Abstract
In this chapter we will explain what cell signaling is, how it works, what it serves for and especially why it is important in the context of cancer. The basics about three key signaling pathways, JAK/STAT, MEK/ERK and PI3K/Akt/mTOR, which are frequently involved in all different types of cancer will be adressed. Many other signaling pathways exist, such as Notch, Wnt, Hedgehog, etc., that also partake in tumorigenesis. We will also address what consist the so-called “signaling therapies”, which are their advantages, understand their potential and have some insight into the mechanisms that explain why they frequently fail. We will tackle some of the strategies aiming at overcoming resistance associated with signaling therapies and their possible caveats. Finally, we hope it will be clear the need for a deep characterization of the cancer patient in order to devise the best targeted therapies.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
After reading this chapter you should understand what cell signaling is, how it works, what it serves for and especially why it is important in the context of cancer. You should know the basics about three key signaling pathways:
-
JAK/STAT,
-
MEK/ERK and
-
PI3K/Akt/mTOR,
and realize that they are very frequently involved in all different types of cancer. You should know, however, that many other signaling pathways exist, such as Notch, Wnt, Hedgehog, etc., that also partake in tumorigenesis. Importantly, you are expected to know in what consist the so-called “signaling therapies”, which are their advantages, understand their potential and have some insight into the mechanisms that explain why they frequently fail. You should also understand some of the strategies aiming at overcoming resistance associated with signaling therapies and their possible caveats. Finally, we hope we will convey the need for a deep characterization of the cancer patient in order to devise the best targeted therapies.
FormalPara Learning ObjectivesAfter reading this chapter, students should be able to:
-
1.
Understand the importance of signal transduction in both physiological and cancer contexts.
-
2.
Enumerate the major post-translational protein modifications observed in signal transduction.
-
3.
Recognize the main elements of the most common signal transduction pathways involved in cancer progression.
-
4.
Pinpoint the reasons for hyper-activation of signal transduction cascades in cancer.
-
5.
Identify the most commonly used therapeutic strategies to target the signaling machinery – “signaling therapies”.
-
6.
Recognize the mechanisms of resistance to signaling therapies, and possible strategies to overcome resistance.
-
Signal transduction and signal transduction pathways
-
Microenvironmental and cell-intrinsic cues
-
Post-translational protein modifications
-
Protein phosphorylation
-
Signaling players
-
Oncogene addiction
-
Signaling therapies
-
Resistance mechanisms
3.1 What Signal Transduction Is and How it Works
Complex biological systems, such as mammals or birds (to name just two of many potential examples), engage mechanisms that allow them to perceive and cope with an ever-changing environment. This ability to respond to external cues in an appropriate manner is essential for their survival and adaptation. At a vastly smaller scale unicellular organisms, such as bacteria, do exactly the same – because this process of perception-integration-response is as essential for their overall fitness as it is for multicellular organisms. So much so that any cell in a human being or in a mouse will have the same exact ability – of transforming signals, mainly from the outside of the cell, into a response (which we broadly call signal transduction). In fact, signal transduction, which occurs in each cell of the body, is largely required for the correct functioning of the whole organism [1].
How does signal transduction work in a cell? Like anything in biology, it is both very simple and very complex. Let’s start by the simple before we move on to the more complex. To perceive the environment a cell needs a receptor, which will enable it to receive an “input”. This receptor may come in many “flavors”: it can be a cytokine receptor, a growth factor receptor, an adhesion molecule, a chemokine receptor and many other receptors that are located at the surface of any cell, in the plasma membrane. Or it can be cell type-specific, such as the T cell receptor, which as the name indicates is present only in T cells of the adaptive immune system; or the many odor receptors that are present only in olfactory neurons. It can even be a hormone receptor, which is not located at the cell surface but rather in the cytoplasm. Whatever the “flavor”, the receptor will allow the cell to receive an “input” signal that triggers a response. This response is mediated by biochemical reactions that are usually referred to as signal transduction pathways or signal transduction cascades, since normally they follow a sequential pattern in which an upstream element impacts an immediately downstream element that subsequently affects another element further downstream. Below, we will describe briefly the biochemical reactions involved in signal transduction. At this stage it suffices to say that protein kinases are the most well-studied and prominent elements in these biochemical cascades. Finally, a certain response (an “output”) should result. This response can range from cell cycle progression, to differentiation, to migration or apoptosis. It frequently involves the modulation of transcription factors and consequent transcriptional regulation of different genes, but in some instances it may not involve gene regulation and it may occur only at the protein level (as it happens with certain apoptotic stimuli that lead to release of cytochrome c from mitochondria and consequent activation of caspases) [1, 2].
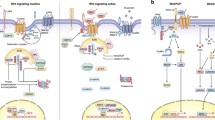
In short, signal transduction is the biochemical process by which a cell perceives a certain cue, and triggers decision-making ultimately leading to a response to the initial cue (◘ Fig. 3.1).
The general organization of cell signaling. Cells have the ability to perceive and receive (environmental) stimuli (input signals, green arrow) via different kind of receptors and other sensors. This information is then integrated and processed to trigger appropriate response(s) (output signals, red arrow) mediated by complex intracellular signaling networks, called signal transduction pathways or cascades, such as JAK/STAT, Ras/MEK/Erk, PI3K/AKT/mTOR, Notch, etc. The cellular outputs may vary from cell cycle progression, to differentiation, to migration or apoptosis, and usually involve modulation at the gene and/or protein levels
Although the majority of the cues will expectedly lie outside of the cell, their nature can vary substantially and include not only cell-cell contact or soluble ligands, but also, for example, the sensing of mechanical forces. In addition, it should be underlined that input signals may originate also within the cell. A notable example is DNA damage, which activates signal transduction pathways that recruit the machinery required for the resolution of the damage and, in case of failure, eventually promote cell death (see ► Chap. 5).
3.2 Major Post-Translational Protein Modifications in Signal Transduction
Signal transduction pathways rely vastly on enzymes that catalyze reactions leading to protein post-translational modifications, including (◘ Table 3.1):
-
ubiquitination,
-
sumoylation,
-
acetylation,
-
methylation,
-
O-glycosylation and
-
phosphorylation.
The latter is, by and large, the most well-studied and characterized. This is not only because it is technically easier to analyze than many of the other modifications but also, and most importantly, because it is the one consistently involved in almost all of the main signal transduction pathways that have been characterized in higher organisms.
Protein phosphorylation is catalyzed by protein kinases, which transfer the terminal phosphate group of ATP to the hydroxyl moiety of an amino acid residue in a target protein. The products of this reaction are a phosphorylated protein and ADP (◘ Fig. 3.2). Since phosphate groups are highly negatively charged, phosphorylation has the ability to modify significantly the charge of the target protein, consequently altering its conformation and thereby affecting its function, association with other proteins and/or cellular localization [3].
Roughly 2% of all eukaryotic genes are protein kinases, and the mammalian kinome includes more than 500 members.
Protein kinases acting within mammalian cells are classified into three categories:
-
1.
Tyrosine-specific protein kinases – phosphorylate tyrosine residues on target proteins
-
2.
Serine/threonine-specific protein kinases – phosphorylate serine or threonine residues
-
3.
Dual-specificity protein kinases – are rare and display the capacity to phosphorylate simultaneously tyrosine and serine/threonine residues [4].
The vast majority of protein phosphorylation occurs on serine residues (80–95%), significantly less on threonines (4–17%) and less than 1% up to 4% on tyrosines. This does not mean phosphorylation of tyrosines is less relevant than of serines or threonines. On the contrary, tyrosine phosphorylation is frequently involved in the most upstream steps of signal transduction, with about half of tyrosine kinases being cell surface receptors and the remaining displaying activity mostly close to the plasma membrane.
Protein tyrosine kinases are subdivided into:
-
1.
Receptor tyrosine kinases (RTKs), if they are cell surface receptors with intrinsic kinase activity, and
-
2.
Receptor-associated or non-receptor tyrosine kinases, if they are cytosolic – normally associated with or recruited by cell surface receptors (◘ Table 3.2) to propagate the signaling to downstream effectors [3, 4].
Of course, dephosphorylation of proteins is also essential for signal transduction. In some cases to propagate the signal and in other cases (perhaps most often) to “reset” the system and allow a certain protein, and the pathway that it pertains, to go back to the pre-stimulus state, which is essential both to prevent pathway permanent activation (that, as we will see below, can put cells at risk for cancer transformation) and to allow for subsequent physiological re-stimulation. The reaction of protein dephosphorylation is catalyzed by protein phosphatases [5]. Although, for simplicity, we will not focus our attention on these in the next sections, it is important to keep in mind that protein phosphatases are as essential as kinases for the homeostasis of the cell as far as signal transduction goes [5].
3.3 Why Signal Transduction Is Important in Cancer?
At this point you should have a grasp of what signal transduction is but may be asking why one should spend time analyzing signaling in the context of cancer. There are several reasons. The most obvious is not difficult to guess, based on the type of cellular responses that are regulated by signal transduction pathways. These include:
-
regulation of cell cycle and proliferation,
-
migration,
-
differentiation,
-
viability versus cell death,
-
metabolism, etc.
It is therefore not surprising that deregulation of signal transduction may have considerable impact on the homeostasis of the cell. For example, aberrant, constitutive activation of a pathway that promotes proliferation in a single cell may lead to a selective advantage that eventually leads to clonal expansion at the expense of other cells. So firstly, signaling pathways are important for cancer simply because they are essential regulators of all the different facets of normal cell physiology. Consequently, signaling players are often oncogenes (or, sometimes, tumor suppressors). An example: the first human proto-oncogene to be cloned, RAS, mutated in around one third of all cancers, encodes a small GTPase essential for activation of Ras/MEK/ERK signaling pathway. In fact, examples can be found in ALL classes of signaling players (◘ Fig. 3.3) [2, 3]. Ligands such cytokines and growth factors (e.g. PDGF, VEGF, Wnt, IL-15) are often overexpressed and involved in aberrant autocrine/paracrine loops that promote malignancy. Their respective receptors, frequently RTKs, are well-known oncogenes (e.g. PDGFR, EGFR, IGFR, Notch, IL7R) often mutated and amplified in cancer cells. Of course, the examples of intracellular signaling constituents known to be altered in cancer are numerous (e.g. PI3Ks, JAKs, Ras, PTEN, etc). Finally, the transcription factors that are regulated by signaling pathways are also frequently oncogenes (e.g. MYC, TAL1) or tumors suppressors (e.g. P53).
Importantly, cancer cells frequently display functional dependence on activation of specific pro-tumoral signaling pathways in a process often referred to as “oncogene addiction”. Why does this happen? Cancer cells are selected for their permanent activation of a certain pathway. This allows them to expand at the expense of their normal counterparts, but it also implicates that the malignant cells depend on the perpetual activation of that pathway for their maintenance [6]. This contrasts with what happens in normal cells that rely only transiently, after a certain physiological stimulus, on the activation of the same pathway. In other words, the “addiction” to a certain oncogenic signaling pathway – a feature not shared by normal cells – constitutes an advantage to the cancer cell but it is its “Achilles’ heel” as well. Evidently this also represents a therapeutic opportunity, because targeting a signaling pathway used by both cancer cells and normal cells may have substantially different consequences in each type – with normal cells expectedly being less sensitive to signaling-specific drugs than transformed, oncogene-addicted, cells.
This leads us to the third reason why signaling is important in cancer: signaling pathways constitute appealing molecular targets for therapeutic intervention, the actual practical reason being that kinases are easily “druggable” (using either ATP-competitive or allosteric inhibitors) and thus often excellent targets for drug development and clinical intervention [7].
Finally, there is a fourth reason placing signaling at the center-spot of tumorigenesis: the crosstalk between cancer cells and their microenvironment relies on the modulation of specific signal transduction pathways. It is now firmly established that a cancer is a complex entity that includes not only the cancer cells themselves but also a multitude of normal cells that include endothelial cells and pericytes, fibroblasts, and diverse immune cells. Therefore, cancer progression depends not only on lesions within the malignant cells but also on interactions between the cancer cells and their surroundings, in a manner that is ultimately beneficial for the tumor. This sort of dialogue occurs via, and is strictly dependent on, modulation of signaling pathways within the cancer cells and the tumor microenvironment [8].
Overall, the involvement of signaling pathways in cancer origin and progression, the dependence of cancer cells on certain pathways and the fact that signaling is also important for the pro-tumoral cross-talk between malignant cells and “normal” cells in their surrounding microenvironment, all contribute to the notion that targeting signaling pathways in a rational manner constitutes a viable therapeutic strategy for cancer intervention. Of course, this requires a profound understanding of the biology and biochemistry underlying signal transduction activation in cancer cells.
3.4 Key Signaling Pathways in Cancer
The list of signal transduction pathways whose involvement in cancer has been demonstrated is growing (◘ Table 3.3). Below, we will focus on three pathways: JAK/STAT, Ras/MEK/Erk, and PI3K/Akt/mTOR, the reason being that, in contrast to the remaining included in ◘ Table 3.3, they are very frequently activated in all types of cancer. Also, they are common effectors of different upstream lesions. For instance, different RTKs and cytokine receptors lead to the activation of STAT, PI3K and MEK signaling. Similarly, the fusion protein BCR/ABL, which is a hallmark of Chronic Myeloid Leukemia (CML), also activates these three signaling pathways. In short, these three critical signal transduction cascades are frequently hyper-activated in cancer as a consequence of gain-of-function mutations in their members, loss-of-function mutations or deletions in negative regulators or activation of upstream receptors. In the latter case, receptor activation can result from gene amplification or gain-of-function mutation, or from autocrine or paracrine aberrant stimulation due to ligand overexpression.
3.4.1 JAK/STAT
This is the pathway canonically activated by cytokines. There are four Janus kinase (JAK) family members (JAK1, JAK2, JAK3 and TYK2), which are tyrosine-specific protein kinases of around 110–130 kDa. With the exception of JAK3, which is largely restricted to hematopoietic cells, JAKs are ubiquitously expressed [9]. The other elements of the pathway are the signal transducers and activators of transcription (STATs), which, as the name indicates, are latent transcription factors located in the cytoplasm. STATs are encoded by seven genes: STAT1-4, STAT5A, STAT5B and STAT6. STATs possess several important domains:
-
an N-terminal domain used for instance for nuclear translocation,
-
a coiled-coil domain involved in protein-protein interactions,
-
a DNA-binding domain,
-
a Src-homology 2 (SH2) domain required for binding to the receptor and subsequent STAT dimerization, and
-
a C-terminal transactivation domain required for transcriptional activity. This transactivation domain is also where a critical tyrosine residue is located that is essential for STAT5 activity [9].
Some STATs (STAT-1, -3, -4 and -5) display one additional serine residue in the transactivation domain that is required for maximal transcriptional activity, which is obviously not a target of JAKs (given the fact that they are strict tyrosine kinases). This serine actually represents a good example of a phenomenon that is frequent in signal transduction: cross-talk between different pathways. In this case the serine residue is known to be phosphorylated by ERKs.
How does JAK/STAT pathway work? It is actually very straightforward. The conformational changes triggered by the binding of a cytokine to its receptor, allow for the recruitment and/or activation of JAKs that associate with the cytoplasmic region of the receptor. The JAKs then transphosphorylate each other, leading to their maximal activity, upon which they phosphorylate tyrosine residues in the receptor itself that constitute binding sites for STATs via their SH2 domains. The recruitment of STATs to the vicinity of JAKs allows the latter to phosphorylate them, promoting STAT homo- or hetero-dimerization and consequent translocation into the nucleus, where they are finally allowed to bind to the promoters of target genes and act as bona fide transcription factors (◘ Fig. 3.4) [10]. Examples of STAT target genes include MYC, CCND1 (encoding Cyclin D1), BCL2 and BCL2L1 (encoding BCL-XL), which are positively involved in induction of proliferation and protection from apoptosis [11,12,13].
Summary of JAK/STAT pathway. STAT proteins are recruited to the cytokine receptor and phosphorylated by JAKs, leading to STAT dimerization and nuclear translocation. In the nucleus, STAT dimers bind DNA and promote transcription of several target genes, such as MYC, CCND1 and Bcl-2-family members BCL2 and BCL2L1, which are positively involved in induction of cell viability, proliferation and cell cycle progression
3.4.2 Ras/MEK/Erk
The MAPK pathway has three major signaling modules, but we are going to focus only on the classical extracellular signal-regulated kinase (Erk) pathway, or simply Ras/MEK/Erk pathway. This pathway is mainly activated by mitogenic growth factors and cytokines [14, 15].
It starts with activation of a Ras guanine nucleotide exchange factor (RasGEF) that triggers the release of GDP from Ras and its association with GTP. This event turns on the small GTPase Ras, which indirectly (through a rather complex mechanism) activates Raf and thereby transduces the signal via a cascade that contains three levels (◘ Fig. 3.5):
-
First Level: Raf, a MAPK kinase kinase (MAPKKK), phosphorylates and thereby activates MEK,
-
Second Level: MEK, a MAPK kinase (MAPKK), phosphorylates and thereby activates Erk,
-
Third Level: Erk, a MAP kinase [15], phosphorylates and activates several downstream targets, including the kinases p90RSK [16], Mnk [17] and Msk [18] and transcription factors such as c-Myc, c-Fos, c-Jun and C/EBPβ [19]. It consequently promotes cell cycle progression by regulating the expression of cyclin D1, p27kip1 and p21cip1 [20].
Summary of Ras/MEK/Erk pathway. Upon ligand binding, Ras protein is turned on by association with a GTP molecule, thus leading to the activation of the phosphorylation cascade MAPKKK-MAPKK-MAPK. Erk1/2 (MAPK) activation leads to phosphorylation and activation of several downstream targets, including Rsk, Msk and Mnk, which are involved in mRNA translation. When translocated into the nucleus, Erk1/2 phosphorylates and regulates the action of different transcription factors such as MYC, FOS, JUN, C/EBPβ
Overall, this pathway promotes cell survival and proliferation and regulates differentiation.
3.4.3 PI3K/Akt/mTOR
The PI3K/Akt/mTOR pathway is a major cellular signaling module known to promote cell growth and survival, to inhibit apoptosis and to control metabolism [21, 22]. There are three different classes (I-III) of the phosphatidylinositol 3-OH-kinases (PI3K), but only the ones belonging to class IA are activated by RTKs.
Class IA PI3Ks form heterodimers of:
-
a p85 regulatory subunit and
-
a p110 catalytic subunit [23].
Upon activation, PI3K catalyzes the conversion of phosphatidylinositol-4,5-bisphosphate (PIP2) into the second messenger phosphatidylinositol-3,4-5-trisphosphate (PIP3) [21,22,23]. This event triggers the recruitment of proteins containing a pleckstrin homology (PH) domain to the vicinity of the plasma membrane. Those proteins include the major downstream effector of the pathway, the serine/threonine kinase Akt/PKB (Protein kinase B), the serine/threonine 3-phosphoinositide dependent protein kinase-1 (PDK1) and the mammalian target of rapamycin complex 2 (mTORC2). Their co-localization elicits Akt phosphorylation by PDK1 and mTORC2 (which acts as PDK2) and consequent Akt full activation. PDK1 phosphorylates T308, whereas mTORC2 phosphorylates S473 [21, 22]. Once activated, Akt is able to activate or repress, through phosphorylation, multiple downstream targets (◘ Fig. 3.6). Without going into too much detail, Akt:
-
inhibits the glycogen synthase kinase-3α/β (GSK3α/β), thus promoting both cell proliferation and viability;
-
inhibits the forkhead box O family of transcription factors (FoxOs), known to promote the transcription of pro-apoptotic genes and cell cycle inhibitors;
-
inhibits Bad and Caspase-9, proteins directly involved in apoptosis;
-
leads to the activation of another complex involving mTOR (mTORC1), which is crucial for cell growth and metabolism [21, 23].
Summary of PI3K/Akt/mTOR pathway. Activated PI3K phosphorylates PIP2 into PIP3, while PTEN antagonizes its action by dephosphorylation of PIP3 into PIP2. The generation of the second messenger PIP3 leads to phosphorylation and activation of Akt by both PDK1 and mTORC2 (acting as PDK2). Activated Akt has several intracellular targets, such as GSK-3, FoxO family members, Bad, Caspase 9 and TSC1/2 complex. Inactivation of the TSC1/2 complex through phosphorylation leads to stabilization and activation of mTORC1, which in turn promotes protein translation via activation of p70S6K and inactivation of 4E-BP1. Further details of this pathway are described in the main text
Downregulation of this pathway is mediated by the tumor suppressor PTEN (Phosphatase and tensin homolog), a lipid phosphatase that dephosphorylates PIP3 into PIP2 (◘ Fig. 3.6) [21,22,23].
Since mTOR will be used to discuss some mechanisms of resistance to signaling therapies, we should go slightly deeper on how it works. As you have guessed by now, mTOR forms two distinct complexes:
-
mTORC1, mainly composed by mTOR, the catalytic subunit, regulatory-associated protein of mTOR (Raptor), mammalian lethal with Sec13 protein 8 (mLST8) and proline-rich AKT substrate 40 kDa (PRAS40);
-
mTORC2, mostly formed by mTOR, rapamycin-insensitive companion of mTOR (Rictor); mammalian stress-activated protein kinase interacting protein (mSIN1) and mLST8 [24].
Importantly, mTORC2 phosphorylates and activates Akt directly, whereas Akt activates mTORC1 by two means: directly, via PRAS40 phosphorylation [25]; and indirectly, by phosphorylation of the tuberous sclerosis complex 2 (TSC2), which promotes destabilization of the heterodimer TSC1/2 and consequent activation of the small GTPase Rheb (◘ Fig. 3.6) [26]. Upon activation, mTORC1 phosphorylates and inactivates the eukaryotic initiation factor 4E (eIF4E)-binding protein 1 (4E-BP1) [27] and activates the p70 ribosomal S6 kinase (p70S6K) [28], leading to an increase in protein translation at the ribosome.
3.5 Targeted Signaling Therapies: The Pros and Cons
Now that we know the very basics of signaling we can better understand its potential to deregulate the balance in a cell towards an excessively proliferative state. In addition, one should recall that aberrant activation of cell surface receptors, such as RTKs, often leads to the concomitant activation of STATs, MEK/ERK and PI3K/Akt/mTOR signaling, amongst other pathways. In other words, the oncogenic potential of RTKs actually lies not on a direct detrimental effect on the cell but precisely on the ability of RTKs to activate downstream, potentially harmful, signaling players.
Of course, the knowledge that constitutive activation of certain signaling pathways is causal in cancer progression, maintenance and, quite often, resistance to conventional chemotherapies, makes them highly attractive as targets for therapeutic intervention. We have already mentioned the relative easiness in developing pharmacological inhibitors for protein kinases. So, one obvious strategy is to use small molecule, cell-permeable inhibitors that can directly target intracellular kinases [3]. We will mention this strategy repeatedly in the following sections. Yet, there is another obvious strategy if one refers to the most upstream elements in a signaling pathway, those that locate at the cell surface. In those cases, the plasma membrane does not constitute an obstacle and therefore antibodies can be used to neutralize and consequently inhibit the activity of the surface receptor, thereby preventing the activation of downstream signaling [3]. This strategy has the benefit of often also triggering an immune response against the (tumor) cells that express the receptor.
Contrary to conventional chemotherapy, these therapies targeting signaling elements (“targeted signaling therapies” or “signaling therapies”) have the advantage of being substantially more selective against the cancer cells and thus, at least to some extent, having the possibility to decrease toxicities towards normal cells and thus side-effects. This increased selectivity results from the fact that some signaling drugs inhibit targets expressed only by the tumor cells, such as the fusion protein BCR-ABL or mutant BRAF, which we will mention in more detail below. The other, less obvious, reason was already mentioned: oncogene addiction, the absolute reliance of cancer cells on a certain pathway in contrast to a normal cell. This does not mean that signaling therapies are bullet-proof. Unfortunately, they do sometimes generate substantial side-effects. This, however, is not the real problem in most cases, but rather the development of resistance. In this regard, targeted therapies can be just as ineffective (if not worse at times) as conventional chemotherapies. Why this happens and how we can tackle this problem is what we will discuss next.
3.6 Why Do Cancer Cells Develop Resistance to Targeted Signaling Therapies?
There are many mechanisms of resistance to signaling therapies. We will briefly mention a few of the most well-known.
3.6.1 Mutational Activation of an Element in the Pathway Downstream from the Target
Let’s start with a simple example. A group of RTKs well-known for their involvement in cancer is the epidermal growth factor receptor (EGFR/ErbB/Her) family, which has four members, the most tumorigenic of which appears to be EGFR2 (or Her2/Neu) [29, 30]. Our example, however, will rely on EGFR1. An effective treatment for colorectal cancer patients whose malignant cells express EGFR1, and consequently display constitutive activation of downstream signaling, involves the administration of a monoclonal antibody, named Cetuximab. It happens that the benefit of Cetuximab administration (in combination with conventional chemotherapy, as compared to conventional chemotherapy alone) is limited to patients with wild type, non-mutated, KRAS [31]. Why is this? Well, remember that RAS, no matter what isoform, is an oncogene and so mutated Ras in the context of cancer means ACTIVATED Ras. Now remember that the other name for signaling pathways is signaling cascades – one element activates the next which activates the next, and so on. Cetuximab inhibits EGFR1, which will no longer activate Ras, which will no longer activate Raf, which will no longer activate MEK, which will no longer activate ERK. Great! But wait a second now. How will Cetuximab inhibit Raf and MEK and ERK if Ras is constitutively activated as a consequence of a mutation and no longer requires EGFR to be turned on (◘ Fig. 3.7)? A minor cancer clone displaying a KRAS mutation will eventually expand and drive resistance to Cetuximab. This notion that a targeted therapy can become obsolete due to the mutational activation of a downstream effector of the target is of utmost importance, and in fact sequencing for RAS mutations is now clinical practice for selecting patients for treatment with anti-EGFR therapies [32].
Mechanisms of acquired resistance to receptor tyrosine kinase inhibitors such as Cetuximab, an anti-EGFR monoclonal antibody. a Cancers that are addicted to RTKs have activation of Ras/MAPK signaling, which promotes cell growth and survival. b Cancers expressing EGFR, and consequently displaying constitutive activation of downstream signaling, are sensitive to the tyrosine kinase inhibitor Cetuximab. Inhibition of the receptor leads to loss of Ras/MAPK signaling, blocking cell growth and survival and promoting apoptosis. c Cancers can become resistant by two means: first, by mutational activation of a downstream effector of the target (in this case Ras); and second, by mutation in other signaling players in other parallel pathways (e.g. PI3K activating mutations)
Of course, it is important to take into account the fact that there are other effector pathways downstream from EGFR. So, knowing that anti-EGFR targeting agents are not effective against all KRAS wild type cases begged the question: how about mutations in other signaling players (downstream from RAS or in other parallel pathways)? For example, what if PI3K is constitutively active? [32]. Although the relevance of mutations in PI3K family members has proved controversial and not generally implemented in the clinic to determine which patients should be treated with cetuximab and equivalent drugs, the fact is that this is another potential mechanism of tumor to escape to therapy. The corollary is that one should have the best characterization of the patient’s mutational profile in order to device the most effective signaling therapies (◘ Fig. 3.7).
Food for thought: there is evidence that JAK inhibitors such as the JAK1/JAK2-specific ruxolitinib may be useful to treat acute lymphoblastic leukemia patients, especially (but not only) those with gain-of-function mutations in the upstream receptor IL-7R or in JAK1 itself [33,34,35]. However, it is also known that some of these patients display STAT5B activating mutations [36]. How sensitive to ruxolitinib will the patients with STAT5B mutations be? Based on pre-clinical studies, the answer is: very little. However, the use of Bcl-2 antagonists may prove useful for those cases, given that STAT5 positively regulates anti-apoptotic Bcl-2 family members that can be inhibited by this class of drugs [35].
3.6.2 Release from a Negative Feedback Loop
So far, we have centered our attention on positive regulation of signaling pathways. However, as mentioned already, all pathways have control mechanisms to prevent permanent activation of signaling pathways for the homeostasis of the cell. We can use PI3K/Akt/mTOR pathway to illustrate how negative feedback loop mechanisms can turn into a problem in this context. You already know that mTOR phosphorylates p70S6K to regulate protein translation. What you may not know yet is that p70S6K is also involved in the negative regulation of PI3K-mediated signaling by phosphorylating and thereby hampering the function of IRS1, an adaptor protein that sometimes “links” cell surface receptors to PI3K and thus contributes to the full activation of PI3K (◘ Fig. 3.6) [32]. To block PI3K/Akt/mTOR pathway activation in cancer cells one can use rapamycin (Sirolimus) or one of its analogs (Temsirolimus), which are highly selective allosteric inhibitors of mTORC1 [37]. This blocks the pathway on a downstream element, so it should be a very effective and smart way of making sure that the cascade is shutdown even if activating mutations occur in different upstream members. This is true. However, the allosteric inhibitors of mTORC1 have a downside: they can also inhibit the negative feedback loop. This means that PI3K and Akt are no longer under the control of this mechanism and therefore their activity can actually increase. This would not be a problem if the pathway was absolutely linear – it would be inhibited at the mTORC1 level. But we know that Akt can regulate several other important players in cell cycle and apoptosis regulation and so easily bypass the effects of mTORC1 inhibitors. This problem can be overcome by the use of the so-called TOR kinase inhibitors, which inhibit mTOR catalytic site and thus affect both mTORC1 and mTORC2 (e.g. Torkinib (PP242), Ku-0063794), or by using dual inhibitors of mTOR and PI3K (e.g. PI-103, NVP-BEZ-235) [37, 38].
3.6.3 Mutations and Amplifications in the Target
Imatinib and related inhibitors constitute the most successful example of signaling therapies. Imatinib is used to treat patients whose malignant cells display BCR/ABL fusion proteins (ABL is a tyrosine kinase that becomes always ON as a result of this fusion) that arise from the reciprocal non-random translocation t(9;22)(q34;q11), and the consequent expression of a minute chromosome 22, called Philadelphia (Ph) chromosome. Virtually all patients with Chronic Myeloid Leukemia (CML) and a fraction of Acute Lymphoblastic Leukemia (ALL) patients are Ph-positive. The use of Imatinib has dramatically improved the survival and prognosis of these patients, especially those with CML [39]. However, resistance develops in some cases [40, 41]. Why is this? Imatinib binds close to the ATP binding site of BCR/ABL, locking it in a closed, self-inhibited conformation. Mutations exist in BCR/ABL itself that cause resistance to Imatinib by shifting its equilibrium toward the open or active conformation [41, 42]. There is also evidence that BCR/ABL locus amplification may occur, leading to higher expression of the protein that contributes to resistance to the inhibitor [42]. In any case, the strategy to overcome resistance normally involves the use of the so-called second-generation BCR/ABL inhibitors (such as Dasatinib), which in fact are less specific than Imatinib and affect also downstream targets of BCR/ABL such Src-family kinases [37, 43]. This means that even in the event of mutations occurring in BCR/ABL there is a safeguard mechanism of inhibition acting downstream. Of course, the potential downside is that the less specific an inhibitor is the more likely it may affect normal cells and cause side-effects.
Similar mechanisms of resistance have been described for another good example of targeted therapies: Vemurafenib – the specific BRAFV600E inhibitor, which demonstrated activity in melanoma patients with this specific mutation. In fact, paradoxical activation of MEK/ERK signaling due to the release from negative feedback loops is also at play in this case, as well as the occurrence (albeit infrequent) of activating ERK mutations.
3.6.4 Other Mechanisms of Resistance
Vemurafenib, inhibitor of BRAFV600E mutation, serves to exemplify other mechanisms of resistance. For example, via triggering of alternative mechanisms of activation of downstream members of the pathway. For example, gain-of-function mutations in MLK, a kinase able to phosphorylate MEK, lead to ERK activation without the requirement for any mutations in ERK itself or any other member of the canonical MEK/ERK pathway. Strange as it may sound given the fact that Ras is upstream of Raf, any event leading to Ras activation may also contribute to resistance to Vemurafenib. This is the consequence of Vemurafenib being a highly specific inhibitor of mutant B-Raf (BRAFV600E). The problem in this case is that there are other Raf isoforms (A-Raf, C-Raf) that can be activated by Ras and propagate the signaling even in the presence of Vemurafenib. Thus, if the cells display (or gain) activating mutations in Ras, or in upstream RTKs, or deletion or loss-of-function mutations in NF1 (a negative regulator of Ras), or aberrant autocrine loops, they will inevitably end-up driving resistance to the inhibitor. What are the strategies to deal with this challenge? Once again, one may try combining drugs – for instance, using MEK inhibitors (such as Trametinib) concomitantly with Vemurafenib to block putative activation of MEK/Erk pathway [44]. Besides the danger of increased toxicities this does not necessarily solve the problem, since some patients will display mechanisms of resistance that involve activation of PI3K/Akt/mTOR signaling rather than MEK/Erk – once again alerting to the necessity of thoroughly characterizing each patient’s mutational profile.
Only tailor-made therapies (i.e. adapted to the molecular characteristics of the tumor and the patient) rationally-designed against actionable targets will have, most likely in combination with other strategies, namely immunotherapies, the potential to effectively cure cancer patients.
Take Home Message
The following points were covered in this chapter:
-
Signaling pathways are essential for cellular homeostasis and their deregulation is often associated with cancer.
-
Signaling pathways are excellent targets for therapeutic intervention in cancer. However, therapies targeting signaling players also have caveats – development of resistance being the most important.
-
There are numerous mechanisms of resistance to signaling therapies, but also many rational strategies to overcome them.
-
In the ongoing fight to minimize the likelihood of development of resistance, to decrease toxicities and to maximize anti-cancer effects, it is of utmost importance to have a deep knowledge of the mechanisms of signal transduction, and the molecular characteristics of the patient’s tumor, so that rationally-designed therapies can be implemented and potential resistance predicted and managed.
Questions
-
1.
Explain briefly what signal transduction is and why it is so important for the cells.
-
2.
Give few examples of input signals and output responses.
-
3.
List the main post-translational protein modifications in signal transduction and briefly explain the most characterized one.
-
4.
List the main categories of human protein kinases and give an example in each category.
-
5.
Why is signaling transduction so important in cancer? Indicate three reasons.
-
6.
Indicate the three signaling pathways frequently activated in all types of cancer and what mainly contributes to their hyper-activation in cancer.
-
7.
What are “signaling therapies” and their advantages?
-
8.
Explain briefly one mechanism of resistance to targeted signaling therapies, using one example.
-
9.
What are the common strategies to overcome resistance to signaling therapies?
References
Pawson T, Nash P (2000) Protein-protein interactions define specificity in signal transduction. Genes Dev 14:1027–1047
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signalling network. Nat Rev Mol Cell Biol 2:127–137. https://doi.org/10.1038/35052073
Adjei AA, Hidalgo M (2005) Intracellular signal transduction pathway proteins as targets for cancer therapy. J Clin Oncol 23:5386–5403. https://doi.org/10.1200/JCO.2005.23.648
Blume-Jensen P, Hunter T (2001) Oncogenic kinase signalling. Nature 411:355–365. https://doi.org/10.1038/35077225
Goldstein BJ (1992) Protein-tyrosine phosphatases and the regulation of insulin action. J Cell Biochem 48:33–42. https://doi.org/10.1002/jcb.240480107
Luo J, Solimini NL, Elledge SJ (2009) Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 136:823–837. https://doi.org/10.1016/j.cell.2009.02.024
Hopkins AL, Groom CR (2002) The druggable genome. Nat Rev Drug Discov 1:727–730. https://doi.org/10.1038/nrd892
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144:646–674. https://doi.org/10.1016/j.cell.2011.02.013
Rane SG, Reddy EP (2000) Janus kinases: components of multiple signaling pathways. Oncogene 19:5662–5679. https://doi.org/10.1038/sj.onc.1203925
O’Shea JJ et al (2015) The JAK-STAT pathway: impact on human disease and therapeutic intervention. Annu Rev Med 66:311–328. https://doi.org/10.1146/annurev-med-051113-024537
Leonard WJ, O’Shea JJ (1998) Jaks and STATs: biological implications. Annu Rev Immunol 16:293–322. https://doi.org/10.1146/annurev.immunol.16.1.293
Bowman T, Garcia R, Turkson J, Jove R (2000) STATs in oncogenesis. Oncogene 19:2474–2488. https://doi.org/10.1038/sj.onc.1203527
Lord JD, McIntosh BC, Greenberg PD, Nelson BH (2000) The IL-2 receptor promotes lymphocyte proliferation and induction of the c-myc, bcl-2, and bcl-x genes through the trans-activation domain of Stat5. J Immunol 164:2533–2541
Roux PP, Blenis J (2004) ERK and p38 MAPK-activated protein kinases: a family of protein kinases with diverse biological functions. Microbiol Mol Biol Rev 68:320–344. https://doi.org/10.1128/MMBR.68.2.320-344.2004
Dhillon AS, Hagan S, Rath O, Kolch W (2007) MAP kinase signalling pathways in cancer. Oncogene 26:3279–3290. https://doi.org/10.1038/sj.onc.1210421
Smith JA, Poteet-Smith CE, Malarkey K, Sturgill TW (1999) Identification of an extracellular signal-regulated kinase (ERK) docking site in ribosomal S6 kinase, a sequence critical for activation by ERK in vivo. J Biol Chem 274:2893–2898
Waskiewicz AJ, Flynn A, Proud CG, Cooper JA (1997) Mitogen-activated protein kinases activate the serine/threonine kinases Mnk1 and Mnk2. EMBO J 16:1909–1920. https://doi.org/10.1093/emboj/16.8.1909
Deak M, Clifton AD, Lucocq LM, Alessi DR (1998) Mitogen- and stress-activated protein kinase-1 (MSK1) is directly activated by MAPK and SAPK2/p38, and may mediate activation of CREB. EMBO J 17:4426–4441. https://doi.org/10.1093/emboj/17.15.4426
Davis RJ (1995) Transcriptional regulation by MAP kinases. Mol Reprod Dev 42:459–467. https://doi.org/10.1002/mrd.1080420414
Kerkhoff E, Rapp UR (1998) Cell cycle targets of Ras/Raf signalling. Oncogene 17:1457–1462. https://doi.org/10.1038/sj.onc.1202185
Barata JT, Cardoso AA, Boussiotis VA (2005) Interleukin-7 in T-cell acute lymphoblastic leukemia: an extrinsic factor supporting leukemogenesis? Leuk Lymphoma 46:483–495. https://doi.org/10.1080/10428190400027852
Song G, Ouyang G, Bao S (2005) The activation of Akt/PKB signaling pathway and cell survival. J Cell Mol Med 9:59–71
Cantley LC (2002) The phosphoinositide 3-kinase pathway. Science 296:1655–1657. https://doi.org/10.1126/science.296.5573.1655
Laplante M, Sabatini DM (2012) mTOR signaling in growth control and disease. Cell 149:274–293. https://doi.org/10.1016/j.cell.2012.03.017
Vander Haar E, Lee SI, Bandhakavi S, Griffin TJ, Kim DH (2007) Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat Cell Biol 9:316–323. https://doi.org/10.1038/ncb1547
Shaw RJ, Cantley LCR (2006) PI(3)K and mTOR signalling controls tumour cell growth. Nature 441:424–430. https://doi.org/10.1038/nature04869
Richter JD, Sonenberg N (2005) Regulation of cap-dependent translation by eIF4E inhibitory proteins. Nature 433:477–480. https://doi.org/10.1038/nature03205
Ma XM, Blenis J (2009) Molecular mechanisms of mTOR-mediated translational control. Nat Rev Mol Cell Biol 10:307–318. https://doi.org/10.1038/nrm2672
Roskoski R Jr (2004) The ErbB/HER receptor protein-tyrosine kinases and cancer. Biochem Biophys Res Commun 319:1–11. https://doi.org/10.1016/j.bbrc.2004.04.150
Rowinsky EK (2004) The erbB family: targets for therapeutic development against cancer and therapeutic strategies using monoclonal antibodies and tyrosine kinase inhibitors. Annu Rev Med 55:433–457. https://doi.org/10.1146/annurev.med.55.091902.104433
Van Cutsem E et al (2009) Cetuximab and chemotherapy as initial treatment for metastatic colorectal cancer. N Engl J Med 360:1408–1417. https://doi.org/10.1056/NEJMoa0805019
Engelman JA (2009) Targeting PI3K signalling in cancer: opportunities, challenges and limitations. Nat Rev Cancer 9:550–562. https://doi.org/10.1038/nrc2664
Flex E et al (2008) Somatically acquired JAK1 mutations in adult acute lymphoblastic leukemia. J Exp Med 205:751–758. https://doi.org/10.1084/jem.20072182
Furqan M, Mukhi N, Lee B, Liu D (2013) Dysregulation of JAK-STAT pathway in hematological malignancies and JAK inhibitors for clinical application. Biomark Res 1:5. https://doi.org/10.1186/2050-7771-1-5
Waibel M et al (2013) Combined targeting of JAK2 and Bcl-2/Bcl-xL to cure mutant JAK2-driven malignancies and overcome acquired resistance to JAK2 inhibitors. Cell Rep 5:1047–1059. https://doi.org/10.1016/j.celrep.2013.10.038
Kontro M et al (2014) Novel activating STAT5B mutations as putative drivers of T-cell acute lymphoblastic leukemia. Leukemia 28:1738–1742. https://doi.org/10.1038/leu.2014.89
Janes MR et al (2010) Effective and selective targeting of leukemia cells using a TORC1/2 kinase inhibitor. Nat Med 16:205–213. https://doi.org/10.1038/nm.2091
Fan QW et al (2006) A dual PI3 kinase/mTOR inhibitor reveals emergent efficacy in glioma. Cancer Cell 9:341–349. https://doi.org/10.1016/j.ccr.2006.03.029
O’Dwyer ME, Mauro MJ, Druker BJ (2002) Recent advancements in the treatment of chronic myelogenous leukemia. Annu Rev Med 53:369–381. https://doi.org/10.1146/annurev.med.53.082901.103853
Shah NP et al (2002) Multiple BCR-ABL kinase domain mutations confer polyclonal resistance to the tyrosine kinase inhibitor imatinib (STI571) in chronic phase and blast crisis chronic myeloid leukemia. Cancer Cell 2:117–125
Shah NP, Sawyers CL (2003) Mechanisms of resistance to STI571 in Philadelphia chromosome-associated leukemias. Oncogene 22:7389–7395. https://doi.org/10.1038/sj.onc.1206942
Gorre ME et al (2001) Clinical resistance to STI-571 cancer therapy caused by BCR-ABL gene mutation or amplification. Science 293:876–880. https://doi.org/10.1126/science.1062538
Kantarjian H et al (2010) Dasatinib versus imatinib in newly diagnosed chronic-phase chronic myeloid leukemia. N Engl J Med 362:2260–2270. https://doi.org/10.1056/NEJMoa1002315
Tran KA et al (2016) MEK inhibitors and their potential in the treatment of advanced melanoma: the advantages of combination therapy. Drug Des Devel Ther 10:43–52. https://doi.org/10.2147/DDDT.S93545
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Barata, J.T., Oliveira, M.L. (2019). Cell Signaling in Cancer. In: Fior, R., Zilhão, R. (eds) Molecular and Cell Biology of Cancer. Learning Materials in Biosciences. Springer, Cham. https://doi.org/10.1007/978-3-030-11812-9_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-11812-9_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-11811-2
Online ISBN: 978-3-030-11812-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)