic networks. In: proceedings of the national academy of sciences of the united states of america, vol. 99 no 23 pp 15112–15117. http://www.ncbi.nlm.nih.gov/pubmed/12415116http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC137552
Shlomi T, Berkman O, Ruppin E (2005) Regulatory on/off minimization of metabolic flux changes after genetic perturbations. In: Proceedings of the national academy of sciences, vol 102, no 21, pp 7695–7700. http://www.pnas.org/content/102/21/7695
Shlomi T, Cabili MN, Herrgård MJ, Palsson B, Ruppin E (2008) Network-based prediction of human tissue-specific metabolism. Nat Biotechnol 26(9):1003–1010. https://doi.org/10.1038/nbt.1487
Smallbone K, Simeonidis E, Swainston N, Mendes P (2010) Towards a genome-scale kinetic model of cellular metabolism. BMC Syst Biol 4:6. https://doi.org/10.1186/1752-0509-4-6
Stanford NJ, Lubitz T, Smallbone K, Klipp E, Mendes P, Liebermeister W (2013) Systematic construction of kinetic models from genome-scale metabolic networks. PLoS One 8(11):e79195. https://doi.org/10.1371/journal.pone.0079195
Strucko T, Zirngibl K, Pereira F, Kafkia E, Mohamed ET, Rettel M et al (2018) Laboratory evolution reveals regulatory and metabolic trade-offs of glycerol utilization in Saccharomyces cerevisiae. Metab Eng 47:73–82. https://doi.org/10.1016/j.ymben.2018.03.006
Suastegui M, Matthiesen JE, Carraher JM, Hernandez N, Rodriguez-Quiroz N, Okerlund A, et al (2016) Combining metabolic engineering and electrocatalysis: application to the production of polyamides from sugar. Angew Chemie Int Ed 55(7):2368–2373. https://doi.org/10.1002/anie.201509653
Sun Z, Meng H, Li J, Wang J, Li Q, Wang Y, et al (2014) Identification of novel knockout targets for improving terpenoids biosynthesis in Saccharomyces cerevisiae. PLoS One 9(11):e112615. https://doi.org/10.1371/journal.pone.0112615
Suthers PF, Zomorrodi A, Maranas CD (2009) Genome-scale gene/reaction essentiality and synthetic lethality analysis. Mol Syst Biol 5:301. https://doi.org/10.1038/msb.2009.56
Swainston N, Smallbone K, Mendes P, Kell D, Paton N (2011) The SuBliMinaL toolbox: automating steps in the reconstruction of metabolic networks. J Integr Bioinf 8(2):186. http://www.ncbi.nlm.nih.gov/pubmed/22095399
Tepper N, Shlomi T (2010) Predicting metabolic engineering knockout strategies for chemical production: accounting for competing pathways. Bioinformatics 26(4):536–543. https://doi.org/10.1093/bioinformatics/btp704
Tomas-Gamisans M, Ferrer P, Albiol J (2018) Fine-tuning the P. pastoris iMT1026 genome-scale metabolic model for improved prediction of growth on methanol or glycerol as sole carbon sources. Microb Biotech 11(1):224–237. http://www.ncbi.nlm.nih.gov/pubmed/29160039. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=PMC5743807
Toro L, Pinilla L, Quintero JC, Rios R (2014) Flux Balance analysis and strain optimization for ethanol production in Saccharomyces cerevisiae. Springer, Cham, pp 177–182. https://doi.org/10.1007/978-3-319-01568-2_26
Tramontano M, Andrejev S, Pruteanu M, Klünemann M, Kuhn M, Galardini M, et al (2018) Nutritional preferences of human gut bacteria reveal their metabolic idiosyncrasies. Nat Microbiol 3(4):514–522. https://doi.org/10.1038/s41564-018-0123-9
Trinh CT, Srienc F (2009) Metabolic engineering of Escherichia coli for efficient conversion of glycerol to ethanol. Appl Environ Microbiol 75(21):6696–6705. https://doi.org/10.1128/AEM.00670-09
Unrean P, Jeennor S, Laoteng K (2016) Systematic development of biomass overproducing Scheffersomyces stipitis for high-cell-density fermentations. Synth Syst Biotechnol 1(1):47–55. https://www.sciencedirect.com/science/article/pii/S2405805X15300211
Unrean P, Trinh CT, Srienc F (2010) Rational design and construction of an efficient E. coli for production of diapolycopendioic acid. Metab Eng 12(2):112–122. https://doi.org/10.1016/j.ymben.2009.11.002
Urbanczik R (2007) Enumerating constrained elementary flux vectors of metabolic networks. IET Syst Biol 1(5):274–279
Varma A, Palsson BO (1994) Stoichiometric flux balance models quantitatively predict growth and metabolic by-product secretion in wild-type Escherichia coli W3110. Appl Environ Microbiol 60(10):3724–3731
Vazquez A, Oltvai ZN (2016) Macromolecular crowding explains overflow metabolism in cells. Sci Rep 6:31007. https://doi.org/10.1038/srep31007
von Kamp A, Klamt S (2014) Enumeration of Smallest Intervention Strategies in Genome-Scale Metabolic Networks. PLoS Comput Biol 10(1):e1003378. https://doi.org/10.1371/journal.pcbi.1003378
von Kamp A, Thiele S, Hädicke O, Klamt S (2017) Use of CellNetAnalyzer in biotechnology and metabolic engineering. J Biotechn 261:221–228. https://doi.org/10.1016/j.jbiotec.2017.05.001
Xu G, Zou W, Chen X, Xu N, Liu L, Chen J (2012) Fumaric acid production in Saccharomyces cerevisiae by in silico aided metabolic engineering. PLoS One 7(12):e52086. https://doi.org/10.1371/journal.pone.0052086
Yang L, Yurkovich JT, King ZA, Palsson BO (2018) Modeling the multi-scale mechanisms of macromolecular resource allocation. 45:8–15. https://doi.org/10.1016/j.mib.2018.01.002
Yu T, Zhou YJ, Huang M, Liu Q, Pereira R, David F et al (2018) Reprogramming yeast metabolism from alcoholic fermentation to lipogenesis. Cell 174(6):1549–1558.e14. https://doi.org/10.1016/j.cell.2018.07.013
Zelezniak A, Sheridan S, Patil KR (2014) Contribution of Network Connectivity in Determining the Relationship between Gene Expression and Metabolite Concentration Changes. PLoS Comput Biol 10(4):e1003572. https://doi.org/10.1371/journal.pcbi.1003572
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Castillo, S., Patil, K.R., Jouhten, P. (2019). Yeast Genome-Scale Metabolic Models for Simulating Genotype–Phenotype Relations. In: Sá-Correia, I. (eds) Yeasts in Biotechnology and Human Health. Progress in Molecular and Subcellular Biology, vol 58. Springer, Cham. https://doi.org/10.1007/978-3-030-13035-0_5
Download citation
DOI: https://doi.org/10.1007/978-3-030-13035-0_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-13034-3
Online ISBN: 978-3-030-13035-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)