Abstract
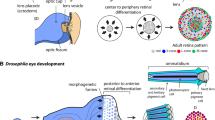
The Drosophila compound eye is a regular structure, in which about 750 units, called ommatidia, are arranged in a highly regular pattern. Eye development proceeds in a stereotypical fashion, where epithelial cells of the eye imaginal discs are specified, recruited, and differentiated in a sequential order that leads to the highly precise structure of an adult eye. Even small perturbations, for example in signaling pathways that control proliferation, cell death, or differentiation, can impair the regular structure of the eye, which can be easily detected and analyzed. In addition, the Drosophila eye has proven to be an ideal model for studying the genetic control of neurodegeneration, since the eye is not essential for viability. Several human neurodegeneration diseases have been modeled in the fly, leading to a better understanding of the function/misfunction of the respective gene. In many cases, the genes involved and their function are conserved between flies and human. More strikingly, when ectopically expressed in the fly eye some human genes without a Drosophila counterpart can induce neurodegeneration, detectable by aberrant phototaxis, impaired electrophysiology, or defects in eye morphology. These defects are often rather subtle alteration in shape, size, or arrangement of the cells, and can be easily scored at the ultrastructural level. This chapter aims to provide an overview regarding the analysis of the retina by various means.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wolff T, Ready DF (1991) Cell death in normal and rough eye mutants of Drosophila. Development 113:825–839
Baumann O, Lutz K (2006) Photoreceptor morphogenesis in the Drosophila compound eye: R1–R6 rhabdomeres become twisted just before eclosion. J Comp Neurol 498:68–79
Zuker CS, Cowman AF, Rubin GM (1985) Isolation and structure of a rhodopsin gene from D. melanogaster. Cell 40:851–858
O’Tousa JE, Baehr W, Martin RL, Hirsh J, Pak WL, Applebury ML (1985) The Drosophila ninaE gene encodes an opsin. Cell 40:839–850
Montell C, Jones K, Zuker C, Rubin G (1987) A second opsin gene expressed in the ultraviolet-sensitive R7 photoreceptor cells of Drosophila melanogaster. J Neurosci 7:1558–1566
Zuker CS, Montell C, Jones K, Laverty T, Rubin GM (1987) A rhodopsin gene expressed in photoreceptor cell R7 of the Drosophila eye: homologies with other signal-transducing molecules. J Neurosci 7:1550–1557
Feiler R, Bjornson R, Kirschfeld K, Mismer D, Rubin GM, Smith DP, Socolich M, Zuker CS (1992) Ectopic expression of ultraviolet-rhodopsins in the blue photoreceptor cells of Drosophila: visual physiology and photochemistry of transgenic animals. J Neurosci 12:3862–3868
Salcedo E, Huber A, Henrich S, Chadwell LV, Chou WH, Paulsen R, Britt SG (1999) Blue- and green-absorbing visual pigments of Drosophila: ectopic expression and physiological characterization of the R8 photoreceptor cell-specific Rh5 and Rh6 rhodopsins. J Neurosci 19:10716–10726
Harris WA, Stark WS, Walker JA (1976) Genetic dissection of the photoreceptor system in the compound eye of Drosophila melanogaster. J Physiol 256:415–439
Reinke R, Krantz DE, Yen D, Zipursky SL (1988) Chaoptin, a cell surface glycoprotein required for Drosophila photoreceptor cell morphogenesis, contains a repeat motif found in yeast and human. Cell 52:291–301
Tomlinson A, Bowtell DD, Hafen E, Rubin GM (1987) Localization of the sevenless protein, a putative receptor for positional information, in the eye imaginal disc of Drosophila. Cell 51:143–150
Campos-Ortega JA, Jürgens G, Hofbauer A (1979) Cell clones and pattern formation: studies on sevenless, a mutant of Drosophila melanogaster. Wilhelm Roux’s Arch Dev Biol 186:27–50
Tomlinson A, Kimmel BE, Rubin GM (1988) Rough, a Drosophila homeobox gene required in photoreceptors R2 and R5 for inductive interactions in the developing eye. Cell 55:771–784
Baker NE, Rubin GM (1989) Effect on eye development of dominant mutations in Drosophila homologue of the EGF receptor. Nature 340:150–153
Miyamoto H, Nihonmatsu I, Kondo S, Ueda R, Togashi S, Hirata K, Ikegami Y, Yamamoto D (1995) Canoe encodes a novel protein containing a GLGF/DHR motif and functions with Notch and scabrous in common developmental pathways in Drosophila. Genes Dev 9:612–625
Basler K, Christen B, Hafen E (1991) Ligand-independent activation of the sevenless receptor tyrosine kinase changes the fate of cells in the developing Drosophila eye. Cell 64:1069–1081
Grzeschik N, Knust E (2005) IrreC/rst-mediated cell sorting during Drosophila pupal eye development depends on proper localisation of DE-cadherin. Development 132:2035–2045
Yang Y, Ballinger D (1994) Mutations in calphotin, the gene encoding a Drosophila photoreceptor cell-specific calcium-binding protein, reveal roles in cellular morphogenesis and survival. Genetics 138:413–421
Mishra M, Oke A, Lebel C, McDonald EC, Plummer Z, Cook TA, Zelhof AC (2010) Pph13 and orthodenticle define a dual regulatory pathway for photoreceptor cell morphogenesis and function. Development 137:2895–2904
Zelhof AC, Koundakjian E, Scully AL, Hardy RW, Pounds L (2003) Mutation of the photoreceptor specific homeodomain gene Pph13 results in defects in phototransduction and rhabdomere morphogenesis. Development 130:4383–4392
Li BX, Satoh AK, Ready DF (2007) Myosin V, Rab11 and dRip11 direct apical secretion and cellular morphogenesis in Drosophila photoreceptor cells. J Cell Biol 177:659–669
Muschalik N, Knust E (2011) Increased levels of the cytoplasmic domain of Crumbs repolarise developing Drosophila photoreceptors. J Cell Sci 124:3715–3725
Richard M, Grawe F, Knust E (2006) DPATJ plays a role in retinal morphogenesis and protects against light-dependent degeneration of photoreceptor cells in the Drosophila eye. Dev Dyn 235:895–907
Johnson K, Grawe F, Grzeschik N, Knust E (2002) Drosophila Crumbs is required to inhibit light-induced photoreceptor degeneration. Curr Biol 12:1675–1680
Hong Y, Ackerman L, Jan LY, Jan Y-N (2003) Distinct roles of Bazooka and Stardust in the specification of Drosophila photoreceptor membrane architecture. Proc Natl Acad Sci U S A 100:12712–12717
Berger S, Bulgakova NA, Grawe F, Johnson K, Knust E (2007) Unravelling the genetic complexity of Drosophila stardust during photoreceptor morphogenesis and prevention of light-induced degeneration. Genetics 176:2189–2200
Pellikka M, Tanentzapf G, Pinto M, Smith C, McGlade CJ, Ready DF, Tepass U (2002) Crumbs, the Drosophila homologue of human CRB1/RP12, is essential for photoreceptor morphogenesis. Nature 416:143–149
Pham H, Yu H, Laski FA (2008) Cofilin/ADF is required for retinal elongation and morphogenesis of the Drosophila rhabdomere. Dev Biol 318:82–91
Matsuo T, Takahashi K, Suzuki E, Yamamoto D (1999) The Canoe protein is necessary in adherens junctions for development of ommatidial architecture in the Drosophila compound eye. Cell Tissue Res 298:397–404
Husain N, Pellikka M, Hong H, Klimentova T, Choe K-M, Clandinin TR, Tepass U (2006) The Agrin/perlecan-related protein eyes shut is essential for epithelial lumen formation in the Drosophila retina. Dev Cell 11:483–493
Zelhof AC, Hardy RW, Becker A, Zuker CS (2006) Transforming the architecture of compound eyes. Nature 443:696–699
Cheli VT, Daniels RW, Godoy R, Hoyle DJ, Kandachar V, Starcevic M, Martinez-Agosto JA, Poole S, DiAntonio A, Lloyd VK, Chang HC, Krantz DE, Dell’Angelica EC (2010) Genetic modifiers of abnormal organelle biogenesis in a Drosophila model of BLOC-1 deficiency. Hum Mol Genet 19:861–878
Pulipparacharuvil S, Akbar MA, Ray S, Sevrioukov EA, Haberman AS, Rohrer J, Kramer H (2005) Drosophila Vps16A is required for trafficking to lysosomes and biogenesis of pigment granules. J Cell Sci 118:3663–3673
Wu CF, Wong F (1977) Frequency characteristics in the visual system of Drosophila: genetic dissection of electroretinogram components. J Gen Physiol 69:705–724
Hardie RC, Postma M (2008) Phototransduction in microvillar photoreceptors of Drosophila and other invertebrates. In: Basbaum AI, Kaneko A, Shephard GM, Westheimer G (eds) The senses: a comprehensive reference. Academic, San Diego, pp 77–130
Pak WL (2010) Why Drosophila to study phototransduction? J Neurogenet 24:55–66
Pak WL, Grossfield J, Whiten V (1969) Non- phototactic mutants in a study of vision of Drosophila. Nature 222:351–354
Hotta Y, Benzer S (1969) Abnormal electroretinograms in visual mutants of Drosophila. Nature 222:354–356
Heisenberg M (1971) Isolation of mutants lacking the optomotor response. Drosoph lnf Serv 112:65–93
Heisenberg M (1997) Genetic approach to neuroethology. Bioessays 19:1065–1073
Franceschini N (1972) Pupil and pseudopupil in the compound eye of Drosophila. In: Wehner R (ed) Information processing in the visual systems of arthropods. Springer, Berlin, pp 75–82
Steele F, O’Tousa JE (1990) Rhodopsin activation causes retinal degeneration in Drosophila rdgC mutant. Neuron 4:883–890
Pichaud F, Desplan C (2001) A new visualization approach for identifying mutations taht affect differentiation and organization of the Drosophila ommatidia. Development 128:815–826
Meyer NE, Joel-Almagor T, Frechter S, Minke B, Huber A (2006) Subcellular translocation of the eGFP-tagged TRPL channel in Drosophila photoreceptors requires activation of the phototransduction cascade. J Cell Sci 119:2592–2603
Pandey UB, Nichols CD (2011) Human disease models in Drosophila melanogaster and the role of the fly in therapeutic drug discovery. Pharmacol Rev 63:411–436
Whitworth AJ (2011) Drosophila models of Parkinson’s disease. Adv Genet 73:1–50
Ambegaokar SS, Roy B, Jackson GR (2010) Neurodegenerative models in Drosophila: polyglutamine disorders, Parkinson disease, and amyotrophic lateral sclerosis. Neurobiol Dis 40:29–39
St. Johnston D (2002) The art and design of genetic screens: Drosophila melanogaster. Nat Rev Genet 31:176–188
Wang T, Montell C (2007) Phototransduction and retinal degeneration in Drosophila. Pflugers Arch 454:821–847
Rogge RD, Karlovich CA, Banerjee U (1991) Genetic dissection of a neurodevelopmental pathway: son of sevenless functions downstream of the sevenless and EGF receptor tyrosine kinases. Cell 64:39–48
Bonfini L, Karlovich CA, Dasgupta C, Banerjee U (1992) The son of sevenless gene product: a putative activator of Ras. Science 255:603–606
Shulman JM, Feany MB (2003) Genetic modifiers of tauopathy in Drosophila. Genetics 165:1233–1242
Garen SH, Kankel DR (1983) Golgi and genetic mosaic analyses of visual system mutants in Drosophila melanogaster. Dev Biol 96:445–466
Becker HJ (1957) Über Röntgenmosaikflecken und Defektmutationen am Auge von Drosophila und die Entwicklungsphysiologie des Auges. Z Indukt Abstamm Vererbungsl 88:333–373
Stern C (1936) Somatic crossing over and segregation in Drosophila melanogaster. Genetics 21:625–730
Thaker HM, Kankel DR (1992) Mosaic analysis gives an estimate of the extent of genomic involvement in the development of the visual system in Drosophila melanogaster. Genetics 131:883–894
Golic KG, Lindquist S (1989) The FLP recombinase of yeast catalyzes site-specific recombination in the Drosophila genome. Cell 59:499–509
Stowers RS, Schwarz TL (1999) A genetic method for generating Drosophila eyes composed exclusively of mitotic clones of a single genotype. Genetics 152:1631–1639
Xu T, Rubin GM (1993) Analysis of genetic mosaics in developing and adult Drosophila tissues. Development 117:1223–1237
Newsome TP, Asling B, Dickson BJ (2000) Analysis of Drosophila photoreceptor axon guidance in eye-specific mosaics. Development 127:851–860
Brand AH, Perrimon N (1993) Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118:401–415
Elliott DA, Brand AH (2008) The GAL4 system: a versatile system for the expression of genes. Methods Mol Biol 420:79–95
McGuire SE, Le PT, Osborn AJ, Matsumoto K, Davis RL (2003) Spatiotemporal rescue of memory dysfunction in Drosophila. Science 302:1765–1768
McGuire SE, Deshazer M, Davis RL (2004) Gene expression systems in Drosophila: a synthesis of time and space. Trends Genet 20:384–391
Fernandez-Funez P, Nino-Rosales ML, de Gouyon B, She WC, Luchak JM, Martinez P, Turiegano E, Benito J, Capovilla M, Skinner PJ, McCall A, Canal I, Orr H, Zoghbi HY, Botas J (2000) Identification of genes that modify ataxin-1-induced neurodegeneration. Nature 408:101–106
Cook T, Zelhof A, Mishra M, Nie J (2011) 800 facets of retinal degeneration. Prog Mol Biol Transl Sci 100:331–368
Lu B (2009) Recent advances in using Drosophila to model neurodegenerative diseases. Apoptosis 14:1008–1020
Bonini NM, Fortini ME (2002) Applications of the Drosophila retina to human disease modeling. In: Moses K (ed) Drosophila eye development. Springer, Heidelberg, pp 257–271
Tepass U, Knust E (1993) Crumbs and stardust act in a genetic pathway that controls the organization of epithelia in Drosophila melanogaster. Dev Biol 159:311–326
Izaddoost S, Nam S-C, Bhat MA, Bellen HJ, Choi K-W (2002) Drosophila crumbs is a positional cue in photoreceptor adherens junctions and rhabdomeres. Nature 416:178–183
Bhat MA, Izaddoost S, Lu Y, Cho KO, Choi KW, Bellen HJ (1999) Discs lost, a novel multi-PDZ domain protein, establishes and maintains epithelial polarity. Cell 96:833–845
Bachmann A, Schneider M, Grawe F, Theilenberg E, Knust E (2001) Drosophila Stardust is a partner of Crumbs in the control of epithelial cell polarity. Nature 414:638–643
Hong Y, Stronach B, Perrimon N, Jan LY, Jan YN (2001) Drosophila Stardust interacts with Crumbs to control polarity of epithelia but not neuroblasts. Nature 414:634–638
Bulgakova NA, Rentsch M, Knust E (2010) Antagonistic functions of Two stardust isoforms in Drosophila photoreceptor cells. Mol Biol Cell 21:3915–3925
Bulgakova NA, Kempkens Ö, Knust E (2008) Multiple domains of Drosophila Stardust differentially mediate localisation of the Crumbs/Stardust complex during photoreceptor development. J Cell Sci 121:2018–2026
Oda H, Uemura T, Harada Y, Iwai Y, Takeichi M (1994) A Drosophila homolog of cadherin associated with armadillo and essential for embryonic cell–cell adhesion. Dev Biol 165:716–726
Riggleman B, Schedl P, Wieschaus E (1990) Spatial expression of the Drosophila segment polarity gene armadillo is posttranscriptionally regulated by wingless. Cell 63:549–560
Karagiosis SA, Ready DF (2004) Moesin contributes an essential structural role in Drosophila photoreceptor morphogenesis. Development 131:725–732
Satoh AK, Li BX, Xia H (2008) Calcium-activated myosin V closes the drosophila pupil. Curr Biol 18:951–955
Lebovitz RM, Takeyasu K, Fambrough DM (1989) Molecular characterization and expression of the (Na+ + K+)-ATPase alpha-subunit in Drosophila melanogaster. EMBO J 8:193–201
Yasuhara JC, Baumann O, Takeyasu K (2000) Localization of Na/K-ATPase in developing and adult Drosophila melanogaster photoreceptors. Cell Tissue Res 300:239–249
Blochlinger K, Bodmer R, Jan LY, Jan YN (1990) Patterns of expression of cut, a protein required for external sensory organ development in wild-type and cut mutant Drosophila embryos. Genes Dev 4:1322–1331
Zipursky SL, Venkatesh TR, Teplow DB, Benzer S (1984) Neuronal development in the Drosophila retina: monoclonal antibodies as molecular probes. Cell 36:15–26
Pielage J, Stork T, Bunse I, Klämbt C (2003) The cell survival gene discs lost encodes a cytoplasmic Codanin-1 like protein, not a homolog of the tight junction PDZ-protein Patj. Dev Cell 5:841–851
Acknowledgments
We thank Michaela Rentsch for help with the electron micrographs, Franziska Friedrich for help in preparing Figs. 3, 4, and 5, and Nagananda Gurudev for the figure of the optical neutralization. Work of E. K. is supported by the Max-Planck Society (MPG) and a grant from the EC (HEALTH-F2-2008-200234).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2012 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Mishra, M., Knust, E. (2012). Analysis of the Drosophila Compound Eye with Light and Electron Microscopy. In: Weber, B., LANGMANN, T. (eds) Retinal Degeneration. Methods in Molecular Biology, vol 935. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-080-9_11
Download citation
DOI: https://doi.org/10.1007/978-1-62703-080-9_11
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-079-3
Online ISBN: 978-1-62703-080-9
eBook Packages: Springer Protocols