Abstract
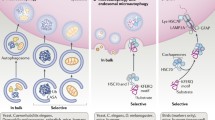
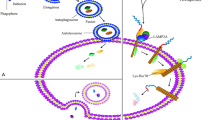
Chaperone-mediated autophagy (CMA) is a selective type of autophagy whereby a specific subset of intracellular proteins is targeted to the lysosome for degradation. These proteins are identified by a chaperone that targets them to lysosomes. There, they are translocated into the organelle lumen through a lysosomal membrane receptor/translocation complex. CMA plays an important role in maintaining cellular proteostasis by eliminating damaged and altered proteins. CMA also participates in the control of the cellular energetic balance through recycling of amino acids resulting from lysosomal proteolysis of the substrate proteins. Lastly, due to the intrinsic protein selectivity of CMA, this type of autophagy exerts regulatory functions by mediating timely degradation of key cellular proteins that participate in processes such as lipid and glucose metabolism, cell cycle, DNA repair, and cellular reprogramming, among others. Dysfunctional CMA occurs with age and has now been described in a growing list of human pathologies such as metabolic disorders, neurodegeneration, cancer, immunodeficiency, and diabetes. In this chapter, we describe current methodologies to quantitatively analyze CMA activity in different experimental models.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kaushik S, Cuervo AM (2012) Chaperone-mediated autophagy: a unique way to enter the lysosome world. Trends Cell Biol 22:407–417
Dice JF (1990) Peptide sequences that target cytosolic proteins for lysosomal proteolysis. Trends Biochem Sci 15:305–309
Chiang H et al (1989) A role for a 70-kilodalton heat shock protein in lysosomal degradation of intracellular proteins. Science 246:382–385
Cuervo AM, Dice JF (1996) A receptor for the selective uptake and degradation of proteins by lysosomes. Science 273:501–503
Bandyopadhyay U et al (2008) The chaperone-mediated autophagy receptor organizes in dynamic protein complexes at the lysosomal membrane. Mol Cell Biol 28:5747–5763
Agarraberes F, Terlecky S, Dice J (1997) An intralysosomal hsp70 is required for a selective pathway of lysosomal protein degradation. J Cell Biol 137:825–834
Bandyopadhyay U et al (2010) Identification of regulators of chaperone-mediated autophagy. Mol Cell 39:535–547
Arias E et al (2015) Lysosomal mTORC2/PHLPP1/Akt Regulate Chaperone-Mediated Autophagy. Mol Cell 59:270–284
Sahu R et al (2011) Microautophagy of cytosolic proteins by late endosomes. Develop Cell 20:131–139
Salvador N et al (2000) Import of a cytosolic protein into lysosomes by chaperone-mediated autophagy depends on its folding state. J Biol Chem 275:27447–27456
Cuervo AM et al (1995) Activation of a selective pathway of lysosomal proteolysis in rat liver by prolonged starvation. Am J Phys 269:C1200–C1208
Wing SS et al (1991) Proteins containing peptide sequences related to KFERQ are selectively depleted in liver and heart, but not skeletal muscle, of fasted rats. Biochem J 275:165–169
Kiffin R et al (2004) Activation of chaperone-mediated autophagy during oxidative stress. Mol Biol Cell 15:4829–4840
Schneider JL, Suh Y, Cuervo AM (2014) Deficient chaperone-mediated autophagy in liver leads to metabolic dysregulation. Cell Metab 20:417–432
Park C, Suh Y, Cuervo AM (2015) Regulated degradation of Chk1 by chaperone-mediated autophagy in response to DNA damage. Nat Commun 6:6823
Valdor R et al (2014) Chaperone-mediated autophagy regulates T cell responses through targeted degradation of negative regulators of T cell activation. Nat Immunol 15:1046–1054
Cuervo AM, Dice JF (2000) Age-related decline in chaperone-mediated autophagy. J Biol Chem 275:31505–31513
Cuervo AM et al (2004) Impaired degradation of mutant alpha-synuclein by chaperone-mediated autophagy. Science 305:1292–1295
Kiffin R et al (2007) Altered dynamics of the lysosomal receptor for chaperone-mediated autophagy with age. J Cell Sci 120:782–791
Orenstein SJ et al (2013) Interplay of LRRK2 with chaperone-mediated autophagy. Nat Neurosci 16:394–406
Anguiano J et al (2013) Chemical modulation of chaperone-mediated autophagy by retinoic acid derivatives. Nat Chem Biol 9:374–382
Berger JJ, Dice JF (1985) Effect of serum deprivation and replacement on proteolysis in cultured human fibroblasts. Prog Clin Biol Res 180:479–481
Massey AC et al (2006) Consequences of the selective blockage of chaperone-mediated autophagy. Proc Natl Acad Sci U S A 103:5905–5910
Massey AC et al (2008) Early cellular changes after blockage of chaperone-mediated autophagy. Autophagy 4:442–456
Koga H et al (2011) A photoconvertible fluorescent reporter to track chaperone-mediated autophagy. Nat Commun 2:386
Cuervo AM, Dice JF, Knecht E (1997) A population of rat liver lysosomes responsible for the selective uptake and degradation of cytosolic proteins. J Biol Chem 272:5606–5615
Finn P et al (2005) Effects of small molecules on chaperone-mediated autophagy. Autophagy 1:141–145
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Juste, Y.R., Cuervo, A.M. (2019). Analysis of Chaperone-Mediated Autophagy. In: Ktistakis, N., Florey, O. (eds) Autophagy. Methods in Molecular Biology, vol 1880. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8873-0_47
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8873-0_47
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8872-3
Online ISBN: 978-1-4939-8873-0
eBook Packages: Springer Protocols