Abstract
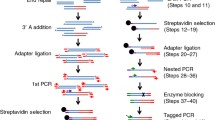
Current techniques for examining the global creation and repair of DNA double-strand breaks are restricted in their sensitivity, and such techniques mask any site-dependent variations in breakage and repair rate or fidelity. We present here a system for analyzing the fate of documented DNA breaks, using the MLL gene as an example, through application of ligation-mediated PCR. Here, a simple asymmetric double-stranded DNA adapter molecule is ligated to experimentally induced DNA breaks and subjected to seminested PCR using adapter- and gene-specific primers. The rate of appearance and loss of specific PCR products allows detection of both the break and its repair. Using the additional technique of inverse PCR, the presence of misrepaired products (translocations) can be detected at the same site, providing information on the fidelity of the ligation reaction in intact cells. Such techniques may be adapted for the analysis of DNA breaks and rearrangements introduced into any identifiable genomic location. We have also applied parallel sequencing for the high-throughput analysis of inverse PCR products to facilitate the unbiased recording of all rearrangements located at a specific genomic location.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Super HJ, McCabe NR, Thirman MJ et al (1993) Rearrangements of the MLL gene in therapy-related acute myeloid leukemia in patients previously treated with agents targeting DNA-topoisomerase II. Blood 82:3705–3711
Rowley JD (1993) Rearrangements involving chromosome band 11Q23 in acute leukaemia. Semin Cancer Biol 4:377–385
Aplan PD, Lombardi DP, Ginsberg AM, Cossman J, Bertness VL, Kirsch IR (1990) Disruption of the human SCL locus by “illegitimate” V-(D)-J recombinase activity. Science 250:1426–1429
Tycko B, Sklar J (1990) Chromosomal translocations in lymphoid neoplasia: a reappraisal of the recombinase model. Cancer Cells 2:1–8
Schlissel MS (1998) Structure of nonhairpin coding-end DNA breaks in cells undergoing V(D)J recombination. Mol Cell Biol 18: 2029–2037
Staley K, Blaschke AJ, Chun J (1997) Apoptotic DNA fragmentation is detected by a semi-quantitative ligation-mediated PCR of blunt DNA ends. Cell Death Differ 4:66–75
Wyllie AH (1980) Glucocorticoid-induced thymocyte apoptosis is associated with endogenous endonuclease activation. Nature 284: 555–556
Richardson C, Jasin M (2000) Frequent chromosomal translocations induced by DNA double-strand breaks. Nature 405:697–700
Vaux DL, Strasser A (1996) The molecular biology of apoptosis. Proc Natl Acad Sci U S A 93:2239–2244
Longtine J, Fox E, Reynolds C, Sklar J (2001) Molecular analysis of DNA rearrangements in leukemias and non-Hodgkin’s lymphomas. Curr Protoc Hum Genet Chapter 10: Unit 10.4
Popescu NC, Zimonjic DB (1997) Molecular cytogenetic characterization of cancer cell alterations. Cancer Genet Cytogenet 93: 10–21
Metzker ML (2010) Sequencing technologies—the next generation. Nat Rev Genet 11:31–46
Tucker T, Marra M, Friedman JM (2009) Massively parallel sequencing: the next big thing in genetic medicine. Am J Hum Genet 85:142–154
Betti CJ, Villalobos MJ, Diaz MO, Vaughan AT (2001) Apoptotic triggers initiate translocations within the MLL gene involving the nonhomologous end joining repair system. Cancer Res 61:4550–4555
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120:621–623
Forrester HB, Yeh RF, Dewey WC (1999) A dose response for radiation-induced intrachromosomal DNA rearrangements detected by inverse polymerase chain reaction. Radiat Res 152:232–238
Forrester HB, Radford IR (1998) Detection and sequencing of ionizing radiation-induced DNA rearrangements using the inverse polymerase chain reaction. Int J Radiat Biol 74: 1–15
Le H, Singh S, Shih SJ et al (2009) Rearrangements of the MLL gene are influenced by DNA secondary structure, potentially mediated by topoisomerase II binding. Genes Chromosomes Cancer 48:806–815
Singh S, Le H, Shih SJ, Ho B, Vaughan AT (2010) Suberoylanilide hydroxamic acid modification of chromatin architecture affects DNA break formation and repair. Int J Radiat Oncol Biol Phys 76:566–573
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Humana Press
About this protocol
Cite this protocol
Singh, S., Shih, SJ., Vaughan, A.T.M. (2014). Detection of DNA Double-Strand Breaks and Chromosome Translocations Using Ligation-Mediated PCR and Inverse PCR. In: Keohavong, P., Grant, S. (eds) Molecular Toxicology Protocols. Methods in Molecular Biology, vol 1105. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-739-6_30
Download citation
DOI: https://doi.org/10.1007/978-1-62703-739-6_30
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-738-9
Online ISBN: 978-1-62703-739-6
eBook Packages: Springer Protocols