Abstract
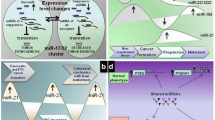
The transcription factor p73 synthesizes a large number of isoforms and presents high structural and functional homology with p53, a well-known tumor suppressor and a famous “Holy Grail” of anticancer targeting. p73 has attracted increasing attention mainly because (a) unlike p53, p73 is rarely mutated in cancer, (b) some p73 isoforms can inhibit all hallmarks of cancer, and (c) it has the ability to mimic oncosuppressive functions of p53, even in p53-mutated cells. These attributes render p73 and its downstream pathways appealing for therapeutic targeting, especially in mutant p53-driven cancers. p73 functions are, at least partly, mediated by microRNAs (miRNAs), which constitute nodal components of p73-governed networks. p73 not only regulates transcription of crucial miRNA genes, but is also predicted to affect miRNA populations in a transcription-independent manner by developing protein-protein interactions with components of the miRNA processing machinery. This combined effect of p73, both in miRNA transcription and maturation, appears to be isoform-dependent and can result in a systemic switch of cell miRNomes toward either an anti-oncogenic or oncogenic outcome. In this review, we combine literature search with bioinformatics approaches to reconstruct the p73-governed miRNA network and discuss how these crosstalks may be exploited to develop next-generation therapeutics.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Abraham CG, Espinosa JM (2015) The crusade against mutant p53: does the COMPASS point to the holy grail? Cancer Cell 28(4):407–408. https://doi.org/10.1016/j.ccell.2015.09.019
Olivier M, Hollstein M, Hainaut P (2010) TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol 2(1):a001008. https://doi.org/10.1101/cshperspect.a001008
Giaretti W, Rapallo A, Sciutto A, Macciocu B, Geido E, Hermsen MA, Postma C, Baak JP, Williams RA, Meijer GA (2000) Intratumor heterogeneity of k-ras and p53 mutations among human colorectal adenomas containing early cancer. Anal Cell Pathol 21(2):49–57
Ren ZP, Olofsson T, Qu M, Hesselager G, Soussi T, Kalimo H, Smits A, Nistér M (2007) Molecular genetic analysis of p53 intratumoral heterogeneity in human astrocytic brain tumors. J Neuropathol Exp Neurol 66(10):944–954. https://doi.org/10.1097/nen.0b013e318156bc05
Logotheti S, Pavlopoulou A, Galtsidis S, Vojtesek B, Zoumpourlis V (2013) Functions, divergence and clinical value of TAp73 isoforms in cancer. Cancer Metastasis Rev 32(3–4):511–534. https://doi.org/10.1007/s10555-013-9424-x
Engelmann D, Meier C, Alla V, Pützer BM (2015) A balancing act: orchestrating amino-truncated and full-length p73 variants as decisive factors in cancer progression. Oncogene 34(33):4287–4299. https://doi.org/10.1038/onc.2014.365
Stiewe T, Zimmermann S, Frilling A, Esche H, Pützer BM (2002) Transactivation-deficient DeltaTA-p73 acts as an oncogene. Cancer Res 62(13):3598–3602
Ming L, Sakaida T, Yue W, Jha A, Zhang L, Yu J (2008) Sp1 and p73 activate PUMA following serum starvation. Carcinogenesis 29(10):1878–1884. https://doi.org/10.1093/carcin/bgn150
Chakraborty J, Banerjee S, Ray P, Hossain DM, Bhattacharyya S, Adhikary A, Chattopadhyay S, Das T, Sa G (2010) Gain of cellular adaptation due to prolonged p53 impairment leads to functional switchover from p53 to p73 during DNA damage in acute myeloid leukemia cells. J Biol Chem 285(43):33104–33112. https://doi.org/10.1074/jbc.M110.122705
John K, Alla V, Meier C, Pützer BM (2011) GRAMD4 mimics p53 and mediates the apoptotic function of p73 at mitochondria. Cell Death Differ 18(5):874–886. https://doi.org/10.1038/cdd.2010.153
Nelson P, Kiriakidou M, Sharma A, Maniataki E, Mourelatos Z (2003) The microRNA world: small is mighty. Trends Biochem Sci 28(10):534–540. https://doi.org/10.1016/j.tibs.2003.08.005
Shen J, Hung MC (2015) Signaling-mediated regulation of MicroRNA processing. Cancer Res 75(5):783–791. https://doi.org/10.1158/0008-5472.CAN-14-2568
Xiao Y, Xu C, Guan J, Ping Y, Fan H, Li Y, Zhao H, Li X (2012) Discovering dysfunction of multiple microRNAs cooperation in disease by a conserved microRNA co-expression network. PLoS One 7(2):e32201. https://doi.org/10.1371/journal.pone.0032201
Schmitz U, Lai X, Winter F, Wolkenhauer O, Vera J, Gupta SK (2014) Cooperative gene regulation by microRNA pairs and their identification using a computational workflow. Nucleic Acids Res 42(12):7539–7552. https://doi.org/10.1093/nar/gku465
Lai X, Gupta SK, Schmitz U, Marquardt S, Knoll S, Spitschak A, Wolkenhauer O, Pützer BM, Vera J (2018) MiR-205-5p and miR-342-3p cooperate in the repression of the E2F1 transcription factor in the context of anticancer chemotherapy resistance. Theranostics 8(4):1106–1120. https://doi.org/10.7150/thno.19904
Nazarov PV, Reinsbach SE, Muller A, Nicot N, Philippidou D, Vallar L, Kreis S (2013) Interplay of microRNAs, transcription factors and target genes: linking dynamic expression changes to function. Nucleic Acids Res 41(5):2817–2831. https://doi.org/10.1093/nar/gks1471
Sengupta D, Bandyopadhyay S (2013) Topological patterns in microRNA-gene regulatory network: studies in colorectal and breast cancer. Mol BioSyst 9(6):1360–1371. https://doi.org/10.1039/c3mb25518b
Hermeking H (2012) MicroRNAs in the p53 network: micromanagement of tumour suppression. Nat Rev Cancer 12(9):613–626. https://doi.org/10.1038/nrc3318
Galtsidis S, Logotheti S, Pavlopoulou A, Zampetidis CP, Papachristopoulou G, Scorilas A, Vojtesek B, Gorgoulis V, Zoumpourlis V (2017) Unravelling a p73-regulated network: the role of a novel p73-dependent target, MIR3158, in cancer cell migration and invasiveness. Cancer Lett 388:96–106. https://doi.org/10.1016/j.canlet.2016.11.036
Bracken CP, Scott HS, Goodall GJ (2016) A network-biology perspective of microRNA function and dysfunction in cancer. Nat Rev Genet 17(12):719–732. https://doi.org/10.1038/nrg.2016.134
Cheng F, Jia P, Wang Q, Lin CC, Li WH, Zhao Z (2014) Studying tumorigenesis through network evolution and somatic mutational perturbations in the cancer interactome. Mol Biol Evol 31(8):2156–2169. https://doi.org/10.1093/molbev/msu167
Zhu Y, Skogerbø G, Ning Q, Wang Z, Li B, Yang S, Sun H, Li Y (2012) Evolutionary relationships between miRNA genes and their activity. BMC Genomics 13:718. https://doi.org/10.1186/1471-2164-13-718
Koufaris C (2016) Human and primate-specific microRNAs in cancer: evolution, and significance in comparison with more distantly-related research models: the great potential of evolutionary young microRNA in cancer research. BioEssays 38(3):286–294. https://doi.org/10.1002/bies.201500135
Niwa R, Slack FJ (2007) The evolution of animal microRNA function. Curr Opin Genet Dev 17(2):145–150. https://doi.org/10.1016/j.gde.2007.02.004
Meunier J, Lemoine F, Soumillon M, Liechti A, Weier M, Guschanski K, Hu H, Khaitovich P, Kaessmann H (2013) Birth and expression evolution of mammalian microRNA genes. Genome Res 23(1):34–45. https://doi.org/10.1101/gr.140269.112
Ory B, Ramsey MR, Wilson C, Vadysirisack DD, Forster N, Rocco JW, Rothenberg SM, Ellisen LW (2011) A microRNA-dependent program controls p53-independent survival and chemosensitivity in human and murine squamous cell carcinoma. J Clin Invest 121(2):809–820. https://doi.org/10.1172/JCI43897
Jacques C, Calleja LR, Baud’huin M, Quillard T, Heymann D, Lamoureux F, Ory B (2016) miRNA-193a-5p repression of p73 controls Cisplatin chemoresistance in primary bone tumors. Oncotarget 7(34):54503–54514. https://doi.org/10.18632/oncotarget.10950
Teng Y, Ren Y, Hu X, Mu J, Samykutty A, Zhuang X, Deng Z, Kumar A, Zhang L, Merchant ML, Yan J, Miller DM, Zhang HG (2017) MVP-mediated exosomal sorting of miR-193a promotes colon cancer progression. Nat Commun 8:14448. https://doi.org/10.1038/ncomms14448
Tran N (2016) Cancer exosomes as miRNA factories. Trends Cancer 2(7):329–331. https://doi.org/10.1016/j.trecan.2016.05.008
Gao Q, Zheng J (2018) microRNA-323 upregulation promotes prostate cancer growth and docetaxel resistance by repressing p73. Biomed Pharmacother 97:528–534. https://doi.org/10.1016/j.biopha.2017.10.040
Jiang X, Li H (2018) MiR-1180-5p regulates apoptosis of Wilms’ tumor by targeting. Onco Targets Ther 11:823–831. https://doi.org/10.2147/OTT.S148684
Kim T, Veronese A, Pichiorri F, Lee TJ, Jeon YJ, Volinia S, Pineau P, Marchio A, Palatini J, Suh SS, Alder H, Liu CG, Dejean A, Croce CM (2011) p53 regulates epithelial-mesenchymal transition through microRNAs targeting ZEB1 and ZEB2. J Exp Med 208(5):875–883. https://doi.org/10.1084/jem.20110235
Piovan C, Palmieri D, Di Leva G, Braccioli L, Casalini P, Nuovo G, Tortoreto M, Sasso M, Plantamura I, Triulzi T, Taccioli C, Tagliabue E, Iorio MV, Croce CM (2012) Oncosuppressive role of p53-induced miR-205 in triple negative breast cancer. Mol Oncol 6(4):458–472. https://doi.org/10.1016/j.molonc.2012.03.003
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G, Vadas MA, Khew-Goodall Y, Goodall GJ (2008) The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10(5):593–601. https://doi.org/10.1038/ncb1722
Knouf EC, Garg K, Arroyo JD, Correa Y, Sarkar D, Parkin RK, Wurz K, O’Briant KC, Godwin AK, Urban ND, Ruzzo WL, Gentleman R, Drescher CW, Swisher EM, Tewari M (2012) An integrative genomic approach identifies p73 and p63 as activators of miR-200 microRNA family transcription. Nucleic Acids Res 40(2):499–510. https://doi.org/10.1093/nar/gkr731
Alla V, Kowtharapu BS, Engelmann D, Emmrich S, Schmitz U, Steder M, Pützer BM (2012) E2F1 confers anticancer drug resistance by targeting ABC transporter family members and Bcl-2 via the p73/DNp73-miR-205 circuitry. Cell Cycle 11(16):3067–3078. https://doi.org/10.4161/cc.21476
Lu Z, Jiao D, Qiao J, Yang S, Yan M, Cui S, Liu Z (2015) Restin suppressed epithelial-mesenchymal transition and tumor metastasis in breast cancer cells through upregulating mir-200a/b expression via association with p73. Mol Cancer 14:102. https://doi.org/10.1186/s12943-015-0370-9
Agostini M, Knight RA (2014) miR-34: from bench to bedside. Oncotarget 5(4):872–881. https://doi.org/10.18632/oncotarget.1825
Meier C, Hardtstock P, Joost S, Alla V, Pützer BM (2016) p73 and IGF1R regulate emergence of aggressive cancer stem-like features via miR-885-5p control. Cancer Res 76(2):197–205. https://doi.org/10.1158/0008-5472.CAN-15-1228
Zhang Y, Liao JM, Zeng SX, Lu H (2011) p53 downregulates Down syndrome-associated DYRK1A through miR-1246. EMBO Rep 12(8):811–817. https://doi.org/10.1038/embor.2011.98
Liao JM, Zhou X, Zhang Y, Lu H (2012) MiR-1246: a new link of the p53 family with cancer and Down syndrome. Cell Cycle 11(14):2624–2630. https://doi.org/10.4161/cc.20809
Batliner J, Buehrer E, Fey MF, Tschan MP (2012) Inhibition of the miR-143/145 cluster attenuated neutrophil differentiation of APL cells. Leuk Res 36(2):237–240. https://doi.org/10.1016/j.leukres.2011.10.006
Rossi M, De Laurenzi V, Munarriz E, Green DR, Liu YC, Vousden KH, Cesareni G, Melino G (2005) The ubiquitin-protein ligase Itch regulates p73 stability. EMBO J 24(4):836–848. https://doi.org/10.1038/sj.emboj.7600444
Sampath D, Calin GA, Puduvalli VK, Gopisetty G, Taccioli C, Liu CG, Ewald B, Liu C, Keating MJ, Plunkett W (2009) Specific activation of microRNA106b enables the p73 apoptotic response in chronic lymphocytic leukemia by targeting the ubiquitin ligase Itch for degradation. Blood 113(16):3744–3753. https://doi.org/10.1182/blood-2008-09-178707
O’Malley BW, Kumar R (2009) Nuclear receptor coregulators in cancer biology. Cancer Res 69(21):8217–8222. https://doi.org/10.1158/0008-5472.CAN-09-2223
Koeppel M, van Heeringen SJ, Kramer D, Smeenk L, Janssen-Megens E, Hartmann M, Stunnenberg HG, Lohrum M (2011) Crosstalk between c-Jun and TAp73alpha/beta contributes to the apoptosis-survival balance. Nucleic Acids Res 39(14):6069–6085. https://doi.org/10.1093/nar/gkr028
Nakano K, Bálint E, Ashcroft M, Vousden KH (2000) A ribonucleotide reductase gene is a transcriptional target of p53 and p73. Oncogene 19(37):4283–4289
Suzuki HI, Yamagata K, Sugimoto K, Iwamoto T, Kato S, Miyazono K (2009) Modulation of microRNA processing by p53. Nature 460(7254):529–533. https://doi.org/10.1038/nature08199
Boominathan L (2010) The tumor suppressors p53, p63, and p73 are regulators of microRNA processing complex. PLoS One 5(5):e10615. https://doi.org/10.1371/journal.pone.0010615
Yeom KH, Lee Y, Han J, Suh MR, Kim VN (2006) Characterization of DGCR8/Pasha, the essential cofactor for Drosha in primary miRNA processing. Nucleic Acids Res 34(16):4622–4629. https://doi.org/10.1093/nar/gkl458
Wang Y, Medvid R, Melton C, Jaenisch R, Blelloch R (2007) DGCR8 is essential for microRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat Genet 39(3):380–385. https://doi.org/10.1038/ng1969
Rupaimoole R, Slack FJ (2017) MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov 16(3):203–222. https://doi.org/10.1038/nrd.2016.246
van Rooij E, Purcell AL, Levin AA (2012) Developing microRNA therapeutics. Circ Res 110(3):496–507. https://doi.org/10.1161/CIRCRESAHA.111.247916
Skourti E, Logotheti S, Kontos CK, Pavlopoulou A, Dimoragka PT, Trougakos IP, Gorgoulis V, Scorilas A, Michalopoulos I, Zoumpourlis V (2016) Progression of mouse skin carcinogenesis is associated with the orchestrated deregulation of mir-200 family members, mir-205 and their common targets. Mol Carcinog 55(8):1229–1242. https://doi.org/10.1002/mc.22365
Villanueva J, Vultur A, Lee JT, Somasundaram R, Fukunaga-Kalabis M, Cipolla AK, Wubbenhorst B, Xu X, Gimotty PA, Kee D, Santiago-Walker AE, Letrero R, D’Andrea K, Pushparajan A, Hayden JE, Brown KD, Laquerre S, McArthur GA, Sosman JA, Nathanson KL, Herlyn M (2010) Acquired resistance to BRAF inhibitors mediated by a RAF kinase switch in melanoma can be overcome by cotargeting MEK and IGF-1R/PI3K. Cancer Cell 18(6):683–695. https://doi.org/10.1016/j.ccr.2010.11.023
Ganju A, Khan S, Hafeez BB, Behrman SW, Yallapu MM, Chauhan SC, Jaggi M (2017) miRNA nanotherapeutics for cancer. Drug Discov Today 22(2):424–432. https://doi.org/10.1016/j.drudis.2016.10.014
Xie Y, Murray-Stewart T, Wang Y, Yu F, Li J, Marton LJ, Casero RA, Oupický D (2017) Self-immolative nanoparticles for simultaneous delivery of microRNA and targeting of polyamine metabolism in combination cancer therapy. J Control Release 246:110–119. https://doi.org/10.1016/j.jconrel.2016.12.017
Wilusz JE, Sunwoo H, Spector DL (2009) Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 23(13):1494–1504. https://doi.org/10.1101/gad.1800909
Huarte M (2015) The emerging role of lncRNAs in cancer. Nat Med 21(11):1253–1261. https://doi.org/10.1038/nm.3981
Zhang A, Xu M, Mo YY (2014) Role of the lncRNA-p53 regulatory network in cancer. J Mol Cell Biol 6(3):181–191. https://doi.org/10.1093/jmcb/mju013
Acknowledgments
This work was supported by the German Cancer Aid, Dr. Mildred Scheel Stiftung [grant 70112353], the German Research Foundation (DFG) [grant PU188/17-1], Wilhelm Sander-Stiftung [grant 2015.036.1], German Federal Ministry of Education and Research (BMBF) grant 0316171 as part of the project eBio:SysMet, and Rostock University Medical Faculty for the project Systems Medicine of Cancer Invasion and Metastasis to B.M.P.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Logotheti, S., Marquardt, S., Pützer, B.M. (2019). p73-Governed miRNA Networks: Translating Bioinformatics Approaches to Therapeutic Solutions for Cancer Metastasis. In: Lai, X., Gupta, S., Vera, J. (eds) Computational Biology of Non-Coding RNA. Methods in Molecular Biology, vol 1912. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8982-9_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8982-9_2
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8981-2
Online ISBN: 978-1-4939-8982-9
eBook Packages: Springer Protocols