Abstract
Although the wide availability of prostate-specific antigen (PSA) has revolutionized prostate cancer (PCa) screening, resulting in a decrease in PCa metastasis and death, the ubiquitous use of PSA screening has also led to overdetection and overtreatment (Schroder et al. N Engl J Med 360(13): 1320–8, 2009). Since all prostate epithelial cells synthesize PSA, an elevated PSA can reflect the presence of cancer but can also be caused by benign prostatic hyperplasia (BPH), infection, and/or chronic inflammation. Therefore, there has been a concerted effort to discover and validate novel PCa biomarkers. This chapter discusses (1) the challenges of PCa biomarker research, the types of PCa biomarkers, and the statistical considerations for biomarker discovery and validation; (2) the isoforms of PSA and their clinical applications; (3) several promising blood-based biomarkers for PCa diagnosis and/or prognostication (i.e., human kallikrein-related peptidase 2, urokinase plasminogen activator, transforming growth factor-beta 1, interleukin-6, endoglin, and prostate cancer specific autoantibodies and alpha-methylacyl-CoA racemase); and (4) the benefit of and need for combining biomarkers into different panels for each disease state.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Benign Prostatic Hyperplasia
- Radical Prostatectomy
- Prostate Biopsy
- Biochemical Recurrence
- Digital Rectal Exam
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Although the wide availability of prostate-specific antigen (PSA) has revolutionized prostate cancer (PCa) screening, resulting in a decrease in PCa metastasis and death, the ubiquitous use of PSA screening has also led to overdetection and overtreatment [1]. Since all prostate epithelial cells synthesize PSA, an elevated PSA can reflect the presence of cancer but can also be caused by benign prostatic hyperplasia (BPH), infection, and/or chronic inflammation. Therefore, there has been a concerted effort to discover and validate novel PCa biomarkers. This chapter discusses (1) the challenges of PCa biomarker research, the types of PCa biomarkers, and the statistical considerations for biomarker discovery and validation; (2) the isoforms of PSA and their clinical applications; (3) several promising blood-based biomarkers for PCa diagnosis and/or prognostication (i.e., human kallikrein-related peptidase 2, urokinase plasminogen activator, transforming growth factor-beta 1, interleukin-6, endoglin, and prostate cancer specific autoantibodies and alpha-methylacyl-CoA racemase); and (4) the benefit of and need for combining biomarkers into different panels for each disease state.
Biomarker Challenge in Prostate Cancer
A PubMed search for “prostate cancer” and “biomarker” or “marker” in the English language yielded 3,159 hits (accessed 7/10/2011). Despite the plethora of biomarkers reported to be “promising,” only one biomarker—total PSA in blood—is routinely used by urologists. Why are PCa biomarkers not living up to their promise? The answer lies not in a lack of pathophysiological understanding, biochemical techniques, or research funding but instead in the inherent difficulties of the pre-analytical, analytical, and post-analytical stages of biomarker discovery and validation [2–4]. Great care must be taken to standardize sample collection and/or storage conditions. The assay(s) employed must be accurate and precise. The number of test samples must be large enough so that the results are statistically significant. It is necessary to be able to generalize the assay to a diverse population and to standardize it for easy commercial use. Finally, the biomarker must provide additional useful clinical information in a cost-effective manner.
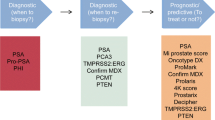
According to the NIH, a biomarker is a characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes, or pharmaceutical responses to a therapeutic intervention [5]. Prostate cancer biomarkers can be categorized into six different functional groups:
-
1.
Detection/screening – This biomarker is used for evaluating patients with either risk factors for or symptoms of PCa.
-
2.
Diagnostic – This biomarker can help classical histopathological characteristics in assessing the presence or absence of cancer.
-
3.
Prognostic – This biomarker is used to predict the outcome of patients based on different risks of recurrence or progression thereby allowing individualized management.
-
4.
Predictive – This biomarker is used to predict whether the treatment (drug or other therapy) will be effective and/or monitor the effectiveness of the treatment. It can help identify the best treatment modality.
-
5.
Therapeutic target – This biomarker can help identify the patients who will benefit from a particular treatment regimen. It identifies the molecular targets of novel therapies and is affected by therapy.
-
6.
Surrogate endpoint – This biomarker is used to substitute for a clinical endpoint and/or to measure clinical benefit, such as mortality due to disease or the recurrence or relapse of disease. Biomarkers can reduce time factors and costs for Phase I and II clinical trials by replacing clinical endpoints.
In 2002, the National Cancer Institute’s Early Detection Research Network (NCI EDRN) developed a five-phase approach to systematic discovery and validation of biomarkers (Fig. 7.1) [2, 6–9]. The schema directs researchers and clinicians in designing and carrying out the biomarker development process and delineates the extensive process necessary from discovery to clinical application. The process is long and costly and a high rate of failure is to be expected. Therefore, each stage of the process must be well considered, particularly study design, patient selection, and statistical analysis.
Most biomarkers do not provide sufficient information to be used independently of other information. The optimal use of biomarkers lies in incorporating them in a model that also includes standard clinical data [2, 3, 7, 10–12]. The model would then be used to provide individual patient care for use in improving diagnosis or treatment. To determine the value of a new biomarker, it is not sufficient to show that it is significantly related to the outcome, statistically significant in a multivariable model including the standard clinical and pathologic factors, or more significant than the standard clinical and pathologic factors [13, 14]. A variable that is statistically significant in a multivariable model might not improve the model’s predictive accuracy. P-value and odds/hazard ratio do not meaningfully describe a biomarker’s ability to classify patients. For a biomarker to be potentially clinically useful, it is necessary to show that adding the biomarker to an existing model based on the most important clinical and pathologic factors improves the predictive accuracy (discrimination and calibration) of the model [2, 13, 15–18].
One major issue with model development is the need for appropriate validation. When one develops a model incorporating a biomarker, a set of patients is used to develop the model. By definition, the model is most accurate in predicting the outcome for this set of patients. Therefore biomarker-based models need to be validated on data not used to develop it. There are two general types of validation: internal validation on the original dataset and external validation on an independent dataset (preferred) [2, 6]. External validation on a different dataset evaluates whether the risk prediction tool can be generalized to wider populations than the original dataset. Biomarkers that provide a continuous score provide potentially more useful information than cut points since risk levels are not truly discrete but a continuum of risk [4, 19]. Finally, methods that incorporate clinical consequences such as decision curve analysis are crucial to the evaluation of biomarkers [14, 18, 20]. This type of analysis allows insight into the consequences of using a biomarker in the clinic. Several methods are available including decision curve analysis, which combines simplicity with efficient computations [21–24]. Decision analytic evaluation should be performed during later stages of research before clinical implementation of the biomarker.
PSA Molecular Isoforms
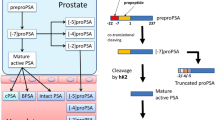
Enhancing the diagnostic accuracy of total PSA (tPSA), particularly specificity, is critical, since higher specificity would reduce the number of biopsies performed in men not affected by PCa. Several different strategies have been investigated, including the use of age-specific tPSA cutoffs, tPSA density, tPSA density of the transition zone, tPSA velocity, and the measurement of various molecular forms of PSA [25–28]. Prostate-specific antigen exists in free and complexed forms in serum. Improvements in measuring PSA isoforms have allowed the measurement of free PSA (fPSA) and its ratio to tPSA [29–31]. Of the fPSA portion, there are three distinct cleavage isoforms: proPSA, BPH-associated PSA (BPSA), and intact free PSA [32]. The precursor of PSA is a 261 amino acid pre-pro-protein. Subsequent processing by human kallikrein 2 (hK2) and other proteases produces the active 237 amino acid mature PSA [32]. Complexed PSA is a measure of how much PSA in serum is bound to α2-macroglobulin, α1-protease inhibitor, or α1-antichymotrypsin. Currently, there is no commercially available assay which specifically measures the complexes of α2-macroglobulin with PSA.
The FDA has approved the use of percent fPSA testing [i.e., (fPSA/tPSA) × 100] as an adjunct to tPSA in men with a serum tPSA concentration between 4 and 10 ng/mL. A higher percent fPSA (%fPSA) value indicates a lower probability of finding PCa on biopsy and raises the likelihood that the elevation in tPSA is due to the presence of BPH [33, 34]. In a multicenter, prospective trial, Catalona et al. reported that when a %fPSA of <25% is used for triggering a sextant prostate biopsy, it yielded a 95% sensitivity for PCa detection and increased the specificity by 20% over PSA alone [33]. In response to the realization that sextant biopsies misclassify up to one-third of patients who have PCa as without cancer, a more recent evaluation of the utility of percent free PSA in patients undergoing extended 10- or 12-core biopsy has suggested a lower diagnostic efficiency of %fPSA [35]. A meta-analysis of 41 studies examining fPSA in patients with tPSA between 4 and 10 ng/mL found that a test cutoff of 20% would lead to 94% sensitivity and 13% specificity and that likelihood ratios exceeded 2.0 only at %fPSA of 7% or less [36], suggesting that %fPSA only improves clinical information at extreme values. While most investigators agree that %fPSA can improve the diagnostic performance of tPSA between 4 and 10 ng/mL, the most appropriate %fPSA cutoff value remains debatable.
The utility of fPSA has also been examined in the tPSA ≤ 4 ng/mL range. Catalona et al. determined that with a %fPSA cutoff of less or equal to 27%, they were able to obtain a sensitivity of 90% and avoid 18% of unnecessary biopsies in men over the age of 50 with a tPSA of 2.6–4.0 ng/mL, with 83% of these cancers being clinically significant [37]. Rowe et al. used a %fPSA cutoff of 20% in patients age 50–65 with tPSA 1.1–3.99 ng/mL and found PCa in 11.3% of the patients biopsied [38]. Pepe et al. used %fPSA thresholds of 15% for patients with tPSA ≤ 2.5 ng/mL and 20% for patients with tPSA between 2.6 and 4 ng/mL and found PCa in 25.6% and 27.4% of the patients, respectively [39]. These studies suggest that %fPSA can help detect PCa in patients with tPSA below 4 ng/mL but the optimal cutoff threshold remains to be determined.
Data on the usefulness of fPSA to predict clinical outcomes is also inconclusive. Graefen and coworkers [40] failed to detect an independent association of preoperative free PSA with biochemical failure in 581 unscreened patients who underwent radical prostatectomy for clinically localized PCa [40]. In contrast, Shariat and colleagues found that lower preoperative serum fPSA is an independent predictor of advanced pathologic features, biochemical progression, and patterns of aggressive disease progression in 402 consecutive men treated with radical prostatectomy for clinically localized PCa who had tPSA levels less than 10 ng/mL [30].
ProPSA
Studies have shown that higher levels of proPSA are associated with PCa. In men with PSA levels between 6.0 and 24.0 ng/ml, the [−2]proPSA fraction was found to be significantly higher in men with PCa [32, 41]. Moreover, authors demonstrated the utility of the proPSA to fPSA ratio for screening patients with PSA levels between 2.5 and 4.0 ng/mL and between 4.0 and 10.0 ng/mL [42]. Elevated proPSA to fPSA ratios have also been associated with pathologic features of aggressive disease and decreased biochemical disease-free survival after radical prostatectomy [43, 44]. A new automated tool using the [−2]proPSA assay with a %fPSA-based artificial neural network was capable of detecting PCa and the PCa subgroup with more aggressive features with higher accuracy than tPSA or %fPSA alone [45]. In a prospective cohort of men enrolled into active surveillance for PCa, serum and tissue levels of proPSA at diagnosis were associated with the need for subsequent treatment [46]. The authors hypothesized that the increase in the ratio of serum proPSA to %fPSA might be driven by increased proPSA production from “premalignant” cells. In a prospective multicenter cancer detection study, the addition of %[−2]proPSA (defined as [−2]proPSA/10/fPSA) to a logistic regression model including clinical and demographic factors, PSA, and fPSA improved the model’s diagnostic accuracy, particularly in the clinically significant 2–10 ng/mL PSA range [47]. In addition, the authors observed that [−2]proPSA may be associated with more aggressive features of PCa [47], suggesting a role for proPSA as a staging and prognostic biomarker.
Molecular forms of PSA may differ in their in vitro stability properties. Therefore, information about the pre-analytical conditions is essential for proper clinical interpretation. For proper measurement of [−2]proPSA, blood samples should be centrifuged within 3 h of blood draw. Serum may be stored at room temperature or refrigerated (+4 °C) for a maximum of 48 h and should be frozen if stored for a longer period. However, two freeze-thaw cycles have no effect on [−2]proPSA stability [48].
BPSA
BPH-associated PSA (BPSA) is formed by the internal cleavage of PSA between Lys182 and Ser183. BPSA is expressed in nodular hyperplasia limited to the transition zone of men with BPH. BPSA can be detected in semen, blood, and prostate, and its levels correlate with transition zone volume and obstructive voiding symptoms [32, 49]. BPSA seems to be a promising marker of BPH since a direct association between its secretion and the volume of the transition zone has been shown to exist [49]. As such, BPSA is a better predictor of prostate enlargement than total and free PSA [50]. In addition, BPSA is not affected by age and is significantly higher in the presence of BPH symptoms. Adjusting the level of fPSA for BPSA resulted in 13–17% improvement in specificity compared to fPSA alone, while maintaining a sensitivity of 90–95% in PCa diagnosis [51].
Intact and Nicked PSA
Intact PSA includes both mature and proPSA single-chain PSA, whereas nicked PSA is PSA that has been internally cleaved between Lys145 and Lys146. The level of intact PSA and ratio of nicked to tPSA have shown potential for improving the discrimination of PCa from BPH [52, 53]. Similar to fPSA, intact PSA levels degrade with freezing, storage, and thawing [52]. The increasing amount of information on these PSA-related markers, together with the clinical parameters, calls for further assessment and integration into diagnostic and prognostic instruments that could serve the daily practice of early detection and screening for PCa.
Emerging Blood-Based Prostate Cancer Biomarkers
Human Kallikrein 2 (hK2)
Human kallikrein-related peptidase 2 is a secreted serine protease from the same gene family as PSA [54]. They share 80% sequence homology and are both primarily expressed in the prostate gland [54]. Despite these structural similarities, hK2 and tPSA differ in their enzymatic activities. The levels of hK2 in prostate tissue, plasma, semen, and serum are less than 2% that of tPSA, although hK2 mRNA transcript expression represents half that of total PSA. Similar to tPSA, serum hK2 is present in two forms in the blood: one bound to various protease inhibitors and the other (preponderant) free in the circulation. Several studies have shown that, when used in conjunction with free and total PSA, serum hK2 could improve the discrimination of men with PCa from men without cancer [55–57]. It has also been suggested that hK2 could predict poor differentiation, extracapsular extension, and biochemical recurrence in patients treated with radical prostatectomy [58–60]. However, this finding has not been validated by other authors [61]. The usefulness of hK2 for the preoperative staging of localized PCa, therefore, remains controversial. The addition of hK2 to three other kallikreins (total, free, and intact PSA) improved the prediction of prostate biopsy results in men with elevated tPSA (increase of predictive accuracy from 68% to 72% to ∼83%) [17]. Considering the risk of PCa at 20%, the number of biopsies would have been reduced by half, missing 3 out of 40 high-grade tumors [17]. This shows that if hK2 is to be used in PCa management, it can only be useful in a panel of biomarkers.
Urokinase Plasminogen Activator (uPA)
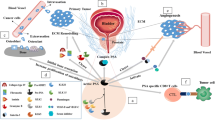
The urokinase plasminogen activation axis represents a potential target for PCa markers by being involved in various phases of tumor development and progression through degradation of the extracellular matrix. The serum protease uPA may play a role in cancer progression by binding to the uPA receptor (uPAR) and consequently converting plasminogen to plasmin, which activates proteases related to the degradation of extracellular matrix proteins [62]. Immunohistochemical staining of radical prostatectomy specimens revealed that overexpression of both uPA and its inhibitor (PAI-1) was associated with aggressive PCa recurrence [63]. In patients with a tPSA level above 2 ng/mL, soluble uPAR and fPSA measured in serum before prostate biopsy improved the regression model accuracy for prediction of PCa [64]. Steuber et al. have shown that uPAR fragments were significant predictors of PCa on biopsy specimens of patients with an elevated PSA [64].
Both uPA and uPAR might also have prognostic value. Elevated circulating levels of uPA and uPAR have been linked to PCa stage and bone metastases [63, 65–67]. In a study of 429 patients treated with radical prostatectomy, preoperative plasma uPA was a strong predictor of biochemical recurrence. Both preoperative uPA and uPAR were associated with features of aggressive biochemical recurrence, such as development of distant metastasis and fast PSA doubling time, suggesting an association with occult metastatic disease at the time of local therapy. Moreover, elevation of plasma uPA and uPAR levels in PCa patients seemed to be partly caused by local release from the prostate. Larger multi-institutional studies are under way to validate the potential role of uPA and uPAR as markers of early metastatic PCa.
Transforming Growth Factor-Beta 1 (TGF-β1) and Interleukin-6 (IL-6)
Transforming growth factor-beta 1 is a growth factor involved in the regulation of several cellular mechanisms including proliferation, immune response, differentiation, and angiogenesis [68]. TGF-β1 has been shown to promote cell progression in PCa models, and its local expression has been associated with higher tumor grade, tumor invasion, and metastasis in PCa patients [69–71]. Several studies have shown that increased levels of circulating TGF-β1 were associated with cancer progression, occult and documented metastasis, and biochemical progression in PCa patients [70, 72, 73].
Interleukin-6 is a cytokine with variable effects on immune and hematopoietic mechanisms. In vitro and in vivo studies have shown that both IL-6 and its receptor (IL-6R) were expressed in PCa [74, 75]. Several authors reported that elevated serum levels of IL-6 and/or IL-6R were associated with metastatic and hormone refractory disease and suggested that IL-6 could predict progression and survival of PCa patients [76–78].
Based on these findings, Kattan and associates developed and internally validated a prognostic model that incorporates plasma TGF-β1 and IL-6R into a standard nomogram for prediction of biochemical recurrence following radical prostatectomy [15]. This combination of serum markers and classical clinical parameters improved the predictive accuracy by a statistically and prognostically substantial margin (increase in predictive accuracy from 75% to 84%). However, before a biomarker can become useful in daily clinical management, it needs to be externally validated in an independent cohort of patients (Fig. 7.1) [2, 6]. Therefore, in a multi-institutional dataset of 423 patients treated with radical prostatectomy, Shariat et al. confirmed that plasma levels of TGF-β1 and IL-6R considerably enhanced the accuracy of the standard preoperative nomogram for the prediction of biochemical recurrence (accuracy of clinical features plus biomarkers 87.9% vs. 71.1% for clinical features alone; p < 0.001). Such prognostic models refine our ability to identify patients at a high risk of biochemical recurrence after radical prostatectomy that may benefit from inclusion into perioperative clinical trials and other intensified follow-up protocols.
Endoglin
Endoglin, or CD 105, is a transmembrane glycoprotein that is typically expressed by human vascular endothelial cells. Functionally, it is a cell-surface coreceptor for TGF-β1 and -β3 [79], which modulates cellular responses to TGF-β in the early steps of endothelial cell proliferation. Its critical role in angiogenesis has prompted investigators to evaluate the role of endoglin in cancer progression and metastasis. In PCa, endoglin is preferentially found on new, immature blood vessels, and immunohistochemical analysis supports an association between endoglin expression and disease progression [80]. Urine levels of endoglin may distinguish patients with PCa and may help in the staging of the disease [81]. In addition, preoperative plasma endoglin levels were found to be associated with metastasis to regional lymph nodes [82], as well as established features of biologically aggressive PCa such as higher pathologic Gleason sum and biochemical recurrence following radical prostatectomy [83]. Use of preoperative plasma endoglin could help decide whether and how extensively to perform a lymphadenectomy, as well as preoperative identification of patients at risk for disease progression. This would help select patients for neoadjuvant and/or adjuvant therapy or enrollment into clinical trials. Moreover, endoglin may be valuable as a surrogate biomarker for occult metastatic disease in patients with presumed organ-confined disease. Further investigation is needed to validate endoglin as a useful biomarker in men with PCa and to elucidate the mechanistic role of this biomarker in the progression of PCa.
Prostate Cancer Specific Autoantibodies and α-Methylacyl-CoA Racemase (AMACR)
Autoantibodies, such as those detected in autoimmune and infectious diseases, can be produced by cancer patients in response to tumor-associated antigens overexpressed in cancerous cells. α-Methylacyl-CoA racemase is an enzyme involved in fat metabolism, which has a strong expression in PCa tissues [84]. Immunostaining, using monoclonal antibodies to AMACR, is often used for the diagnosis of PCa, given its high diagnostic accuracy (sensitivity of 97% and specificity of 92%) [85]. A humoral response to tumor-related autoantibodies can be detected in the serum through amplification with high affinity antibodies and T cells. Autoantibodies to AMACR have been detected in the blood of PCa patients, and a recent study showed that they could help distinguish cancerous from healthy patients with more accuracy than PSA [86, 87]. Other autoantibodies to antigens expressed in PCa (Huntington-interacting protein 1, protasomes) have also been detected, and it has been reported that their combination could improve the screening performance, reaching a specificity of 97% [88]. Using a so-called “immunomics” technique, Wang et al. analyzed the overall humoral response against specific tumoral antigens in PCa and were able to identify multiple antigens [89]. With this panel of autoantibodies, they could detect PCa with a sensitivity of 81.6% and a specificity of 88.2%, which was more accurate than PSA alone. Prior et al. found that combining AMACR, MMP-2, and methylation of GSTP1/RASSF1A with PSA led to a significant improvement in PCa detection over PSA alone with specificities up to 96.6% [90]. Additional studies are needed to validate the potential prognostic value of autoantibodies and AMACR in PCa.
Combination of Multiple Biomarkers for Improved Cancer Detection and/or Prognostication
In the course of validating new biomarkers for PCa, a model combining the new biomarker with PSA is often used and shown to be superior to PSA alone. By combining a panel of biomarkers with varied individual sensitivities and specificities, it is possible to create a model with improved predictive accuracy. Multiple biomarkers are more likely to capture the complex biological potential of the heterogeneous prostate cancer population. For example, Cao et al. combined PCA3, TMPRSS2-ERG, Annexin A3, and sarcosine into a multimodality biomarker panel that outperforms any single biomarker that functions robustly in patients with PSA 4–10 ng/mL [91]. Additional studies are required to optimize and validate this panel. A biomarker may reflect disruption of a biochemical pathway by a particular mechanism. Given the complexity of the molecular abnormalities associated with PCa, it is improbable that a single marker can accurately segregate tumors of similar clinicopathologic phenotypes into distinct prognostic categories. Therefore, combinations of independent, yet complementary markers, may provide a more accurate prediction of outcomes compared to a single marker [27]. The future of cancer profiling relies on the combination of a panel of complimentary biomarkers that can give accurate molecular staging and indicate the likelihood of aggressive behavior [7, 11, 20, 92].
The group of Vickers and Lilja has developed a statistical model that predicts prostate biopsy outcomes based on age, digital rectal exam (DRE), and a panel of four kallikrein markers—tPSA, fPSA, intact PSA, and hK2. Using data from the randomized prostate cancer screening trial in Göteborg, Sweden [one center of the European Randomized Study of Prostate Cancer Screening (ERSPC)], they estimated that, for every 1,000 previously unscreened men with elevated total PSA, use of the model to determine biopsy would reduce biopsy rates by 573, while missing only a small number of cancers (31 out of 152 low-grade cancers and 3 out of 40 high-grade cancers) [17]. These findings were subsequently replicated in independent cohorts, reducing the number of biopsies by 50% and recommending against biopsy primarily in men with low-grade cancer [93, 94]. These findings have also been verified in men who recently have undergone previous screening, with resultant improvements in predictive accuracy [95, 96]. Gupta et al. demonstrated that the panel of four kallikrein markers can predict the outcome of prostate biopsy in men who had previously undergone prostate biopsy during previous screening [97]. This model, in addition to age and DRE, substantially improved the predictive accuracy of a base model (comprising of total PSA, age, and DRE), for both low- and high-grade cancers.
Shariat et al. found that the addition of a panel comprised of preoperative plasma levels of TGF-β1, soluble IL-6R, IL-6, endoglin, vascular endothelial growth factor (VEGF), and vascular cell adhesion molecule-1 (VCAM-1 or CD 106) [67, 68, 70, 82, 83, 98–100] improved the predictive accuracy of the Kattan preoperative nomogram [10] by 15.0% (i.e., 71.6–86.6%) [7, 20]. This increase substantially exceeds accuracy gains obtained from the consideration of detailed pathologic descriptors of PCa at radical prostatectomy. Svatek et al. confirmed the strong predictive value of preoperative levels of the candidate biomarkers after adjusting for the effect of postoperative features [92]. The addition of preoperative levels of the candidate biomarkers improved the accuracy of the base model (i.e., tPSA, surgical margin status, extracapsular extension, seminal vesicles invasion, lymph node involvement, and pathologic Gleason sum) for prediction of biochemical recurrence by a statistically and prognostically significant margin (79–86%, p < 0.001). Predictive tools integrating biomarker levels could constitute the new standard for counseling patients regarding their risk of recurrence following curative therapy and for designing clinical trials to test neoadjuvant and/or adjuvant treatment strategies in high-risk patients. However, while prediction of biochemical recurrence is important, prediction of response to therapy as well as metastasis and survival is more important for the management of PCa patients [101].
Conclusion
In the PSA era of PCa diagnosis, PCa screening remains controversial due to the risk of overdiagnosis, overtreatment, and the inability to differentiate aggressive tumors. Therefore, new biomarkers are greatly needed to improve the sensitivity and specificity of PCa diagnosis. A substantial amount of effort and funding has been and continues to be invested in the search for new biomarkers and nomograms to improve PCa diagnosis and prognostication by a clinically significant margin. A panel including multiple biomarkers utilizing blood-based, protein-based, gene-based, and/or urine-based modalities in combination with multiple forms of PSA may be necessary to obtain optimal predictive accuracy for PCa diagnosis and prognostication.
Editorial Commentary:
It seems like barely a week goes by when a representative or scientific liaison of some company approaches me regarding new or improved biomarkers or prediction methodologies. The consistent finding is that they seem to all passionately believe that they bring immense value by providing the clinician with more information. Although that concept is intuitively appealing, it has become clear that, “no, I don’t need more information—I need actionable information.” Despite impressive statistics suggesting a value to the role of the markers discussed in this chapter and other technologies on the horizon, so far none of these candidates has delivered significant value to the clinician or patient, so none has developed broad acceptance in the diagnostic toolbox.
Nevertheless, further work in this arena is undeniably worthwhile based on inadequacy of the king of all markers—PSA. A number of investigators are performing exciting work in this arena, and we are hopeful that this will change soon. Concepts such as the use of panels as described, plus further investigation to define markers that can truly predict—not just give “more information”—are one of the more open fields of discovery at this point in time.
–J. Stephen Jones
References
Schroder FH, et al. Screening and prostate-cancer mortality in a randomized European study. N Engl J Med. 2009;360(13):1320–8.
Shariat SF, et al. Beyond prostate-specific antigen: new serologic biomarkers for improved diagnosis and management of prostate cancer. Rev Urol. 2004;6(2): 58–72.
Shariat SF, Karakiewicz PI. Perspectives on prostate cancer biomarkers. Eur Urol. 2008;54(1):8–10.
Shariat SF, et al. Tumor markers in prostate cancer I: blood-based markers. Acta Oncol. 2011;50 (Suppl 1): 61–75.
Atkinson AJ, et al. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin Pharmacol Ther. 2001;69(3):89–95.
Bensalah K, Montorsi F, Shariat SF. Challenges of cancer biomarker profiling. Eur Urol. 2007;52(6): 1601–9.
Bensalah K, et al. New circulating biomarkers for prostate cancer. Prostate Cancer Prostatic Dis. 2008;11(2):112–20.
Verma M, Srivastava S. New cancer biomarkers deriving from NCI early detection research. Recent Results Cancer Res. 2003;163:72–84. discussion 264–6.
Winget MD, et al. Development of common data elements: the experience of and recommendations from the early detection research network. Int J Med Inform. 2003;70(1):41–8.
Shariat SF, et al. An updated catalog of prostate cancer predictive tools. Cancer. 2008;113(11):3075–99.
Shariat SF, et al. New blood-based biomarkers for the diagnosis, staging and prognosis of prostate cancer. BJU Int. 2008;101(6):675–83.
Shariat SF, et al. Plasminogen activation inhibitor-1 improves the predictive accuracy of prostate cancer nomograms. J Urol. 2007;178(4 Pt 1):1229–36. discussion 1236–7.
Kattan MW. Judging new markers by their ability to improve predictive accuracy. J Natl Cancer Inst. 2003;95(9):634–5.
Vickers AJ, Kattan MW, Daniel S. Method for evaluating prediction models that apply the results of randomized trials to individual patients. Trials. 2007;8:14.
Kattan MW, et al. The addition of interleukin-6 soluble receptor and transforming growth factor beta1 improves a preoperative nomogram for predicting biochemical progression in patients with clinically localized prostate cancer. J Clin Oncol. 2003;21(19): 3573–9.
Shariat SF, et al. Improved prediction of disease relapse after radical prostatectomy through a panel of preoperative blood-based biomarkers. Clin Cancer Res. 2008;14(12):3785–91.
Vickers AJ, et al. A panel of kallikrein markers can reduce unnecessary biopsy for prostate cancer: data from the European Randomized Study of prostate cancer screening in Goteborg, Sweden. BMC Med. 2008;6:19.
Vickers AJ, et al. Systematic review of statistical methods used in molecular marker studies in cancer. Cancer. 2008;112(8):1862–8.
Thompson IM, et al. Operating characteristics of prostate-specific antigen in men with an initial PSA level of 3.0 ng/ml or lower. JAMA. 2005;294(1):66–70.
Shariat SF, et al. Multiple biomarkers improve prediction of bladder cancer recurrence and mortality in patients undergoing cystectomy. Cancer. 2008;112(2): 315–25.
Vickers AJ, Elkin EB. Decision curve analysis: a novel method for evaluating prediction models. Med Decis Making. 2006;26(6):565–74.
Vickers AJ, et al. Extensions to decision curve analysis, a novel method for evaluating diagnostic tests, prediction models and molecular markers. BMC Med Inform Decis Mak. 2008;8:53.
Vickers AJ. Decision analysis for the evaluation of diagnostic tests, prediction models and molecular markers. Am Stat. 2008;62(4):314–20.
Elkin EB, Vickers AJ, Kattan MW. Primer: using decision analysis to improve clinical decision making in urology. Nat Clin Pract Urol. 2006;3(8):439–48.
Lilja H, Ulmert D, Vickers AJ. Prostate-specific antigen and prostate cancer: prediction, detection and monitoring. Nat Rev Cancer. 2008;8(4):268–78.
Schroder FH, et al. Early detection of prostate cancer in 2007. Part 1: PSA and PSA kinetics. Eur Urol. 2008;53(3):468–77.
Shariat SF, Karam JA, Roehrborn CG. Blood biomarkers for prostate cancer detection and prognosis. Future Oncol. 2007;3(4):449–61.
Shariat SF, et al. Inventory of prostate cancer predictive tools. Curr Opin Urol. 2008;18(3):279–96.
Lilja H. Significance of different molecular forms of serum PSA. The free, noncomplexed form of PSA versus that complexed to alpha 1-antichymotrypsin (Review). Urol Clin North Am. 1993;20(4):681–6.
Shariat SF, et al. Pre-operative percent free PSA predicts clinical outcomes in patients treated with radical prostatectomy with total PSA levels below 10 ng/ml. Eur Urol. 2006;49(2):293–302.
Catalona WJ, et al. Comparison of percent free PSA, PSA density, and age-specific PSA cutoffs for prostate cancer detection and staging. Urology. 2000;56(2): 255–60.
Mikolajczyk SD, et al. Free prostate-specific antigen in serum is becoming more complex. Urology. 2002;59(6):797–802.
Catalona WJ, et al. Use of the percentage of free prostate-specific antigen to enhance differentiation of prostate cancer from benign prostatic disease: a prospective multicenter clinical trial. JAMA. 1998;279(19):1542–7.
Woodrum DL, et al. Interpretation of free prostate specific antigen clinical research studies for the detection of prostate cancer. J Urol. 1998;159(1):5–12.
Canto EI, et al. Effects of systematic 12-core biopsy on the performance of percent free prostate specific antigen for prostate cancer detection. J Urol. 2004;172(3):900–4.
Lee R, et al. A meta-analysis of the performance characteristics of the free prostate-specific antigen test. Urology. 2006;67(4):762–8.
Catalona WJ, Smith DS, Ornstein DK. Prostate cancer detection in men with serum PSA concentrations of 2.6 to 4.0 ng/mL and benign prostate examination. Enhancement of specificity with free PSA measurements. JAMA. 1997;277(18):1452–5.
Rowe EW, et al. Prostate cancer detection in men with a ‘normal’ total prostate-specific antigen (PSA) level using percentage free PSA: a prospective screening study. BJU Int. 2005;95(9):1249–52.
Pepe P, et al. Prevalence and clinical significance of prostate cancer among 12,682 men with normal digital rectal examination, low PSA levels (< or =4 ng/ml) and percent free PSA cutoff values of 15 and 20%. Urol Int. 2007;78(4):308–12.
Graefen M, et al. Percent free prostate specific antigen is not an independent predictor of organ confinement or prostate specific antigen recurrence in unscreened patients with localized prostate cancer treated with radical prostatectomy. J Urol. 2002;167(3):1306–9.
Mikolajczyk SD, et al. A truncated precursor form of prostate-specific antigen is a more specific serum marker of prostate cancer. Cancer Res. 2001;61(18): 6958–63.
Sokoll LJ, et al. Proenzyme psa for the early detection of prostate cancer in the 2.5–4.0 ng/ml total psa range: preliminary analysis. Urology. 2003;61(2):274–6.
Catalona WJ, et al. Serum pro-prostate specific antigen preferentially detects aggressive prostate cancers in men with 2 to 4 ng/ml prostate specific antigen. J Urol. 2004;171(6 Pt 1):2239–44.
Catalona WJ, et al. Serum pro prostate specific antigen improves cancer detection compared to free and complexed prostate specific antigen in men with prostate specific antigen 2 to 4 ng/ml. J Urol. 2003;170(6 Pt 1):2181–5.
Stephan C, et al. A [−2]proPSA-based artificial neural network significantly improves differentiation between prostate cancer and benign prostatic diseases. Prostate. 2009;69(2):198–207.
Makarov DV, et al. Pro-prostate-specific antigen measurements in serum and tissue are associated with treatment necessity among men enrolled in expectant management for prostate cancer. Clin Cancer Res. 2009;15(23):7316–21.
Sokoll LJ, et al. A prospective, multicenter, National Cancer Institute early detection research network study of [−2]proPSA: improving prostate cancer detection and correlating with cancer aggressiveness. Cancer Epidemiol Biomarkers Prev. 2010;19(5): 1193–200.
Semjonow A, et al. Pre-analytical in-vitro stability of [−2]proPSA in blood and serum. Clin Biochem. 2010;43(10–11):926–8.
Mikolajczyk SD, et al. “BPSA,” a specific molecular form of free prostate-specific antigen, is found predominantly in the transition zone of patients with nodular benign prostatic hyperplasia. Urology. 2000;55(1):41–5.
Canto EI, et al. Serum BPSA outperforms both total PSA and free PSA as a predictor of prostatic enlargement in men without prostate cancer. Urology. 2004;63(5):905–10. discussion 910–1.
Stephan C, et al. Benign prostatic hyperplasia-associated free prostate-specific antigen improves detection of prostate cancer in an artificial neural network. Urology. 2009;74(4):873–7.
Nurmikko P, et al. Discrimination of prostate cancer from benign disease by plasma measurement of intact, free prostate-specific antigen lacking an internal cleavage site at Lys145-Lys146. Clin Chem. 2001;47(8):1415–23.
Steuber T, et al. Association of free-prostate specific antigen subfractions and human glandular kallikrein 2 with volume of benign and malignant prostatic tissue. Prostate. 2005;63(1):13–8.
Yousef GM, Diamandis EP. The new human tissue kallikrein gene family: structure, function, and association to disease. Endocr Rev. 2001;22(2):184–204.
Nam RK, et al. Serum human glandular kallikrein-2 protease levels predict the presence of prostate cancer among men with elevated prostate-specific antigen. J Clin Oncol. 2000;18(5):1036–42.
Becker C, et al. Clinical value of human glandular kallikrein 2 and free and total prostate-specific antigen in serum from a population of men with prostate-specific antigen levels 3.0 ng/mL or greater. Urology. 2000;55(5):694–9.
Becker C, et al. Discrimination of men with prostate cancer from those with benign disease by measurements of human glandular kallikrein 2 (HK2) in serum. J Urol. 2000;163(1):311–6.
Haese A, et al. Human glandular kallikrein 2 levels in serum for discrimination of pathologically organ-confined from locally-advanced prostate cancer in total PSA-levels below 10 ng/ml. Prostate. 2001;49(2): 101–9.
Kwiatkowski MK, et al. In prostatism patients the ratio of human glandular kallikrein to free PSA improves the discrimination between prostate cancer and benign hyperplasia within the diagnostic “gray zone” of total PSA 4 to 10 ng/mL. Urology. 1998;52(3):360–5.
Recker F, et al. The importance of human glandular kallikrein and its correlation with different prostate specific antigen serum forms in the detection of prostate carcinoma. Cancer. 1998;83(12):2540–7.
Kurek R, et al. Prognostic value of combined “triple”-reverse transcription-PCR analysis for prostate-specific antigen, human kallikrein 2, and prostate-specific membrane antigen mRNA in peripheral blood and lymph nodes of prostate cancer patients. Clin Cancer Res. 2004;10(17):5808–14.
Duffy MJ. Urokinase-type plasminogen activator: a potent marker of metastatic potential in human cancers. Biochem Soc Trans. 2002;30(2):207–10.
Gupta A, et al. Predictive value of the differential expression of the urokinase plasminogen activation axis in radical prostatectomy patients. Eur Urol. 2009;55(5):1124–33.
Steuber T, et al. Free PSA isoforms and intact and cleaved forms of urokinase plasminogen activator receptor in serum improve selection of patients for prostate cancer biopsy. Int J Cancer. 2007;120(7): 1499–504.
Hienert G, et al. Urokinase-type plasminogen activator as a marker for the formation of distant metastases in prostatic carcinomas. J Urol. 1988;140(6): 1466–9.
Miyake H, et al. Elevation of serum levels of urokinase-type plasminogen activator and its receptor is associated with disease progression and prognosis in patients with prostate cancer. Prostate. 1999;39(2): 123–9.
Shariat SF, et al. Association of the circulating levels of the urokinase system of plasminogen activation with the presence of prostate cancer and invasion, progression, and metastasis. J Clin Oncol. 2007;25(4):349–55.
Shariat SF, et al. Preoperative plasma levels of transforming growth factor beta(1) (TGF-beta(1)) strongly predict progression in patients undergoing radical prostatectomy. J Clin Oncol. 2001;19(11): 2856–64.
Truong LD, et al. Association of transforming growth factor-beta 1 with prostate cancer: an immunohistochemical study. Hum Pathol. 1993;24(1):4–9.
Shariat SF, et al. Association of pre- and postoperative plasma levels of transforming growth factor beta(1) and interleukin 6 and its soluble receptor with prostate cancer progression. Clin Cancer Res. 2004;10(6): 1992–9.
Shariat SF, et al. Tissue expression of transforming growth factor-beta1 and its receptors: correlation with pathologic features and biochemical progression in patients undergoing radical prostatectomy. Urology. 2004;63(6):1191–7.
Shariat SF, et al. Preoperative plasma levels of transforming growth factor beta(1) strongly predict clinical outcome in patients with bladder carcinoma. Cancer. 2001;92(12):2985–92.
Ivanovic V, et al. Elevated plasma levels of TGF-beta 1 in patients with invasive prostate cancer. Nat Med. 1995;1(4):282–4.
Hobisch A, et al. Interleukin-6 regulates prostate-specific protein expression in prostate carcinoma cells by activation of the androgen receptor. Cancer Res. 1998;58(20):4640–5.
Giri D, Ozen M, Ittmann M. Interleukin-6 is an autocrine growth factor in human prostate cancer. Am J Pathol. 2001;159(6):2159–65.
Michalaki V, et al. Serum levels of IL-6 and TNF-alpha correlate with clinicopathological features and patient survival in patients with prostate cancer. Br J Cancer. 2004;90(12):2312–6.
Nakashima J, et al. Serum interleukin 6 as a prognostic factor in patients with prostate cancer. Clin Cancer Res. 2000;6(7):2702–6.
Stark JR, et al. Circulating prediagnostic interleukin-6 and C-reactive protein and prostate cancer incidence and mortality. Int J Cancer. 2009;124(11):2683–9.
Cheifetz S, et al. Endoglin is a component of the transforming growth factor-beta receptor system in human endothelial cells. J Biol Chem. 1992;267(27): 19027–30.
Wikstrom P, et al. Endoglin (CD105) is expressed on immature blood vessels and is a marker for survival in prostate cancer. Prostate. 2002;51(4):268–75.
Fujita K, et al. Endoglin (CD105) as a urinary and serum marker of prostate cancer. Int J Cancer. 2009;124(3):664–9.
Karam JA, et al. Use of preoperative plasma endoglin for prediction of lymph node metastasis in patients with clinically localized prostate cancer. Clin Cancer Res. 2008;14(5):1418–22.
Svatek RS, et al. Preoperative plasma endoglin levels predict biochemical progression after radical prostatectomy. Clin Cancer Res. 2008;14(11):3362–6.
Rubin MA, et al. Alpha-Methylacyl coenzyme A racemase as a tissue biomarker for prostate cancer. JAMA. 2002;287(13):1662–70.
Jiang Z, et al. Alpha-methylacyl-CoA racemase: a multi-institutional study of a new prostate cancer marker. Histopathology. 2004;45(3):218–25.
Sreekumar A, et al. Humoral immune response to alpha-methylacyl-CoA racemase and prostate cancer. J Natl Cancer Inst. 2004;96(11):834–43.
Cardillo MR, et al. Can p503s, p504s and p510s gene expression in peripheral-blood be useful as a marker of prostatic cancer? BMC Cancer. 2005;5:111.
Bradley SV, et al. Serum antibodies to huntingtin interacting protein-1: a new blood test for prostate cancer. Cancer Res. 2005;65(10):4126–33.
Wang X, et al. Autoantibody signatures in prostate cancer. N Engl J Med. 2005;353(12):1224–35.
Prior C, et al. Use of a combination of biomarkers in serum and urine to improve detection of prostate cancer. World J Urol. 2010;28(6):681–6.
Cao DL, et al. A multiplex model of combining gene-based, protein-based, and metabolite-based with positive and negative markers in urine for the early diagnosis of prostate cancer. Prostate. 2011;71(7):700–10.
Svatek RS, et al. Pre-treatment biomarker levels improve the accuracy of post-prostatectomy nomogram for prediction of biochemical recurrence. Prostate. 2009;69(8):886–94.
Vickers A, et al. Reducing unnecessary biopsy during prostate cancer screening using a four-kallikrein panel: an independent replication. J Clin Oncol. 2010;28(15):2493–8.
Benchikh A, et al. A panel of kallikrein markers can predict outcome of prostate biopsy following clinical work-up: an independent validation study from the European Randomized Study of prostate cancer screening, France. BMC Cancer. 2010;10:635.
Vickers AJ, et al. Impact of recent screening on predicting the outcome of prostate cancer biopsy in men with elevated prostate-specific antigen: data from the European Randomized Study of prostate cancer screening in Gothenburg, Sweden. Cancer. 2010;116(11):2612–20.
Vickers AJ, et al. A four-kallikrein panel predicts prostate cancer in men with recent screening: data from the European Randomized Study of screening for prostate cancer, Rotterdam. Clin Cancer Res. 2010;16(12):3232–9.
Gupta A, et al. A four-kallikrein panel for the prediction of repeat prostate biopsy: data from the European Randomized Study of prostate cancer screening in Rotterdam, Netherlands. Br J Cancer. 2010;103(5):708–14
Shariat SF, et al. Plasma levels of interleukin-6 and its soluble receptor are associated with prostate cancer progression and metastasis. Urology. 2001;58(6):1008–15.
Shariat SF, et al. Association of preoperative plasma levels of vascular endothelial growth factor and soluble vascular cell adhesion molecule-1 with lymph node status and biochemical progression after radical prostatectomy. J Clin Oncol. 2004;22(9):1655–63.
Shariat SF, et al. External validation of a biomarker-based preoperative nomogram predicts biochemical recurrence after radical prostatectomy. J Clin Oncol. 2008;26(9):1526–31.
Shariat SF, et al. Detection of clinically significant, occult prostate cancer metastases in lymph nodes using a splice variant-specific rt-PCR assay for human glandular kallikrein. J Clin Oncol. 2003;21(7):1223–31.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Wang, L.C., Scherr, D.S., Shariat, S.F. (2013). Blood-Based Tumor Markers for Prostate Cancer. In: Jones, J. (eds) Prostate Cancer Diagnosis. Current Clinical Urology. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-188-2_7
Download citation
DOI: https://doi.org/10.1007/978-1-62703-188-2_7
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-187-5
Online ISBN: 978-1-62703-188-2
eBook Packages: MedicineMedicine (R0)