Abstract
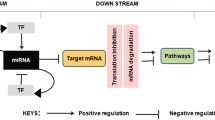
This chapter is devoted to illustrate the usage of state-of-the-art methodologies for miRNA regulatory network construction and analysis. Advantages in understanding the role of miRNAs in regulating gene expression are increasing the possibility of developing targeted therapies and drugs. This new possibility can be exploited by gaining new knowledge through analyzing interactions between a specific miRNA and a targeted gene.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Flynt AS et al (2008) Biological principles of microRNA-mediated regulation: shared themes amid diversity. Nat Rev Genet 9(11):831–842. https://doi.org/10.1038/nrg2455nn
Gurtan AM et al (2013) The role of miRNAs in regulating gene expression networks. J Mol Biol 425(19):3582–3600. https://doi.org/10.1016/j.jmb.2013.03.007
Hobert O (2008) Gene regulation by transcription factors and microRNAs. Science 319(5871):1785–1786. https://doi.org/10.1126/science.1151651
Baglioni M, Russo F, Geraci F, Rizzo M, Rainaldi G, Pellegrini M (2015) A new method for discovering disease-specific MiRNA-target regulatory networks. PLoS One 10(4). https://doi.org/10.1371/journal.pone.0122473
Russo F, Belling K, Jensen AB, Scoyni F, Brunak S, Pellegrini M (2017) MicroRNAs, regulatory networks, and comorbidities: decoding complex systems. In: Dalmay T (ed) MicroRNA detection and target identification, Methods in molecular biology, vol 1580. Humana Press, New York, NY, pp 281–295
R Core Team (2017) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Cava C, Colaprico A, Bertoli G et al (2017) SpidermiR: An R/Bioconductor Package for Integrative Analysis with miRNA Data. Int J Mol Sci 18(2):274. https://doi.org/10.3390/ijms18020274
Pajak M, Simpson TI (2016) miRNAtap: miRNAtap: microRNA targets–aggregated predictions. R package version 112.0
Bonnici V et al (2018) http://arenaidb.ba.itb.cnr.it. Accessed 30 Apr 2018
Fan Y, Siklenka K, Arora SK, Ribeiro P, Kimmins S, Xia J (2016) miRNet - dissecting miRNA-target interactions and functional associations through network-based visual analysis. Nucleic Acids Res 44(Web Server issue):W135–W141. https://doi.org/10.1093/nar/gkw288
Bisognin A, Sales G, Coppe A, Bortoluzzi S, Romualdi C (2012) MAGIA2: from miRNA and genes expression data integrative analysis to microRNA–transcription factor mixed regulatory circuits. Nucleic Acids Res 40(Web Server issue):W13–W21. https://doi.org/10.1093/nar/gks460n
Huber W, Carey VJ, Gentleman R et al (2015) Orchestrating high-throughput genomic analysis with Bioconductor. Nat Methods 12(2):115–121. https://doi.org/10.1038/nmeth.3252
RStudio Team (2015) RStudio: integrated development for R. RStudio, Inc., Boston, MA. URL http://www.rstudio.com/
Alexa A, Rahnenfuhrer J (2016) topGO: enrichment analysis for gene ontology. R package version 2.30.1
Colaprico A, Silva TC, Olsen C et al (2016) TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res 44(8):e71. https://doi.org/10.1093/nar/gkv1507
Rukov JL, Wilentzik R, Jaffe I, Vinther J, Shomron N (2014) Pharmaco-miR: linking microRNAs and drug effects. Brief Bioinform 15(4):648–659. https://doi.org/10.1093/bib/bbs082
Warde-Farley D, Donaldson SL, Comes O et al (2010) The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res 38(Web Server issue):W214–W220. https://doi.org/10.1093/nar/gkq537
Grillari J, Hackl M, Grillari-Voglauer R (2010) miR-17–92 cluster: ups and downs in cancer and aging. Biogerontology 11(4):501–506. https://doi.org/10.1007/s10522-010-9272-9
Hossain A, Kuo MT, Saunders GF (2006) Mir-17-5p regulates breast cancer cell proliferation by inhibiting translation of AIB1 mRNA. Mol Cell Biol 26(21):8191–8201. https://doi.org/10.1128/MCB.00242-06
Agarwal V, Bell GW, Nam J-W, Bartel DP (2015) Predicting effective microRNA target sites in mammalian mRNAs. elife 4:e05005. https://doi.org/10.7554/eLifhbe.05005
Karagkouni D, Paraskevopoulou MD, Chatzopoulos S et al (2018) DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA–gene interactions. Nucleic Acids Res 46(Database issue):D239–D245. https://doi.org/10.1093/nar/gkx1141
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS (2004) Human MicroRNA Targets. PLoS Biol 2(11):e363. https://doi.org/10.1371/journal.pbio.0020363
Krek A, Grün D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, MacMenamin P, da Piedade I, Gunsalus KC, Stoffel M, Rajewsky N (2005) Combinatorial microRNA target predictions. Nat Genet 37(5):495–500. https://doi.org/10.1038/ng1536
Wong N, Wang X (2015) miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res 43(D1):D146–D152. https://doi.org/10.1093/nar/gku1104
Carlson M (2017) org.Hs.eg.db: Genome wide annotation for Human. R package version 3.5.0
Ashburner M, Ball CA, The Gene Ontology Consortium et al (2000) Gene Ontology: tool for the unification of biology. Nat Genet 25(1):25–29. https://doi.org/10.1038/75556
The Gene Ontology Consortium (2017) Expansion of the gene ontology knowledgebase and resources. Nucleic Acids Res 245(Database issue):D331–D338. https://doi.org/10.1093/nar/gkw1108
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45(Database issue):D353–D361. https://doi.org/10.1093/nar/gkw1092
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44(Database issue):D457–D462. https://doi.org/10.1093/nar/gkv1070
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Sales G, Coppe A, Bisognin A, Biasiolo M, Bortoluzzi S, Romualdi C (2010) MAGIA, a web-based tool for miRNA and genes integrated analysis. Nucleic Acids Res 38(Web Server issue):W352–W359. https://doi.org/10.1093/nar/gkq423
Kertesz M et al (2007) The role of site accessibility in microRNA target recognition. Nat Genet 39(10):1278–1284. https://doi.org/10.1038/ng2135
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Huang DW, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13. https://doi.org/10.1093/nar/gkn923
Pajak M and Simpson TI (2016) miRNAtap.db: Data for miRNAtap. R package version 0.99.10
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Mensi, A., Bonnici, V., Caligola, S., Giugno, R. (2019). Construction and Analysis of miRNA Regulatory Networks. In: Laganà, A. (eds) MicroRNA Target Identification. Methods in Molecular Biology, vol 1970. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-9207-2_9
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9207-2_9
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-9206-5
Online ISBN: 978-1-4939-9207-2
eBook Packages: Springer Protocols