Abstract
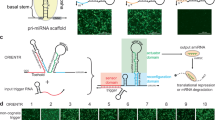
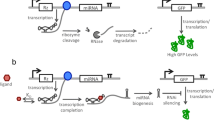
MicroRNAs (miRNAs) offer powerful tools for targeted gene silencing in almost all eukaryotes. These tools have received considerable attention for their utility in both fundamental genetic studies and as therapeutic agents. Rendering individual microRNAs responsive to endogenous or exogenously applied molecules (or ligands) can improve the stringency of silencing and can mediate autonomous control. This chapter describes the construction of ligand-responsive miRNAs that undergo reduced processing and subsequent gene silencing when bound by the recognized ligand. Following a simple set of rules, the engineered microRNAs can be readily modified to target different sequences and to bind different ligands. Individual miRNAs also can be incorporated into the same transcript for tunable, multi-gene silencing.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Gu S, Kay MA (2010) How do miRNAs mediate translational repression? Silence 1:11
Alvarez-Garcia I, Miska EA (2005) MicroRNA functions in animal development and human disease. Development 132:4653–4662
Leisner M, Bleris L, Lohmueller J, Xie Z, Benenson Y (2010) Rationally designed logic integration of regulatory signals in mammalian cells. Nat Nanotechnol 5:666–670
Dickins RA, Hemann MT, Zilfou JT, Simpson DR, Ibarra I, Hannon GJ, Lowe SW (2005) Probing tumor phenotypes using stable and regulated synthetic microRNA precursors. Nat Genet 37:1289–1295
Tuerk C, Gold L (1990) Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249:505–510
Ellington AD, Szostak JW (1990) In vitro selection of RNA molecules that bind specific ligands. Nature 346:818–822
Hermann T, Patel DJ (2000) Adaptive recognition by nucleic acid aptamers. Science 287:820–825
Jenison RD, Gill SC, Pardi A, Polisky B (1994) High-resolution molecular discrimination by RNA. Science 263:1425–1429
Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, Sohn SY, Cho Y, Zhang BT, Kim VN (2006) Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125:887–901
Zeng Y, Cullen BR (2005) Efficient processing of primary microRNA hairpins by Drosha requires flanking nonstructured RNA sequences. J Biol Chem 280:27595–27603
Zeng Y, Yi R, Cullen BR (2005) Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J 24:138–148
Beisel CL, Chen YY, Culler SJ, Hoff KG, Smolke CD (2011) Design of small molecule-responsive microRNAs based on structural requirements for Drosha processing. Nucleic Acids Res 39:2981–2994
Petri S, Meister G (2013) siRNA design principles and off-target effects. Methods Mol Biol 986:59–71
An CI, Trinh VB, Yokobayashi Y (2006) Artificial control of gene expression in mammalian cells by modulating RNA interference through aptamer-small molecule interaction. RNA 12:710–716
Acknowledgments
This work was supported by North Carolina State University (start-up funds to CLB), the National Science Foundation (fellowship to RJB), the National Institutes of Health (RC1GM091298), and the Defense Advanced Research Projects Agency (HR0011-11-2-0002).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Beisel, C.L., Bloom, R.J., Smolke, C.D. (2014). Construction of Ligand-Responsive MicroRNAs that Operate Through Inhibition of Drosha Processing. In: Ogawa, A. (eds) Artificial Riboswitches. Methods in Molecular Biology, vol 1111. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-755-6_19
Download citation
DOI: https://doi.org/10.1007/978-1-62703-755-6_19
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-754-9
Online ISBN: 978-1-62703-755-6
eBook Packages: Springer Protocols