Abstract
RNA interference (RNAi) is a process of sequence-specific posttranscriptional gene silencing induced by double-strand RNA, and this phenomenon has been shown to function in higher organisms including mammals, and methods that exploit RNAi mechanisms have been developing. Recently, RNAi induced by short interfering siRNAs has been experimentally introduced as a cancer therapy and is expected to be developed as a nucleic acid-based medicine. Moreover, RNAi technology is used in biomarker-based screening, which is a new screening method based on transcriptional profiling to identify the specific transcriptional activities altered by the compounds of interest. In this chapter, we briefly review the mechanism of RNAi and discuss in detail some of the most recent findings concerning the administration of potential nucleic acid-based drugs. We next discuss several current clinical trials of RNAi therapies against cancers. Finally, we introduce a new high-throughput screening method based on transcriptional profiling for drug discovery. Current studies and clinical trials demonstrate that RNAi technology could establish a novel and promising therapeutic tool against cancers.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
13.1 Introduction
RNA interference (RNAi) is a process of sequence-specific posttranscriptional gene silencing induced by double-strand RNA (dsRNA), and this phenomenon was discovered in Caenorhabditis elegans (C. elegans) [1]. RNAi has been shown to function in higher organisms including mammals, and methods that exploit RNAi mechanisms have been developing. Aberrant expression of endogenous normal or mutant genes occurs in pathological conditions, resulting in alterations in signal pathways, cellular proliferation, and apoptosis. Posttranscriptional gene regulation by RNAi controls these alterations positively or negatively, and consequently RNAi has now been well established as a method for experimental analyses of gene function in vitro. Recently, short interfering RNA (siRNA), which induces RNAi, has been experimentally introduced as a cancer therapy and is expected to be developed as a nucleic acid-based medicine, and several clinical trials of RNAi therapies against cancers are ongoing. To develop nuclear medicine against cancers, we have two important issues to overcome: one is to select suitable gene targets and another is to develop effective drug delivery systems (DDSs). DDSs are divided into two categories: viral vector-based carriers and nonviral-based carriers. Although viral vectors are the most powerful tools for transfection so far, especially retroviral and lentiviral vectors randomly integrate into host cells’ DNA and those might induce insertional mutagenesis [2–4]. The use of nonviral DDSs including cationic liposomes [5, 6] and atelocollagen [7, 8] is preferred because it offers greater safety for clinical application than does the use of viral DDSs.
In addition to the development of a nucleic acid-based medicine, RNAi is put to practical use for a high-throughput screening for development of molecular targeting agents. The alternation of the related gene transcripts which are investigated after the knockdown of the targeted gene transcript by RNAi is compared with that of gene transcripts treated by compounds with unknown functions. The compounds which demonstrate the resemble alternation are recognized as molecular target compounds for the interested gene [9–11]. In this chapter, we discuss the application of RNAi for the development of medicine against cancers.
13.2 Mechanisms of RNAi
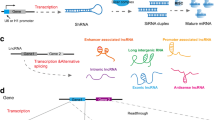
RNAi processes can be roughly divided into the initiation phase and the effector phase. In the initiation phase, following introduction of dsRNA into a target cell, dsRNA encounters a dsDNA-specific RNAse III family ribonuclease Dicer. Dicer is a modular enzyme and is composed of an N-terminal helicase domain, an RNA-binding Piwi/Argonaute/Zwille (PAZ) domain, two tandem RNAse III domains, and a dsRNA-binding domain [12]. Dicer acts to produce both siRNAs and microRNAs (miRNAs) [13–16]. dsRNA is processed into shorter lengths of 21–23 nucleotides (nts) dsRNAs, termed siRNAs by the ribonuclease activity of Dicer. dsRNA precursors are sequentially processed by the two RNAse III domains of Dicer and cleaved into smaller dsRNAs with 3′ dinucleotide overhangs [12]. In the biogenesis of miRNA, pre-miRNA is also processed into a miRNA duplex (Biogenesis of miRNA is discussed below).
In the second effector phase, smaller dsRNAs enter into an RNA-induced silencing complex (RISC) assembly pathway [17]. RISC is ribonucleoprotein complex that contains Argonaute (Ago) proteins, siRNAs or miRNAs, and complementary mRNAs. Ago is a family of proteins characterized by the presence of a PAZ domain and a PIWI domain [18]. The PAZ domain of Ago protein is likely to engage siRNA or miRNA, and the PIWI domain adopts an RNAse H-like structure that can catalyze the cleavage of the guide strand. The dsRNA is unwound by ATP-dependent RNA helicase activity to form two single strands of RNA. dsRNA is unwounded by ATP-dependent RNA helicase activity to form two single strands of RNA. The guide (antisense) strand directs silencing targeted mRNA, and the other strand is called the passenger (sense) strand. Ago2 protein binds the guide strand and cleaves its targeted RNA at the phosphodiester bond which is positioned between nucleotides 10 and 11. The cleaved products are rapidly degraded because of its unprotected ends, and the passenger strand is also degraded. After dissociation of cleaved mRNAs from siRNA, the RISC encounters and cleaves mRNA, resulting in decrease of expression of the target gene (Fig. 13.1).
Mechanisms of RNA interference. Synthesized short interference RNA (siRNA) or double-strand (ds) RNA is introduced into a target cell. The dsRNA is processed into siRNA length of 21–23 nucleotides by Dicer (initiation phase). siRNA then enters an RNA-induced silencing complex (RISC) assembly pathway. The dsRNA unwinds to form two single strands of RNA. The passenger strand rapidly degrades and the guide strand binds and cleaves the target mRNA, resulting in mRNA degradation (effector phase)
13.3 Target Genes for Cancer Therapy
The RNAi technology in the clinical setting has relied on localized drug delivery first. This reason is that the localized administration could maintain higher concentrations of siRNAs in the targeted diseases. However, thanks to the development of DDSs (see Refs. [19, 20]), RNAi has recently been evaluated as a therapeutic strategy for cancer treatment. To develop nuclear medicine against cancers, suitable gene targets should be selected (Table 13.1). The definition of cancers is cell proliferation without normal regulation, and one of the most important characteristics of cancers is to bereave the host’s life with their malignant behaviors. Such targets include anti-apoptotic proteins, cell cycle regulators, transcription factors, signal transduction proteins, and factors associated with malignant biological behaviors of cancer cells, all of these genes are associated with the poor prognosis of cancer patients.
Among such suitable genes, BCL2 protein is one of the anti-apoptotic members of BCL family proteins and contributes to resistance to apoptosis against external stimuli, including cytotoxic agents. BCL2 participates in tumorigenesis and progression and its overexpression in tumor cells correlates with the poor prognosis of the cancer patients [21–24]. Many studies have demonstrated that siRNA treatment against BCL2 inhibited the proliferation of tumor cells [5, 25–27]. Intravenous administration of synthetic BCL2 siRNA, using a cationic or pegylated cationic liposome, suppressed tumor progression in a xenograft mouse model, and BCL2 siRNA treatment significantly elongated the survival of cancer-bearing mice [5, 27]. Oblimersen sodium is a 18-mer phosphorothioate antisense oligonucleotide designed to bind to the first six codons of the human BCL2 mRNA [28]. Though this nucleic acid medicine is an antisense oligonucleotide, it has been also used in a substantial number of clinical trials against several types of cancers [29–33]. These observations indicate that BCL2 is a suitable target for cancer therapy.
Signal transduction molecules are other candidates for RNAi. Member of the signal transduces and activator of transcription (STAT) family act as key components of cytokine signaling pathways that regulate gene expression. Among STAT family, STAT3 is most strongly implicated in carcinogenesis. Its constitutively active form is detected in variety of cancers and dysregulates the downstream target genes of cell proliferation [34] and survival [35, 36]. RNAi therapy against STAT3 demonstrates the inhibition of tumor progression as well as invasion [37–40].
Bcr-Abl fusion protein, which is created by the molecular consequence of the t(9; 22) translocation, is a constitutively active tyrosine kinase that causes Philadelphia (Ph)-positive leukemias [41]. Imatinib mesylate (IM; Gleevec™, Glivec™) was developed as a first-generation tyrosine kinase inhibitor (TKI), and its emergence has dramatically changed the outcomes of therapies against Ph-positive leukemia, especially chronic myelogenous leukemia (CML) [42–45]. Moreover, several second generation TKIs developed to overcome resistance to IM have yielded excellent outcomes [46–49]. These clinical observations demonstrated that targeting Bcr-Abl protein is a promising strategy to eliminate Bcr-Abl-positive leukemic cells. The approach to downregulate the expression of Bcr-Abl mRNA by RNAi was investigated in vitro [50–53]. Koldehoff et al. reported the in vivo administration of synthetic Bcr-Abl siRNA with cationic liposomes in a patient with recurrent Ph-positive CML resistant to IM [54]. This patient had a high level of Bcr-Abl transcripts and subcutaneous nodule, and she was treated with 10 μg/kg of Bcr-Abl siRNA intravenously by a bolus injection and 300 μg iRNA was directly injected into CML node. The level of Bcr-Abl mRNA transcript was drastically decreased; however, no obvious effects were observed after the second and third courses. Although this report was not constructed as a clinical trial, these observations are worth noting for developing nuclear medicine against CML.
β-catenin is a downstream protein of the canonical Wnt signaling pathway that has been shown to play an important role in the process of development, proliferation, and differentiation [55]. In the absence of Wnt signals, adenomatous polyposis coli (APC), Axin, glycogen synthase kinase-3β (GSK3β), and casein kinase 1α (CK1α) form a complex called the “β-catenin destruction complex.” GSK3β and CK1α target serine/threonine residues at the N terminus of β-catenin for phosphorylation [56]. Phosphorylated β-catenin is recognized and polyubiquitinated by β-transducin repeat-containing protein (β-TrCP), a component of a ubiquitin ligase complex, targeting β-catenin for degradation by the 26S proteasome [57, 58]. On the other hand, the binding of Wnt ligands to Frizzled (Fz) receptors and the low-density lipoprotein receptor-related protein 5/6 (LRP5/6) co-receptors induces the phosphorylation of Disheveled (Dvl) and prevents GSK3β-dependent phosphorylation of β-catenin. Stabilized β-catenin translocates into the nucleus and interacts with T cell factor (TCF)/lymphocyte enhancer factor (LEF). In the absence of β-catenin, TCF/LEF, which interacts with Groucho and histone deacetylase (HDAC), acts as a repressor of the transcription [59]. The β-catenin/TCF complex regulates the transcription of a number of genes associated with cell proliferation and apoptosis, as well as the expression of growth factors. Typical β-catenin/TCF target genes that are associated with cell proliferation are c-myc and cyclin D1. The c-myc oncogene regulates cell cycle progression and apoptosis. Cyclin D1 activates cyclin-dependent kinases leading to cell cycle progression. Recently, this pathway has been focused on as it is involved in cancer development. Aberrant activation of Wnt/β-catenin signaling is observed in many human cancers. Genetic mutations of Wnt signaling pathway components are primarily responsible for this aberrant activation and cause β-catenin to escape the degradation process and lead to nuclear stabilized β-catenin accumulation [60]. Treatment of siRNAs against β-catenin successfully suppressed the proliferation of colon cancer cells and myeloma cells by inducing caspase-dependent apoptosis [61–63]. Thus, β-catenin represents a suitable target for RNAi therapy.
Molecules controlling cell division are also useful targets for cancer therapy. Polo-like kinases (PLKs) belong to the family of serine/threonine kinases. PLK family has identified PLK-1, PLK-2 (SNK), PLK-3 (FNK), and PLK-4 (SAK) in mammalians so far and PLKs function as regulators of both cell cycle progression and cellular response to DNA damage. PLK-1 is the best characterized among them to date. PLK-1 regulates cell division at several points in the mitotic phase: mitotic entry through CDK1 activation, bipolar spindle formation, chromosome alignment, segregation of chromosomes, and cytokinesis [64]. Whereas PLK-1 is scarcely detectable in most adult tissues [65, 66], PLK-1 is overexpressed in cancerous tissues [65], and many reports have described that PLK-1 is overexpressed in cancerous tissues and that PLK-1 expression levels were tightly correlated with histological grades of tumors, clinical stages, and prognosis of the patients.
Inhibition of PLK-1 activity in cancer cells induces mitotic arrest and tumor cell apoptosis. Depletion of PLK-1 mRNA also inhibits the functions of PLK-1 protein in DNA damages and spindle formation and causes the inhibition of the cell proliferation in a time- and a dose-dependent manner. PLK-1 siRNA treatment induces an arrest at the G2/M phase in the cell cycle with the increase of CDC2/Cyclin B1 and the transfected cells had dumbbell-like and misaligned nuclei. Moreover, the caspase activation was induced in these cells [6, 67, 68]. These observations indicate that PLK-1 could be an excellent target for cancer therapy.
Other candidate siRNA targets are molecules that define the malignant behavior of cancerous cells. The vascular endothelial growth factor (VEGF)/VEGF receptor (VEGFR) axis plays an important role in angio- and lymphangiogenesis. VEGF family has seven members. Among them, VEGF-A stimulates angiogenesis in tumor masses, enhances the permeability of the blood vessels, and promotes the motility of cancer cells, which results in metastases [69, 70]. The previous investigations reveal that VEGF-A depletion successfully prevents metastasis of cancers [71, 72]. In contrast to VEGF-A, VEGF-C and VEGF-D are associated with tumor lymphangiogenesis and lymph node metastasis. Depletion of VEGF-C/D inhibits metastasis of cancers [73, 74]. Another example of the molecule associated with metastasis is the urokinase-type plasminogen activator (u-PA). u-PA binds to u-PA receptor (u-PAR), and this molecule activates plasminogen and matrix metalloproteases, which enhances the degradation of basement membranes and extracellular matrices and promotes metastases [75, 76]. Data using a mouse model demonstrated that the administration of u-PAR inhibited metastasis and progression of oral squamous cell carcinoma [77]. These molecules associated with metastasis will also be attractive targets of RNAi therapy.
13.4 microRNAs
microRNAs (miRNAs), as the name suggests, are very short RNAs consisted of 21 nts. Those short RNAs regulate target gene expression through translation repression or mRNA degradation, and consequently miRNAs involve diverse biological processes in eukaryocytes. miRNAs are derived from stem-loop-structured primary miRNAs (pri-miRNAs) by the cleavage activity of Drocha, a nuclear-localized member of the RNAse III family, to yield short precursor miRNAs called pre-miRNAs. Pre-miRNAs comprising 70–90 nts exhibit a hairpin structure with a 5′-phosphate and a 3′-2 nts overhang. After translocation from the nucleus to the cytoplasm by Exprtin-5 pre-miRNAs are processed by Dicer into miRNAs of 21 nts. miRNAs as well as siRNAs enter into RISC assembly pathway. Unlike siRNAs, the mature miRNAs often have a partially complementary sequence to the target mRNAs, and a single miRNA might bind to numerous target genes. Therefore, a single miRNA has diverse functions including proliferation, differentiation, and apoptosis [78].
One of the mechanisms of carcinogenesis is the imbalance of oncogenes and tumor suppressor genes caused by several factors including carcinogen. miRNAs affect gene expression by regulating the translation of mRNAs into proteins. In many cancers, some kinds of miRNAs negatively regulate tumor suppressor. miRs-15/16 are downregulated in chronic lymphocytic leukemia (CLL). miR-15a and mir16-2 recognize target sites on the 3′UTR of BCL-2, an anti-apoptotic oncogene [79]. These miRNAs regulate BCL-2 expression in normal cells. However, these are deleted in patients with CLL. On the contrary, other kinds of miRNAs regulate carcinogenesis and tumor progression. Mir-17-92 cluster is overexpressed in lung cancer tissues [80] and its target genes are PTEN and RB2 [81]. These observations indicate that the overexpression of this miR-17-93 cluster induces the carcinogenesis in lung tissues. Anti-miRNA oligonucleotides (AMOs) can suppress the miRNA activity [82], and recently MAOs are developed as nucleic acid medicines [83–86]. miRNAs regulating anti-apoptosis and cell proliferation are also suitable target molecules against cancers.
13.5 Preclinical Application of RNAi
Before the clinical trials for RNAi therapy, preclinical studies are performed. We introduce two applications of PLK-1 siRNA for cancer therapy. One application is an intravesical treatment against urinary bladder cancers. PLK-1 protein is overexpressed in urinary bladder tumors, and moreover PLK-1 expression levels are correlated with histological grades of tumors, clinical stages, and prognosis of the patients [6]. Superficial urinary bladder cancers are approximately 70 % of urinary bladder cancers at initial diagnosis. After resected transurethrally, Bacillus Calmette-Guerin (BCG), mitomycin C, and Adriamycin are administered intravesically to prevent the recurrence of or diminish the residual cancers [87]. However, half of superficial cancers recur, and consequently novel intravesical treatment should be developed. Clinical trials of RNAi therapy often rely on localized drug delivery because maintenance of higher siRNAs concentrations is necessary for efficacy against the targeted diseases. The urinary bladder which is closed to the urethra is considered as a “putative” in vitro space. In accordance with the unique idea, the efficacy of intravesical therapy of PLK-1 siRNA against urinary bladder cancers was investigated. Bladder cancer-bearing mice were established by the implantation of luciferase (Luc)-labeled UM-UC-3 bladder cancer cells into the murine bladder cavity through the urethra. After the engraftment of cancer cells in the bladder was evaluated by using the in vivo imaging system (IVIS) of bioluminescence imaging (BLI) [88], cancer-bearing mice were treated with PLK-1 siRNA/cationic liposome complexes. Tumor progression was significantly suppressed by the intravesical treatment of PLK-1 siRNA [6].
Another application is a systemic administration of siRNAs against liver metastatic tumors of lung cancers. Distant metastasis is one of the life-threatening factors in lung cancer patients. Despite the development of new molecular targeting agents [89, 90], current therapies are not sufficient to cure or manage the patients with distant metastasis [91, 92]. Therefore, novel therapies should be developed. Kawata et al. investigated the effects of PLK-1 siRNA on the liver metastasis of lung cancers in an orthotopic liver metastatic mouse model. Spleens were exposed to allow direct intrasplenic injections of Luc-labeled A549 non-small cell lung cancer cells. After the removal of spleens, the Luc-labeled A549 cell engraftment was confirmed by using IVIS, and then PLK-1 siRNA/atelocollagen complexes were administered by intravenous injection for 10 days. On day 35, mice treated with PLK-1 siRNA/atelocollagen complex showed the significant suppression of tumor growth compared to mice treated with nonsense siRNA/atelocollagen complex or PBS/atelocollagen complex which showed extensive metastases in the livers. These findings indicate that PLK-1 siRNA/atelocollagen complex is an attractive therapeutic tool for further development as a treatment against liver metastasis of lung cancer [8].
13.6 Adverse Effects of RNAi
Although RNAi shows excellent specificity in gene silencing, several adverse effects are brought in in vivo application. One probable adverse effect is activation of immune reaction. Mammalian immune cells express family of Toll-like receptors (TLRs), which play an essential role in innate immune responses. TLRs recognize microbial ligands including bacterial lipopolysaccharide, lipopeptides, or viral and bacterial RNA and DNA. Among 13 TLRs, TLR7 and TLR8 recognize ssRNA sequence-dependently and produce interferons (IFNs) and inflammatory cytokines such as IL-12 and TNF-α through the activation of NF-κB and IFN regulatory factor (IRF)-7. For this immune response, the length of single-strand RNA (ssRNA) is important and 16–19 nt ssRNA induces IFN production although 12 nt ssRNAs contains the immunostimulatory motif (GUCCUUAA) [93]. The administration of siRNAs into mammalian cells activates the immune systems also sequence-independently. siRNAs induce dsRNA-activated protein kinase (PKR) autophosphorylation and PKR produces IFNs through the activation of NK-κB and IRF-3. TLR3 recognizes unmethylated CpG DNA but not ssRNA. dsRNA directly binds to TLR3 and this signaling pathway is activated sequence-independently [94]. Interestingly, although the receptors recognizing a ssRNA containing a CpG motif and a 6 nt poly-(G) run at the 3′ end are still unknown, a ssRNA activates monocytes [95]. TLR 9, which expresses in endosomes, recognizes CpG oligodeoxynucleotides (ODNs). Purified recombinant TLR 9 binds CpG ODNs directly in a sequence- and pH-dependent manner [96]. Thus, the activation of immune response by siRNAs is dependent on their sequence and chemical nature, implying that chemical modifications of siRNAs might prevent the immune activation. The 2′ position of nucleotides is within TLR-7-interacting sequences and 2′ O-methyl or 2′ fluoro modification abrogate immune response. Furthermore, the uridine or guanosine modification is most effective [97]. Locked nucleic acid modifications of the 3′ of 5′ termini of the sense strand of siRNAs can reduce the immunostimulatory effects [93]. siRNAs conjugated to cholesterol have no significant activation of immune system and improve the distribution of siRNA to the targeted organ including the liver. Systemic administration of cholesterol-conjugated apolipoprotein B siRNAs induces a decrease of apolipoprotein B expression in liver and jejunum of mice, resulting in a decrease in cholesterol levels without the activation of immune systems [98].
Besides perfect complementarity of siRNAs in target RNA sequence, partially complementary sequences in unintended RNAs induce gene silencing (off-target effect). This effect is induced by the sequence complementarity in the seed region of siRNAs or short-hairpin RNAs (shRNAs) [99]. Moreover, the 7 nt motif complementary to 2–8 nt at the 5′ end of antisense strands of siRNAs has been shown to be a key determinant in directing off-target effects [100]. There are several ways to control the off-target effects. The in silico screening of siRNA constructs are useful for optimization to prevent the off-target effects, and several groups have been developing algorithm [101, 102]. Chemical modification is also useful. For example, the O-methyl modification of the 2′-position of the ribose within the seed region of siRNAs reduces the off-target effect [103]. Asymmetrically designed siRNAs reduce off-target effects compared to symmetric siRNAs. Sun et al. designed asymmetric RNA duplexes of various lengths with overhangs at the 3′ and 5′ ends of the antisense strand to target genes. All siRNAs against target genes were designed to match the same 19 nt sequence. The asymmetric siRNAs effectively induced gene silencing of targeted genes without silencing of nontargeted genes [104].
shRNAs can also induce stable gene silencing. Consequently, it is possible that long-term silencing by shRAN overexpression causes fatal adverse effects. Because shRNA is processed through the miRNA pathway, the miRNA maturation is blocked in response to shRNA concentration. Grimm et al. demonstrated that the sustained high-level shRNA expression in the liver of mice by AAV vector downregulated liver-derived miRNAs, resulting in hepatic injury and death. Morbidity was associated with the downregulation of liver-derived miRNAs [105]. They speculated that saturation of Exportin-5 whose function is nuclear transport inhibited the miRNA maturation pathway. On the contrary, Constein et al. demonstrated that the administration of synthesized siRNAs induced acute and long-term gene silencing without interrupting the endogenous miRNA biogenesis [106]. As mentioned by Grimm et al. [105], higher expression of shRNAs by viral vector might influence the miRNA biogenesis. Considering these findings, careful modification and formulation of siRNAs could avoid the competition between siRNA and miRNA.
13.7 Clinical Trials of RNAi Towards Cancer Therapies
siRNA cancer therapies have been conducted in clinical settings, but few clinical trials for cancer therapy are ongoing (Table 13.2). Alnylam Pharmaceuticals is developing ALN-VSP01 targeting kinase spindle protein and VEGF, and conducting a Phase I study in patients with advanced tumors with liver involvement. Calando Pharmaceuticals is conducting a Phase I study of CALAA-01 in patients with solid tumors refractory to standard-of-care therapies. CALAA-01 is composed of RRM2 siRNA and CDP nanoparticles called Rondel™, and CALAA-01 has been proven safe and effective in mice and nonhuman primates’ studies. Clinical studies using LNAs are also ongoing. Santaris Pharma has developed LNA against Bcl-2, SPC2996, for use in an ongoing Phase I/II study in patients with relapsed or refractory chronic lymphocytic leukemia is ongoing. Enzon Pharmaceuticals has developed a LNA against hypoxia-inducible factor-1α and a Phase I/II study in patients with advanced solid tumors or lymphoma is ongoing. National Cancer Institute and Tekmira Pharmaceuticals are conducting clinical trials on PLK-1 RNA interference against solid tumor or lymphoma. As clinical trials of cancer therapies have just started, their outcomes are expected.
13.8 Biomarker-Based Screening
RNA interference technology is also used in the field of drug discovery. The biomarker-based screening is a new high-throughput screening method based on transcriptional profiling and identifies the specific transcriptional activities altered by the compounds of interest. PGX Health, A division of Clinical Data Inc. (formerly Avalon Pharmaceuticals, MD, USA) assessed the transcriptional response of a colon cancer cell line to treatment with β-catenin siRNA using full-genome microarray analysis [9]. Nine biomarkers were selected for their potential as indicators for cancer therapy. A library of 90,000 individual compounds was screened to identify compounds that showed a similar expression pattern to the siRNA (Fig. 13.2). Finally, the compound LC-363 was detected based on its ability to mimic the effect of β-catenin knockdown. The effect of AV-65, one of LC-363 compound series, on MM cells and CML cells was investigated. AV-65 inhibited the proliferation of MM and CML cells by promoting the degradation of β-catenin and inhibiting β-catenin/TCF transcriptional activity. AV-65 decreased the expression of c-myc, cyclin D1, and survivin, which resulted in the inhibition of tumor cell proliferation through the apoptotic pathway [10, 11]. Moreover, AV-65 treatment prolonged the survival of orthotopic MM-bearing mice [11]. A clinical study with this compound series in solid and hematopoietic malignancies will be carried out in the future.
Biomarker-based screening using RNA interference. This assay proceeds in two steps: the first step consists of setting up the signature of siRNA against target gene. The second step involves screening for compounds with the similar expression patterns. Consequently, hit compounds that inhibit the downstream signal of the target gene
13.9 Conclusion
RNAi therapy against cancers has just started and the outcomes are expected. However, it should be warranted to establish the pharmacokinetics and pharmacodynamics of siRNAs on the administration for the potential approval of siRNA as a tool for cancer therapy. Moreover, to maximize efficacy and to minimize adverse effects of RNAi, it should be determined whether siRNAs are best administered alone or in combination with chemotherapeutic agents [107], and whether it is better to administer a single specific siRNA or multiple specific siRNAs [108–110].
In conclusion, RNAi therapy represents a powerful strategy against cancers and may offer a novel and attractive therapeutic option. The success of RNAi depends on the suitable selection of target genes. Besides developing nucleic acid-based medicine, RNAi technology is applied into the field of drug discovery. We anticipate that RNAi technology could establish a novel and promising therapeutic tool against cancers.
References
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391(6669):806–811
Check E (2002) A tragic setback. Nature 420(6912):116–118
Hacein-Bey-Abina S, von Kalle C, Schmidt M, Le Deist F, Wulffraat N, McIntyre E, Radford I, Villeval JL, Fraser CC, Cavazzana-Calvo M, Fischer A (2003) A serious adverse event after successful gene therapy for X-linked severe combined immunodeficiency. N Engl J Med 348(3):255–256
Nguyen T, Menocal EM, Harborth J, Fruehauf JH (2008) RNAi therapeutics: an update on delivery. Curr Opin Mol Ther 10(2):158–167
Yano J, Hirabayashi K, Nakagawa S, Yamaguchi T, Nogawa M, Kashimori I, Naito H, Kitagawa H, Ishiyama K, Ohgi T, Irimura T (2004) Antitumor activity of small interfering RNA/cationic liposome complex in mouse models of cancer. Clin Cancer Res 10(22):7721–7726
Nogawa M, Yuasa T, Kimura S, Tanaka M, Kuroda J, Sato K, Yokota A, Segawa H, Toda Y, Kageyama S, Yoshiki T, Okada Y, Maekawa T (2005) Intravesical administration of small interfering RNA targeting PLK-1 successfully prevents the growth of bladder cancer. J Clin Invest 115(4):978–985. doi:10.1172/JCI23043
Sano A, Maeda M, Nagahara S, Ochiya T, Honma K, Itoh H, Miyata T, Fujioka K (2003) Atelocollagen for protein and gene delivery. Adv Drug Deliv Rev 55(12):1651–1677
Kawata E, Ashihara E, Kimura S, Takenaka K, Sato K, Tanaka R, Yokota A, Kamitsuji Y, Takeuchi M, Kuroda J, Tanaka F, Yoshikawa T, Maekawa T (2008) Administration of PLK-1 small interfering RNA with atelocollagen prevents the growth of liver metastases of lung cancer. Mol Cancer Ther 7(9):2904–2912. doi:10.1158/1535-7163.MCT-08-0473
Bol D, Ebner R (2006) Gene expression profiling in the discovery, optimization and development of novel drugs: one universal screening platform. Pharmacogenomics 7(2):227–235. doi:10.2217/14622416.7.2.227
Nagao R, Ashihara E, Kimura S, Strovel JW, Yao H, Takeuchi M, Tanaka R, Hayashi Y, Hirai H, Padia J, Strand K, Maekawa T (2011) Growth inhibition of imatinib-resistant CML cells with the T315I mutation and hypoxia-adaptation by AV65 – a novel Wnt/beta-catenin signaling inhibitor. Cancer Lett 312(1):91–100. doi:10.1016/j.canlet.2011.08.002
Yao H, Ashihara E, Strovel JW, Nakagawa Y, Kuroda J, Nagao R, Tanaka R, Yokota A, Takeuchi M, Hayashi Y, Shimazaki C, Taniwaki M, Strand K, Padia J, Hirai H, Kimura S, Maekawa T (2011) AV-65, a novel Wnt/beta-catenin signal inhibitor, successfully suppresses progression of multiple myeloma in a mouse model. Blood Cancer J 1:e43
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409(6818):363–366
Grishok A, Pasquinelli AE, Conte D, Li N, Parrish S, Ha I, Baillie DL, Fire A, Ruvkun G, Mello CC (2001) Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 106(1):23–34
Hutvagner G, Zamore PD (2002) A microRNA in a multiple-turnover RNAi enzyme complex. Science 297(5589):2056–2060
Kolb FA, Zhang H, Jaronczyk K, Tahbaz N, Hobman TC, Filipowicz W (2005) Human dicer: purification, properties, and interaction with PAZ PIWI domain proteins. Methods Enzymol 392:316–336
Murchison EP, Hannon GJ (2004) MiRNAs on the move: miRNA biogenesis and the RNAi machinery. Curr Opin Cell Biol 16(3):223–229
Agrawal N, Dasaradhi PV, Mohmmed A, Malhotra P, Bhatnagar RK, Mukherjee SK (2003) RNA interference: biology, mechanism, and applications. Microbiol Mol Biol Rev 67(4):657–685
Parker JS, Barford D (2006) Argonaute: a scaffold for the function of short regulatory RNAs. Trends Biochem Sci 31(11):622–630
Whitehead KA, Langer R, Anderson DG (2009) Knocking down barriers: advances in siRNA delivery. Nat Rev Drug Discov 8(2):129–138. doi:10.1038/nrd2742
Oh YK, Park TG (2009) siRNA delivery systems for cancer treatment. Adv Drug Deliv Rev 61(10):850–862. doi:10.1016/j.addr.2009.04.018
Iqbal J, Neppalli VT, Wright G, Dave BJ, Horsman DE, Rosenwald A, Lynch J, Hans CP, Weisenburger DD, Greiner TC, Gascoyne RD, Campo E, Ott G, Muller-Hermelink HK, Delabie J, Jaffe ES, Grogan TM, Connors JM, Vose JM, Armitage JO, Staudt LM, Chan WC (2006) BCL2 expression is a prognostic marker for the activated B-cell-like type of diffuse large B-cell lymphoma. J Clin Oncol 24(6):961–968
Lauwers GY, Scott GV, Karpeh MS (1995) Immunohistochemical evaluation of bcl-2 protein expression in gastric adenocarcinomas. Cancer 75(9):2209–2213
Pezzella F, Turley H, Kuzu I, Tungekar MF, Dunnill MS, Pierce CB, Harris A, Gatter KC, Mason DY (1993) bcl-2 protein in non-small-cell lung carcinoma. N Engl J Med 329(10):690–694
Sinicrope FA, Hart J, Michelassi F, Lee JJ (1995) Prognostic value of bcl-2 oncoprotein expression in stage II colon carcinoma. Clin Cancer Res 1(10):1103–1110
Fu GF, Lin XH, Han QW, Fan YR, Xu YF, Guo D, Xu GX, Hou YY (2005) RNA interference remarkably suppresses bcl-2 gene expression in cancer cells in vitro and in vivo. Cancer Biol Ther 4(8):822–829
Ruckert F, Samm N, Lehner AK, Saeger HD, Grutzmann R, Pilarsky C (2010) Simultaneous gene silencing of Bcl-2, XIAP and survivin re-sensitizes pancreatic cancer cells towards apoptosis. BMC Cancer 10:379
Sonoke S, Ueda T, Fujiwara K, Sato Y, Takagaki K, Hirabayashi K, Ohgi T, Yano J (2008) Tumor regression in mice by delivery of Bcl-2 small interfering RNA with pegylated cationic liposomes. Cancer Res 68(21):8843–8851
Klasa RJ, Gillum AM, Klem RE, Frankel SR (2002) Oblimersen Bcl-2 antisense: facilitating apoptosis in anticancer treatment. Antisense Nucleic Acid Drug Dev 12(3):193–213
Advani PP, Paulus A, Masood A, Sher T, Chanan-Khan A (2011) Pharmacokinetic evaluation of oblimersen sodium for the treatment of chronic lymphocytic leukemia. Expert Opin Drug Metab Toxicol 7(6):765–774. doi:10.1517/17425255.2011.579105
Chanan-Khan AA, Niesvizky R, Hohl RJ, Zimmerman TM, Christiansen NP, Schiller GJ, Callander N, Lister J, Oken M, Jagannath S (2009) Phase III randomised study of dexamethasone with or without oblimersen sodium for patients with advanced multiple myeloma. Leuk Lymphoma 50(4):559–565
Galatin PS, Advani RH, Fisher GA, Francisco B, Julian T, Losa R, Sierra MI, Sikic BI (2011) Phase I trial of oblimersen (Genasense(R)) and gemcitabine in refractory and advanced malignancies. Invest New Drugs 29(5):971–977
Rudin CM, Salgia R, Wang X, Hodgson LD, Masters GA, Green M, Vokes EE (2008) Randomized phase II study of carboplatin and etoposide with or without the bcl-2 antisense oligonucleotide oblimersen for extensive-stage small-cell lung cancer: CALGB 30103. J Clin Oncol 26(6):870–876
Sternberg CN, Dumez H, Van Poppel H, Skoneczna I, Sella A, Daugaard G, Gil T, Graham J, Carpentier P, Calabro F, Collette L, Lacombe D (2009) Docetaxel plus oblimersen sodium (Bcl-2 antisense oligonucleotide): an EORTC multicenter, randomized phase II study in patients with castration-resistant prostate cancer. Ann Oncol 20(7):1264–1269
Bromberg J (2002) Stat proteins and oncogenesis. J Clin Invest 109(9):1139–1142. doi:10.1172/JCI15617
Konnikova L, Simeone MC, Kruger MM, Kotecki M, Cochran BH (2005) Signal transducer and activator of transcription 3 (STAT3) regulates human telomerase reverse transcriptase (hTERT) expression in human cancer and primary cells. Cancer Res 65(15):6516–6520. doi:10.1158/0008-5472.CAN-05-0924
Masuda M, Suzui M, Yasumatu R, Nakashima T, Kuratomi Y, Azuma K, Tomita K, Komiyama S, Weinstein IB (2002) Constitutive activation of signal transducers and activators of transcription 3 correlates with cyclin D1 overexpression and may provide a novel prognostic marker in head and neck squamous cell carcinoma. Cancer Res 62(12):3351–3355
Xu Y, Li X, Zhang S, Shen D, Li H, Wu Y, Qiu Y, Ji Y, Chen F (2012) Targeting Stat3 suppresses growth of U251 cell-derived tumours in nude mice. J Clin Neurosci 19(3):443–446. doi:10.1016/j.jocn.2011.04.017
Gao L, Zhang L, Hu J, Li F, Shao Y, Zhao D, Kalvakolanu DV, Kopecko DJ, Zhao X, Xu DQ (2005) Down-regulation of signal transducer and activator of transcription 3 expression using vector-based small interfering RNAs suppresses growth of human prostate tumor in vivo. Clin Cancer Res 11(17):6333–6341. doi:10.1158/1078-0432.CCR-05-0148
Fan Y, Zhang YL, Wu Y, Zhang W, Wang YH, Cheng ZM, Li H (2008) Inhibition of signal transducer and activator of transcription 3 expression by RNA interference suppresses invasion through inducing anoikis in human colon cancer cells. World J Gastroenterol 14(3):428–434
Ling X, Arlinghaus RB (2005) Knockdown of STAT3 expression by RNA interference inhibits the induction of breast tumors in immunocompetent mice. Cancer Res 65(7):2532–2536. doi:10.1158/0008-5472.CAN-04-2425
Sawyers CL (1999) Chronic myeloid leukemia. N Engl J Med 340(17):1330–1340. doi:10.1056/NEJM199904293401706
O’Brien SG, Guilhot F, Larson RA, Gathmann I, Baccarani M, Cervantes F, Cornelissen JJ, Fischer T, Hochhaus A, Hughes T, Lechner K, Nielsen JL, Rousselot P, Reiffers J, Saglio G, Shepherd J, Simonsson B, Gratwohl A, Goldman JM, Kantarjian H, Taylor K, Verhoef G, Bolton AE, Capdeville R, Druker BJ, Investigators I (2003) Imatinib compared with interferon and low-dose cytarabine for newly diagnosed chronic-phase chronic myeloid leukemia. N Engl J Med 348(11):994–1004. doi:10.1056/NEJMoa022457
Sawyers CL, Hochhaus A, Feldman E, Goldman JM, Miller CB, Ottmann OG, Schiffer CA, Talpaz M, Guilhot F, Deininger MW, Fischer T, O’Brien SG, Stone RM, Gambacorti-Passerini CB, Russell NH, Reiffers JJ, Shea TC, Chapuis B, Coutre S, Tura S, Morra E, Larson RA, Saven A, Peschel C, Gratwohl A, Mandelli F, Ben-Am M, Gathmann I, Capdeville R, Paquette RL, Druker BJ (2002) Imatinib induces hematologic and cytogenetic responses in patients with chronic myelogenous leukemia in myeloid blast crisis: results of a phase II study. Blood 99(10):3530–3539
Talpaz M, Silver RT, Druker BJ, Goldman JM, Gambacorti-Passerini C, Guilhot F, Schiffer CA, Fischer T, Deininger MW, Lennard AL, Hochhaus A, Ottmann OG, Gratwohl A, Baccarani M, Stone R, Tura S, Mahon FX, Fernandes-Reese S, Gathmann I, Capdeville R, Kantarjian HM, Sawyers CL (2002) Imatinib induces durable hematologic and cytogenetic responses in patients with accelerated phase chronic myeloid leukemia: results of a phase 2 study. Blood 99(6):1928–1937
Druker BJ, Guilhot F, O’Brien SG, Gathmann I, Kantarjian H, Gattermann N, Deininger MW, Silver RT, Goldman JM, Stone RM, Cervantes F, Hochhaus A, Powell BL, Gabrilove JL, Rousselot P, Reiffers J, Cornelissen JJ, Hughes T, Agis H, Fischer T, Verhoef G, Shepherd J, Saglio G, Gratwohl A, Nielsen JL, Radich JP, Simonsson B, Taylor K, Baccarani M, So C, Letvak L, Larson RA, Investigators I (2006) Five-year follow-up of patients receiving imatinib for chronic myeloid leukemia. N Engl J Med 355(23):2408–2417. doi:10.1056/NEJMoa062867
Kantarjian H, Giles F, Wunderle L, Bhalla K, O’Brien S, Wassmann B, Tanaka C, Manley P, Rae P, Mietlowski W, Bochinski K, Hochhaus A, Griffin JD, Hoelzer D, Albitar M, Dugan M, Cortes J, Alland L, Ottmann OG (2006) Nilotinib in imatinib-resistant CML and Philadelphia chromosome-positive ALL. N Engl J Med 354(24):2542–2551. doi:10.1056/NEJMoa055104
Talpaz M, Shah NP, Kantarjian H, Donato N, Nicoll J, Paquette R, Cortes J, O’Brien S, Nicaise C, Bleickardt E, Blackwood-Chirchir MA, Iyer V, Chen TT, Huang F, Decillis AP, Sawyers CL (2006) Dasatinib in imatinib-resistant Philadelphia chromosome-positive leukemias. N Engl J Med 354(24):2531–2541. doi:10.1056/NEJMoa055229
Kimura S, Ashihara E, Maekawa T (2006) New tyrosine kinase inhibitors in the treatment of chronic myeloid leukemia. Curr Pharm Biotechnol 7(5):371–379
Khoury HJ, Cortes JE, Kantarjian HM, Gambacorti-Passerini C, Baccarani M, Kim DW, Zaritskey A, Countouriotis A, Besson N, Leip E, Kelly V, Brummendorf TH (2012) Bosutinib is active in chronic phase chronic myeloid leukemia after imatinib and dasatinib and/or nilotinib therapy failure. Blood 119(15):3403–3412. doi:10.1182/blood-2011-11-390120
Li MJ, McMahon R, Snyder DS, Yee JK, Rossi JJ (2003) Specific killing of Ph+ chronic myeloid leukemia cells by a lentiviral vector-delivered anti-bcr/abl small hairpin RNA. Oligonucleotides 13(5):401–409. doi:10.1089/154545703322617087
Scherr M, Battmer K, Winkler T, Heidenreich O, Ganser A, Eder M (2003) Specific inhibition of bcr-abl gene expression by small interfering RNA. Blood 101(4):1566–1569. doi:10.1182/blood-2002-06-1685
Rangatia J, Bonnet D (2006) Transient or long-term silencing of BCR-ABL alone induces cell cycle and proliferation arrest, apoptosis and differentiation. Leukemia 20(1):68–76. doi:10.1038/sj.leu.2403999
Arthanari Y, Pluen A, Rajendran R, Aojula H, Demonacos C (2010) Delivery of therapeutic shRNA and siRNA by Tat fusion peptide targeting BCR-ABL fusion gene in Chronic Myeloid Leukemia cells. J Control Release 145(3):272–280. doi:10.1016/j.jconrel.2010.04.011
Koldehoff M, Steckel NK, Beelen DW, Elmaagacli AH (2007) Therapeutic application of small interfering RNA directed against bcr-abl transcripts to a patient with imatinib-resistant chronic myeloid leukaemia. Clin Exp Med 7(2):47–55. doi:10.1007/s10238-007-0125-z
Wodarz A, Nusse R (1998) Mechanisms of Wnt signaling in development. Annu Rev Cell Dev Biol 14:59–88. doi:10.1146/annurev.cellbio.14.1.59
Liu C, Li Y, Semenov M, Han C, Baeg GH, Tan Y, Zhang Z, Lin X, He X (2002) Control of beta-catenin phosphorylation/degradation by a dual-kinase mechanism. Cell 108(6):837–847
Latres E, Chiaur DS, Pagano M (1999) The human F box protein beta-Trcp associates with the Cul1/Skp1 complex and regulates the stability of beta-catenin. Oncogene 18(4):849–854. doi:10.1038/sj.onc.1202653
Kitagawa M, Hatakeyama S, Shirane M, Matsumoto M, Ishida N, Hattori K, Nakamichi I, Kikuchi A, Nakayama K, Nakayama K (1999) An F-box protein, FWD1, mediates ubiquitin-dependent proteolysis of beta-catenin. EMBO J 18(9):2401–2410. doi:10.1093/emboj/18.9.2401
Roose J, Molenaar M, Peterson J, Hurenkamp J, Brantjes H, Moerer P, van de Wetering M, Destree O, Clevers H (1998) The Xenopus Wnt effector XTcf-3 interacts with Groucho-related transcriptional repressors. Nature 395(6702):608–612. doi:10.1038/26989
Moon RT, Kohn AD, De Ferrari GV, Kaykas A (2004) WNT and beta-catenin signalling: diseases and therapies. Nat Rev Genet 5(9):691–701. doi:10.1038/nrg1427
Ashihara E, Kawata E, Nakagawa Y, Shimazaski C, Kuroda J, Taniguchi K, Uchiyama H, Tanaka R, Yokota A, Takeuchi M, Kamitsuji Y, Inaba T, Taniwaki M, Kimura S, Maekawa T (2009) beta-catenin small interfering RNA successfully suppressed progression of multiple myeloma in a mouse model. Clin Cancer Res 15(8):2731–2738. doi:10.1158/1078-0432.CCR-08-1350
Verma UN, Surabhi RM, Schmaltieg A, Becerra C, Gaynor RB (2003) Small interfering RNAs directed against beta-catenin inhibit the in vitro and in vivo growth of colon cancer cells. Clin Cancer Res 9(4):1291–1300
Liyan W, Xun S, Xiangwei M (2011) Effect of beta -catenin siRNA on proliferation and apoptosis of hepatoma cell line SMMC-7721 and HepG-2. Hepatogastroenterology 58(110–111):1757–1764. doi:10.5754/hge11017
Barr FA, Sillje HH, Nigg EA (2004) Polo-like kinases and the orchestration of cell division. Nat Rev Mol Cell Biol 5(6):429–440. doi:10.1038/nrm1401
Lake RJ, Jelinek WR (1993) Cell cycle- and terminal differentiation-associated regulation of the mouse mRNA encoding a conserved mitotic protein kinase. Mol Cell Biol 13(12):7793–7801
Hamanaka R, Maloid S, Smith MR, O’Connell CD, Longo DL, Ferris DK (1994) Cloning and characterization of human and murine homologues of the Drosophila polo serine-threonine kinase. Cell Growth Differ 5(3):249–257
Liu X, Erikson RL (2003) Polo-like kinase (Plk)1 depletion induces apoptosis in cancer cells. Proc Natl Acad Sci U S A 100(10):5789–5794. doi:10.1073/pnas.1031523100
Kawata E, Ashihara E, Maekawa T (2011) RNA interference against polo-like kinase-1 in advanced non-small cell lung cancers. J Clin Bioinform 1(1):6. doi:10.1186/2043-9113-1-6
Yancopoulos GD, Davis S, Gale NW, Rudge JS, Wiegand SJ, Holash J (2000) Vascular-specific growth factors and blood vessel formation. Nature 407(6801):242–248. doi:10.1038/35025215
Ellis LM (2004) Angiogenesis and its role in colorectal tumor and metastasis formation. Semin Oncol 31(6 Suppl 17):3–9. doi:10.1053/j.seminoncol.2004.11.028
Wang S, Liu H, Ren L, Pan Y, Zhang Y (2008) Inhibiting colorectal carcinoma growth and metastasis by blocking the expression of VEGF using RNA interference. Neoplasia 10(4):399–407
Salva E, Kabasakal L, Eren F, Ozkan N, Cakalagaoglu F, Akbuga J (2012) Local delivery of chitosan/VEGF siRNA nanoplexes reduces angiogenesis and growth of breast cancer in vivo. Nucleic Acid Ther 22(1):40–48. doi:10.1089/nat.2011.0312
Chen Z, Varney ML, Backora MW, Cowan K, Solheim JC, Talmadge JE, Singh RK (2005) Down-regulation of vascular endothelial cell growth factor-C expression using small interfering RNA vectors in mammary tumors inhibits tumor lymphangiogenesis and spontaneous metastasis and enhances survival. Cancer Res 65(19):9004–9011. doi:10.1158/0008-5472.CAN-05-0885
Lin W, Jiang L, Chen Y, She F, Han S, Zhu J, Zhou L, Tang N, Wang X, Li X (2012) Vascular endothelial growth factor-D promotes growth, lymphangiogenesis and lymphatic metastasis in gallbladder cancer. Cancer Lett 314(2):127–136. doi:10.1016/j.canlet.2011.09.004
Andreasen PA, Kjoller L, Christensen L, Duffy MJ (1997) The urokinase-type plasminogen activator system in cancer metastasis: a review. Int J Cancer 72(1):1–22
Dass K, Ahmad A, Azmi AS, Sarkar SH, Sarkar FH (2008) Evolving role of uPA/uPAR system in human cancers. Cancer Treat Rev 34(2):122–136
Zhou H, Tang Y, Liang X, Yang X, Yang J, Zhu G, Zheng M, Zhang C (2009) RNAi targeting urokinase-type plasminogen activator receptor inhibits metastasis and progression of oral squamous cell carcinoma in vivo. Int J Cancer 125(2):453–462
Cho WC (2010) MicroRNAs in cancer – from research to therapy. Biochim Biophys Acta 1805(2):209–217. doi:10.1016/j.bbcan.2009.11.003
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M, Wojcik SE, Aqeilan RI, Zupo S, Dono M, Rassenti L, Alder H, Volinia S, Liu CG, Kipps TJ, Negrini M, Croce CM (2005) miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 102(39):13944–13949. doi:10.1073/pnas.0506654102
Hayashita Y, Osada H, Tatematsu Y, Yamada H, Yanagisawa K, Tomida S, Yatabe Y, Kawahara K, Sekido Y, Takahashi T (2005) A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 65(21):9628–9632. doi:10.1158/0008-5472.CAN-05-2352
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115(7):787–798
Krutzfeldt J, Kuwajima S, Braich R, Rajeev KG, Pena J, Tuschl T, Manoharan M, Stoffel M (2007) Specificity, duplex degradation and subcellular localization of antagomirs. Nucleic Acids Res 35(9):2885–2892. doi:10.1093/nar/gkm024
Park JK, Kogure T, Nuovo GJ, Jiang J, He L, Kim JH, Phelps MA, Papenfuss TL, Croce CM, Patel T, Schmittgen TD (2011) miR-221 silencing blocks hepatocellular carcinoma and promotes survival. Cancer Res 71(24):7608–7616. doi:10.1158/0008-5472.CAN-11-1144
Sun L, Yan W, Wang Y, Sun G, Luo H, Zhang J, Wang X, You Y, Yang Z, Liu N (2011) MicroRNA-10b induces glioma cell invasion by modulating MMP-14 and uPAR expression via HOXD10. Brain Res 1389:9–18. doi:10.1016/j.brainres.2011.03.013
Shi SJ, Zhong ZR, Liu J, Zhang ZR, Sun X, Gong T (2012) Solid lipid nanoparticles loaded with anti-microRNA oligonucleotides (AMOs) for suppression of microRNA-21 functions in human lung cancer cells. Pharm Res 29(1):97–109. doi:10.1007/s11095-011-0514-6
Meng W, Jiang L, Lu L, Hu H, Yu H, Ding D, Xiao K, Zheng W, Guo H, Ma W (2012) Anti-miR-155 oligonucleotide enhances chemosensitivity of U251 cell to taxol by inducing apoptosis. Cell Biol Int. doi:10.1042/CBI20100918
Gee J, Sabichi AL, Grossman HB (2002) Chemoprevention of superficial bladder cancer. Crit Rev Oncol Hematol 43(3):277–286
Nogawa M, Yuasa T, Kimura S, Kuroda J, Sato K, Segawa H, Yokota A, Maekawa T (2005) Monitoring luciferase-labeled cancer cell growth and metastasis in different in vivo models. Cancer Lett 217(2):243–253. doi:10.1016/j.canlet.2004.07.010
Sharma SV, Bell DW, Settleman J, Haber DA (2007) Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 7(3):169–181. doi:10.1038/nrc2088
Oxnard GR, Miller VA (2010) Use of erlotinib or gefitinib as initial therapy in advanced NSCLC. Oncology 24(5):392–399
Bremnes RM, Sundstrom S, Aasebo U, Kaasa S, Hatlevoll R, Aamdal S, Norweigian Lung Cancer Study G (2003) The value of prognostic factors in small cell lung cancer: results from a randomised multicenter study with minimum 5 year follow-up. Lung Cancer 39(3):303–313
Hoang T, Xu R, Schiller JH, Bonomi P, Johnson DH (2005) Clinical model to predict survival in chemonaive patients with advanced non-small-cell lung cancer treated with third-generation chemotherapy regimens based on eastern cooperative oncology group data. J Clin Oncol 23(1):175–183. doi:10.1200/JCO.2005.04.177
Hornung V, Guenthner-Biller M, Bourquin C, Ablasser A, Schlee M, Uematsu S, Noronha A, Manoharan M, Akira S, de Fougerolles A, Endres S, Hartmann G (2005) Sequence-specific potent induction of IFN-alpha by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nat Med 11(3):263–270. doi:10.1038/nm1191
Trinchieri G, Sher A (2007) Cooperation of Toll-like receptor signals in innate immune defence. Nat Rev Immunol 7(3):179–190. doi:10.1038/nri2038
Marques JT, Williams BR (2005) Activation of the mammalian immune system by siRNAs. Nat Biotechnol 23(11):1399–1405. doi:10.1038/nbt1161
Rutz M, Metzger J, Gellert T, Luppa P, Lipford GB, Wagner H, Bauer S (2004) Toll-like receptor 9 binds single-stranded CpG-DNA in a sequence- and pH-dependent manner. Eur J Immunol 34(9):2541–2550. doi:10.1002/eji.200425218
Judge AD, Bola G, Lee AC, MacLachlan I (2006) Design of noninflammatory synthetic siRNA mediating potent gene silencing in vivo. Mol Ther 13(3):494–505. doi:10.1016/j.ymthe.2005.11.002
Soutschek J, Akinc A, Bramlage B, Charisse K, Constien R, Donoghue M, Elbashir S, Geick A, Hadwiger P, Harborth J, John M, Kesavan V, Lavine G, Pandey RK, Racie T, Rajeev KG, Rohl I, Toudjarska I, Wang G, Wuschko S, Bumcrot D, Koteliansky V, Limmer S, Manoharan M, Vornlocher HP (2004) Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 432(7014):173–178. doi:10.1038/nature03121
Jackson AL, Burchard J, Schelter J, Chau BN, Cleary M, Lim L, Linsley PS (2006) Widespread siRNA “off-target” transcript silencing mediated by seed region sequence complementarity. RNA 12(7):1179–1187. doi:10.1261/rna.25706
Lin X, Ruan X, Anderson MG, McDowell JA, Kroeger PE, Fesik SW, Shen Y (2005) siRNA-mediated off-target gene silencing triggered by a 7 nt complementation. Nucleic Acids Res 33(14):4527–4535. doi:10.1093/nar/gki762
Yamada T, Morishita S (2005) Accelerated off-target search algorithm for siRNA. Bioinformatics 21(8):1316–1324. doi:10.1093/bioinformatics/bti155
Park YK, Park SM, Choi YC, Lee D, Won M, Kim YJ (2008) AsiDesigner: exon-based siRNA design server considering alternative splicing. Nucleic Acids Res 36(Web Server issue):W97–W103. doi:10.1093/nar/gkn280
Jackson AL, Burchard J, Leake D, Reynolds A, Schelter J, Guo J, Johnson JM, Lim L, Karpilow J, Nichols K, Marshall W, Khvorova A, Linsley PS (2006) Position-specific chemical modification of siRNAs reduces “off-target” transcript silencing. RNA 12(7):1197–1205. doi:10.1261/rna.30706
Sun X, Rogoff HA, Li CJ (2008) Asymmetric RNA duplexes mediate RNA interference in mammalian cells. Nat Biotechnol 26(12):1379–1382. doi:10.1038/nbt.1512
Grimm D, Streetz KL, Jopling CL, Storm TA, Pandey K, Davis CR, Marion P, Salazar F, Kay MA (2006) Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441(7092):537–541. doi:10.1038/nature04791
John M, Constien R, Akinc A, Goldberg M, Moon YA, Spranger M, Hadwiger P, Soutschek J, Vornlocher HP, Manoharan M, Stoffel M, Langer R, Anderson DG, Horton JD, Koteliansky V, Bumcrot D (2007) Effective RNAi-mediated gene silencing without interruption of the endogenous microRNA pathway. Nature 449(7163):745–747. doi:10.1038/nature06179
von Bueren AO, Shalaby T, Oehler-Janne C, Arnold L, Stearns D, Eberhart CG, Arcaro A, Pruschy M, Grotzer MA (2009) RNA interference-mediated c-MYC inhibition prevents cell growth and decreases sensitivity to radio- and chemotherapy in childhood medulloblastoma cells. BMC Cancer 9:10. doi:10.1186/1471-2407-9-10
Shi XH, Liang ZY, Ren XY, Liu TH (2009) Combined silencing of K-ras and Akt2 oncogenes achieves synergistic effects in inhibiting pancreatic cancer cell growth in vitro and in vivo. Cancer Gene Ther 16(3):227–236. doi:10.1038/cgt.2008.82
Honma K, Iwao-Koizumi K, Takeshita F, Yamamoto Y, Yoshida T, Nishio K, Nagahara S, Kato K, Ochiya T (2008) RPN2 gene confers docetaxel resistance in breast cancer. Nat Med 14(9):939–948. doi:10.1038/nm.1858
Mu P, Nagahara S, Makita N, Tarumi Y, Kadomatsu K, Takei Y (2009) Systemic delivery of siRNA specific to tumor mediated by atelocollagen: combined therapy using siRNA targeting Bcl-xL and cisplatin against prostate cancer. Int J Cancer 125(12):2978–2990. doi:10.1002/ijc.24382
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Ashihara, E., Maekawa, T. (2013). RNA Interference for Oncology: Clinical Prospects Beyond the Hype. In: Danquah, M., Mahato, R. (eds) Emerging Trends in Cell and Gene Therapy. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-417-3_13
Download citation
DOI: https://doi.org/10.1007/978-1-62703-417-3_13
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-416-6
Online ISBN: 978-1-62703-417-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)