Abstract
Cancer is the product of natural selection happening within our bodies. Our cells can be seen as individuals in populations, able to mature, reproduce with mutation, and die. All it takes is chance (and external factors such as smoking habits) for the emergence of mutant cells that divide uncontrollably and take over the whole tissue, creating aggressive tumors. If we understand the roots of this process, we will be able to develop better therapies that target the uniqueness of each cancer. In this chapter we will discuss how cancer is modulated both by somatic and germline evolution and we will examine the consequences of interpreting tumor progression as an adaptive process.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
We will discuss how natural selection is not restricted to populations of organisms but can in fact happen in cells and tissues. We will see how there is an antagonism between genes selected during somatic evolution and those that were selected during human evolution. We will glimpse at how the forces of natural selection shape tumors and their diversity. At the end of the chapter we will have an idea how each cancer is the result of a unique evolutionary process that depends on the host (patient) but also on the environment and on chance.
FormalPara Learning ObjectivesAfter reading this chapter students should be able to:
-
1.
Name/describe the basic ingredients for evolution to occur.
-
2.
Distinguish between somatic and germline evolution.
-
3.
View cancer as an outcome of natural selection acting within tissues.
-
4.
Understand that not all diversity is advantageous for the tumor, i.e., there are driver and passenger mutations.
-
5.
Glimpse into the origins of the enormous diversity we observe in tumors.
-
6.
Understand why each cancer is a unique disease.
-
7.
Consider how we can use tools from evolutionary biology to fight cancer.
-
Mutation – change in the DNA sequence.
-
Selection – differences in the ability of a cell or organism to reproduce.
-
Fitness – measure of reproductive ability (selection). It is often equated to growth rate.
-
Genetic drift – changes in the frequency of mutations caused by chance.
-
Selective sweep – rapid increase in frequency of a mutation due to selection.
-
Antagonistic pleiotropy – pleiotropy is the observation that one gene can affect multiple traits. It is antagonistic when those traits affect natural selection in opposite directions.
-
Epistasis – the fact that mutations have different effects depending on the rest of the genome. It comes mostly from the interaction between proteins.
-
Drivers and passengers – Driver mutations are mutations that cause a selective sweep. Passenger mutations are mutations that are present in the same genome as drivers and hence also increase in frequency. Usually they do not contribute to fitness.
2.1 Basic Ingredients of an Evolutionary Process
Evolution is inescapable in replicating biological systems: whenever there are entities capable of self-replication, subject to mutations that cause phenotypic change, there will be evolution. These entities can be organisms or groups of cells, including those that compose the somatic tissues of animals. In general, the changes that take place within somatic tissues are not represented in the germline. So, by definition, evolution within a somatic lineage is finite. However, the changes produced by somatic evolution may have terrible consequences to the individual.
In this chapter, we will review how somatic evolution can lead to cancer, particularly to very aggressive cancers. We will see how evolution can make each cancer unique and therefore not conform to a solution (therapy) that fits all.
2.2 Multi-cellular Organisms: The Clash of Two Evolutionary Processes
Throughout the lifetime of a multi-cellular organism, each cell goes through thousands or millions of cell divisions. In each division there is a high chance that at least one mutation will occur (for example, an average of eight mutations per cell division in human fibroblasts [1]). These mutations may be innocuous, i.e., have no effect on the behavior of the cell, or they can significantly alter its phenotype. Because of the latter type of mutations, eukaryotic cells have a number of checkpoints that make sure the cell cycle stops once a major error is detected. That error is then either repaired or the cell dies. Moreover, there are external mechanisms such as the action of the immune system which detect cells that are “misbehaving” and eliminate them. All of these systems are the result of natural selection acting at the level of the organism. They evolved and were maintained because they make sure all the cells behave in such a way as to maximize the survival and reproduction of the organism. Hence, the genome of a species is shaped by natural selection to maximize the survival of the germline!
Multi-cellular organisms thus evolve in two very different ways and at different time-scales: somatic evolution at the time scale of days and germline evolution at the time-scale of generations (decades in humans). Moreover, somatic evolution is self-contained. In general, whatever evolved within an organism dies with it. Curious exceptions have been found such as the rare transmissible cancers, as facial tumors of the Tasmanian devil [2] and a type of leukemia-like disease in clams [3]. In these two examples, cancer cells are transmitted horizontally from individual to individual, much like an infectious disease. But these are the exceptions that prove the rule. In general, somatic evolution is finite while germline evolution goes on forever unless the population goes extinct. In addition to the difference in timescales, there are also major differences on the selective pressures acting on soma and germline. In the somatic tissue, the cells that are over-represented will be those that proliferate faster. Fast proliferation can lead to organ-malfunction and cancer which shorten the life expectancy and the ability of the organism to leave progeny. Therefore, mutations that increase cellular proliferation will be selected-for in the somatic tissue but selected-against in the germline.
Due to this conflict between somatic and germline evolution, the germline acquired genes which reduce the probability of evolution within the soma. The most well-known are tumor-suppressor genes which you will read about in the other chapters of this book. These are genes which kill the cells when they behave abnormally. One notable example of the acquisition of tumor-suppressor genes in the germline is the Elephant. These are large animals and therefore it takes many somatic cell divisions to produce an adult. For this reason, there should be a general positive correlation between cancer incidence and body size. However, that is not the case, an observation known as Peto’s paradox [4]. The solution that natural selection found for the Elephants was the acquisition of extra copies of the tumor suppressor gene p53 [5]. Moreover, during evolution, Elephants reactivated a gene that had been made silent (a pseudogene) in the Paenungulate ancestor and which also has tumor-suppressor activities [6]. There are other mechanisms such as physical barriers to proliferation. Since we know these mechanisms can be very effective, one can ask, “why do animals still have cancer?”.
As stated above, adaptive evolution maximizes the probability of survival of the germline. However, once an organism has produced its offspring, there is no longer evolutionary pressure to keep it alive. Let’s imagine an allele (a variant of a gene) that increases the production of sperm. This allele will increase fertility, its carriers will have plenty of offspring and soon the allele will dominate the population. Now let’s say that this allele, in addition to increasing sperm count, also leads to a moderate increase in mutation rate in skin cells. This increases the chances of skin cancer. But if the tumors only occur after the reproductive period, by the time the individuals die of skin cancer, their children are already part of the population. In other words, the allele that caused the increase in skin cancer already spread to the next generation and it does not matter that the individual dies as long as the germ cell has passed. It is due to this problem of time scale that genes that are deleterious late in life, but neutral or beneficial early in life, survive. ◘ Figure 2.1 is a schematic representation of this decrease in the strength of natural selection as time passes.
Schematic representation of the strength of natural selection acting on phenotypes as a function of the age of the individuals expressing those phenotypes. In either panel the axes reflect there two measures and the line shows how natural selection is very strong up to the age of reproduction and becomes much weaker after that. Panel a represents a feature which is neutral early in life but deleterious later. Mutations causing this phenotype may remain in the population because natural selection is too weak to remove them. Panel b represents the antagonistic pleiotropy hypothesis whereby features which are beneficial early and deleterious late will be selected for because natural selection is strongest when they show their benefit. (Adapted from Fabian and Flatt [7])
There are two types of mutations that escape natural selection because of the timing of their actions:
-
Those that are neutral early in life but deleterious after reproduction;
-
And those that are beneficial early in life but deleterious later on, as in our imaginary example above.
There are then two competing hypotheses:
-
Mutation-accumulation hypothesis – that says most mutations that cause ageing-related diseases are of the first type
-
Antagonistic-pleiotropy hypothesis – that says the second type is more prevalent.
As often, reality is probably a mixture of the two. In humans and even in other mammals, it is difficult to identify these types of mutations but there are a few hints that they might exist. In mice, two short alleles of the gene coding for TP53 are described to both confer resistance to cancer and reduce lifespan [8]. Similarly, a study in a human population before access to contraception shows higher fertility in carriers of BRCA1 and BRCA2 mutations [9]. Both of these alleles are associated with increased rates of cancer. You will learn about the mode of action of BRCA as well as TP53 in other chapters of this book (► Chap. 5).
Taking all of this into account, it is easy to imagine that cancer is one of those diseases that was not counter-selected in our ancestors because it occurred well past the age of reproduction. This explains why humans are still plagued by it. It is nevertheless a very particular disease in that cancer in itself is an evolutionary process. It is as if each of us has in our body populations of cells that are fighting for survival, despite the fact that they will never be transmitted to the next generation of humans. Alas, natural selection is extremely short sighted.
2.3 Intra-tumor Diversity and Evolution
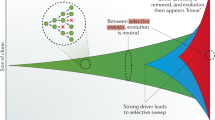
Much like any other evolving population, tumors are very diverse within themselves. Decades ago, before the advent of large-scale sequencing, there was an idea that cancer resulted from the sequential take-over of clonal populations, much like in a population of bacteria. One cell would become mutated and increase its numbers, taking over the population in what is known as a selective sweep. Once that was done, a second mutation would come along and begin its own rise in frequency and so on. This turned out not to be the case, neither for cancer, nor for bacteria. In fact, several mutants may arise at the same time and begin increasing in frequency. Since they cannot recombine and be put in the same genome, they are destined to either compete (in which case only one will win) or cooperate. In either case, for some time that may be quite long, there will be co-existence of both variants. Now, given the large mutation rate, there can be several of these mutants. All increasing the growth rate and all either competing or cooperating, or both. Moreover, in addition to these mutations that increase the growth rate, there are also those that are either neutral or deleterious (decreasing the growth rate). Under normal circumstances, these mutations would either be purged or increase very slowly. However, when combined with mutations with strong positive effects they will hitchhike and become quite frequent as well. This is where the distinction between driver and passenger mutations comes in. Drivers are those mutations that cause the tumor to increase in size or in survival and passengers are those that do not do anything to the tumor. ◘ Figure 2.2 shows an example of how driver mutations compete and how passengers (or hitchhikers) can increase in frequency as well.
Representation of the frequency of mutations in a tumor over time. The colors represent different mutations and the area shows their frequency at each time point. The grey bubbles are passenger mutations that appear and only have a chance of increasing in frequency if one of the cells carrying them also acquires a driver mutation. Note how certain mutations that are beneficial such as p53−/− appear and are lost if they have to compete against stronger mutations such as p16−/−. (Adapted from Maley and Reid [10])
It is now known that there are many drivers and passengers within a tumor. However, successful therapies must target drivers and not passengers. Therefore, identifying the drivers is one of the most important yet challenging tasks of modern day’s genetics. To make matters more complicated, passenger mutations can interact with drivers and with each other and contribute to tumor progression by conveying different phenotypic traits. For instance, imagine a tumor that has a drug efflux pump as a passenger mutation; while the tumor is growing, and the patient is not under therapy, the pump does not give any advantage. However, when therapy starts the clones that have this passenger mutation can become selected. Hence, the definition of driver and passenger, while very useful, it is not absolute.
2.4 Sources of Diversity and the Impact of Mutations on Fitness
Errors are inevitable in any replicating system. Mutations are errors in DNA which are not repaired by the cell. Indeed, cells have a number of mechanisms to repair errors in DNA (see ► Chap. 5). These include, for example, mismatch repair proteins, which detect mismatches between the two DNA strands and correct them. Still these systems are not perfect and allow a certain number of mutations to pass to the next cell generation. We call the mutation rate, the number of mutations that occur per cell, per generation. Mutation rates can be expressed per nucleotide or per genome. Mutation rates vary between cells and between organisms and it is not known what selects them to be a certain number. For the germline it is thought that the mutation rate is as low as it can be without compromising the normal functioning of the cell – if the cell spends all its resources in repair, it cannot do other essential functions such as metabolism. For somatic tissues, the mutation rate depends on the type of cell. Fibroblasts, for example, have a rate 100-fold higher than the germline [1]. The precise reason for this is not known. One trivial hypothesis relates to the antagonistic pleiotropy described above. Somatic cells need to replicate fast in order for development to occur. Moreover, processes such as wound healing require quick and efficient production of new cells. Diverting energy to speed means there is less time and fewer resources for repair. There has to be a fine balance between speed and repair. Any mutation happening in the germline or in early development will mean the death of the organism. However, mutations occuring in differentiated cells like adult fibroblasts will have little impact on reproduction and hence these cells have evolved to spend less resources on repair and more on proliferation. In addition to spontaneous mutations, the exposure of humans to chemicals, radiation or other external conditions which increase the DNA error rate also leads to increase in the mutation rate. The two most well-known examples are smoking, which increases the probability of a number of cancers, partially due to an increase in mutation rate [11]; and skin cancer, where the appearance of tumors is clearly related to exposure to radiation from the sun, which also leads to an increased mutation rate [12].
2.5 Mutation Spectra and the Fitness Effects of Mutations
Independently of what causes them, there are different types of mutations. There are single nucleotide changes (SNPs – the ‘P’ stands for polymorphism and is reminiscent of a time when mutation rates were estimated from polymorphic DNA variants in populations), insertions and deletions, structural variants such as inversions, translocations, duplications and large deletions, whole chromosome gains and losses. Indeed, when looking at cancers, all of these mutations are visible [11].
Having more mutations is not in itself a good thing for the cancer cell. Being good, or bad as we saw earlier, depends a lot on the level of organization we look at. In this case, a mutation is good for cancer if it increases the survival and the growth rate of cancer cells. In other words, a mutation is good for cancer if it increases its fitness. Most mutations are neutral or deleterious, i.e., they do not affect fitness, or they decrease it. This is true even for cancer (or so we think). The rationale behind this statement is that when we make a random change to a functioning object, we very rarely make it better. Imagine hitting a functioning car with a very large hammer. What are the odds it will go faster? In addition to these arguments, there is also evidence form bacteria, yeast, Caenorhabditis elegans among others, which were grown in the lab in the absence of natural selection. Under these conditions their fitness decreases, indicating that most spontaneous mutations that affect fitness are deleterious. See for example Kibota, T. T., & Lynch, M. (1996) [13].
2.6 Constraints to Evolution/Genetic Drift
So far, I have equated evolution with adaptation. However, evolution is not always positive. Evolution is a change in the frequency of genetic variants with time. It does not mean that it has to be positive. And indeed, not all the diversity we observe is beneficial. In fact, we expect that some of the mutations we observe in cancer are deleterious (i.e., they decrease the tumor’s fitness). These mutations may be passengers that increased in frequency because they happened to be in the same genome as a beneficial mutation; or they might have increased in frequency by chance, i.e., via the process of genetic drift. Any process that slows down natural selection will increase genetic drift. In general, genetic drift is the result of either a high mutation rate or of demographic constraints such as small population sizes or the presence of spatial structure. In the case of a high mutation rate, genetic drift becomes stronger than natural selection when deleterious mutations happen faster than natural selection can eliminate them. One approach to eliminate cancer is actually to increase its mutation rate [14]. The problem is that a moderate increase in mutation rate actually aids cancer by producing more driver mutations. Therefore, any such therapy must ensure the increase is large enough. The other way to increase genetic drift is by interfering with demography. One option is to reduce the number of cells that compose the tumor. If the number of cells is small enough, the chance of producing driver mutations is low and instead it is easier for deleterious mutations to increase in frequency. In the same way, the existence of spatial structure (e.g., in epithelia) also leads to an increase in genetic drift. Any driver mutation that comes along will take longer to take-over the whole population because it can only compete with a few neighboring cells at a time. It is as if the total population size was smaller [15]. Many tumors come from epithelia. The most likely reason is the extent of cell turnover in these tissues, which probably leads to an increased mutation rate. Evolutionary theory predicts that, if not for the spatial structure, these tumors would be even more numerous and aggressive.
The role of spatial structure in preventing somatic evolution was likely co-opted by germline evolution to prevent tumors in highly proliferative tissues such as the intestinal epithelium. There, cell division is restricted to a small niche in the crypts. As cells divide, they migrate through the villi and are eventually shed. This means that any mutation acquired during this process will be pushed away and out of the tissue, even if it divides very quickly. ◘ Figure 2.3 shows a schematic representation of the intestinal epithelium and of the path cells take during normal tissue function.
The organization of the epithelial tissue. The only cells that remain in the tissue are located in the crypt. The rest are pushed away as these cells divide and eventually are shed into the external environment where they die (anoikis). Any mutation that occurs outside the crypt will be lost, even if it leads to increased proliferation. (Adapted from Barker [16])
2.7 Why Cancer Is Not a Single Disease and Why We Have Not Cured It: An Evolutionist’s Perspective
Just as no two humans are alike, no two cancers are alike, even within the same patient. Cancer is the result of chance, selection, genetic and biophysical constraints. Whether or not a particular mutation contributes to cancer or not is highly dependent on the genetic background (a phenomenon known as epistasis). Therefore, a tumor in one individual will have a different set of driver mutations than a tumor in someone else. In the same manner, chance events will be very different between any two patients. All of this points to a fact that medical doctors have been aware of for some time: each tumor is a unique disease that must be studied and treated individually. In fact, nowadays tumor genotyping is becoming common practice and new tumor biomarkers are helping to improve clinical outcomes by directing therapies. However, most biomarkers are not 100% prognostic. This is because we are still far from predicting the output of the interaction of the different mutations (plus the intronic mutations, epigenetic modifications, etc.), making it unpredictable to know which is, the best therapy for each individual patient. This is why different functional tests are being developed to challenge patient tumor cells with the different therapy options to select the best one, just like antibiograms for bacterial infections. These tests include xenografting patient tumor cells into animal models like zebrafish and mouse, generating organoids or cultivating tumor explants in the presence of the therapeutic options and then evaluating their response. Let’s hope one of these tests work.
Take Home Message
This chapter covers the following points:
-
Cells within tissues evolve in much the same way as populations of organisms. This is known as somatic evolution.
-
Somatic evolution occurs in a shorter timescale than organismal (germline) evolution. Moreover, the phenotypes that are selected in one are often opposite. For example, somatic evolution may select for fast cell division and low repair mechanisms while the opposite will be selected for in the germline.
-
Somatic evolution is finite, and the mutations selected during this process will die with the organism.
-
Germline evolution is not able to eliminate all mutations that cause disease. Namely, for diseases that only happen after the age of reproduction (like cancer), natural selection is not very effective.
-
Mutations may accumulate in the germline that make the individual prone to cancer, either because they have no effect on the reproductive output, or because they actually increase fertility early in life.
-
A tumor is the result of a very complex evolutionary process where both natural selection and genetic drift are at play. Not all mutations found in a cancer contribute to the tumor.
-
Each tumor is unique just as each human is unique. Therapies should take this into account.
Questions
-
1.
What are the minimum requirements for evolution?
-
2.
Give some example of genes that may be subject to antagonistic pleiotropy.
-
3.
How would you identify mutations that are causative of a tumor?
References
Milholland B, Dong X, Zhang L, Hao X, Suh Y, Vijg J (2017) Differences between germline and somatic mutation rates in humans and mice. Nat Commun 8:15183. https://doi.org/10.1038/ncomms15183. PubMed PMID: 28485371; PubMed Central PMCID:PMC5436103
Mozos E, Méndez A, Gómez-Villamandos JC, Martín De Las Mulas J, Pérez J (1996) Immunohistochemical characterization of canine transmissible venereal tumor. Vet Pathol 33(3):257–263
Metzger MJ, Villalba A, Carballal MJ, Iglesias D, Sherry J, Reinisch C, Muttray AF, Baldwin SA, Goff SP (2016) Widespread transmission of independent cancer lineages within multiple bivalve species. Nature 534(7609):705–709. Epub 2016 Jun 22. PubMed PMID: 27338791; PubMed Central PMCID: PMC4939143
Peto R (2015) Quantitative implications of the approximate irrelevance of mammalian body size and lifespan to lifelong cancer risk. Philos Trans R Soc Lond Ser B Biol Sci 370(1673):pii: 20150198. https://doi.org/10.1098/rstb.2015.0198. PubMed PMID: 26056360; PubMed Central PMCID: PMC4581039
Sulak M, Fong L, Mika K, Chigurupati S, Yon L, Mongan NP, Emes RD, Lynch VJ (2016) TP53 copy number expansion is associated with the evolution of increased body size and an enhanced DNA damage response in elephants. elife 5:pii:e11994. https://doi.org/10.7554/eLife.11994
Vazquez JM, Sulak M, Chigurupati S, Lynch VJ (2018) A zombie LIF gene in elephants is upregulated by TP53 to induce apoptosis in response to DNA damage. Cell Rep 24(7):1765–1776. https://doi.org/10.1016/j.celrep.2018.07.042
Fabian D, Flatt T (2011) The evolution of aging. Nat Educ Knowl 3(10):9
Ungewitter E, Scrable H (2009) Antagonistic pleiotropy and p53. Mech Ageing Dev 130(1–2):10–17. https://doi.org/10.1016/j.mad.2008.06.002. Epub 2008 Jul 1. Review. PubMed PMID: 18639575; PubMed Central PMCID: PMC2771578
Smith KR, Hanson HA, Hollingshaus MS (2013) BRCA1 and BRCA2 mutations and female fertility. Curr Opin Obstet Gynecol 25(3):207–213. https://doi.org/10.1097/GCO.0b013e32835f1731. Review. PubMed PMID: 23411475; PubMed Central PMCID: PMC4010322
Maley CC, Reid BJ (2005) Natural selection in neoplastic progression of Barrett’s esophagus. Semin Cancer Biol 15(6):474–483. Review
de Bruin EC, McGranahan N, Mitter R, Salm M, Wedge DC, Yates L, Jamal-Hanjani M, Shafi S, Murugaesu N, Rowan AJ, Grönroos E, Muhammad MA, Horswell S, Gerlinger M, Varela I, Jones D, Marshall J, Voet T, Van Loo P, Rassl DM, Rintoul RC, Janes SM, Lee SM, Forster M, Ahmad T, Lawrence D, Falzon M, Capitanio A, Harkins TT, Lee CC, Tom W, Teefe E, Chen SC, Begum S, Rabinowitz A, Phillimore B, Spencer-Dene B, Stamp G, Szallasi Z, Matthews N, Stewart A, Campbell P, Swanton C (2014) Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346(6206):251–256. https://doi.org/10.1126/science.1253462. PubMed PMID: 25301630; PubMed Central PMCID: PMC4636050
Berneburg M, Gattermann N, Stege H, Grewe M, Vogelsang K, Ruzicka T, Krutmann J (1997) Chronically ultraviolet-exposed human skin shows a higher mutation frequency of mitochondrial DNA as compared to unexposed skin and the hematopoietic system. Photochem Photobiol 66(2):271–275
Kibota TT, Lynch M (1996) Estimate of the genomic mutation rate deleterious to overall fitness in E. coli. Nature 381(6584):694–696
Prindle MJ, Fox EJ, Loeb LA (2010) The mutator phenotype in cancer: molecular mechanisms and targeting strategies. Curr Drug Targets 11(10):1296–1303. Review. PubMed PMID: 20840072; PubMed Central PMCID: PMC4073693
Martens EA, Kostadinov R, Maley CC, Hallatschek O (2011) Spatial structure increases the waiting time for cancer. New J Phys 13:pii: 115014. Epub 2011 Nov 28. PubMed PMID: 22707911; PubMed Central PMCID: PMC3375912
Barker N (2014) Adult intestinal stem cells: critical drivers of epithelial homeostasis and regeneration. Nat Rev Mol Cell Biol 15(1):19–33. https://doi.org/10.1038/nrm3721. Epub 2013 Dec 11. Review
Further Reading
Leroi AM, Bartke A, De Benedictis G, Franceschi C, Gartner A, Gonos ES, Fedei ME, Kivisild T, Lee S, Kartaf-Ozer N, Schumacher M, Sikora E, Slagboom E, Tatar M, Yashin AI, Vijg J, Zwaan B (2005) What evidence is there for the existence of individual genes with antagonistic pleiotropic effects? Mech Ageing Dev 126(3):421–429. Review
Lipinski KA, Barber LJ, Davies MN, Ashenden M, Sottoriva A, Gerlinger M (2016) Cancer evolution and the limits of predictability in precision cancer medicine. Trends Cancer 2(1):49–63. https://doi.org/10.1016/j.trecan.2015.11.003. Epub 2016 Jan 29. Review. PubMed PMID: 26949746; PubMed Central PMCID: PMC4756277
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Perfeito, L. (2019). Cancer as an Evolutionary Process. In: Fior, R., Zilhão, R. (eds) Molecular and Cell Biology of Cancer. Learning Materials in Biosciences. Springer, Cham. https://doi.org/10.1007/978-3-030-11812-9_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-11812-9_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-11811-2
Online ISBN: 978-3-030-11812-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)