Abstract
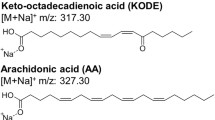
LC-MS/MS with multiple reaction monitoring (MRM) is a powerful tool for targeted metabolomics analysis including screening and quantification of known metabolites. Given the complexity of biological samples, the difference in ionization efficiency, and signal intensity of each metabolite, isotopically labeled internal standards are often used for accurate quantification. In this chapter, we describe a detailed protocol for the quantitative analysis of polyunsaturated fatty acids (PUFAs) and their oxidized products (oxylipins) by LC-MS/MS-MRM with isotope dilution. PUFAs are very susceptible to oxidation by both enzymatic and nonenzymatic pathways. Free PUFAs and corresponding oxylipins, known as bioactive lipids, are involved in many processes with varying biological functions depending on their chemical structure and concentration. Accurate quantification is thus becoming crucial to understanding the role of these bioactive lipids in health, disease(s), and other settings.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Guma M, Tiziani S, Firestein GS (2016) Metabolomics in rheumatic diseases: desperately seeking biomarkers. Nat Rev Rheumatol 12:269–281
Hill CB, Bacic A, Roessner U (2014) LC-MS profiling to link metabolic and phenotypic diversity in plant mapping populations. Methods Mol Biol 1198:29–41
Qi Y, Song Y, Gu H et al (2014) Global metabolic profiling using ultra-performance liquid chromatography/quadrupole time-of-flight mass spectrometry. Methods Mol Biol 1198:15–27
Ruiz-Canela M, Hruby A, Clish CB et al (2017) Comprehensive Metabolomic profiling and incident cardiovascular disease: a systematic review. J Am Heart Assoc 6
Woodside JV, Draper J, Lloyd A et al (2017) Use of biomarkers to assess fruit and vegetable intake. Proc Nutr Soc 76:308–315
Begou O, Gika HG, Wilson ID et al (2017) Hyphenated MS-based targeted approaches in metabolomics. Analyst 142:3079–3100
Lu W, Bennett BD, Rabinowitz JD (2008) Analytical strategies for LC-MS-based targeted metabolomics. J Chromatogr B Analyt Technol Biomed Life Sci 871:236–242
Wei R, Li G, Seymour AB (2014) Multiplexed, quantitative, and targeted metabolite profiling by LC-MS/MRM. Methods Mol Biol 1198:171–199
Wei R, Li G, Seymour AB (2010) High-throughput and multiplexed LC/MS/MRM method for targeted metabolomics. Anal Chem 82:5527–5533
Savolainen OI, Sandberg AS, Ross AB (2016) A simultaneous metabolic profiling and quantitative multimetabolite Metabolomic method for human plasma using gas-chromatography tandem mass spectrometry. J Proteome Res 15:259–265
Maclennan MS, Kok MGM, Soliman L et al (2018) Capillary electrophoresis-mass spectrometry for targeted and untargeted analysis of the sub-5kDa urine metabolome of patients with prostate or bladder cancer: a feasibility study. J Chromatogr B Analyt Technol Biomed Life Sci 1074-1075:79–85
Pannkuk EL, Laiakis EC, Authier S et al (2016) Targeted metabolomics of nonhuman primate serum after exposure to ionizing radiation: potential tools for high-throughput Biodosimetry. RSC Adv 6:51192–51202
Zhu J, Djukovic D, Deng L et al (2014) Colorectal cancer detection using targeted serum metabolic profiling. J Proteome Res 13:4120–4130
Gasperotti M, Masuero D, Guella G et al (2014) Development of a targeted method for twenty-three metabolites related to polyphenol gut microbial metabolism in biological samples, using SPE and UHPLC-ESI-MS/MS. Talanta 128:221–230
Datta PK, Deshmane S, Khalili K et al (2016) Glutamate metabolism in HIV-1 infected macrophages: role of HIV-1 Vpr. Cell Cycle 15:2288–2298
Masoodi M, Volmer DA (2014) Comprehensive quantitative determination of PUFA-related bioactive lipids for functional lipidomics using high-resolution mass spectrometry. Methods Mol Biol 1198:221–232
Fu X, Felcyn JR, Odem-Davis K et al (2016) Bioactive lipids accumulate in stored red blood cells despite leukoreduction: a targeted metabolomics study. Transfusion 56:2560–2570
Zimring JC, Slichter S, Odem-Davis K et al (2016) Metabolites in stored platelets associated with platelet recoveries and survivals. Transfusion 56:1974–1983
Wang Y, Armando AM, Quehenberger O et al (2014) Comprehensive ultra-performance liquid chromatographic separation and mass spectrometric analysis of eicosanoid metabolites in human samples. J Chromatogr A 1359:60–69
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Fu, X., Anderson, M., Wang, Y., Zimring, J.C. (2019). LC-MS/MS-MRM-Based Targeted Metabolomics for Quantitative Analysis of Polyunsaturated Fatty Acids and Oxylipins. In: D'Alessandro, A. (eds) High-Throughput Metabolomics. Methods in Molecular Biology, vol 1978. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9236-2_7
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9236-2_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9235-5
Online ISBN: 978-1-4939-9236-2
eBook Packages: Springer Protocols