Abstract
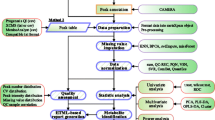
NMR data from large studies combining multiple cohorts is becoming common in large-scale metabolomics. The data size and combination of cohorts with diverse properties leads to special problems for data processing and analysis. These include alignment, normalization, detection and removal of outliers, presence of strong correlations, and the identification of unknowns. Nonetheless, these challenges can be addressed with suitable algorithms and techniques, leading to enhanced data sets ripe for further data mining.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Albanes D, Moore S, Ulrich C, Stolzenberg-Solomon R, Poole E, Temprosa M et al (2017) COnsortium for METabolomics studies (COMETS): leveraging resources to accelerate scientific discovery. FASEB J 30(1):lb129
Soininen P, Kangas AJ, Wurtz P, Suna T, Ala-Korpela M (2015) Quantitative serum nuclear magnetic resonance metabolomics in cardiovascular epidemiology and genetics. Circ Cardiovasc Genet 8(1):192–206
Elliott P, Posma JM, Chan Q, Garcia-Perez I, Wijeyesekera A, Bictash M et al (2015) Urinary metabolic signatures of human adiposity. Sci Transl Med 7(285):285ra62
Holmes E, Loo RL, Stamler J, Bictash M, Yap IK, Chan Q et al (2008) Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 453(7193):396–400
Keun HC, Ebbels TM, Antti H, Bollard ME, Beckonert O, Schlotterbeck G et al (2002) Analytical reproducibility in (1)H NMR-based metabonomic urinalysis. Chem Res Toxicol 15(11):1380–1386
Dumas ME, Maibaum EC, Teague C, Ueshima H, Zhou B, Lindon JC et al (2006) Assessment of analytical reproducibility of (1)H NMR spectroscopy based metabonomics for large-scale epidemiological research: the INTERMAP study. Anal Chem 78(7):2199–2208
Dona AC, Jimenez B, Schafer H, Humpfer E, Spraul M, Lewis MR et al (2014) Precision high-throughput proton NMR spectroscopy of human urine, serum, and plasma for large-scale metabolic phenotyping. Anal Chem 86(19):9887–9894
Viant MR, Bearden DW, Bundy JG, Burton IW, Collette TW, Ekman DR et al (2009) International NMR-based environmental metabolomics intercomparison exercise. Environ Sci Technol 43(1):219–225
Jimenez B, Holmes E, Heude C, Tolson RFM, Harvey N, Lodge SL et al (2018) Quantitative lipoprotein subclass and low molecular weight metabolite analysis in human serum and plasma by 1H NMR spectroscopy in a multilaboratory trial. Anal Chem 90(20):11962–11971
Karaman I, Ferreira DL, Boulange CL, Kaluarachchi MR, Herrington D, Dona AC et al (2016) Workflow for integrated processing of multicohort untargeted 1H NMR metabolomics data in large-scale metabolic epidemiology. J Proteome Res 15(12):4188–4194
Chambers JC, Obeid OA, Refsum H, Ueland P, Hackett D, Hooper J et al (2000) Plasma homocysteine concentrations and risk of coronary heart disease in UK Indian Asian and European men. Lancet 355(9203):523–527
Bild DE, Bluemke DA, Burke GL, Detrano R, Diez Roux AV, Folsom AR et al (2002) Multi-ethnic study of atherosclerosis: objectives and design. Am J Epidemiol 156(9):871–881
Ikram MA, Brusselle GGO, Murad SD, van Duijn CM, Franco OH, Goedegebure A et al (2018) The Rotterdam study: 2018 update on objectives, design and main results. Eur J Epidemiol 32(9):807–850
Beckonert O, Keun HC, Ebbels TM, Bundy J, Holmes E, Lindon JC et al (2007) Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat Protoc 2(11):2692–2703
Karakach TK, Wentzell PD, Walter JA (2009) Characterization of the measurement error structure in 1D 1H NMR data for metabolomics studies. Anal Chim Acta 636(2):163–174
Tredwell GD, Bundy JG, De Iorio M, Ebbels TMD (2016) Modelling the acid/base 1H NMR chemical shift limits of metabolites in human urine. Metabolomics 12(10):1–10
Pearce JTM, Athersuch TJ, Ebbels TMD, Lindon JC, Nicholson JK, Keun HC (2008) Robust algorithms for automated chemical shift calibration of 1D 1H NMR spectra of blood serum. Anal Chem 80(18):7158–7162
Veselkov K, Lindon J, Ebbels T, Volynkin V, Crockford D, Holmes E et al (2009) Recursive segment-wise peak alignment of biological 1H NMR spectra for improved metabolic biomarker recovery. Anal Chem 81(1):56–66
Blaise BJ, Shintu L, Elena B, Emsley L, Dumas M-E, Toulhoat P (2009) Statistical recoupling prior to significance testing in nuclear magnetic resonance based metabonomics. Anal Chem 81(15):6242–6251
Sousa SAA, Magalh+úes A, Ferreira MMC (2013) Optimized bucketing for NMR spectra: three case studies. Chemom Intell Lab Syst 122(0):93–102
Hao J, Astle W, De Iorio M, Ebbels TM (2012) BATMAN--an R package for the automated quantification of metabolites from nuclear magnetic resonance spectra using a Bayesian model. Bioinformatics 28(15):2088–2090
Ravanbakhsh S, Liu P, Bjordahl TC, Mandal R, Grant JR, Wilson M et al (2015) Accurate, fully-automated NMR spectral profiling for metabolomics. PLoS One 10(5):e0124219
Dieterle F, Ross A, Schlotterbeck G, Senn H (2006) Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in H-1 NMR metabonomics. Anal Chem 78(13):4281–4290
Kohl S, Klein M, Hochrein J, Oefner P, Spang R, Gronwald W (2012) State-of-the art data normalization methods improve NMR-based metabolomic analysis. Metabolomics 8(1):146–160
Craig A, Cloarec O, Holmes E, Nicholson JK, Lindon JC (2006) Scaling and normalization effects in NMR spectroscopic metabonomic data sets. Anal Chem 78(7):2262–2267
Sysi-Aho M, Katajamaa M, Yetukuri L, Orešič M (2007) Normalization method for metabolomics data using optimal selection of multiple internal standards. BMC Bioinformatics 8(1):1–17
Bro R, Smilde AK (2014) Principal component analysis. Anal Methods 6(9):2812–2831
van Velzen EJJ, Westerhuis JA, Van Duynhoven JPM, Van Dorsten FA, Hoefsloot HCJ, Jacobs DM et al (2008) Multilevel data analysis of a crossover designed human nutritional intervention study. J Proteome Res 7(10):4483–4491
Karaman İ, Nørskov NP, Yde CC, Hedemann MS, Bach Knudsen KE, Kohler A (2015) Sparse multi-block PLSR for biomarker discovery when integrating data from LC–MS and NMR metabolomics. Metabolomics 11(2):367–379
Couto Alves A, Rantalainen M, Holmes E, Nicholson JK, Ebbels TMD (2009) Analytic properties of statistical total correlation spectroscopy (STOCSY) based information recovery in 1H NMR metabolic data sets. Anal Chem 81(6):2075–2084
Chadeau-Hyam M, Ebbels TM, Brown IJ, Chan Q, Stamler J, Huang CC et al (2010) Metabolic profiling and the metabolome-wide association study: significance level for biomarker identification. J Proteome Res 9(9):4620–4627
Castagne R, Boulange CL, Karaman I, Campanella G, Santos Ferreira DL, Kaluarachchi MR et al (2017) Improving visualization and interpretation of metabolome-wide association studies: an application in a population-based cohort using untargeted (1)H NMR metabolic profiling. J Proteome Res 16(10):3623–3633
Filntisi A, Fotakis C, Asvestas P, Matsopoulos GK, Zoumpoulakis P, Cavouras D (2017) Automated metabolite identification from biological fluid 1H NMR spectra. Metabolomics 13(12):146
Takis PG, Schäfer H, Spraul M, Luchinat C (2017) Deconvoluting interrelationships between concentrations and chemical shifts in urine provides a powerful analysis tool. Nat Commun 8(1):1662
Tardivel PJC, Canlet C, Lefort G, Tremblay-Franco M, Debrauwer L, Concordet D et al (2017) ASICS: an automatic method for identification and quantification of metabolites in complex 1D 1H NMR spectra. Metabolomics 13(10):109
Ludwig C, Viant MR (2009) Two-dimensional J-resolved NMR spectroscopy: review of a key methodology in the metabolomics toolbox. Phytochem Anal 21(1):22–32
Dona AC, Kyriakides M, Scott F, Shephard EA, Varshavi D, Veselkov K et al (2016) A guide to the identification of metabolites in NMR-based metabonomics/metabolomics experiments. Comput Struct Biotechnol J 14:135–153
Cloarec O, Dumas ME, Craig A, Barton RH, Trygg J, Hudson J et al (2005) Statistical total correlation spectroscopy: an exploratory approach for latent biomarker identification from metabolic 1H NMR data sets. Anal Chem 77(5):1282
Robinette SL, Lindon JC, Nicholson JK (2013) Statistical spectroscopic tools for biomarker discovery and systems medicine. Anal Chem 85(11):5297–5303
Posma JM, Garcia-Perez I, De Iorio M, Lindon JC, Elliott P, Holmes E et al (2012) Subset optimization by reference matching (STORM): an optimized statistical approach for recovery of metabolic biomarker structural information from 1H NMR spectra of biofluids. Anal Chem 84(24):10694–10701
Sumner LW, Amberg A, Barrett D, Beale MH, Beger R, Daykin CA et al (2007) Proposed minimum reporting standards for chemical analysis. Metabolomics 3:211–221
Acknowledgments
I.K. and T.E. acknowledge support from the EU PhenoMeNal project (Horizon 2020, 654241). I.K. acknowledges support from the UK Dementia Research Institute, which is supported by the MRC, the Alzheimer’s Society and Alzheimer’s Research UK. T.E. and G.G. acknowledge support by National Institutes of Health (R01HL133932).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ebbels, T.M.D., Karaman, I., Graça, G. (2019). Processing and Analysis of Untargeted Multicohort NMR Data. In: Gowda, G., Raftery, D. (eds) NMR-Based Metabolomics. Methods in Molecular Biology, vol 2037. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9690-2_25
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9690-2_25
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9689-6
Online ISBN: 978-1-4939-9690-2
eBook Packages: Springer Protocols