Abstract
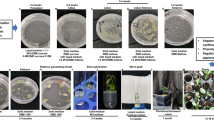
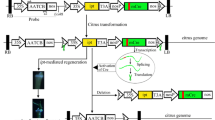
Conventional breeding of citrus types demands a long-term effort due to their complex reproductive biology and long juvenile period. As a compelling alternative, genetic engineering of mature tissues allows the insertion of specific traits into specific elite cultivars, including well-known and widely grown varieties and rootstocks, thus reducing the time and costs involved in improving and evaluating them. Conventional breeding for resistance to CTV in citrus varieties has been largely unsuccessful as well as cloning of the genes conferring resistance to specific citrus types. RNA interference (RNAi), based on producing dsRNAs (usually using intron-hairpin constructs) highly homologous to specific CTV sequences to trigger RNA silencing, has been employed to produce virus-resistant transgenic citrus plants. The most successful construct has been an intron-hairpin vector carrying full-length, untranslatable versions of the genes p25, p20, and p23 from the virus. Using it, we have generated full resistance against CTV in Mexican lime. Moreover, this strategy is applicable to all those citrus varieties amenable to mature transformation, including sweet oranges, sour oranges, mandarins, Citrus macrophylla, and limes.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Moreno P, Ambrós S, Albiach-Martí MR et al (2008) Citrus tristeza virus: a pathogen that changed the course of the citrus industry. Mol Plant Pathol 9:251–268
Yoshida Y (1985) Inheritance of susceptibility to Citrus tristeza virus in trifoliate orange. Bull Fruit Tree Res Stn 12:17–25
Yoshida T (1993) Inheritance of immunity to Citrus tristeza virus of trifoliate orange in some citrus intergeneric hybrids. Bull Fruit Tree Res Stn 25:33–43
Gmitter FG, Xiao SY, Huang S et al (1996) A localized linkage map of the Citrus tristeza virus resistance gene region. Theor Appl Genet 92:688–695
Fire A, Xu S, Montgomery MK et al (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Smith NA, Singh SP, Wang MB et al (2000) Total silencing by intron-spliced hairpin RNAs. Nature 407:319–320
Soler N, Fagoaga C, Chiibi S et al (2011) RNAi-mediated protection against Citrus tristeza virus in transgenic Citrus plants. In: Erdmann V, Barciszewski J (eds) Non coding RNAs in plants. RNA Technologies. Springer, Berlin, Heidelberg
Soler N, Plomer M, Fagoaga C et al (2012) Transformation of Mexican lime with an intron-hairpin construct expressing untranslatable versions of the genes coding for the three silencing suppressors of Citrus tristeza virus confers complete resistance to the virus. Plant Biotechnol J 10:597–608
Fagoaga C, López C, Hermoso de Mendoza A et al (2006) Post-transcriptional gene silencing of the p23 silencing suppressor of Citrus tristeza virus confers resistance to the virus in transgenic Mexican lime. Plant Mol Biol 60:153–165
Ruiz-Ruiz S, Navarro B, Gisel A et al (2011) Citrus tristeza virus infection induces the accumulation of viral small RNAs (21–24-nt) mapping preferentially at the 3′-terminal region of the genomic RNA and affects the host small RNA profile. Plant Mol Biol 75:607–619
Lu R, Folimonov A, Shintaku M et al (2004) Three distinct suppressors of RNA silencing encoded by a 20-kb viral RNA genome. Proc Natl Acad Sci U S A 101:15742–15747
Peña L, Cervera M, Fagoaga C et al (2008) Citrus. In: Kole C, Hall TC (eds) Compendium of transgenic crop plants: tropical and subtropical fruits and nuts. Blackwell Publishing, Oxford, pp 1–62
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassay with tobacco tissue cultures. Physiol Plant 15:473–479
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: b-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Domínguez A, Hermoso de Mendoza A, Guerri J et al (2002) Pathogen-derived resistance to Citrus tristeza virus (CTV) in transgenic Mexican lime (Citrus aurantifolia (Christ.) swing.) plants expressing its p25 coat protein gene. Mol Breed 10:1–10
Cambra M, Garnsey SM, Permar TA et al (1990) Detection of Citrus tristeza virus (CTV) with a mixture of monoclonal antibodies. Phytopathology 80:103
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Soler, N. et al. (2019). Methods for Producing Transgenic Plants Resistant to CTV. In: Catara, A., Bar-Joseph, M., Licciardello, G. (eds) Citrus Tristeza Virus. Methods in Molecular Biology, vol 2015. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9558-5_17
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9558-5_17
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9557-8
Online ISBN: 978-1-4939-9558-5
eBook Packages: Springer Protocols