Abstract
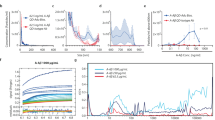
Alzheimer’s disease (AD) pathology is characterized by the presence of extracellular amyloid beta (Aβ), tau hyperphosphorylation, and neuroinflammation. One striking feature in the disease is the clustering of microglia around Aβ plaques. These cells exhibit a highly and chronically activated phenotype, performing a variety of functions, phagocytosis being one of them. Since Aβ phagocytosis by microglia could represent a key aspect in the pathogenesis and progression of the disease, robust and comprehensive methods to evaluate this process are needed. Here, we describe a detailed flow cytometry-based protocol for the analysis of Aβ phagocytosis in vivo.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Winblad B, Amouyel P, Andrieu S et al (2016) Defeating Alzheimer’s disease and other dementias: a priority for European science and society. Lancet Neurol 15:455–532
Sasaguri H, Nilsson P, Hashimoto S et al (2017) APP mouse models for Alzheimer’s disease preclinical studies. EMBO J 36:2473–2487
Baruch K, Deczkowska A, Rosenzweig N et al (2016) PD-1 immune checkpoint blockade reduces pathology and improves memory in mouse models of Alzheimer’s disease. Nat Med 22:135–137
Heneka MT, Carson MJ, Khoury JE et al (2015) Neuroinflammation in Alzheimer’s disease. Lancet Neurol 14:388–405
Heneka MT, Kummer MP, Stutz A et al (2013) NLRP3 is activated in Alzheimer’s disease and contributes to pathology in APP/PS1 mice. Nature 493:674–678
Guerreiro R, Wojtas A, Bras J et al (2013) TREM2 variants in Alzheimer’s disease. N Engl J Med 368:117–127
Hollingworth P, Harold D, Sims R et al (2011) Common variants at ABCA7, MS4A6A/MS4A4E, EPHA1, CD33 and CD2AP are associated with Alzheimer’s disease. Nat Genet 43:429–435
Datta M, Staszewski O, Raschi E et al (2018) Histone deacetylases 1 and 2 regulate microglia function during development, homeostasis, and neurodegeneration in a context-dependent manner. Immunity 48:514–529.e6
Keren-shaul H, Spinrad A, Weiner A et al (2017) Article a unique microglia type associated with restricting development of alzheimer ’ s disease article a unique microglia type associated with restricting development of alzheimer ’ s disease. Cell 169(7):1276–1290.e17
Davalos D, Grutzendler J, Yang G et al (2005) ATP mediates rapid microglial response to local brain injury in vivo. Nat Neurosci 8:752–758
Hickman SE, Kingery ND, Ohsumi TK et al (2013) The microglial sensome revealed by direct RNA sequencing. Nat Neurosci 16:1896–1905
Condello C, Yuan P, Schain A, Grutzendler J (2015) Microglia constitute a barrier that prevents neurotoxic protofibrillar abeta42 hotspots around plaques around plaques. Nat Commun 6:6176
Lucin KM, O’Brien CE, Bieri G et al (2013) Microglial beclin 1 regulates retromer trafficking and phagocytosis and is impaired in Alzheimer’s disease. Neuron 79:873–886
Paolicelli RC, Bolasco G, Pagani F et al (2011) Synaptic pruning by microglia is necessary for normal brain development. Science 333:1456–1458
Czirr E, Castello NA, Mosher KI et al (2017) Microglial complement receptor 3 regulates brain Aβ levels through secreted proteolytic activity. J Exp Med 214(4):1081–1092
Bradshaw EM, Chibnik LB, Keenan BT et al (2013) CD33 Alzheimer’s disease locus: altered monocyte function and amyloid biology. Nat Neurosci 16:848–850
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Tejera, D., Heneka, M.T. (2019). In Vivo Phagocytosis Analysis of Amyloid Beta. In: Garaschuk, O., Verkhratsky, A. (eds) Microglia. Methods in Molecular Biology, vol 2034. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9658-2_21
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9658-2_21
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9657-5
Online ISBN: 978-1-4939-9658-2
eBook Packages: Springer Protocols