Abstract
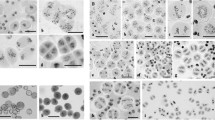
During meiosis, accurate segregation of chromosomes requires the formation of bivalents at metaphase I. In autopolyploids, there are more than two copies of each chromosome with the same chance to form chiasmata at meiosis. This leads to the formation of multivalent configurations in which chiasma quantification is rather complicated. Here, we present an improved cytological protocol, including fluorescence in situ hybridization, to obtain high quality spreads of metaphase I chromosomes from Arabidopsis thaliana autotetraploids. This method allows an accurate analysis of the different meiotic configurations and enables the assessment of the number of chiasmata formed by each tetrasome (group of four homologs).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Masterson J (1994) Stomatal size in fossil plants: evidence for polyploidy in majority of angiosperms. Science 264:421–424. https://doi.org/10.1126/science.264.5157.421
Pozo JC, Ramirez-Parra E (2014) Deciphering the molecular bases for drought tolerance in Arabidopsis autotetraploids. Plant Cell Environ 37:2722–2737. https://doi.org/10.1111/pce.12344
Chao DY, Dilkes B, Luo H et al (2013) Polyploids exhibit higher potassium uptake and salinity tolerance in Arabidopsis. Science 341:658–659. https://doi.org/10.1126/science.1240561
Comai L (2005) The advantages and disadvantages of being polyploid. Nat Rev Genet 6:836–846. https://doi.org/10.1038/nrg1711
Sybenga J (1975) Meiotic configurations: a source of information for estimating genetic parameters. Springer, Berlin
Santos JL, Alfaro D, Sanchez-Moran E et al (2003) Partial Diploidization of meiosis in Autotetraploid Arabidopsis thaliana. Genetics 165:1533–1540
Laibach F (1907) Zur Frage nach der Individualität der Chromosomen im Pflanzenreich. Beih Bot Centralbl 22:191–210
Steinitz-Sears LM (1963) Chromosome studies in Arabidopsis thaliana. Genetics 48:483–490
Fransz P, Armstrong S, Alonso-Blanco C et al (1998) Cytogenetics for the model system Arabidopsis thaliana. Plant J 13:867–876. https://doi.org/10.1046/j.1365-313X.1998.00086.x
Sanchez Moran E, Armstrong SJ, Santos JL et al (2001) Chiasma formation in Arabidopsis thaliana accession Wassileskija and in two meiotic mutants. Chromosom Res 9:121–128. https://doi.org/10.1023/A:1009278902994
Sanchez-Moran E, Armstrong SJ, Santos JL et al (2002) Variation in chiasma frequency among eight accessions of Arabidopsis thaliana. Genetics 162:1415–1422
López E, Pradillo M, Oliver C et al (2012) Looking for natural variation in chiasma frequency in Arabidopsis thaliana. J Exp Bot 63:887–894. https://doi.org/10.1093/jxb/err319
Koorneef M, Fransz P, de Jong H (2003) Cytogenetic tools for Arabidopsis thaliana. Chromosom Res 11:183–194
Bouharmont J, Van De Hende J (1968) Inheritance of lethal chlorophyll mutants in tetraploid Arabidopsis thaliana. Arabidopsis Inform Serv 5:25–26
Bouharmont J (1969) Evolution of chromosome numbers in Arabidopsis polyploids. Chromosomes Today 2:197–201
Morris PC, Altmann T (1994) Tissue culture and transformation. In: Meyerowitz EM, Somerville CR (eds) Cold Spring Harbor laboratory press. Cold Spring Harbor, New York
Heslop-Harrison JS, Maluszynska J (1994) The molecular cytogenetics of Arabidopsis. In: Meyerowitz EM, Sommerville CR (eds) Cold Spring Harbor laboratory press. Cold Spring Harbor, New York
Weiss H, Maluszynska J (2000) Chromosomal rearrangement in autotetraploid plants of Arabidopsis thaliana. Hereditas 133:255–261. https://doi.org/10.1111/j.1601-5223.2000.00255.x
Gerlach WL, Bedbrook JR (1979) Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res 7:1869–1885. https://doi.org/10.1093/nar/7.7.1869
Campell BR, Song Y, Posch TE et al (1992) Sequence and organization of 5S ribosomal RNA-encoding genes of Arabidopsis thaliana. Gene 112:225–228. https://doi.org/10.1016/0378-1119(92)90380-8
Yu Z, Haage K, Streit VE et al (2009) A large number of tetraploid Arabidopsis thaliana lines, generated by a rapid strategy, reveal high stability of neo-tetraploids during consecutive generations. Theor Appl Genet 118:1107–1119. https://doi.org/10.1007/s00122-009-0966-9
Acknowledgments
The authors acknowledge the support of the European Union by the advanced grant Marie Curie Initial Training Network (ITN) “COMREC” (Grant agreement number: 606956). This work has also been partially supported by a grant from the Ministerio de Economía y Competitividad of Spain (AGL2015-67349-P).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Parra-Nunez, P., Pradillo, M., Santos, J.L. (2020). How to Perform an Accurate Analysis of Metaphase I Chromosome Configurations in Autopolyploids of Arabidopsis thaliana. In: Pradillo, M., Heckmann, S. (eds) Plant Meiosis. Methods in Molecular Biology, vol 2061. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9818-0_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9818-0_3
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9817-3
Online ISBN: 978-1-4939-9818-0
eBook Packages: Springer Protocols